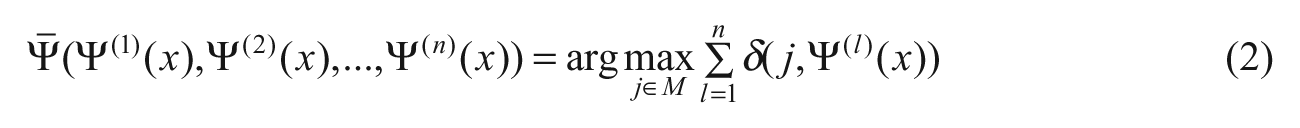Abstract
Pattern recognition and machine learning methods provide an attractive approach for building decision support systems. Classification trees are frequently used algorithms for such tasks owing to their intuitive structure and effectiveness. It has been shown that for complex medical data, combining a number of base classifiers improves their overall accuracy. Classification tree ensembles have a certain number of free parameters to set, which can significantly affect their performance. In recent years such ensembles were often used by practitioners without a mathematical background (e.g. physicians), who may be unaware of how to obtain the optimal settings. Therefore, it is difficult for them to choose the satisfactory properties, while in most of the cases the default parameters proposed for them are not necessarily the most efficient. The aim of this article is to ascertain which types of combined tree classifiers give the best performance for medical decision support and which parameters should be chosen for them. A set of rules for end-users on how to tune their ensembles is proposed.
Keywords
Introduction
Machine learning is an attractive approach for building decision support systems. 1 In such tasks the quality of the knowledge base plays the key role in establishing satisfactory final performance. The following problems are often encountered:
domain experts cannot formulate the rules for decision problems because they might not have the knowledge needed to develop effective algorithms (e.g. human face recognition from images);
incomplete, unbalanced or noisy data owing to the mechanic fault or high cost of information acquisition if often required to be accommodated;
it is not possible to ascertain which classifier will behave better than the others—known as the ‘no free lunch’ theorem 2 ;
the number of classifiers to test is very high and performing a complex investigation would be too time consuming.
The latter two problems are addressed in the current article. Recently, pattern recognition and machine learning have become interdisciplinary areas of research, attracting researchers from different backgrounds, as well as users of classification techniques to automate the decision-making process. 3 Yet, owing to the high complexity of the pattern recognition problem and the great variety of methods and their settings, one may find the task bewildering when attempting to choose an appropriate procedure. 4 This is especially the case in medicine, where decision support systems play an important role.5–7 It is extremely difficult or nearly impossible to guarantee in all cases that individual experts in pattern recognition would agree on the best methods to tune a given system for a specific problem, owing, in part, to the numerous methods that are available. As a consequence, many users, in this case physicians, are often left to their own judgement, which may be lacking in some cases. This article aims to present a set of complex tests of performance that will highlight the strengths and weaknesses of the chosen classification methods. Here, the focus is on the decision trees owing to their intuitive nature and good performance over a variety of problems. 8 Additionally, it is chosen to investigate the behaviour of tree ensembles. 9 Combining basic classifiers is considered as one of the most promising trends in pattern recognition. 10 Classifier ensembles applied for the analysis of complex data often increase the final accuracy, especially in the case of weak predictors, such as decision trees. 11 Additionally, the ensembles, compared with the canonical classifiers, have more free parameters that must be set, therefore making their proper tuning a challenging problem. It is the aim that this article will help practitioners, offering the so-needed advice for using ensembles, based on real data tests.
The content of this article is arranged as follows: in the next section the related work in the field of classification tree ensembles is presented; then, the theoretical framework behind pattern recognition, classification trees and tree ensembles methodologies is briefly presented. After that, the setting-up of computer experiments is discussed, followed by the results and a thorough discussion, and a set of guidance notes and advice on how to tune the tree ensembles. Finally, we present the conclusions.
Related work
Decision trees and their ensembles represent a group of popular methods for medical data classification and are often used in decision support systems. They are constantly being developed by academic researchers. Šprogar et al. 12 thoroughly described how the idea of creating medical decision support systems has changed over last few decades. Ordonez 13 showed that in the task of disease prediction, decision trees often behave better than explicit association rules. Rodriguez et al. 14 studied the methods of functional trees ensembles, yet without giving a clear indication of how they should be tuned for certain medical problems.
The Random Forest algorithm is probably the most popular ensemble algorithm created in recent years. Wu et al. 15 showed that the Random Forest algorithm outperforms most of the canonical classifiers for the prediction of ovarian cancer. It is widely used in bioinformatics, as it can also serve at the same time as a classifier and a feature selector, as shown in Díaz-Uriarte et al. 16 One of the novel usages of this algorithm is the application in problems with unbalanced data representation, which occur often when sufficient samples of pathological examples cannot be collected. One of the first works to report such usage was the paper by Khalilia et al. 17
The Rotation Forest algorithm is a recently introduced method for creating a classifier ensemble. While it is not frequently used, possibly because of its relatively recent emergence, there are some reports of its successful implementation in the medical domain. For example, Ozcift et al. 18 presented the behaviour of 30 different machine-learning algorithms combined using the Rotation Forest algorithm for the medical domain. It is noted, however, that the tests were conducted on a very small number of datasets. In different work, Ozcift 19 presented a combination of feature selection with both Support Vector Machine and Rotation Forest for Parkinson disease diagnosis. Dehzangi et al. 20 reported the successful usage of Rotation Forest for the prediction of protein folds. One of the main drawback of Rotation Forest is that it often does not show a good performance for small datasets with a very high number of features. Therefore, alternative ensemble methods were proposed for such cases, such as those presented by Krawczyk, 21 based on the combination of feature selection and a random subspace method. Another approach for complex data analysis was proposed by Wilk et al. 22 Here, the authors used the one-class classification framework and created the ensemble with the usage of fuzzy logic.
Background
Pattern recognition task
The aim of pattern recognition 23 is to classify a given object to one of the predefined categories on the basis of an observation of the features which describe it. All data concerning the object and its attributes are presented as a feature vector x X. The pattern recognition algorithm Ψ maps the feature space X to the set of class labels M:
The mapping (1) is established based on the examples included in a learning set or rules given by experts. The learning set consists of learning examples, i.e. observations of the features describing an object and its correct classification. Let us assume that there are n classifiers
where δ(j,i) = 0 if i ≠ j, otherwise δ(j,i) = 1. The presented approach is based on the 0-1 loss function. It is worth mentioning that recently a new model of the loss function based on fuzzy logic was proposed. 24 It was also tuned for the implementation in a decision tree algorithm. 25
Decision tree induction
Most algorithms, such as C4.5 given by Quinlan 26 or Alternative Decision Tree (ADTree), are based on the ‘Top Down Induction of Decision Tree’ (TDIDT). 27 The central idea of the TDIDT algorithm is the selection of ‘the best’ attribute, i.e. which attribute to test at each node in the tree. The family of algorithms based on the ID3 method (e.g. C4.5) uses the information gain that measures how well the given attribute separates the training examples according to the target classification. The future implementations of decision tree induction algorithms use measures based on the previously defined information gain (e.g. information ratio).
REPTree and CART
All decision tree algorithms tested in this article are constructed based on the information gain. The main difference among them is the pruning process. A fast decision tree learner uses reduced-error pruning (REPTree) 28 and requires a validation set. Each subtree of a given decision tree is substituted with the best possible ‘leaf’ and then tested using the validation set. If the error is lower than or equal to the previous tree structure, then the subtree is replaced by a leaf. The process continues for all subtrees in the tree.
CART
29
is a class of decision trees that implements minimal cost-complexity pruning. As in the case of REPTree, it also replaces each subtree with a best possible leaf. The difference is, however, that the performance of the modified tree can be lower if it brings about a significant reduction of the tree structure, i.e. the number of leaves. This can be more formally written as follows: for a given decision tree
where α is some parameter (yet to be determined). For a substitution of a subtree
where
Bagging
Bagging (or bootstrap aggregating) is an ensemble meta-algorithm developed by Breiman. 30 It is based on creating a set of new bootstrap object samples from the original dataset and training one classifier on each of them. This assures that each of the classifiers was created on a diverse, heterogeneous dataset.
A dataset is given consisting of
Random forest
The Random Forest approach was introduced by Breiman. 31 It is, to some extent, an extension of the bootstrap aggregation 30 algorithm. It is similar to the previously presented Bagging algorithm as it also uses new subsets to create heterogeneous classifiers. Yet, in this case, the subspaces do not only consist of bootstrap samples of objects, but also of randomly chosen features. Whereas in Bagging each classifier is trained on the same features but on different objects, in Random Forest each tree is trained on different objects and different features.
Let
Rotation Forest
The Rotation Forest is an algorithm for creating classifier ensembles using feature extraction proposed by Rodríguez et al. 32 As a base classifier the decision tree is used. Additionally, it implements principal component analysis (PCA) to increase accuracy and diversity of the ensemble.
The algorithm works as follows: let
Experimental investigation
The main idea behind the experiment is to show that in many practical cases, the results given by ensemble classifiers with standard configurations could be significantly enhanced by choosing appropriate parameter values. The goal is to give some insight on the performance of the presented combined tree classifiers and the effect of different parameter settings on the final accuracy. This can be viewed as an aid for dealing with practical implementation issues in the domain of medical decision support systems. The focus is not on the settings of the base classifiers (single decision trees) and classifier fuser, but rather it is shown that more attention should be paid to the design and tuning of the classifier ensembles. 33 The default fusion methods proposed by the software are used. Because of the focus on medical decision support, all datasets represent real life medical problems. In total, 15 different data sets are chosen, all of which are publically available at the UC Irvine repository. 34 Each dataset applies to a distinct medical condition or a group of diseases—most of which are commonly encountered in an everyday medical practice.
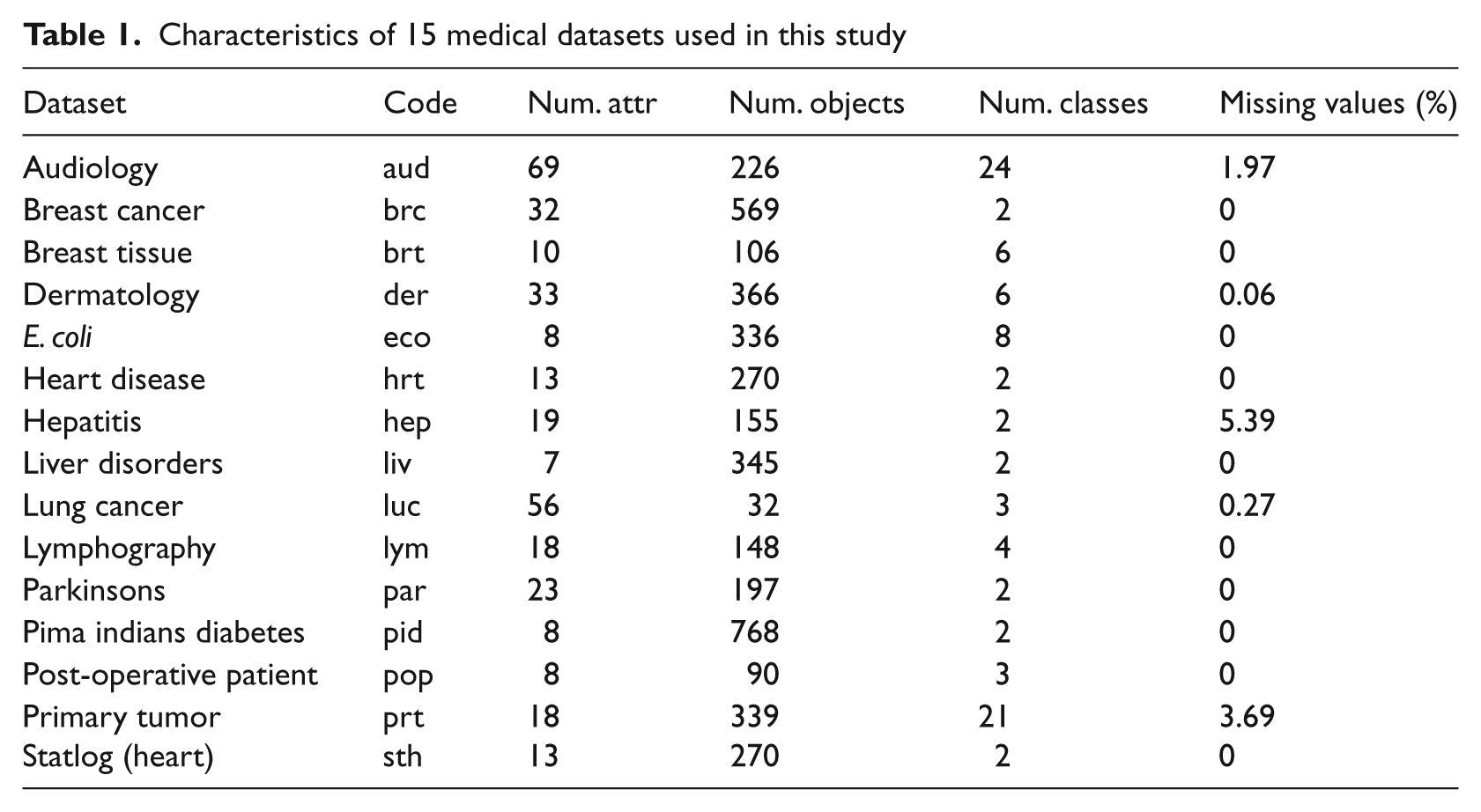
The datasets depict different forms of classification problems. The number of instances range from small datasets (such as lung cancer) with 32 objects to quite large datasets, i.e. more than 500 for breast Cancer and pima indians diabetes. The number of attributes also differs significantly from 8 (E. coli) to 69 (audiology). The attributes are mostly real, binary and nominal values. Five of the tested datasets have missing values, which for some classification methods may have a significant impact on the overall performance. The detailed descriptions of the datasets under consideration are presented in Table 1.
Characteristics of 15 medical datasets used in this study
For the purpose of these experiments two popular ensemble methods are chosen: Random Forest and Bagging. Additionally, a quite recently introduced method known as the Rotation Forest is considered. For the two former algorithms a base classifier must be chosen. In this article the focus is on decision trees, so the base classifiers tested are the three methods from this group of algorithms. These are as follows: C4.5 (the most popular decision tree algorithm), CART and REPTree. Each ensemble method has a unique set of parameters that can be changed and its effect on the outcomes measured.
In the case of Bagging, the percentage of the whole dataset in a training subset was studied. Here, if the parameter is equal to 100 then the training subset has the same size as the full training set. However, they are not the same, because the subset is generated by uniformly sampling the training set with replacement, which leads to duplicates. This can have a negative effect on the performance. To deal with this problem, smaller values of this parameter are investigated, namely 70% and 40%. These are arbitrarily chosen values drawn from experience in working in the machine learning domain. The differences between them in most cases are very clear and, therefore, these values give a clear indication of the influence of this parameter on the overall accuracy. Smaller values could also influence the diversity of the base classifiers by training them on potentially fewer identical instances.
In the Rotation Forest approach two parameters were studied—the number and size of the disjoined subsets of the feature space. The two parameters are mutually exclusive. Both parameters have been tested using the same values, namely 2 and 4.
The influence on the performance of two important factors is now quantified. The number of base classifiers is tested for five values, namely 10, 20, 40, 80 and 120 trees. Also, the depth of each of the base trees is set to one of three values: two arbitrary levels, the default level and twice the default level.
To provide a detailed comparison between the methods a statistical significance test is used. It allows to compare the tested classifiers and to ascertain whether their differences are, indeed, statistically significant. For this purpose, a Combined 5 × 2 cv F Test 35 is used. It repeats five-time twofold cross-validation so that in each of the folds the size of the training and testing sets is equal. This test is conducted by comparison of all versus all. As a test score the probability of rejecting the null hypothesis is adopted, i.e. that classifiers have the same error rates. As an alternative hypothesis, it is conjectured that tested classifiers have different error rates. A small difference in the error rate implies that the different algorithms construct two similar classifiers with similar error rates; thus, the hypothesis should not be rejected. For a large difference, the classifiers have different error rates and the hypothesis should be rejected. Therefore, two classifiers differ in a statistically significant way if the null hypothesis considering them is rejected. The combined version of this test takes a majority vote over the ten possible 5 × 2cv F test results. The benefit of using such a modification is that the combined test has a lower type II error, i.e. a lower probability of rejecting the hypothesis when the classifiers have similar error rates. At the same time, it offers higher power such that there is a larger probability of rejecting when the tested classifiers are different.
All the classification results were obtained with the use of Weka, 36 open source software for data analysis.
Results
In this section the results obtained by experimental investigations are presented. In each of the following tables the best results for a dataset are marked by the grey field. Additionally, the classifiers that showed statistical differences in the Combined 5 × 2 cv F Test are in bold.
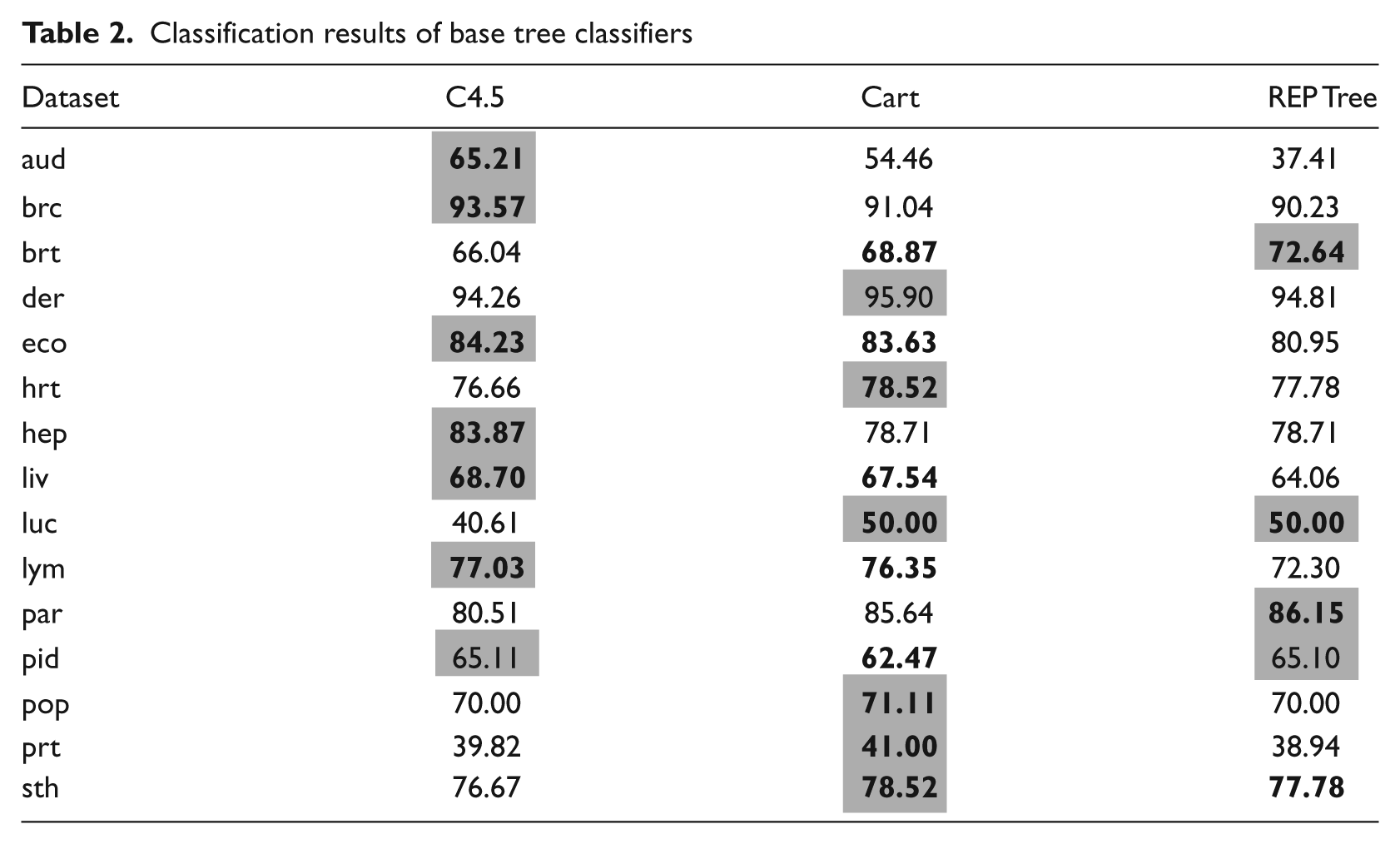
As a base for the further comparison the results of the base tree classifiers are shown in Table 2.
Classification results of base tree classifiers
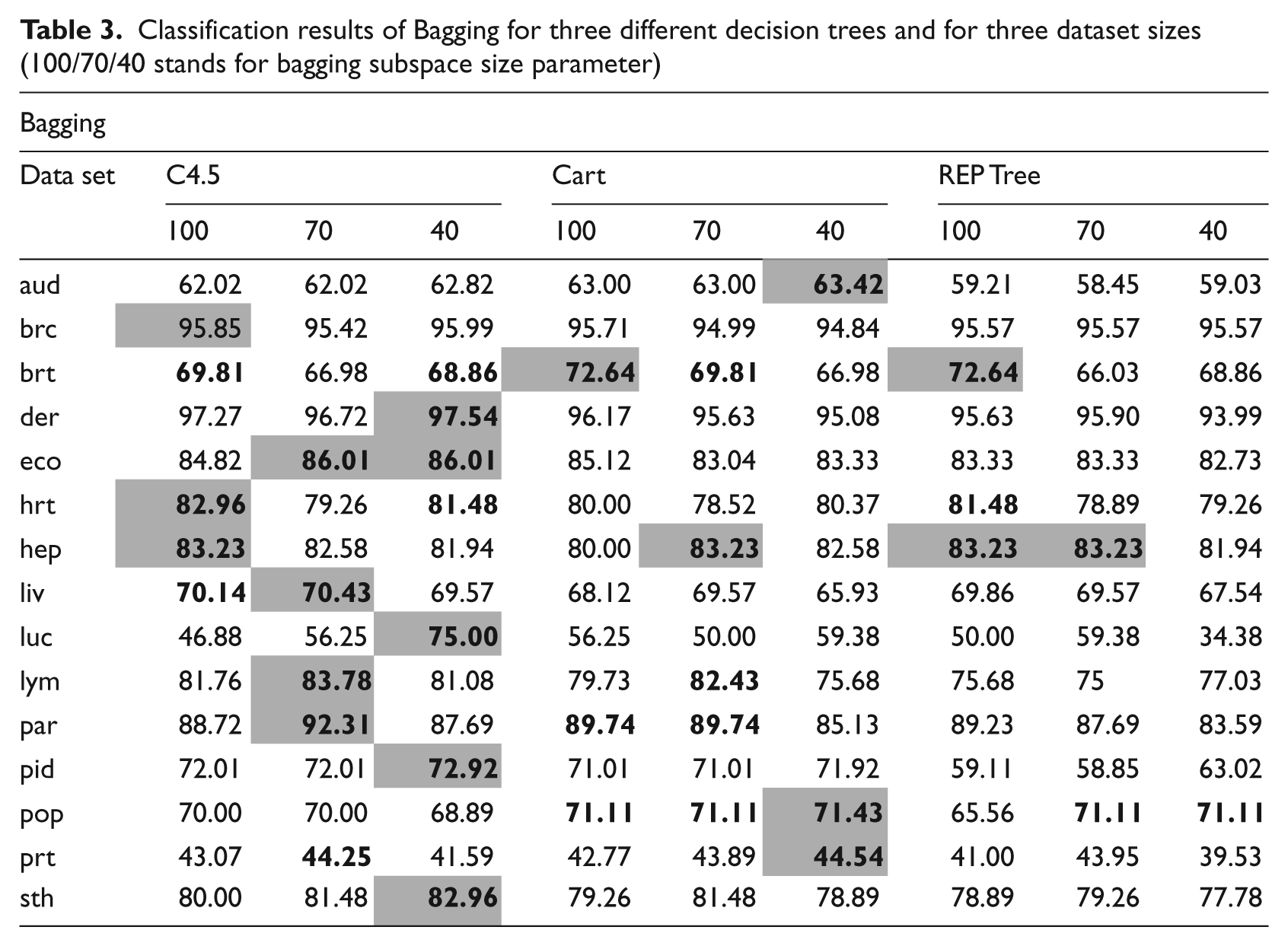
Tables 3–5 present the classification accuracy over 10 datasets for Bagging, Rotation Forest and Random Forest respectively. Table 6 presents the comparison between the best results from three investigated methods.
Classification results of Bagging for three different decision trees and for three dataset sizes (100/70/40 stands for bagging subspace size parameter)
Performance of the Rotation Forest algorithm for three different decision tree classifiers
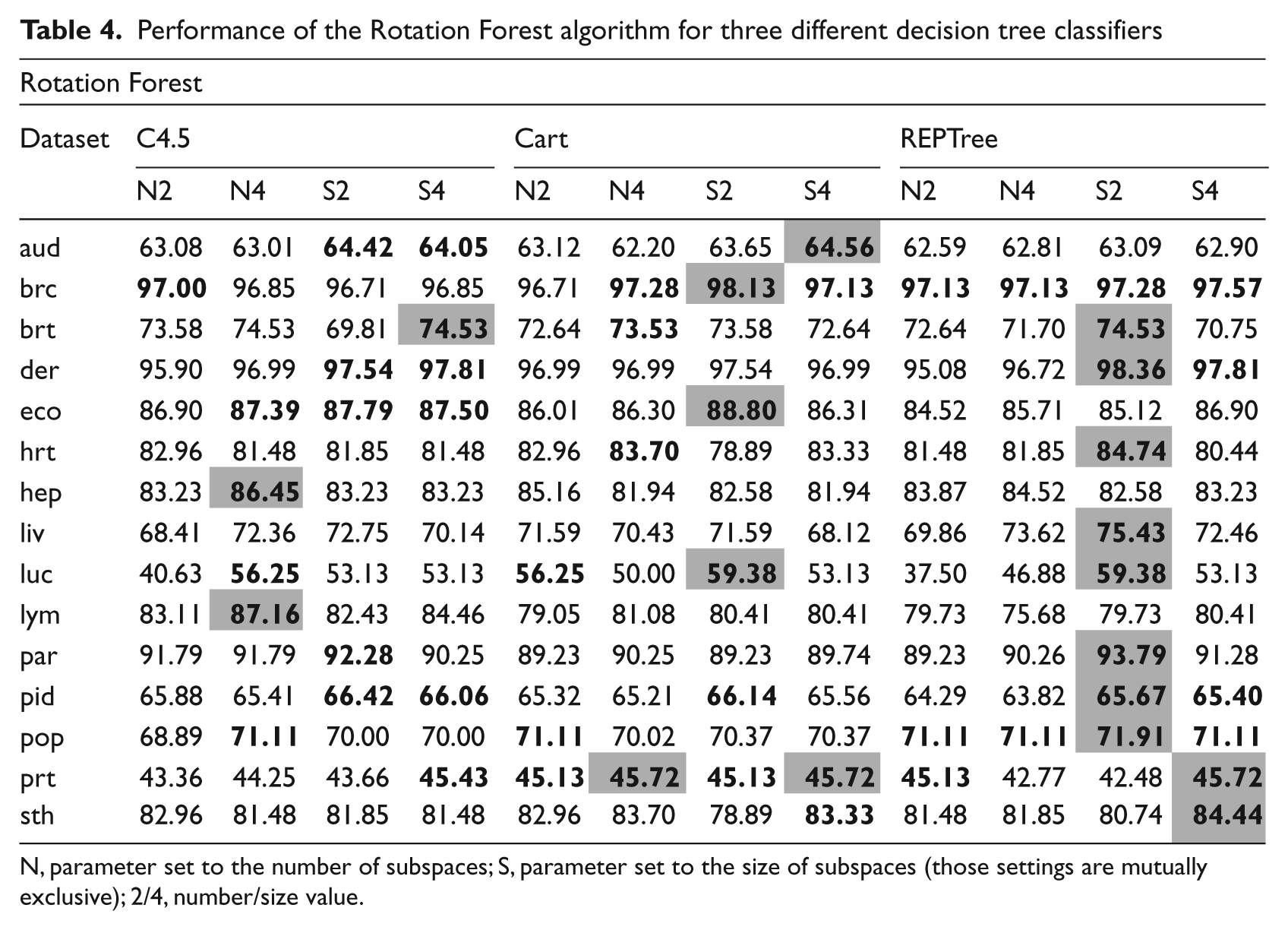
N, parameter set to the number of subspaces; S, parameter set to the size of subspaces (those settings are mutually exclusive); 2/4, number/size value.
Performance of the Random Forest algorithm for different tree depth and ensemble size
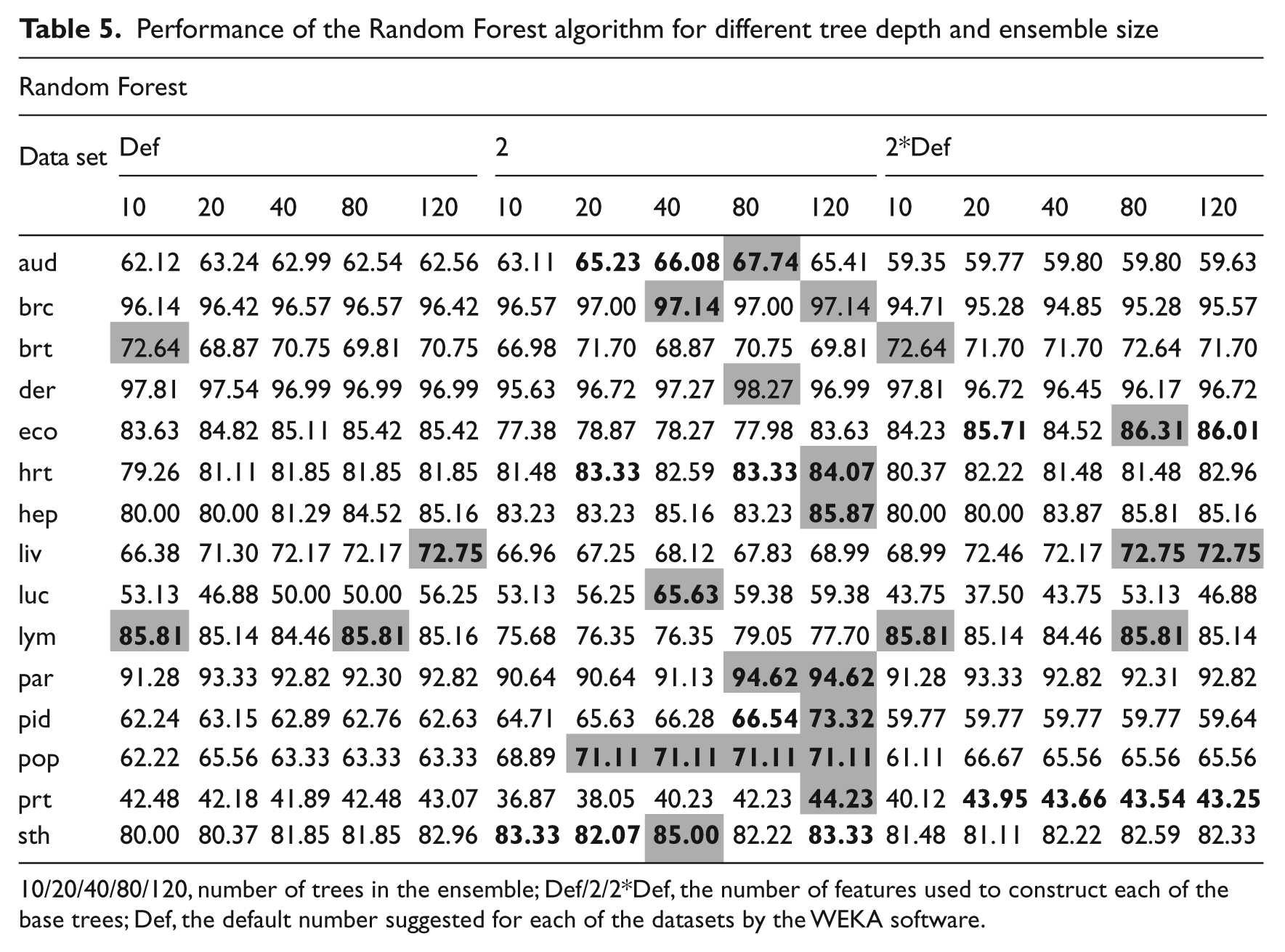
10/20/40/80/120, number of trees in the ensemble; Def/2/2*Def, the number of features used to construct each of the base trees; Def, the default number suggested for each of the datasets by the WEKA software.
Comparison of the best results for three ensemble methods under consideration
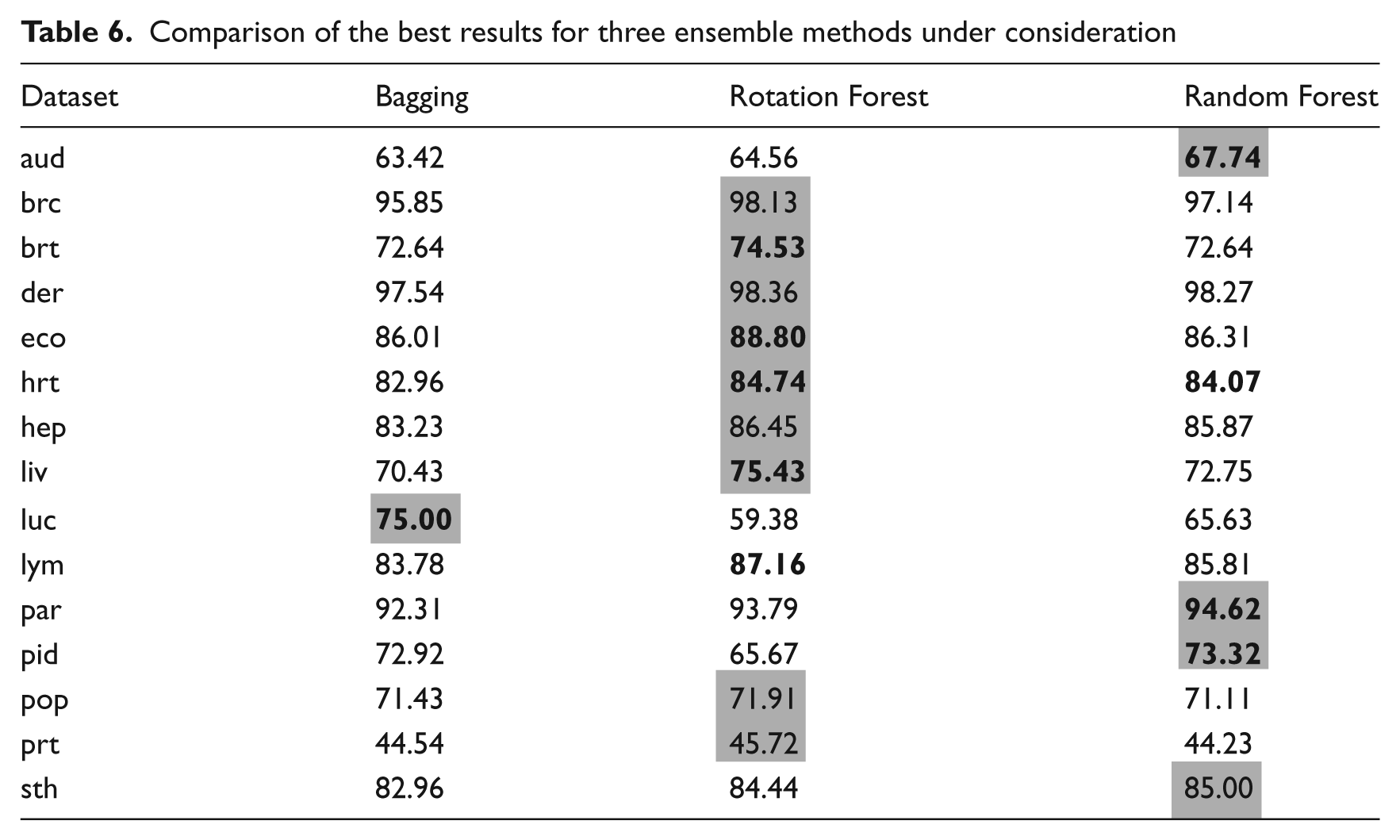
Discussion
Analysing the results from the experiments allows several interesting conclusions to be drawn. It is worth noting that none of the base classifiers outperformed the ensemble methods. This may be owing to the complex nature of the medical data. It is observed that when attempting to use a single classifier trying to model the whole decision space the result is that it tends to either overfit or to have too large a generalisation. In contrast, it is found that ensemble methods, with different areas of competence for each of the classifiers, can overcome this difficulty. The smallest accuracy gain occurred in the case of the breast tissue dataset (<2%) and the largest in the case of the lung cancer dataset (25%). This clearly shows that using tree ensembles is appropriate and the accuracy gain can compensate for the longer classifier training time.
When comparing the best results from the three different classifier ensemble types, it can be seen clearly that Bagging has produced the most inferior. Only in the case of the lung cancer dataset did it outperform the remaining two algorithms. In the case of other datasets the differences between Random Forest and Rotation Forest are very small. For four cases, Random Forest returned the best results, while Rotation Forest was superior for nine. It is worth noting that differences between them were not always statistically significant. This leads to one conclusion that both of these algorithms present good predictive power. However, the Rotation Forest execution time is much greater than that of Random Forest.
In case of the Bagging algorithm, the size of the subspaces and choice of the base classifier could be adjusted. From the experiments it can be seen that use of a REPTree algorithm for Bagging is not a promising direction. For all datasets it was outperformed by the C4.5 and CART. Only for the hepatitis dataset did it return good results, but, at the same time, the two Forest algorithms returned identical results. Therefore, it can be stated that the REPTree should not be used for Bagging. In fact, it was found that C4.5 displayed the highest accuracy for the Bagging procedure. For 8 out of 10 datasets this combination returned the best results. CART proved to be useful in the breast tissue and audiology datasets, but in the latter one it was not statistically significant. In most cases of the Bagging test the smaller size of the subspaces, namely 70% and, especially, 40% returned the best results. The subspace set to 70% returned best results for 6 out of 15 datasets and when set to 40% the best for 8 out of 15.
In the case of the Rotation Forest algorithm, the number or the size of the disjoined subsets of the feature space and choice of the base classifier can be manipulated. For this approach the REPTree returned the best results, surprisingly outperforming in more than half the other two base classifiers considered, i.e. 10 out of 15 cases. This can be explained by the pruning nature of the REPTree. Rotation Forest uses the PCA method for feature extraction and it was shown many times that only a small number of principal components contain most of the discriminant information. Therefore, REPTree allows quick discarding of the irrelevant nodes from the tree, resulting in the highest performance. In second place was the CART algorithm (three best and one equal performance). The worst performance was obtained from the use of C4.5 (one best and one equal). This is in complete contrast to the Bagging approach and it shows the differences between these ensemble methods. As for the other free parameters the best results were given by the size of the subspaces set to 2 for 11 out of 15 datasets. When size was set to four better results were obtained only in 3 cases out of 15. However, experimental results showed that there is little to be gained in changing the number of the subsets. Fixing this parameter returned good results in only 2 cases out of 15. It may be concluded that, this approach should be discouraged.
In the case of the Random Forest algorithm, the number of trees in the ensemble and the number of features, which are randomly chosen for tree induction, can be manipulated. The approach with very small trees, consisting only of two random features, outperformed significantly the other Random Forest methods considered. Additionally, it worked well with larger ensembles (40–120). This stems from the fact that a larger number of weak predictors suggest a more generalised model and at the same time introduces the much-needed diversity into the base classifier pool. 9 When considering larger trees the probability that they are constructed of similar features increases and, therefore, the diversity decreases.
A set of rules/guidelines for end-users and practitioners on how to use the ensembles is now formulated.
Users should concentrate on the Random Forest and Rotation Forest algorithms. Those two approaches are most likely to deliver the best results. If the user does not have time to run the exhaustive test, then Bagging should be avoided.
For Random Forests using numerous ensembles of small trees is most likely to return the best results. Small ensembles of large trees should be avoided. This approach is recommended when there are several features contained within the data.
For the Rotation Forest, the REPTree as a base algorithm returns the best results, especially when the dataset under consideration consists of a large number of objects. The size parameter is worth exploring, while fixing the number of subsets has little positive effect on the classification. This is important, as the implementation of the Rotation Forest provided in the Java language presented in Rodríguez et al. 32 allows only a change of one of these parameters.
Random Forest gives the best performance for datasets with a high number of features. This is owing to the fact that it serves at the same time as a classifier and a feature selector and is able to cope with high-dimensionality problems.
Rotation Forest is able to cope well with problems described by a low number of classes. This is considered to be owing to the use of PCA, which dismisses some of the information (albeit of least importance). In case of numerous classes Random Forest should be tested first.
It should be borne in mind that the tests reported here represent only a small part of a wide variety of medical data. It is challenging to give a clear indication of which algorithm will always return the optimal results. Experience suggests, however, that it is advisable for more than one setting to be tested before the final implementation. Through the experimental trails presented it has been shown that it is possible to significantly reduce the number of settings that need to be taken into consideration.
Conclusions
The article has dealt with a problem of choosing an optimal set of parameters for the decision tree ensembles used in medical decision support tasks. It has been shown that the default settings proposed by the available open source data analysis software do not always give the best results. Making use of default settings only imposes limitations on the performance of such pattern recognition system. It has been demonstrated that there is a need for a practical guide into the manual tuning of the ensemble’s parameters, which allows maximising the efficiency of their data analysis and gives greater insight into the mechanism of combining tree classifiers. Indeed, this was one of the premises which prompted the work, whereby propositions have been evaluated on the basis of computer experiments and carried out on diverse benchmark datasets. On the basis of the experimental results obtained and observations thereof, recommendations for practical implementation of selected methods have been formulated. It is recognised, of course, that one type of setting can never outperform all others on all datasets ever encountered. It has been shown, however, that for the dataset considered, a significant number of settings which never gave good results could be discarded immediately. Another important finding is that, for the remaining pool of settings, it has been shown that certain dependencies exist between the type of data and the type of classifiers that could be used. Consequently, the proposed guidelines allow the practitioner to simplify the task so that it can be flexibly tuned to the data at hand. It is considered that the proposed concept can be useful for tackling real-life medical decision support system problems whereby physicians and clinicians are required to routinely analyse data using standard available pattern recognition software.
Footnotes
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.