Abstract

Commentary on: Wang et al. Development of a polygenic risk score to improve detection of peripheral artery disease. Vasc Med. 2022;27:219-227.
Risk stratification is an important initial step in managing common diseases. Although algorithms for risk stratification are available for several cardiovascular diseases, their predictive accuracy is suboptimal. 1 Genome-wide association studies (GWAS) have identified hundreds of common susceptibility variants for cardiovascular diseases, enabling the use of polygenic risk scores for disease risk prediction. 2 A polygenic risk score is a weighted sum of risk alleles that is typically normally distributed in a population, with those at the tails of distributions having significantly different risk than those in the middle. 3 Because a polygenic risk score is largely orthogonal to conventional risk factors, it can provide incremental risk predictive information.4,5 For example, a polygenic risk score for coronary heart disease can be incorporated into available risk equations, leading to reclassification between low, intermediate, and high-risk groups and influencing clinical decision-making regarding screening or drug therapy.4,5
Peripheral artery disease (PAD) is a relatively common form of atherosclerotic cardiovascular disease that affects nearly 5% of US adults and more than 200 million people worldwide. 6 Diabetes and smoking are the most powerful risk factors for PAD, but heritable factors also predispose to PAD, as demonstrated by twin and sibling studies.7,8 GWAS for PAD have lagged behind those for coronary heart disease, but recent work by several groups has led to the discovery of multiple genetic variants associated with PAD.8 –10 Polygenic risk scores could enhance risk stratification for PAD and identify patients who have an increased risk of prevalent as well as incident PAD, with implications for screening and treatment.
In this issue of Vascular Medicine, Wang and colleagues 11 generated a polygenic risk score for PAD from the Million Veteran Program (MVP) cohort and tested it on an independent cohort of UK Biobank participants. The MVP cohort was more diverse in ancestry and older in age than the UK Biobank training/validation cohort. The authors used the MVP GWAS summary statistics for deriving a polygenic risk score and then ‘tuned’ it in a subset of UK Biobank PAD cases and controls. 9 Finally, a separate UK Biobank cohort was used to validate the best performing polygenic risk score. Two methods were used to calculate polygenic risk scores. The first method 12 selected single nucleotide variants (SNVs) with a p-value lower than a certain threshold and that were not correlated with each other. The second method included SNVs across the genome, 13 modifying the regression coefficients using a Bayesian approach.
Wang and colleagues report the heritability of PAD to be modest, at 11%. 11 Those in the top decile of the polygenic risk score had nearly double the odds of having PAD compared to the remaining 90%. Although the area under the receiver operating curve increased minimally from 0.797 to 0.804, there was significant net reclassification (Table 1). The polygenic risk score performed better in younger than older adults, in men than in women, and in European ancestry individuals than in non-European ancestry individuals. 11 The polygenic risk score for PAD had pleiotropic effects and was associated with greater odds of coronary heart disease, heart failure, and cerebrovascular disease. One can foresee several clinical applications of a polygenic risk score for PAD, including targeting screening ankle–brachial index measurements to adults with a high polygenic risk score. Additionally, if measured earlier in life, a patient with a high score would be encouraged to adopt/maintain healthy lifestyle choices and potentially start on statin therapy to lower the risk of atherosclerotic cardiovascular disease in the future.
Metrics for a polygenic risk score (PRS) for peripheral artery disease (PAD) reported by Wang et al. 11
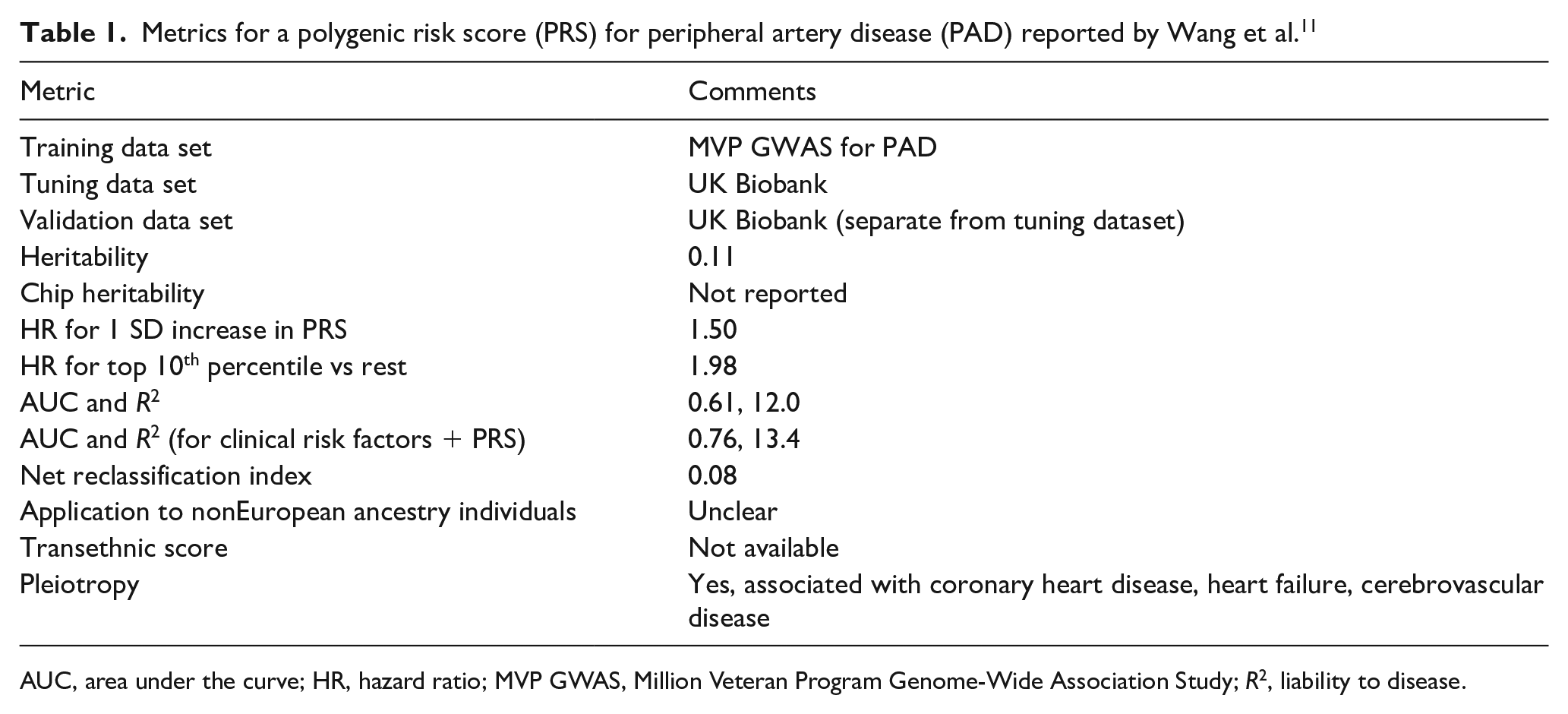
AUC, area under the curve; HR, hazard ratio; MVP GWAS, Million Veteran Program Genome-Wide Association Study; R2, liability to disease.
In 2021, two groups, the ClinGen Complex Disease Working Group and PGS Catalog joined to create a set of reporting standards including ‘predictive ability’, ‘calibration’, and ‘discrimination’ of risk models that include a polygenic risk score.14,15 Potential metrics to evaluate polygenic risk scores in a validation sample include effect size (hazard ratio for 1 SD increase or for being in the top 5th percentile, for example), the contribution of the polygenic risk score to disease liability (pseudo R2), and reclassification indices. The metrics reported by Wang et al. are summarized in Table 1.
Two consortia funded by the National Human Genome Research Institute are addressing major knowledge gaps in the field of polygenic risk scores. The eMERGE Network (https://www.genome.gov/Funded-Programs-Projects/Electronic-Medical-Records-and-Genomics-Network-eMERGE) is investigating how polygenic risk scores for common diseases can be implemented in the clinical setting and lead to a change in outcomes. The PRIMED Network (https://www.genome.gov/Funded-Programs-Projects/Polygenic-Risk-Score-Diversity-Consortium) will develop methods to adapt polygenic risk scores to different ancestry groups as well as admixed groups such as Latinx and African Americans. Most polygenic risk scores are derived from European ancestry populations and there is an urgent need to develop robust polygenic risk scores for diverse ancestry groups to avoid worsening health care disparities. 16
In conclusion, the work by Wang and colleagues 11 adds PAD to the list of cardiovascular diseases for which a polygenic risk score has been reported and demonstrates incremental (but modest) predictive power of the score for PAD. Additional work will be needed to eventually use such a polygenic risk score in the clinic, including increasing the numbers and diversity of cases and controls in GWAS of PAD and the use of novel population genetics methods to boost predictive power of the score in diverse ancestry groups.
Footnotes
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Dr Kullo is funded as part of the NHGRI-supported eMERGE (Electronic Records and Genomics, U01HG006379) and PRIMED (Polygenic Risk Methods in Diverse Populations, U01HG011710) Networks and by K24 HL137010 from the NHLBI.
