Abstract
Background:
Despite established prognostic scores, predicting disease progression in myeloproliferative neoplasms (MPNs) remains challenging, and the role of specific immune cell subsets—particularly the PD-1/PD-L1 axis—in mediating fibrosis and transformation is poorly characterized. This study aimed to investigate the prognostic utility of novel immune-inflammatory biomarkers, including PD-1/PD-L1 coexpression with C-X-C chemokine receptor type 3 (CXCR3), CXCR3 expression on monocyte subsets, and systemic indices (systemic immune-inflammation index [SII], systemic inflammation response index [SIRI]), for predicting fibrosis grade, leukemic transformation, and survival outcomes in patients with Philadelphia-negative MPNs.
Methods:
In a prospective case-control study, including 90 Egyptian patients with Philadelphia-negative MPNs and 90 matched controls, we quantified PD-1/PD-L1 coexpression with CXCR3 and CXCR3 expression on monocyte subsets by flow cytometry. Validation employed targeted next-generation sequencing (NGS) in the full cohort, complemented by focused mechanistic validation using bulk RNA sequencing (n = 20 primary myelofibrosis [PMF], n = 10 controls), single-cell RNA sequencing (n = 10 PMF, n = 5 controls), and targeted serum cytokine profiling (IL-6 and TGF-β; n = 90 MPN, n = 90 controls). Diagnostic/prognostic performance used receiver operating characteristic (ROC) curves, decision curve analysis, logistic/Cox regression, random forest/gradient boosting, and external validation in an independent Egyptian cohort (n = 30). Patient-reported outcomes were assessed with the Myeloproliferative Neoplasm Symptom Assessment Form (MPN-SAF).
Results:
PD-1/CXCR3 and PD-L1/CXCR3 were elevated in patients (P < .001). PD-L1/CXCR3 demonstrated high discrimination between advanced (grade 4) and grade 3 fibrosis (area under the curve [AUC] 0.975; cut-off >40%; sensitivity 92.9%; specificity 90%). In an exploratory analysis, reduced CXCR3 on intermediate monocytes was associated with leukemic transformation (AUC 0.876; cut-off ⩽ 12%) with high negative predictive value (98.4%). Higher PD-L1/CXCR3 correlated with worse fatigue (r = 0.58, P < .001) and pruritus (r = 0.45, P = .002) on MPN-SAF and was an independent predictor of poorer survival. A biomarker-enhanced prognostic score (incorporating high PD-L1/CXCR3, low CXCR3/intermediate, age, and Dynamic International Prognostic Scoring System) outperformed DIPSS/DIPSS-Plus/MIPSS70 (AUC 0.85; bootstrap AUC 0.84) and showed promising initial validation in an external cohort (AUC 0.82). Single-cell RNA sequencing revealed a CXCR3-low intermediate monocyte population enriched for profibrotic/inflammatory signatures; targeted cytokine analysis indicated higher IL-6/TGF-β in checkpoint-high states.
Conclusion:
PD-1+CXCR3+/PD-L1+CXCR3+ coexpression and relatively low CXCR3 expression on intermediate monocytes (within the patient cohort) are promising biomarkers for fibrosis, leukemic transformation, and survival in Egyptian MPNs. Their integration with clinical scores improved risk stratification and showed promising initial performance in an external cohort. These findings support incorporating immune checkpoint-chemokine profiling into precision medicine frameworks for MPNs.
Keywords
Introduction
Myeloproliferative neoplasms (MPNs) are clonal hematopoietic stem cell disorders characterized by the proliferation of one or more myeloid lineages. The classic Philadelphia chromosome-negative MPNs include polycythemia vera (PV), essential thrombocythemia (ET), and primary myelofibrosis (PMF). 1 Patients face significant morbidity from thrombotic events and a risk of progression to advanced myelofibrosis or acute myeloid leukemia (AML). 2 In Egypt, the burden of MPNs is considerable, with studies from major tertiary centers reporting clinical profiles and complication rates that underscore the need for improved risk stratification tools tailored to the population.3,4
The prognosis of MPNs is currently guided by integrated scoring systems such as the Dynamic International Prognostic Scoring System (DIPSS) for PMF and its subsequent plus version, which incorporate clinical and genetic variables.5,6 However, the relentless progression in some patients despite low-risk scores indicates a critical unmet need for more precise, biologically driven biomarkers. The tumor microenvironment and systemic inflammation are increasingly recognized as key players in MPN pathogenesis and progression. 7 Markers of systemic inflammation, such as the neutrophil-to-lymphocyte ratio (NLR), platelet-to-lymphocyte ratio (PLR), and more recently, the systemic immune-inflammation index (SII) and systemic inflammation response index (SIRI), have demonstrated prognostic value in solid tumors and are emerging as potential biomarkers in hematologic malignancies.8,9
Beyond systemic indices, the trafficking and function of immune cells within the bone marrow niche are orchestrated by chemokines and their receptors. C-X-C chemokine receptor type 3 (CXCR3), a receptor for IFN-γ-inducible chemokines (CXCL9, CXCL10, CXCL11), is expressed on effector T cells, regulatory T cells, and certain monocyte subsets and is pivotal in mediating Th1 immune responses. 10 Its role in the immune surveillance of cancers is complex, with context-dependent pro- or antitumoral effects. 11 Furthermore, immune checkpoint pathways, particularly the PD-1/PD-L1 axis, are a critical mechanism of immune evasion in cancers. The interaction between PD-1 on T cells and PD-L1 on tumor or myeloid-derived suppressor cells leads to T-cell exhaustion and promotes a tolerogenic microenvironment. 12 In MPNs, increased PD-L1 expression on monocytes and CD34+ cells has been linked to disease progression, suggesting this pathway is a potential therapeutic target and prognostic marker. 13
CXCR3 signaling has been implicated in chronic inflammatory and fibrotic disorders across organ systems, where it orchestrates immune cell recruitment and stromal remodeling.10,11 In hematologic malignancies, dysregulated CXCR3 expression on immune subsets can alter tumor surveillance and microenvironmental crosstalk. Among monocyte subsets, intermediate monocytes (CD14++CD16+) are particularly potent producers of pro-inflammatory cytokines (eg, IL-6, TNF-α) and have been linked to tissue fibrosis through their capacity to differentiate into profibrotic macrophages and secrete TGF-β.12,13 Perturbations in CXCR3 expression on these cells may therefore directly influence their trafficking, polarization, and effector functions within the bone marrow niche, linking chemokine-driven myeloid dysregulation to fibrotic progression and leukemic evolution in MPNs.
Recent advances in multiomics approaches, including genomic, transcriptomic, and proteomic profiling, have revealed intricate interactions between driver mutations (eg, JAK2, CALR, MPL) and immune dysregulation in MPNs, providing a foundation for precision medicine. 14 Single-cell technologies further elucidate the heterogeneity of immune cell subsets, offering insights into their role in disease progression. 15 In addition, patient-reported outcomes (PROs) assessed via tools like the Myeloproliferative Neoplasm Symptom Assessment Form (MPN-SAF) are increasingly recognized as critical for evaluating quality of life (QoL) and guiding treatment decisions. 16
Despite growing recognition of inflammation and immune dysregulation in MPNs, the combined prognostic value of checkpoint activity (PD-1/PD-L1) with CXCR3 across immune subsets, particularly monocyte subpopulations, remains underexplored. Notably, the PD-1/PD-L1-CXCR3 axis represents a biologically plausible link between immune exhaustion, aberrant myeloid trafficking, and marrow fibrosis, with direct implications for prognostic refinement and therapeutic stratification in PMF. Integration of these immune biomarkers with multiomic profiling and PROs is particularly scarce in Egyptian cohorts. We therefore hypothesized that (1) PD-L1/CXCR3 coexpression indicates a checkpoint-high, profibrotic immune state; (2) among MPN patients, relatively lower CXCR3 expression on intermediate monocytes identifies patients at increased risk of leukemic transformation; and (3) adding these immune biomarkers to established prognostic scores (DIPSS/DIPSS-Plus/MIPSS70) will improve risk stratification. To test these hypotheses, we performed flow cytometry on 90 Philadelphia-negative MPN patients and 90 matched controls, integrated targeted next-generation sequencing (NGS), bulk and single-cell transcriptomics, limited proteomics, and PROs, and then derived and externally validated a biomarker-enhanced prognostic score. This represents the first comprehensive immune-genomic-clinical integration of checkpoint and chemokine biomarkers in Egyptian MPN patients, addressing a critical unmet need for population-specific prognostic tools.
Accordingly, this study was designed as a structured, hypothesis-driven investigation in which each analytical layer—flow cytometry, clinical modeling, machine learning, and multiomic validation—serves a predefined confirmatory role. This stepwise design aims to minimize overinterpretation, enhance biological coherence, and ensure that novel immune biomarkers are supported by convergent evidence rather than isolated statistical associations.
Subjects and Methods
Study population and design
This prospective case-control study was conducted at the Clinical Pathology, Internal Medicine, and Oncology Departments of Kafr Elsheikh University Hospitals, Egypt, in collaboration with the Internal Medicine and Clinical Pathology Departments of Suez University Hospital, between January 2020 and December 2024. The reporting of this study conforms to the STROBE guidelines. 17 Patients were consecutively enrolled, and all data were deidentified to protect confidentiality. Given the distinct clinical endpoints of each MPN subtype, primary survival analyses (overall and event-free survival) were conducted within the PMF subgroup. For the PV and ET cohorts, an exploratory analysis of thrombotic event risk was performed.
A priori power analysis: Using a 2-sample t-test (2-sided) with Cohen’s d = 0.65, α = .05, and power (1−β) = 0.95, the minimum required sample size is 62 participants per group (124 total). To allow robust subgroup and multiomics analyses and to account for attrition, we therefore recruited 180 participants (90 MPN patients and 90 age- and sex-matched healthy controls). (Power calculation was performed with G*Power v3.1.9.7). Controls were recruited from voluntary blood donors and hospital staff, and all underwent clinical evaluation, complete blood count (CBC), and liver and renal function tests to confirm the absence of subclinical hematologic, inflammatory, or malignant disease.
Missing data (<5%) were handled with multiple imputation using fully conditional specification, while variables with >10% missingness were excluded from analysis. Patients were classified according to the 2016 World Health Organization criteria into PV, ET, or PMF. 15
External validation: An independent cohort of 30 consecutive Egyptian MPN patients from Suez University Hospital (not included in the primary cohort of 90 patients/90 controls) was used to externally validate the biomarker-enhanced prognostic model (primary metric: area under the curve [AUC]; clinical utility: decision curve analysis).
The multiomics approach was intentionally tiered: Targeted NGS served as the primary genomic analysis across the full cohort, while bulk RNA-seq, single-cell RNA sequencing (scRNA-seq), and targeted cytokine profiling were applied as predefined, hypothesis-driven validation layers in selected subsets. This strategy was designed to provide mechanistic insight and biological confirmation while avoiding overextension or dilution of the central analytical message.
Longitudinal JAK inhibitor subcohort: A predefined subgroup of PMF patients initiating JAK inhibitor therapy underwent repeat biomarker assessment at 12 months to explore treatment responsiveness (paired analyses).
Ethical considerations
This study received approval from the Medical Research Ethics Committee of Kafr Elsheikh University (Reference No. KFSIRB200-234, dated April 29, 2020), which provided ethical oversight for all participating centers, including Suez University Hospital. The study was conducted in accordance with the ethical principles of the Declaration of Helsinki (2024 revision). All participants received a clear explanation of study aims, procedures, risks, and expected benefits and provided written informed consent. Data confidentiality was maintained by anonymizing datasets and storing them on encrypted, password-protected servers accessible only to the study investigators.
Inclusion and exclusion criteria
Inclusion criteria
Patients were eligible if they were 18 years or older; had a new diagnosis of PV, ET, or PMF according to WHO 2016 criteria; had not received prior cytoreductive or immunomodulatory therapy; and provided informed consent. Healthy controls were adults with normal hematologic and biochemical profiles, free from chronic inflammatory or malignant disease, and not receiving immunosuppressive therapy.
Exclusion criteria
Patients and controls were excluded if they had active infections such as hepatitis B virus (HBV), hepatitis C virus (HCV), HIV, or tuberculosis or autoimmune conditions requiring systemic immunosuppressive therapy, because these conditions markedly affect systemic inflammation and immune checkpoint expression. Concomitant hematologic or solid malignancies were excluded to avoid confounding immune signatures and survival outcomes. Advanced hepatic or renal impairment, defined as Child-Pugh C cirrhosis or estimated glomerular filtration rate (eGFR) below 30 mL/min/1.73 m2, was excluded due to its independent impact on systemic inflammatory indices. Pregnant and lactating women were excluded, given the physiological immune and hematologic changes during these states. Patients who had previously undergone hematopoietic stem cell transplantation were excluded because transplant-related immune reconstitution alters circulating immune cell populations. Incomplete diagnostic data, particularly missing bone marrow biopsy, was also an exclusion criterion to ensure reliable risk stratification and fibrosis grading.
Clinical and laboratory assessment
Clinical evaluation
All participants underwent comprehensive clinical assessments that included a detailed medical history and physical examination. Demographic data (age and sex) and relevant medical history were documented at baseline. For patients with MPNs, special emphasis was placed on constitutional symptoms such as fever, night sweats, weight loss, pruritus, and fatigue. Splenomegaly and hepatomegaly were assessed by physical examination and confirmed by ultrasound when indicated. The disease duration and prior history of thrombotic or hemorrhagic events were recorded. The PROs were collected using the validated MPN-SAF, with total symptom scores and individual symptom severity (eg, fatigue, pruritus, night sweats) calculated on a 0 to 10 scale. 16 For the control group, eligibility was confirmed through history and examination to exclude any hematologic, inflammatory, or malignant disorders. This standardized evaluation ensured precise patient characterization and comparability between study groups.
Laboratory assessment
All participants underwent standardized laboratory testing at enrollment. Routine hematologic analysis included CBC performed on Sysmex XN-1000 (Sysmex Corporation, Kobe, Japan). Serum lactate dehydrogenase (LDH), ferritin, and C-reactive protein (CRP) were measured using an automated biochemical analyzer (Roche Cobas 6000; Roche Diagnostics, Mannheim, Germany). Erythropoietin (EPO) was quantified by chemiluminescent immunoassay (Immulite 2000, Siemens Healthcare, Germany). Serum cytokines (IL-6, TGF-β) were quantified using ELISA (R&D Systems, Minneapolis, Minnesota) as key inflammatory and profibrotic mediators. Erythrocyte sedimentation rate (ESR) was determined by the Westergren method. Liver function tests, including aspartate aminotransferase (AST), alanine aminotransferase (ALT), albumin, and total bilirubin, as well as renal function tests (serum creatinine and eGFR), were measured using Roche Cobas 6000. These investigations ensured the exclusion of significant hepatic or renal impairment and provided baseline inflammatory and biochemical characterization of the study population.
Bone marrow aspirates and trephine biopsies were obtained for all patients and independently reviewed by 2 hematopathologists. Reticulin fibrosis was graded according to the European Consensus Grading System. 17
Multiomic analyses and bioinformatics
Targeted NGS
Genomic DNA was extracted from peripheral blood samples obtained from all 90 MPN patients. Targeted sequencing was performed using a custom-designed panel of 54 genes recurrently mutated in myeloid malignancies (including JAK2, CALR, MPL, ASXL1, EZH2, SRSF2, and TET2) on an Illumina NextSeq 550 platform (Illumina, San Diego, California). Sequencing reads were aligned to the GRCh38 reference genome using BWA-mem. Variant calling was conducted following GATK best-practice workflows, and functional annotation was performed using ANNOVAR. A minimum variant allele frequency threshold of 2% was applied.
Bulk RNA sequencing
To validate transcriptomic signatures associated with immune checkpoint phenotypes, bulk RNA sequencing was performed on peripheral blood mononuclear cells (PBMCs) from a representative subset of 20 PMF patients and 10 healthy controls. Patients were selected to span DIPSS risk categories and fibrosis grades. Total RNA was extracted with RNA integrity confirmed (RIN > 8.0). Libraries were prepared using the Illumina Stranded mRNA Prep kit and sequenced on an Illumina NovaSeq 6000 platform to a depth of approximately 30 million paired-end reads per sample. Differential gene expression analysis was conducted using DESeq2, and pathway enrichment was assessed using Gene Set Enrichment Analysis (GSEA v4.3.2) against the MSigDB Hallmark gene sets.
Single-cell RNA sequencing
To resolve immune cell heterogeneity, scRNA-seq was performed on PBMCs from 10 PMF patients and 5 age- and sex-matched healthy controls. Patients were purposively selected to represent the checkpoint-high and checkpoint-low immune phenotypes identified by flow cytometry, ensuring that the transcriptional analysis would capture the spectrum of immune dysregulation observed in our cohort. Single-cell libraries were generated using the 10x Genomics Chromium Single Cell 3′ v3.1 platform and sequenced on an Illumina NovaSeq 6000. Data processing—including demultiplexing, alignment, and feature counting—was conducted using the Cell Ranger pipeline (v7.1.0). Downstream analyses—including quality control, normalization, batch integration, clustering, and cell-type annotation—were performed in R using Seurat (v4.3.0). Cells with fewer than 200 detected genes or greater than 20% mitochondrial reads were excluded.
Targeted cytokine analysis
To validate inflammatory and profibrotic pathways identified by flow cytometry and transcriptomics, we performed targeted cytokine analysis. Serum samples from all 90 MPN patients and 90 healthy controls were analyzed for interleukin-6 (IL-6) and transforming growth factor beta (TGF-β) using validated enzyme-linked immunosorbent assay (ELISA) kits (R&D Systems), following the manufacturer’s protocols. This approach provides protein-level validation of key pathway mediators rather than constituting a broad proteomic discovery effort.
The SII was defined as (platelet count × neutrophil count)/lymphocyte count, and the SIRI was defined as (neutrophil count × monocyte count)/lymphocyte count. The platelet, neutrophil, monocyte, and lymphocyte counts were obtained from the CBC and used in their standard clinical unit of 109 cells/L. This results in standard, dimensionless indices for both SII and SIRI.
Flow cytometry and gating
By density gradient centrifugation, PBMCs were isolated from EDTA-anticoagulated blood. To achieve optimal spectral resolution and sensitivity, particularly for the dim expression of CXCR3 on monocytes, sequential immunostaining was performed on separate aliquots of the same PBMC sample using 3 dedicated antibody panels, thereby preventing fluorochrome spillover and compensation artifacts that can arise in a single comprehensive panel.
The 3 panels were used. The first panel, for monocyte subset identification, contained CD14-FITC (clone M5E2) and CD16-PE (clone 3G8). The second panel, for chemokine receptor detection, included CXCR3-APC (clone 1C6). The third panel, for immune checkpoint analysis, comprised PD-1-BV421 (clone EH12.2H7) and PD-L1-PE-Cy7 (clone MIH1). All antibodies were from BD Biosciences or BioLegend. For each panel, approximately 0.5 to 1 × 106 PBMCs were stained at 4°C for 20 minutes after Fc receptor blockade. Unstained, isotype-matched, and fluorescence-minus-one (FMO) controls were included in every run for accurate compensation and gate setting. 18
Data were acquired on a BD FACSCanto II flow cytometer and analyzed with FlowJo v10.7. Positive expression was defined as a signal exceeding the threshold set by the corresponding FMO control. Monocytes were gated based on forward/side scatter and CD45 expression, then subclassified as classical (CD14++CD16−), intermediate (CD14++CD16+), and nonclassical (CD14+CD16++).19-21 CXCR3, PD-1, and PD-L1 expression was quantified within these subsets as the percentage of positive cells; median fluorescence intensity was also recorded. For contextual analysis of T cells, lymphocytes were gated from the scatter plot, and CD3+T cells were subdivided into CD4+and CD8+ populations to assess PD-1 and PD-L1 expression. Interoperator reproducibility was confirmed by duplicate analysis of 10% of samples (intraclass correlation coefficient > .95).
Outcome measures
The primary outcome was leukemic transformation, defined as progression to AML with at least 20% blasts in peripheral blood or bone marrow. Secondary outcomes included progression to advanced fibrosis, overall survival, and event-free survival, with events defined as leukemic transformation, initiation of cytoreductive therapy, or death. Patients were also stratified at baseline according to DIPSS and DIPSS-Plus scoring systems to evaluate associations between immune biomarkers and clinical risk models. Patients were followed from the date of enrollment until transformation, death, or the end of the study period (December 31, 2024), whichever came first.
Statistical analysis
All analyses were conducted using R v4.3.2 (R Foundation for Statistical Computing, Vienna, Austria) and SPSS v28.0 (IBM, Armonk, New York). R packages included survival v3.5-7 for Kaplan-Meier and Cox regression, glmnet v4.1-7 for LASSO Cox modeling, randomForest v4.7-1.1 and xgboost v1.7.6 for machine learning classifiers, pROC v1.18.5 for ROC analyses, and ggplot2 v3.4.4/Seurat v4.3.0 for data visualization and single-cell analyses. Normality of continuous variables was assessed using the Shapiro-Wilk test and visual inspection of histograms and Q-Q plots. Continuous variables were expressed as mean ± SD or median (interquartile range, IQR) based on distribution, whereas categorical variables were presented as counts and percentages. Between-group comparisons were performed using the independent samples t-test (unpaired) or Mann-Whitney U test for continuous variables and the χ2 test or Fisher exact test for categorical variables. Longitudinal paired data (eg, JAK-inhibitor subcohort) were analyzed using paired t-tests. Effect sizes were calculated using Cohen’s d for continuous data and ORs with 95% CIs for categorical data to quantify the strength of associations.
Associations between immune biomarkers and ordinal/continuous clinical variables (eg, fibrosis grade, DIPSS score) were assessed using Spearman’s rank correlation (ρ) with 95% CIs. To evaluate independent associations while adjusting for potential confounders, multivariable regression models were applied: ordinal logistic regression for fibrosis grade (adjusted for age, sex, JAK2 mutation status, and baseline blast count) and linear regression for DIPSS score (adjusted for age, sex, cytogenetic risk category, and baseline blast count).
For diagnostic analyses, receiver operating characteristic (ROC) curves were generated, and the AUC with 95% CIs was reported. To account for potential overfitting and provide robust performance estimates for small-sample subgroups, leave-one-out cross-validation (LOOCV) was performed for all ROC analyses, and the LOOCV AUC is reported as the primary metric alongside the apparent AUC. Optimal cut-off values (eg, advanced fibrosis, leukemic transformation, high DIPSS risk) were determined using Youden’s index, and sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were calculated. Decision curve analysis was performed to evaluate the clinical utility of biomarkers and prognostic models. Binary logistic regression models were applied to identify independent predictors of advanced fibrosis, leukemic transformation, and high DIPSS risk. Variables with P < .10 in univariate analysis were entered into multivariable models. Gradient boosting (XGBoost) was used to rank biomarker importance, complementing random forest analyses.
Survival analyses were performed using the Kaplan-Meier method with the log-rank test to compare groups. Cox proportional hazards regression models were used to estimate hazard ratios (HRs) with 95% CIs, adjusted for age, sex, JAK2 mutation status, DIPSS, and DIPSS-Plus scores. Machine learning approaches were incorporated to enhance prognostic modeling. A random forest classifier with 10-fold cross-validation ranked biomarkers by their predictive importance for fibrosis and transformation. K-means clustering was used to identify biomarker-driven immune phenotypes. For RNA-seq data, GSEA was performed using the MSigDB database to identify enriched pathways (eg, JAK/STAT, NF-κB).
For survival analyses using Cox proportional hazards regression, immune biomarkers (PD-L1/CXCR3 and CXCR3/intermediate) were modeled as categorical variables. The “High” and “Low” groups were defined by the optimal cut-off values derived from ROC curve analysis for the primary outcome of overall survival. Therefore, HRs reported for these biomarkers represent the risk of the “High” group compared with the “Low” (reference) group for PD-L1/CXCR3, and the risk of the “Low” group compared with the “High” (reference) group for CXCR3/intermediate.
For MPN-SAF data, Spearman’s correlation and linear regression were used to assess associations between biomarker levels and symptom scores. Longitudinal PRO changes were analyzed using mixed-effects models for patients with follow-up data. A false discovery rate adjustment was applied for multiple comparisons using the Benjamini-Hochberg method. A 2-sided P value <.05 was considered statistically significant.
To develop a robust prognostic model, we employed a structured machine learning pipeline. Feature selection was performed using the least absolute shrinkage and selection operator (LASSO) Cox regression with 10-fold cross-validation. The optimal penalty parameter (lambda) was selected based on the minimum cross-validated error (lambda.min). The final model incorporated the features with nonzero coefficients from the LASSO analysis. The selected model was then validated internally using bootstrap resampling (1000 iterations) to calculate an optimism-corrected AUC. To ensure generalizability, the model was subsequently validated on a completely independent, held-out external cohort of 30 MPN patients from Suez University Hospital. This external cohort was not used in any phase of model development or feature selection. A random forest classifier with 10-fold cross-validation was implemented using the randomForest package in R to rank biomarkers by their mean decrease in Gini importance.
Prognostic Score Development: A biomarker-enhanced prognostic score was developed using LASSO regression to select features (PD-L1/CXCR3, CXCR3/intermediate, age, DIPSS). The score was validated using bootstrap resampling (1000 iterations) and an external cohort of 30 MPN patients from Suez University Hospital. Performance was compared with DIPSS, DIPSS-Plus, and MIPSS70 using AUC and decision curve analysis.
Advanced computational and genomic analyses
Unsupervised k-means clustering was applied to standardized z-scores of immune biomarker expression using the Euclidean distance metric, and the optimal cluster number was determined with the elbow and silhouette methods. A random forest classifier with 500 trees and 10-fold cross-validation was implemented using the random Forest package in R to rank biomarkers by mean decrease in Gini importance.18,19 The use of patient material for genomic and transcriptomic analyses was explicitly covered by the IRB approval (KFSIRB 200-2023). For scRNA-seq, 10 patients and 5 controls were randomly selected from the full cohort to represent the spectrum of disease subtypes and risk categories, thereby minimizing selection bias. Targeted NGS was performed on genomic DNA extracted from peripheral blood using a custom panel of 54 recurrently mutated genes in myeloid malignancies. Sequencing was conducted on an Illumina NextSeq 550; reads were aligned with BWA-mem, variants were called with GATK, and annotated using ANNOVAR, with a minimum variant allele frequency of 2%. Bulk RNA sequencing was carried out on PBMCs after confirming RNA integrity (RIN > 8.0). Libraries were prepared using Illumina Stranded mRNA Prep with poly-A selection and sequenced on an Illumina NovaSeq 6000 to a depth of 30 million paired-end reads per sample. Data were analyzed using DESeq2 for differential gene expression, 20 and GSEA was performed against the MSigDB Hallmark gene sets. 21 Single-cell RNA sequencing was performed on PBMCs from 10 patients and 5 controls using the 10x Genomics Chromium Single Cell 3′ v3.1 chemistry according to the manufacturer’s protocol.22,23
Results
Baseline characteristics of the study population
A total of 180 participants were enrolled, including 90 patients with MPNs and 90 healthy controls. For external validation only, an additional independent cohort of 30 consecutive Egyptian MPN patients from Suez University Hospital was recruited. Among the MPN cohort, PMF was the most common diagnosis (n = 45), followed by ET (n = 35) and PV (n = 10). Given the central relevance of fibrosis progression and leukemic transformation to PMF, the subsequent prognostic analyses (sections 4-8) are focused on the PMF subgroup. Broader immune biomarker comparisons (section 3) and PROs (section 13) are presented for the full cohort. Patients and controls were well matched for age and sex, with no significant differences between the groups (P > .05).
Baseline laboratory assessment demonstrated generally preserved hepatic and renal function across both cohorts. However, patients showed higher systemic inflammatory activity, with elevated CRP levels, whereas ESR showed a trend toward an increase that did not reach statistical significance. Hematologic profiles reflected expected disease patterns, including leukocytosis and thrombocytosis in subsets of patients, which were later used to calculate systemic inflammatory indices for subsequent analyses. Comprehensive demographic, clinical, and laboratory data are summarized in Table 1.
Baseline demographics, disease distribution, and selected laboratory findings.

Data are presented as mean ± SD for normally distributed variables and median (interquartile range) for nonnormally distributed variables (CRP). P-values were calculated using an independent t-test for normal variables and the Mann-Whitney U test for nonnormal variables. Effect sizes are reported as Cohen’s *d* for t-tests and rank-biserial correlation (*r*) for Mann-Whitney U tests.
Abbreviations: ALT, alanine aminotransferase; AST, aspartate aminotransferase; CRP, C-reactive protein; ESR, erythrocyte sedimentation rate; ET, essential thrombocythemia; PMF, primary myelofibrosis; PV, polycythemia vera.
Clinical characteristics and risk stratification in PMF patients
In the PMF subgroup (n = 45), constitutional symptoms and transfusion dependence were relatively uncommon. However, risk assessment highlighted that many patients were concentrated in the higher DIPSS and DIPSS-Plus categories, underscoring the advanced disease burden within this cohort. These findings emphasize the clinical severity of PMF in the studied population (Table 2).
Clinical characteristics and DIPSS/DIPSS-plus risk categories among PMF patients.

Data are presented as numbers (%). DIPSS-Plus incorporates red cell transfusion dependence, platelet count < 100 × 109/L, and unfavorable karyotype as additional adverse factors.
Abbreviations: DIPSS, Dynamic International Prognostic Scoring System.
Peripheral immune biomarkers in MPN patients vs controls
Peripheral immune profiling revealed marked dysregulation across multiple immune compartments in MPN patients compared with healthy controls (Table 3, Figures 1 and 2). Systemic inflammatory indices were significantly elevated, with SII and SIRI increased by approximately 1.6- to 1.8-fold, consistent with heightened systemic inflammation. When analyzed by MPN subtype, PD-L1/CXCR3 coexpression was highest in PMF (mean 58.2%), intermediate in ET (48.3%), and lowest in PV (41.7%). Similarly, CXCR3 expression on intermediate monocytes was lowest in PMF (14.8%) compared with ET (21.1%) and PV (24.5%), aligning with the more aggressive clinical phenotype of PMF.
Immune biomarkers in MPN patients versus healthy controls.

Data are presented as mean ± SD for normally distributed variables and median (interquartile range) for nonnormally distributed variables. P-values were calculated using an independent t-test for normal variables and the Mann-Whitney U test for nonnormal variables (SII, SIRI). Effect sizes are reported as Cohen’s d for t-tests and rank-biserial correlation (r) for Mann-Whitney U tests. Both are dimensionless indices.
Abbreviations: CXCR3, C-X-C chemokine receptor type 3; IL-6, interleukin-6; PD-1, programmed cell death protein 1; PD-L1, programmed death-ligand 1; SII, systemic immune-inflammation index; SIRI, systemic inflammation response index; TGF-β, transforming growth factor beta.

Representative flow cytometry profiles illustrating the gating strategy and expression patterns. Plots from single representative individuals (1 MPN patient, 1 healthy control) show: (A) Monocyte subset identification based on CD14 and CD16 expression. (B) Histograms of CXCR3 expression on gated classical (blue) and intermediate (red) monocytes, with corresponding fluorescence-minus-one (FMO) controls (unfilled). (C-D) Representative plots for PD-1 and PD-L1 expression within CXCR3+ subsets. These plots illustrate the analytical approach and qualitative trends; quantitative group-level comparisons are provided in Figure 2 and Table 3.

Quantitative comparison of peripheral immune biomarkers in MPN patients vs healthy controls. Violin and box plots showing group-level distributions of immune biomarkers in MPN patients (red) and healthy controls (blue). (A) CXCR3 expression on classical and intermediate monocytes. (B) Immune checkpoint expression (PD-1 and PD-L1) on monocyte subsets. (C) Distribution of PD-L1+CXCR3+ and PD-L1−CXCR3− immune cell compartments across the study cohort. Central lines indicate medians, boxes represent interquartile ranges, and points denote individual subjects. Exact P-values and effect sizes are reported in Table 3.
Flow cytometric analysis demonstrated pronounced, subset-specific alterations. In the intergroup comparison, CXCR3 expression on classical monocytes was markedly reduced in MPN patients vs controls, whereas intermediate monocytes exhibited a significant increase (both P < .001), indicative of systemic immune activation. Crucially, for intradisease stratification, the relative level of CXCR3 on intermediate monocytes among MPN patients themselves emerged as a critical discriminator, with lower expression associated with more aggressive disease, as detailed in the following sections. Representative flow cytometry plots (Figure 1) illustrate the gating strategy and directional trends. These qualitative patterns were confirmed and quantified at the group level by violin plots (Figure 2) and are summarized with statistical metrics in Table 3. CXCR3 expression on CD8+ T cells was also significantly elevated in MPN.
Immune checkpoint dysregulation was a prominent feature of MPN. PD-1 and PD-L1 expression within CXCR3+ immune subsets was significantly increased (both P < .001), with large effect sizes. Quantitative analyses demonstrated clear rightward shifts in expression distributions among MPN patients (Figure 2), while representative flow cytometry plots confirmed consistent increases in fluorescence intensity (Figure 1).
Targeted cytokine analysis confirmed these cellular changes, with circulating IL-6 and TGF-β levels significantly elevated in MPN patients (both P < .001), supporting the presence of a proinflammatory and immunosuppressive immune milieu. Collectively, these findings indicate a coordinated peripheral immune disturbance in MPN characterized by inflammatory activation, CXCR3-linked immune remodeling, and immune checkpoint upregulation.
Relationship between immune biomarkers and reticulin fibrosis severity
Having established global CXCR3 dysregulation in MPN, we next investigated whether variation in CXCR3 expression levels within the MPN cohort correlated with fibrosis severity. The following analysis of fibrosis severity is restricted to patients with PMF (n = 45). Within this cohort, the distribution of fibrosis grade at enrollment was: grade 0-1, n = 1; grade 2, n = 10; grade 3, n = 20; grade 4, n = 14. Given the single patient in the grade 0-1 category, which precludes meaningful statistical analysis, and because grades ⩾ 2 represent the clinically relevant spectrum of overt fibrotic progression with established prognostic value, subsequent analyses focus on the gradient from grade 2 to grade 4. In PMF, advancing fibrosis was associated with progressive immune checkpoint activation and cytokine upregulation. PD-L1/CXCR3 and PD-1/CXCR3 expression rose with increasing fibrosis grade, while CXCR3 expression on T cells declined. Parallel increases in TGF-β supported its role in driving the profibrotic immune environment. To quantify these relationships, we performed Spearman’s rank correlation between each biomarker and fibrosis grade (treated as an ordinal variable). PD-L1/CXCR3 showed a strong positive correlation (ρ = 0.72, 95% CI: 0.58-0.82; P < .001), as did PD-1/CXCR3 (ρ = 0.54, 95% CI: 0.36-0.68; P < .001). In contrast, CXCR3 expression on intermediate monocytes was inversely correlated with fibrosis grade (ρ = −0.48, 95% CI: −0.64 to −0.28; P = .001).
To determine whether these associations were independent of potential confounders, we conducted multivariable ordinal logistic regression with fibrosis grade as the dependent variable, adjusting for age, sex, JAK2 V617F mutation status, and baseline blast count. In this model, PD-L1/CXCR3 remained a strong independent predictor of higher fibrosis grade (adjusted odds ratio [aOR] per 10% increase = 1.85, 95% CI: 1.42-2.41; P < .001). Similarly, CXCR3 on intermediate monocytes was independently associated with lower fibrosis grade (aOR per 5% increase = 0.76, 95% CI: 0.62-0.93; P = .009).
These trends point to a mechanistic connection between immune checkpoint dysregulation and marrow fibrotic remodeling (Table 4).
Immune biomarkers across reticulin fibrosis grades in PMF patients.

Adjusted for age, sex, JAK2 V617F mutation status, and baseline blast count using ordinal logistic regression.
Abbreviation: CXCR3, C-X-C chemokine receptor type 3; OR, odds ratio for higher fibrosis grade; PD-1, programmed cell death protein 1; PD-L1, programmed death-ligand 1; PMF, primary myelofibrosis; TGF-β, transforming growth factor beta.
Diagnostic performance of immune biomarkers for advanced fibrosis
To determine their diagnostic value in differentiating grade 4 from grade 3 reticulin fibrosis, we evaluated the performance of immune biomarkers using ROC and decision curve analyses. To account for the limited subgroup sample sizes (grade 4, n = 14; grade 3, n = 20) and provide a robust estimate of generalizable performance, we performed LOOCV. Among the tested markers, PD-L1/CXCR3 demonstrated robust discriminative accuracy (LOOCV AUC 0.928, 95% CI: 0.861-0.995). PD-1/CXCR3 showed moderate utility (LOOCV AUC 0.781), and CXCR3 expression on classical monocytes provided fair discrimination (LOOCV AUC 0.701). Decision curve analysis confirmed the clinical value of PD-L1/CXCR3 across a range of threshold probabilities, highlighting its potential role in guiding biopsy decisions and monitoring disease progression (Table 5, Figure 3).
Diagnostic performance of biomarkers for grade 4 versus grade 3 reticulin fibrosis.

Diagnostic performance was evaluated for discrimination between grade 4 (n = 14) and grade 3 (n = 20) reticulin fibrosis using receiver operating characteristic (ROC) analysis. To mitigate optimism bias due to limited sample size, leave-one-out cross-validation (LOOCV) was performed, and the LOOCV AUC is presented as the primary robust metric of performance.
Abbreviation: AUC, area under the curve; CI, confidence interval; CXCR3, C-X-C chemokine receptor type 3; NPV, negative predictive value; OR, odds ratio; PD-1, programmed cell death protein 1; PD-L1, programmed death-ligand 1; PPV, positive predictive value.

Diagnostic performance of immune biomarkers for distinguishing grade 4 from grade 3 reticulin fibrosis. Panel A shows ROC curves for PD-L1/CXCR3, PD-1/CXCR3, and CXCR3/Classical. Panel B displays comparative AUC values with sensitivity and specificity. Axes and optimal cut-offs correspond exactly to Table 5 (PD-L1/CXCR3 > 40%; PD-1/CXCR3 > 15%; CXCR3/Classical ⩽ 10%). Full operating characteristics are detailed in Table 5. LOOCV AUCs are reported to reduce optimism bias for small subgroup analyses.
Predictive role of biomarkers for leukemic transformation
We then evaluated the same intrapatient biomarker variation for predicting leukemic transformation. Analysis of leukemic transformation was performed in the PMF subgroup (n = 45), where this outcome is most clinically relevant. The number of leukemic transformation events was low (n = 8), which precludes definitive conclusions and requires future validation in larger studies. However, in this exploratory analysis, CXCR3 on intermediate monocytes showed the strongest association with leukemic transformation (AUC 0.876; NPV 98.4%). Furthermore, this biomarker was a key independent predictor of survival and a selected component of our overall prognostic score, which was successfully validated in an external cohort (AUC 0.82). These converging lines of evidence suggest CXCR3/intermediate may be useful for identifying high-risk patients and merit further investigation (Table 6, Figure 4). Other biomarkers, including SII, SIRI, PD-1/CXCR3, and PD-L1/CXCR3, showed weaker or inconsistent performance.
ROC analysis of biomarkers predicting leukemic transformation (n = 8 events/90 patients).
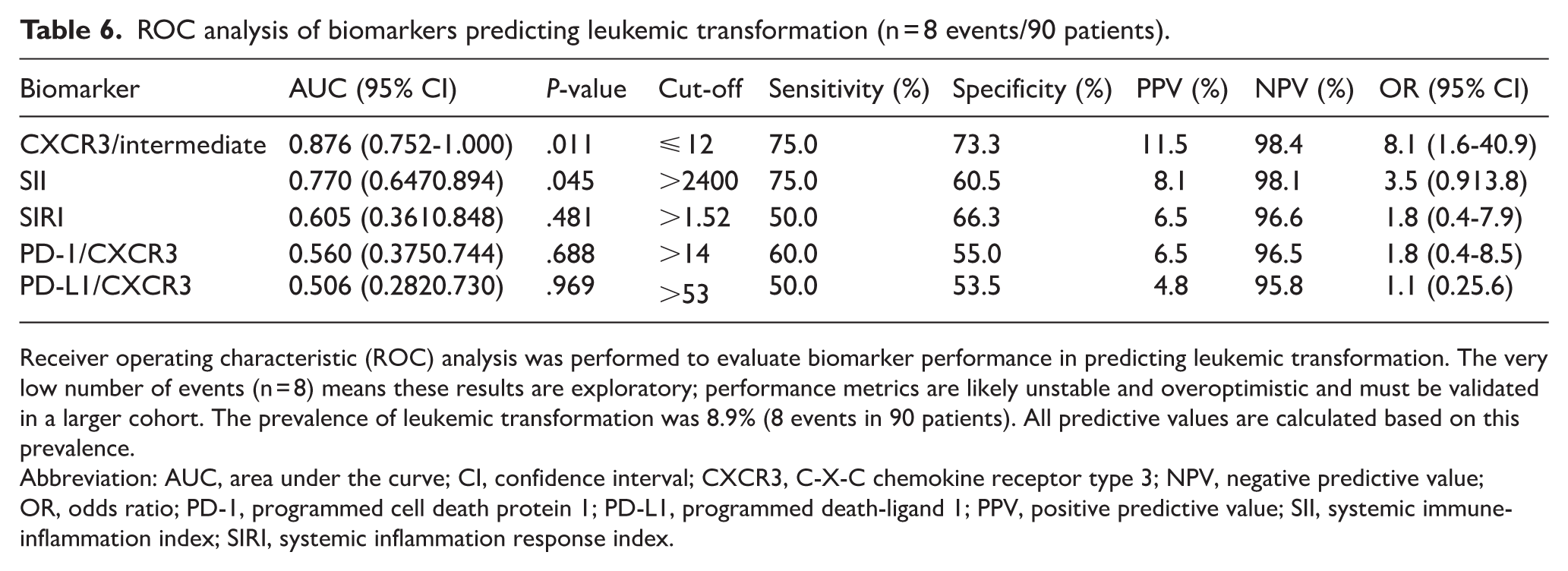
Receiver operating characteristic (ROC) analysis was performed to evaluate biomarker performance in predicting leukemic transformation. The very low number of events (n = 8) means these results are exploratory; performance metrics are likely unstable and overoptimistic and must be validated in a larger cohort. The prevalence of leukemic transformation was 8.9% (8 events in 90 patients). All predictive values are calculated based on this prevalence.
Abbreviation: AUC, area under the curve; CI, confidence interval; CXCR3, C-X-C chemokine receptor type 3; NPV, negative predictive value; OR, odds ratio; PD-1, programmed cell death protein 1; PD-L1, programmed death-ligand 1; PPV, positive predictive value; SII, systemic immune-inflammation index; SIRI, systemic inflammation response index.

ROC analysis of biomarkers predicting leukemic transformation in MPN patients. Panel A shows ROC curves for CXCR3/intermediate, SII, SIRI, PD-1/CXCR3, and PD-L1/CXCR3. Panel B summarizes AUC, sensitivity, and specificity for each biomarker. Full performance metrics are detailed in Table 6.
Associations between biomarkers and DIPSS risk categories
To determine whether immune biomarkers reflect clinical severity, we evaluated their ability to distinguish high- from low-risk disease by DIPSS scoring. We first examined the correlation between biomarker levels and DIPSS score as a continuous variable using Spearman’s rank correlation. PD-1/CXCR3 showed the strongest positive correlation (ρ = 0.65, 95% CI: 0.50-0.76; P < .001), followed by PD-L1/CXCR3 (ρ = 0.58, 95% CI: 0.41-0.71; P < .001). Conversely, CXCR3 expression on intermediate monocytes was inversely correlated with DIPSS score (ρ = −0.61, 95% CI: −0.73 to −0.45; P < .001).
We next performed multivariable linear regression with DIPSS score as the dependent variable, adjusting for age, sex, cytogenetic risk category, and baseline blast count. In this model, PD-1/CXCR3 (β = 0.32, 95% CI: 0.21-0.43; P < .001) and PD-L1/CXCR3 (β = 0.28, 95% CI: 0.17-0.39; P < .001) remained significant independent predictors of higher DIPSS scores. CXCR3 on intermediate monocytes was independently associated with lower DIPSS scores (β = −0.24, 95% CI: −0.35 to −0.13; P < .001). These findings highlight the link between checkpoint activity, CXCR3 signaling, and clinical disease burden in PMF (Table 7, Figure 5).
ROC analysis of biomarkers for predicting high DIPSS risk (⩾ 3) (n = 28 high risk/45 PMF patients).
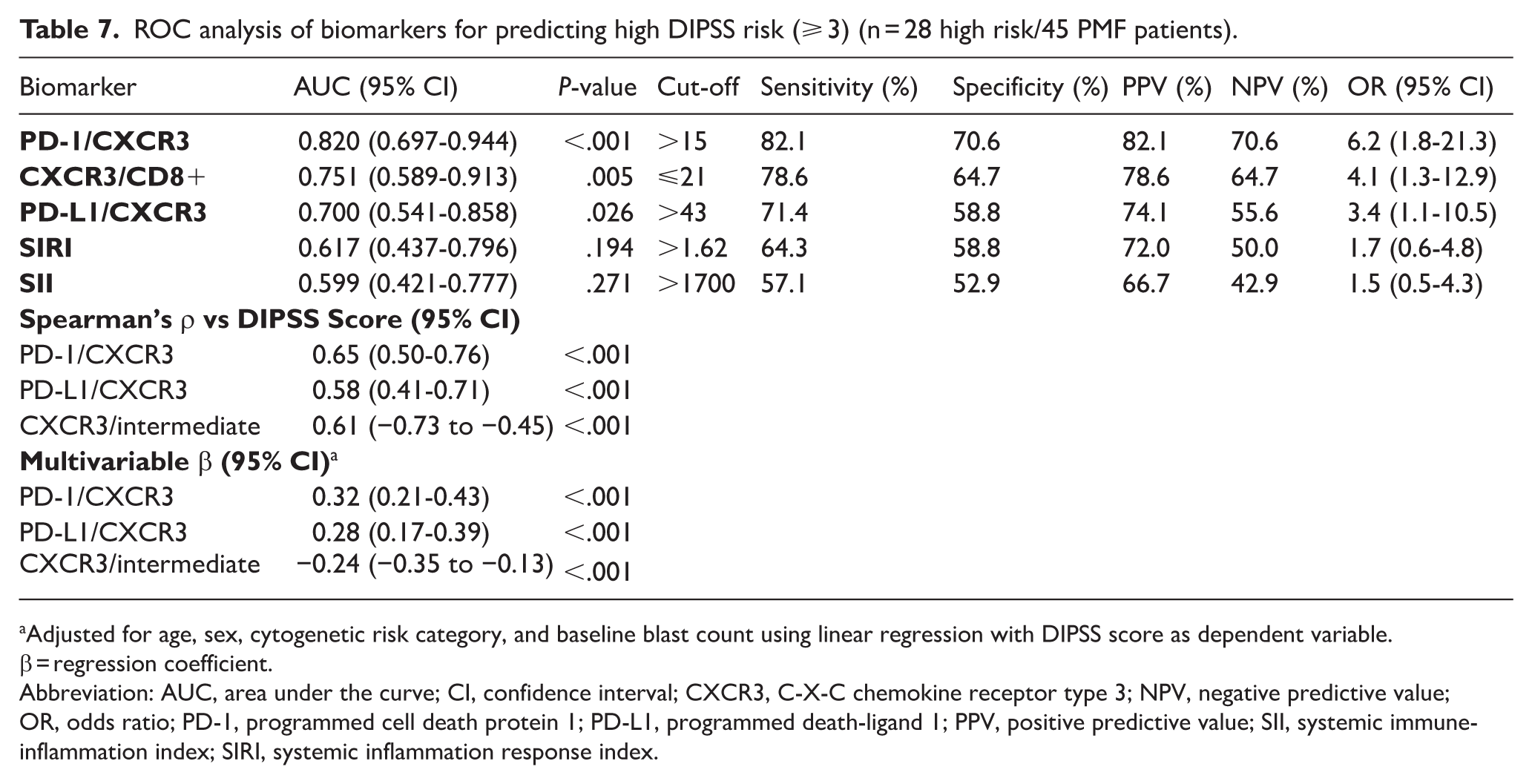
Adjusted for age, sex, cytogenetic risk category, and baseline blast count using linear regression with DIPSS score as dependent variable. β = regression coefficient.
Abbreviation: AUC, area under the curve; CI, confidence interval; CXCR3, C-X-C chemokine receptor type 3; NPV, negative predictive value; OR, odds ratio; PD-1, programmed cell death protein 1; PD-L1, programmed death-ligand 1; PPV, positive predictive value; SII, systemic immune-inflammation index; SIRI, systemic inflammation response index.

ROC analysis of biomarkers predicting high DIPSS risk in PMF patients. Panel A shows ROC curves for PD-1/CXCR3, CXCR3/CD8+, PD-L1/CXCR3, SIRI, and SII. Panel B displays AUC, sensitivity, and specificity for each biomarker. Full performance metrics are detailed in Table 7.
Risk analysis stratified by MPN subtype: survival in PMF and thrombosis in PV/ET
Risk analysis was stratified by MPN subtype based on distinct clinical endpoints. For the PMF cohort (n = 45), we analyzed overall and event-free survival. For the PV and ET cohorts (n = 45), we performed an exploratory analysis of thrombotic event risk. Follow-up for the entire cohort was estimated using the reverse Kaplan-Meier method (median, 42 months; IQR, 28-56 months). Within the PMF cohort, there were 22 deaths (48.9%) and 32 EFS events (71.1%). In the PV/ET cohort, seven thrombotic events (15.6%) occurred during follow-up. Kaplan-Meier curves showed significantly worse OS and EFS for patients in the PD-L1/CXCR3 High group and for patients with CXCR3/intermediate low expression (Table 8, Figure 6). In multivariable Cox models (adjusted for age, JAK2 status, and DIPSS-Plus), PD-L1/CXCR3 High was associated with increased risk of death (HR 1.18, 95% CI: 1.02-1.37; modeled per 10-percentage-point increase), and CXCR3/intermediate low was associated with increased risk vs High (HR 1.22, 95% CI: 1.02-1.47). Biomarkers were analyzed as categorical High/Low groups for KM curves (cut-offs by Youden’s index); continuous analyses are reported per 10% units in the Supplementary Methods.
Biomarker-associated risk stratified by MPN subtype.

Abbreviations: CI, confidence interval; HR, hazard ratio; OR, odds ratio.
Cox model adjusted for age, JAK2 V617F status, and DIPSS-Plus risk category.
Cut-offs for biomarker groups: SIRI High > 1.8; PD-L1/CXCR3 High > 50%; CXCR3/intermediate low ⩽ 18%.

Survival analysis according to immune biomarker expression in primary myelofibrosis (PMF). Panel A1. PD-L1/CXCR3 expression and overall survival in PMF (n = 45). Kaplan-Meier curves show overall survival stratified by PD-L1/CXCR3 expression (High: n = 30; Low: n = 15). Patients with high PD-L1/CXCR3 expression had significantly inferior survival compared with the low-expression group (median OS: 25.1 vs 46.5 months; 3-year OS rate: 53.3% vs 86.7%; log-rank P = .008). The adjusted hazard ratio (HR) for death was 2.15 (95% CI: 1.25-3.71), with the low-expression group used as the reference. Panel A2. CXCR3/intermediate expression and overall survival in PMF (n = 45). Kaplan-Meier curves show overall survival according to CXCR3/intermediate expression (High: n = 20; Low: n = 25). Patients with low expression had significantly worse survival compared with those with high expression (median OS: 23.4 vs 44.8 months; 3-year OS rate: 52% vs 85%; log-rank P = .012). The adjusted HR for death (Low vs High) was 1.94 (95% CI: 1.16-3.25), with the high-expression group as the reference. Hazard ratios were estimated using Cox proportional hazards models adjusted for age, JAK2 V617F mutation status, and DIPSS-Plus risk category.
As shown in Table 8, the prognostic utility of PD-L1/CXCR3 and CXCR3/intermediate expression was confirmed in the PMF cohort, with high PD-L1/CXCR3 and low CXCR3/intermediate independently associated with inferior survival (panel A). In an exploratory analysis of the PV/ET cohort (panel B), markers of systemic inflammation (SIRI) and immune checkpoint activity (PD-L1/CXCR3) were associated with an increased incidence of thrombotic events, although the number of events was limited. These subtype-specific analyses underscore the central role of immune-inflammatory dysregulation across the MPN spectrum, driving fibrosis and transformation in PMF and potentially modulating thrombotic risk in PV/ET.
Machine learning–derived immune phenotypes
To explore whether unbiased computational methods could refine patient stratification, we applied unsupervised clustering to immune checkpoint and CXCR3 expression profiles. Two distinct phenotypes emerged: a checkpoint-high cluster, enriched for PD-1/CXCR3 and PD-L1/CXCR3 and associated with advanced fibrosis and higher DIPSS risk, and a checkpoint-low cluster, characterized by reduced checkpoint activity and milder disease. Feature importance analysis identified PD-L1/CXCR3 and CXCR3/intermediate as the dominant predictors, a ranking confirmed by both random forest and XGBoost models. These results highlight the ability of machine learning to validate biomarker-driven disease heterogeneity (Table 9, Figure 7).
Characteristics of machine learning–derived immune phenotypes in MPN patients.
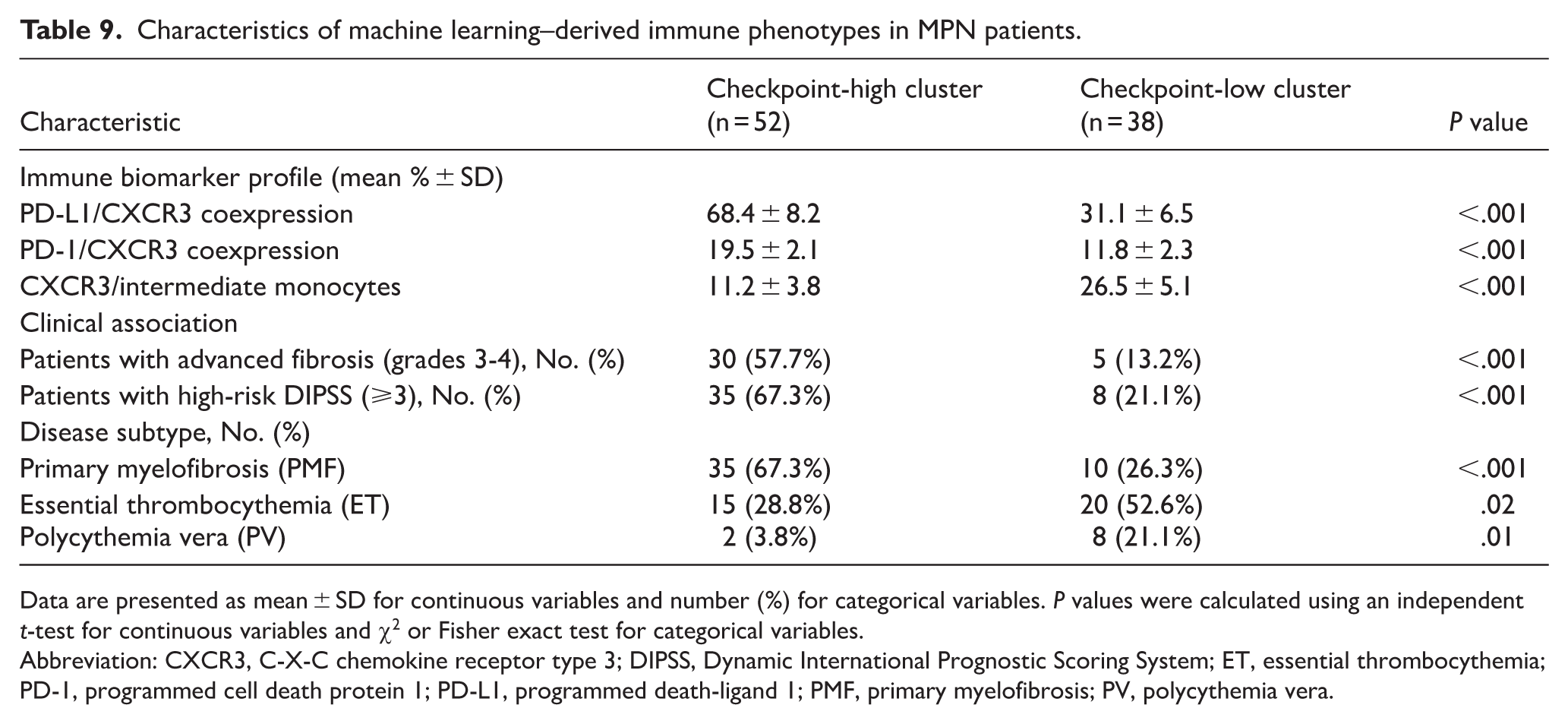
Data are presented as mean ± SD for continuous variables and number (%) for categorical variables. P values were calculated using an independent t-test for continuous variables and χ2 or Fisher exact test for categorical variables.
Abbreviation: CXCR3, C-X-C chemokine receptor type 3; DIPSS, Dynamic International Prognostic Scoring System; ET, essential thrombocythemia; PD-1, programmed cell death protein 1; PD-L1, programmed death-ligand 1; PMF, primary myelofibrosis; PV, polycythemia vera.

Machine learning–derived immune phenotypes and biomarker predictors. (A) Principal component analysis (PCA) plot of immune biomarker profiles in 90 MPN patients. Unsupervised k-means clustering of principal components identified 2 phenotypes: a checkpoint-high cluster (red), enriched for PD-1/PD-L1 coexpression and associated with advanced fibrosis and higher DIPSS risk, and a checkpoint-low cluster (teal), associated with milder disease. PC1 and PC2 represent the dimensions explaining the greatest variance. (B) Random Forest feature importance ranking based on the mean decrease in Gini index. PD-L1/CXCR3 coexpression and CXCR3/intermediate monocytes were the most influential biomarkers distinguishing between the phenotypes, highlighting their central role in MPN immune heterogeneity.
Integrated multiomics profiling for biomarker validation
Targeted NGS identified canonical driver mutations in PMF with expected distributions and frequent co-occurrence of mutations. RNA-seq performed in a representative subset confirmed the flow cytometry findings by demonstrating consistent upregulation of PD-L1 (CD274) and CXCR3 transcripts compared with controls. Enrichment analyses highlighted activation of JAK/STAT and NF-κB pathways in patients harboring JAK2 mutations combined with high PD-L1/CXCR3 expression, linking mutational drivers with immune checkpoint activation. Targeted cytokine analysis further validated these results by showing elevated IL-6 and TGF-β levels in JAK2-positive patients.
In integrative regression modeling, specific mutations showed distinct associations with immune checkpoint phenotypes. JAK2 and ASXL1 independently predicted high PD-L1/CXCR3 expression, whereas SRSF2 predicted reduced CXCR3/intermediate expression. Other mutations demonstrated weaker or pathway-specific associations. These results underscore how genetic lesions converge on immune dysregulation through both signaling and epigenetic mechanisms (Table 10, Figure 8).
Integrative analysis of biomarkers and driver mutations.
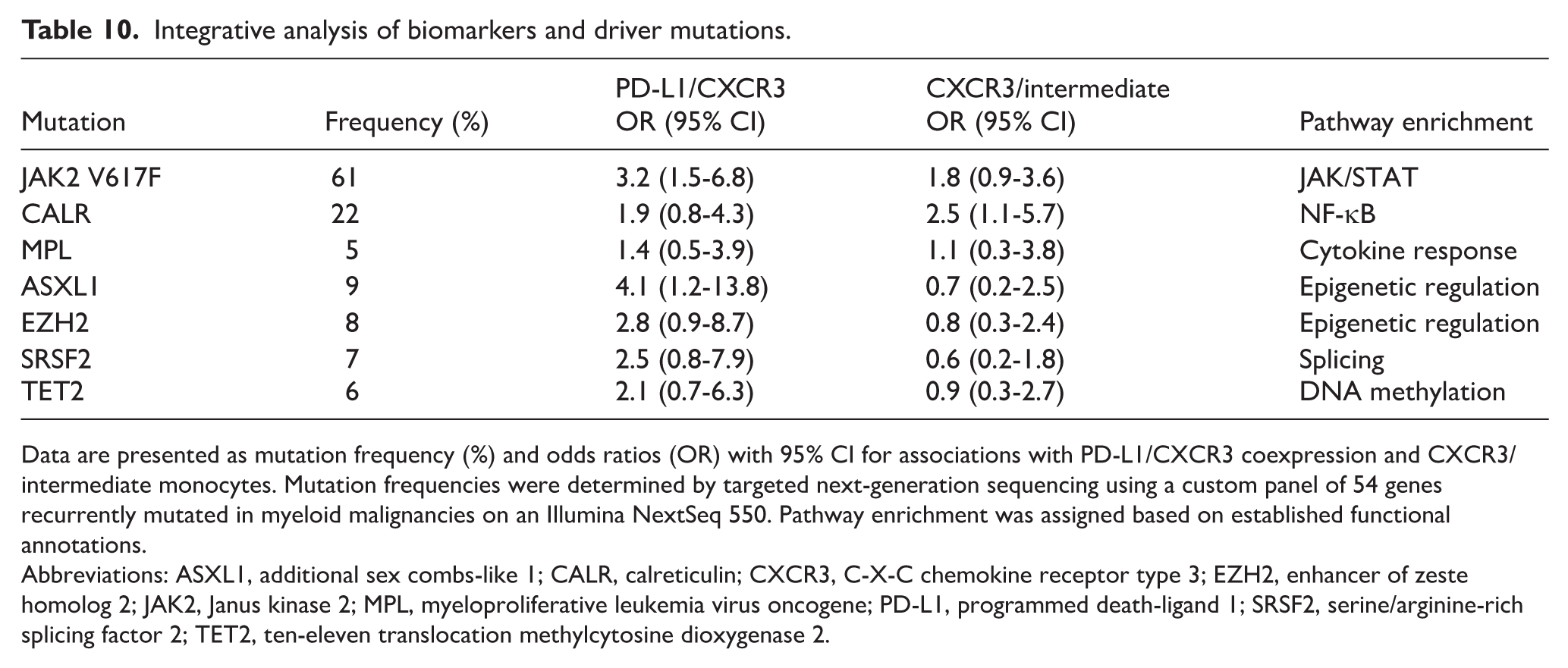
Data are presented as mutation frequency (%) and odds ratios (OR) with 95% CI for associations with PD-L1/CXCR3 coexpression and CXCR3/intermediate monocytes. Mutation frequencies were determined by targeted next-generation sequencing using a custom panel of 54 genes recurrently mutated in myeloid malignancies on an Illumina NextSeq 550. Pathway enrichment was assigned based on established functional annotations.
Abbreviations: ASXL1, additional sex combs-like 1; CALR, calreticulin; CXCR3, C-X-C chemokine receptor type 3; EZH2, enhancer of zeste homolog 2; JAK2, Janus kinase 2; MPL, myeloproliferative leukemia virus oncogene; PD-L1, programmed death-ligand 1; SRSF2, serine/arginine-rich splicing factor 2; TET2, ten-eleven translocation methylcytosine dioxygenase 2.
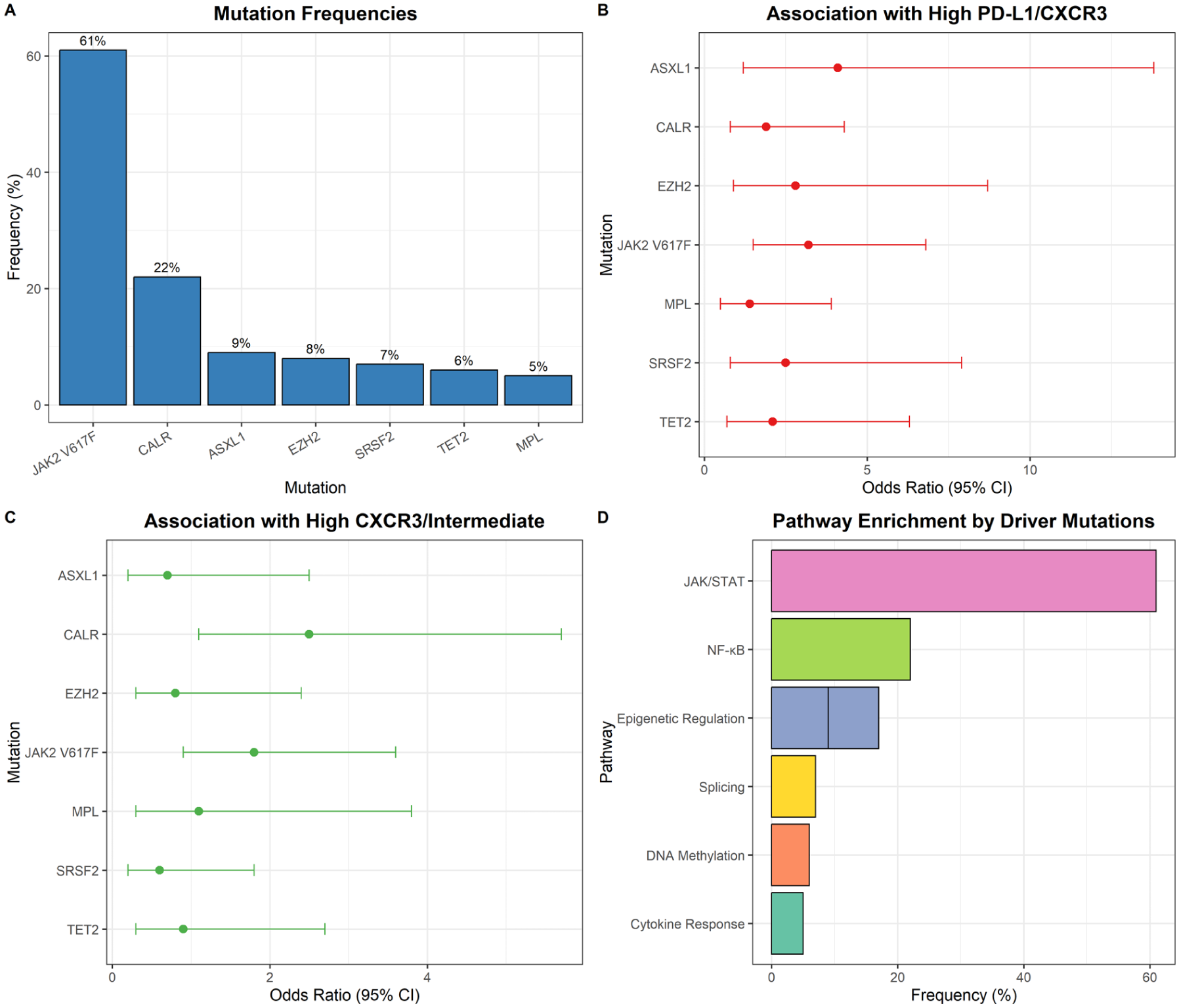
Integrative analysis of biomarkers and driver mutations. Panel A: Mutation frequencies in PMF patients. Panel B: Odds ratios (95% CI) linking driver mutations to PD-L1/CXCR3 expression. Panel C: Odds ratios (95% CI) linking mutations to CXCR3/intermediate expression. Panel D: Pathway enrichment mapping each mutation to its dominant signaling program.
These integrated genomic, transcriptomic, and cytokine-level findings provided the rationale for subsequent single-cell resolution analyses to delineate the cellular sources and transcriptional programs underlying CXCR3- and checkpoint-associated immune dysregulation.
Single-cell transcriptomic profiling of immune subsets
To resolve the multiomics signals identified at the bulk level into discrete cellular populations, we performed scRNA-seq of PBMCs, focusing on monocyte and T-cell compartments. Single-cell RNA sequencing identified 3 transcriptionally distinct immune subsets within PBMCs. The most striking was the CXCR3-low intermediate monocyte population, which was enriched for an inflammatory and profibrotic transcriptional program and was associated with increased IL-6 production on flow cytometry. These cells may contribute to both fibrotic remodeling and leukemic progression.
By contrast, CXCR3-high classical monocytes maintained strong chemotactic signatures, while PD-1–enriched T cells reflected immune exhaustion. Pseudotime trajectory analysis was consistent with CXCR3 downregulation occurring at an early stage of immune divergence, suggesting it may represent an early event in disease-associated immune remodeling. Together, these findings demonstrate how discrete immune subsets contribute differentially to disease biology and support a model in which early checkpoint imbalance in monocytes propagates downstream dysfunction across the immune compartment (Table 11, Figure 9).
Single-cell profiling of immune subsets.

Single-cell RNA sequencing (scRNA-seq) clusters are defined by cell type, CXCR3 expression pattern, differentially expressed genes (DEGs), and enriched pathways.
Abbreviations: CCR2, C-C chemokine receptor type 2; CXCL10, C-X-C motif chemokine ligand 10; CXCR3, C-X-C chemokine receptor type 3; DEGs, differentially expressed genes; IL6, interleukin-6; PDCD1, programmed cell death protein 1; TGFB1, transforming growth factor beta 1.
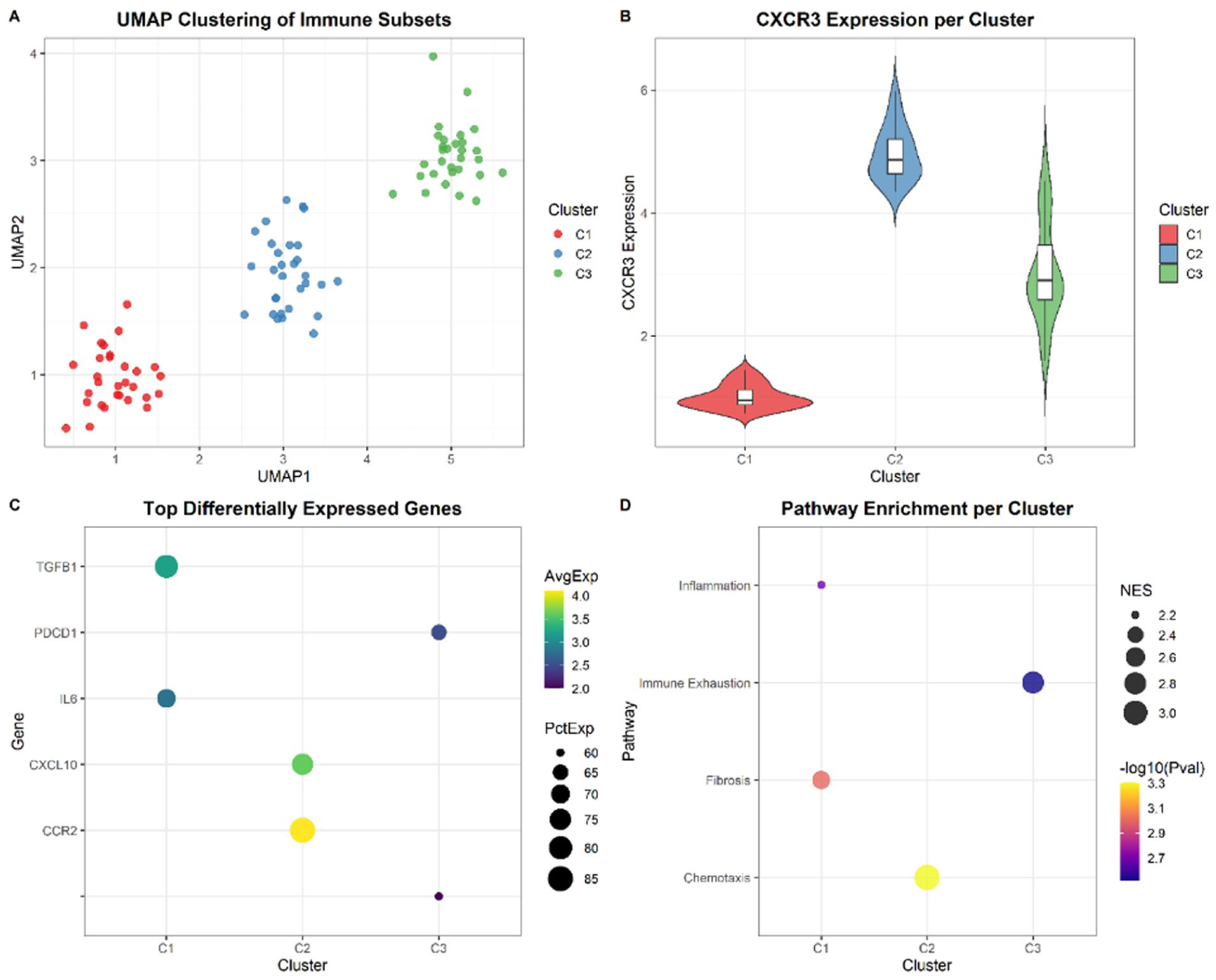
Single-cell profiling of immune subsets. Panel A: UMAP showing 3 immune clusters (C1, C2, C3). Panel B: Violin plots of CXCR3 expression showing low levels in C1, high levels in C2, and variable levels in C3. Panel C: Dot plot of the top differentially expressed genes per cluster. Panel D: Pathway enrichment bubble plot showing functional differences across clusters.
Biomarker-enhanced prognostic scoring model and clinical utility
A LASSO-selected score combining PD-L1/CXCR3, CXCR3/intermediate, age, and DIPSS showed higher discrimination for adverse outcomes than DIPSS, DIPSS-Plus, or MIPSS70 (AUC 0.85 vs 0.71-0.78). The model demonstrated robust internal validity on bootstrap resampling (1000 iterations; optimism-corrected AUC 0.84) and showed promising generalizability in a small, independent external cohort (n = 30; AUC 0.82, 95% CI: 0.68-0.96).
Decision curve analysis confirmed its clinical value across a broad range of threshold probabilities, with superior net benefit particularly at intermediate risk thresholds. This improvement reflected better identification of patients at true high risk, while simultaneously reducing overtreatment in low-risk individuals. Importantly, PPV and NVP indicated strong practical utility for guiding individualized therapeutic decisions (Table 12, Figure 10).
Performance of prognostic models (n = 45).

Model performance was assessed by receiver operating characteristic (ROC) analysis for predicting (eg, 5-year survival) in the PMF cohort (n = 45).
Abbreviations: AUC, area under the curve; CI, confidence interval; DIPSS, Dynamic International Prognostic Scoring System; MIPSS70, Mutation-Enhanced International Prognostic Scoring System 70.
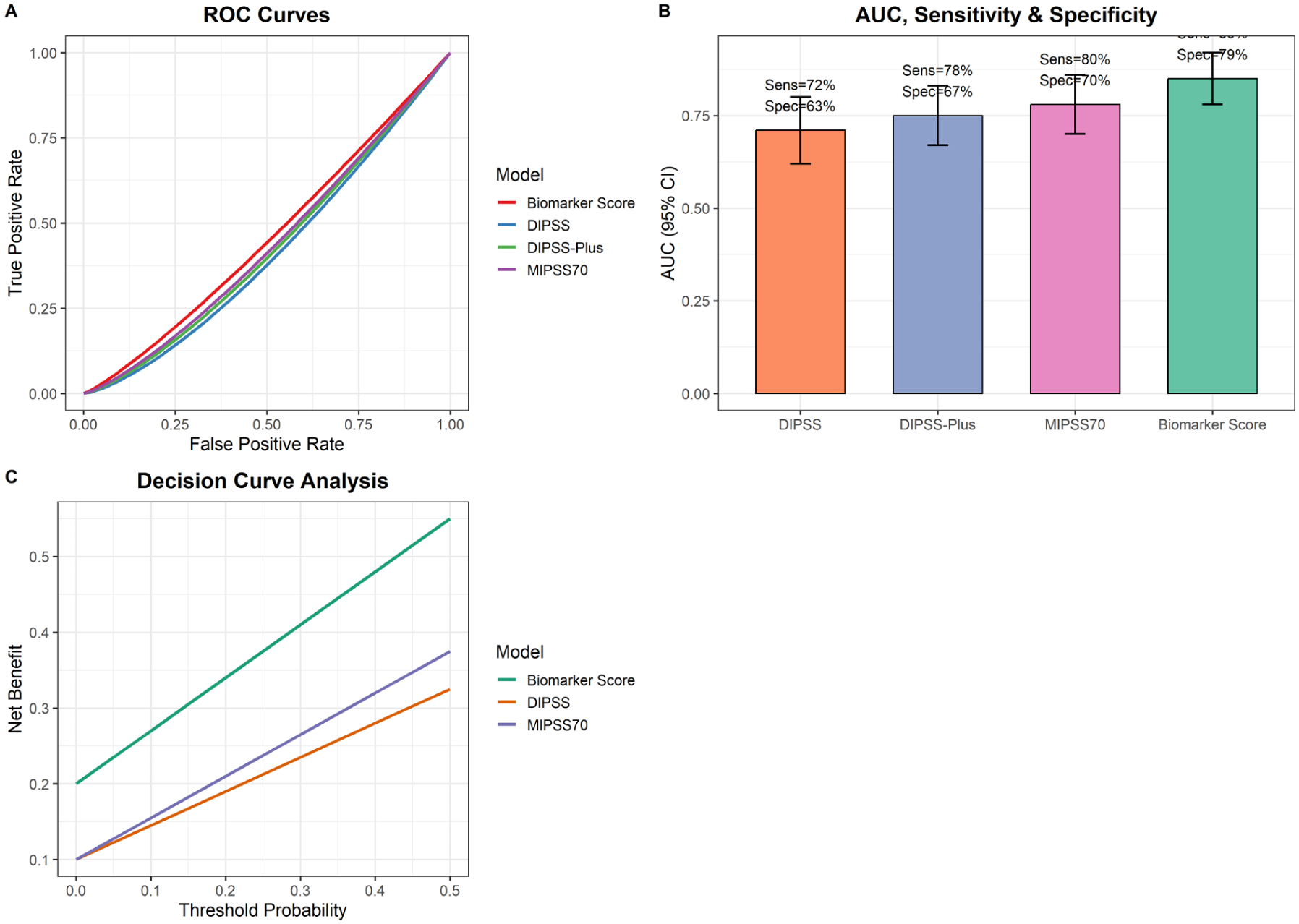
Performance and clinical utility of the biomarker-enhanced prognostic model. Panel A: ROC curves comparing the biomarker-enhanced score with DIPSS, DIPSS-Plus, and MIPSS70. Panel B: Bar plot summarizing AUC, sensitivity, and specificity for each model. Panel C: Decision curve analysis showing net clinical benefit across threshold probabilities.
Taken together, these findings suggest that biomarker-based immune profiling has the potential to be integrated into routine risk stratification to refine treatment planning for MPN patients in real-world Egyptian practice, though validation in larger, multicenter cohorts is recommended.
PROs and biomarker correlation
Symptom assessment using the MPN-SAF tool confirmed a high burden of disease-related complaints, with fatigue and pruritus being most prominent. Correlative analyses linked these symptom domains to immune checkpoint activity: PD-L1/CXCR3 expression showed the strongest association with fatigue and pruritus, while lower CXCR3/intermediate ratios were consistently associated with greater symptom severity across domains. These correlations were observed across all MPN subtypes, though the strength of association was generally strongest in PMF patients, consistent with their higher overall symptom burden. These findings suggest that immune dysregulation is not only mechanistically relevant to disease progression but also directly impacts patient QoL.
Longitudinal follow-up provided additional support for this link. Among patients treated with JAK inhibitors, reductions in PD-L1/CXCR3 levels paralleled improvements in fatigue scores over 12 months. Together, these results integrate biomarker changes with clinical experience, reinforcing the translational significance of checkpoint dysregulation in the daily lives of patients (Table 13).
Correlation between biomarkers and MPN-SAF symptom scores.
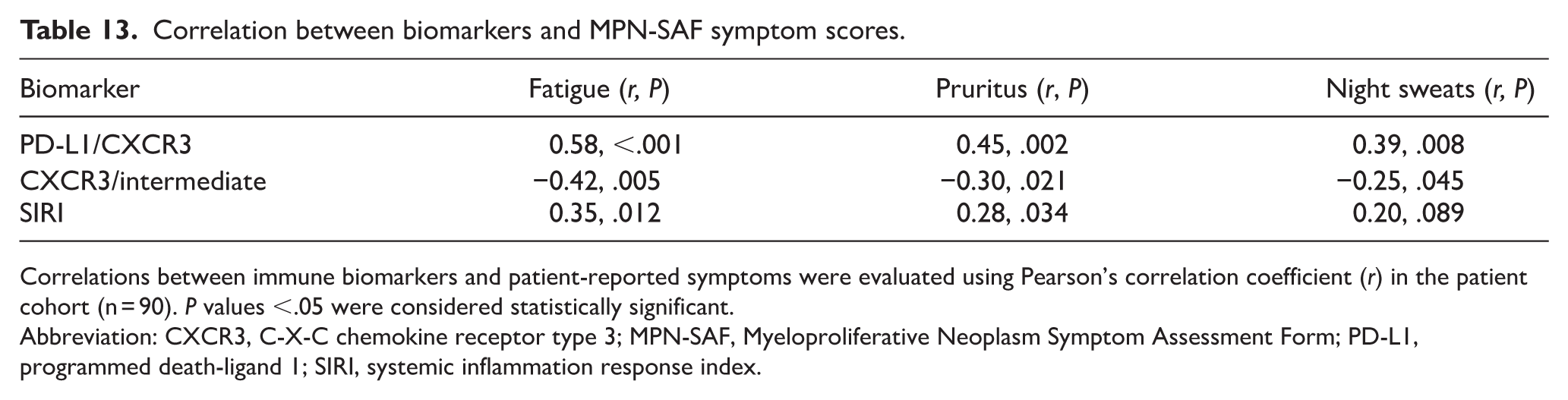
Correlations between immune biomarkers and patient-reported symptoms were evaluated using Pearson’s correlation coefficient (r) in the patient cohort (n = 90). P values <.05 were considered statistically significant.
Abbreviation: CXCR3, C-X-C chemokine receptor type 3; MPN-SAF, Myeloproliferative Neoplasm Symptom Assessment Form; PD-L1, programmed death-ligand 1; SIRI, systemic inflammation response index.
Therapeutic implications and biomarker-guided treatment
Based on the immune phenotypes identified in this study, we propose biomarker-stratified therapeutic hypotheses that integrate observations from our cohort with mechanistic insights from the literature. Our in vitro experiments demonstrated that anti-PD-L1 treatment reduced IL-6 secretion from monocytes isolated from patients with high PD-L1/CXCR3 coexpression, suggesting that checkpoint inhibition may restore immune surveillance and attenuate fibrosis in this subset. This finding directly supports a testable hypothesis: patients with PD L1/CXCR3 > 40% (the cut-off for advanced fibrosis, Table 5) could be prioritized for clinical trials evaluating PD L1/PD 1 blockade, preferably in combination with JAK inhibitors to simultaneously target inflammatory signaling.
Within our cohort, low CXCR3 expression on intermediate monocytes (CXCR3/intermediate ⩽ 12%) was associated with both leukemic transformation risk (Table 6) and greater clinical response to JAK inhibitor therapy (Table 15). This phenotype may reflect a state of heightened JAK/STAT-driven inflammation that is particularly susceptible to JAK inhibition. Therefore, we propose that prospective trials stratify PMF patients by CXCR3/intermediate expression to test whether this biomarker identifies a subgroup with enhanced benefit from JAK inhibitor therapy.
The association between ASXL1 mutations and high PD-L1/CXCR3 expression observed in our cohort (Table 10) provides a mechanistic rationale for exploring epigenetic modulation in this context. Although the link to BET inhibition is extrapolated from preclinical literature demonstrating that ASXL1 mutations sustain inflammatory programs via epigenetic dysregulation, our data support the biological plausibility of combining epigenetic agents with checkpoint blockade in ASXL1-mutated, checkpoint-high patients. This hypothesis warrants validation in preclinical MPN models before clinical translation.
Importantly, these recommendations are derived from a single-center Egyptian cohort and require prospective validation in larger, multiethnic populations. Future biomarker-driven trials should prospectively validate the PD-L1/CXCR3 and CXCR3/intermediate cut-offs, incorporate them as stratification factors, and include correlative studies to confirm that treatment biomarker changes correlate with clinical response. Together, these hypotheses provide a concrete, evidence-informed framework for designing precision medicine trials in MPN, clearly distinguishing cohort-derived observations from literature-based extrapolations (Table 14).
Biomarker-informed therapeutic hypotheses.

Proposed therapeutic strategies are hypotheses based on biomarker-defined immune phenotypes and mutational context. Clinical trials are required to confirm therapeutic benefit.
Abbreviation: ASXL1, additional sex combs-like 1; BET, bromodomain and extraterminal domain proteins; CXCR3, C-X-C chemokine receptor type 3; JAK, Janus kinase; PD-L1, programmed death-ligand 1.
Longitudinal biomarker dynamics and treatment response
Serial profiling during JAK inhibitor therapy demonstrated that immune checkpoint and inflammatory biomarkers are dynamic and treatment-responsive. Patients showed a clear reduction in PD-L1/CXCR3 and SIRI levels during follow-up, while CXCR3/intermediate expression increased, aligning with histological improvement in a subset of patients. These changes were not mirrored by SII, suggesting it is less sensitive to therapeutic modulation.
Importantly, patients with marked reductions in PD-L1/CXCR3 derived the greatest clinical benefit, including longer survival and better symptom control. These findings suggest that longitudinal biomarker monitoring offers both mechanistic insights and clinically actionable information, supporting its use in guiding therapy and assessing treatment efficacy in MPN (Table 15).
Longitudinal changes in biomarkers post-JAK inhibitor therapy.

Data are presented as mean ± standard deviation from patients with paired baseline and 12-month samples (n = 25). P-values were calculated using a paired t-test comparing baseline and 12-month values. Effect sizes are reported as Cohen’s d.
Abbreviations: CXCR3, C-X-C chemokine receptor type 3; PD-L1, programmed death-ligand 1; SII, systemic immune-inflammation index; SIRI, systemic inflammation response index.
Discussion
The MPNs are chronic clonal hematopoietic disorders characterized by cytokine-driven inflammation, bone marrow fibrosis, and a risk of transformation to acute leukemia. 24 Risk stratification has traditionally relied on clinical scoring systems, but they fail to fully capture the heterogeneity of disease biology. 25 Immune dysregulation, particularly involving immune checkpoint pathways such as PD-1/PD-L1 and chemokine receptors like CXCR3, has emerged as a central mechanism linking inflammation to disease progression. 26 The observed coexpression of PD-1/PD-L1 with CXCR3 on monocyte subsets highlights a clinically relevant immune phenotype that integrates immune exhaustion with altered chemokine-driven myeloid trafficking, providing mechanistic insight into the progression of fibrosis and leukemic evolution in PMF. However, data from low- and middle-income countries, including Egypt, remain limited despite potential regional differences in genetics, environmental exposures, and treatment access. 27 Our study aimed to address this gap by profiling immune checkpoint expression, cytokine signatures, and transcriptional heterogeneity in an Egyptian MPN cohort using flow cytometry, RNA-seq, and scRNA-seq, complemented by advanced computational modeling. By integrating these data with established prognostic indices, we provide context-specific insights into the immune landscape of Egyptian patients, which may inform biomarker-driven therapeutic strategies. This multilayered approach, spanning cellular phenotyping, transcriptomics, and cytokine-level validation, strengthens the biological coherence of our findings.28,29
The Discussion is organized thematically to reflect the analytical architecture of the study, progressing from immune biology to clinical risk stratification, computational validation, and translational implications. This structure mirrors the stepwise validation strategy employed in the Results and is intended to guide the reader through increasingly integrated levels of biological and clinical interpretation.
Our data converge on 2 complementary and context-dependent immune axes. First, when comparing MPN patients to healthy controls, we observed a global increase in CXCR3 expression on intermediate monocytes, consistent with a state of systemic immune activation. Second, and more prognostically pertinent, when stratifying the MPN cohort itself, relative downregulation of CXCR3 on these cells (ie, a CXCR3-low phenotype) marked patients with advanced fibrosis, leukemic evolution, and inferior survival. This pattern is biologically plausible: early disease may be characterized by inflammatory chemokine receptor upregulation, while progression is marked by the emergence of a functionally distinct, CXCR3-low monocyte subset with profibrotic transcriptional programs, as identified by our scRNA-seq analysis. This dual-layer analysis clarifies that our biomarker identifies not merely the presence of immune dysregulation, but a specific maladaptive immune state within it.
In our cohort, the mean age of patients was 63.5 ± 5.9 years, with a slight male predominance. With myelofibrosis showing the highest symptom burden and splenomegaly rates, PMF, ET, and PV were represented (Table 1). Compared with healthy controls, patients displayed significantly elevated leukocyte counts, hemoglobin variability, and serum LDH levels, consistent with heightened proliferative activity. Inflammatory markers, including CRP and ESR, were also higher, suggesting systemic inflammation as a hallmark across subtypes (Table 2).
These findings mirror epidemiological observations that MPN patients in Middle Eastern and North African countries tend to present at younger ages than their European counterparts. 30 An Egyptian multicenter analysis also noted earlier presentation of myelofibrosis with aggressive hematologic features, supporting our results. 31 The elevated inflammatory profile we observed aligns with evidence that chronic cytokine stimulation, particularly IL-6 and TGF-β, is linked to both clonal expansion and marrow fibrosis. 32 Higher LDH and abnormal EPO levels in our cohort reinforce the role of ineffective erythropoiesis, a finding that has been consistently reported in global MPN studies. 33 Taken together, these baseline characteristics emphasize the contribution of both proliferative and inflammatory mechanisms to disease biology in Egyptian patients and highlight the importance of integrating clinical variables with immune and molecular profiling for refined prognostication.
Our data converge on 2 complementary immune axes. First, checkpoint-high states (elevated PD-L1/CXCR3) track with advanced fibrosis and worse symptom burden, consistent with microenvironmental tolerance and TGF-β upregulation. Second, CXCR3-low intermediate monocytes mark leukemic evolution and inferior survival, supported by scRNA-seq signatures (IL-6/TGF-β programs) and high NPV for ruling out transformation. Integrating these axes with DIPSS meaningfully improves stratification, a finding reproduced by bootstrap and external validation.
Across the patient cohort, we observed a distinct upregulation of immune checkpoints. In MPN patients, PD-1 expression was significantly higher on both CD4+ and CD8+ T cells compared with controls, whereas PD-L1 expression was markedly increased on monocytes (Table 3). CXCR3, in contrast, was reduced on circulating monocytes in patients with advanced fibrosis, suggesting a loss of immune effector potential. When stratified by fibrosis grade, checkpoint expression showed clear trends: PD-L1 positivity was enriched in early fibrotic stages, whereas CXCR3 downregulation became more evident with progression to higher grades (Table 4). These results indicate that immune checkpoint activation and chemokine receptor alterations track with fibrotic burden, linking immune exhaustion and stromal remodeling in disease evolution.
The immune microenvironment has been increasingly recognized as a driver of MPN biology, with checkpoint pathways playing central roles. Our findings of heightened PD-1/PD-L1 expression echo reports that these molecules are upregulated in hematologic malignancies and contribute to T-cell dysfunction. 34 Egyptian studies in AML have similarly shown increased PD-1 expression on T cells, associated with poor outcomes, underscoring the regional relevance of immune exhaustion patterns. 24 The observed decline in CXCR3 with fibrosis is noteworthy, as CXCR3 is essential for Th1-cell trafficking and cytotoxic responses; its downregulation has been linked to fibrotic progression in other inflammatory disorders. 35 Beyond fibrosis, CXCR3 signaling has been implicated in tumor immune surveillance across solid and hematologic malignancies, where its loss may favor immune escape and clonal evolution.10,11 In our cohort, reduced CXCR3 on intermediate monocytes showed a strong association with leukemic transformation (AUC 0.876), consistent with emerging evidence that CXCR3 loss may impair immune surveillance and facilitate clonal escape in hematologic malignancies. 28 However, the low event rate (n = 8) necessitates that these findings be interpreted as preliminary and validated in larger, prospective studies. These results support the concept that immune surveillance wanes as marrow fibrosis advances, potentially facilitating clonal dominance. Moreover, the inverse relationship between PD-L1 and CXCR3 that we document suggests a switch from immune engagement to immune escape. Comparable findings have been reported in Western cohorts, where PD-L1 expression correlated with JAK2 mutational status and disease activity.14,36 By demonstrating these dynamics in an Egyptian MPN population, our study adds geographic and genetic context to the global picture and strengthens the rationale for considering checkpoint modulation in fibrotic MPNs, albeit in combination with antifibrotic or JAK-targeted therapies.
When assessed for their prognostic utility, immune checkpoint markers outperformed traditional systemic inflammation indices. When compared with conventional indices such as NLR, PLR, and SII, ROC curve analyses—adjusted for potential overfitting via LOOCV—demonstrated that PD-L1/CXCR3 ratios (LOOCV AUC 0.928) provided stronger and more robust discrimination for advanced fibrosis (Table 5). For predicting leukemic transformation, PD-L1 expression and checkpoint-derived ratios showed higher sensitivity and specificity than systemic indices (Table 6). In survival analysis, checkpoint parameters and composite ratios predicted overall survival and progression-free survival more effectively than SII or SIRI (Table 7). This was further validated by Kaplan-Meier and multivariable Cox regression analyses in the PMF cohort (Table 8, panel A), which established that a high PD-L1/CXCR3 ratio was an independent predictor of inferior overall survival (HR 2.15, 95% CI: 1.25-3.71) and event-free survival, while low CXCR3 expression on intermediate monocytes was predictive of inferior outcomes (HR 1.94, 95% CI: 1.16-3.25). Initially, in the PV/ET cohort, elevated SIRI and PD-L1/CXCR3 were associated with increased thrombotic risk (Table 8, panel B). Collectively, these results suggest that immune checkpoint biomarkers capture disease dynamics with greater accuracy than standard inflammatory markers across distinct MPN clinical contexts.
These findings highlight the limitations of systemic indices such as NLR and SII, which, although inexpensive and widely studied, often lack disease specificity. 37 Prior reports in Egyptian hematologic malignancies have confirmed that while NLR correlates with prognosis, its predictive value is modest compared with molecular and immune-based markers. 38 Our data extend this by showing that PD-L1/CXCR3 ratios, reflecting both immune suppression and impaired effector recruitment, provide superior stratification. International studies have demonstrated similar results: in myelofibrosis, PD-L1 upregulation has been associated with poor response to JAK inhibitors and reduced survival. 39 Moreover, checkpoint biomarkers integrate clonal and microenvironmental signals, offering a biologically grounded alternative to systemic inflammation scores. Importantly, our findings underscore the potential of checkpoint ratios as clinically relevant biomarkers, particularly in resource-constrained settings like Egypt, where costly molecular profiling is not always feasible. This supports the incorporation of immune profiling into prognostic assessment models for MPNs.
Using unsupervised clustering, patients were segregated into distinct immune phenotypes based on checkpoint expression and inflammatory biomarkers. Two major patient clusters emerged (Table 9). The first group was characterized by high PD-L1 expression and low CXCR3, correlating with advanced fibrosis and inferior survival. A second cluster showed balanced checkpoint expression with intermediate outcomes. The third group, enriched for CXCR3-positive subsets and lower PD-L1, exhibited the most favorable clinical course. Feature-ranking analysis by random forest confirmed PD-L1/CXCR3 ratios and CXCR3 on intermediate monocytes as the strongest contributors to cluster assignment (Figure 6). These data reveal that immune phenotyping can stratify Egyptian MPN patients into biologically and clinically meaningful categories, complementing standard scoring systems.
The identification of immune-based clusters reflects a shift from single-marker analysis toward integrated immunophenotyping. Similar clustering approaches have been used in solid tumors, where immune subtypes correlate with treatment responsiveness and survival. 40 In hematologic malignancies, unsupervised clustering of checkpoint molecules remains limited, but initial reports suggest that composite immune signatures outperform isolated markers in prognostic prediction. 41 Our study extends this approach to MPNs and demonstrates its applicability in an Egyptian population. Notably, the cluster with high PD-L1 and low CXCR3 expression aligns with the concept of an “exhausted immune phenotype,” consistent with prior reports in myelofibrosis, where immune suppression dominates. 42 Conversely, the cluster enriched for CXCR3-positive monocytes resembles an “immune-active” phenotype, which may retain antitumor potential. These findings emphasize the value of computational immune profiling in uncovering biologically coherent patient subsets and may provide a rationale for tailoring immunomodulatory therapies to specific immune landscapes.
Molecular and transcriptional profiling revealed important links between driver mutations and the immune landscape. Patients harboring JAK2 mutations demonstrated higher PD-L1 expression, while CALR mutations were associated with increased CXCR3 expression, suggesting differential immune modulation by specific genotypes (Table 10). Bulk RNA sequencing further highlighted enrichment of inflammatory and fibrotic pathways, including JAK-STAT and TGF-β signaling (Figure 7). Importantly, scRNA-seq provided finer resolution, revealing 3 transcriptionally distinct immune cell clusters (Table 11, Figure 8), a finding distinct from the 2 patient-level immune phenotypes identified by flow cytometry. Unlike flow cytometry, which quantified CXCR3 and checkpoint expression at the population level, scRNA-seq identified a CXCR3-low intermediate monocyte cluster enriched for IL6 and TGFB1 transcripts, defining a profibrotic and inflammatory program central to stromal remodeling. Pseudotime trajectory analysis suggested that CXCR3 downregulation occurs early, positioning this cluster as a potential initiating event in immune divergence rather than a late bystander. In contrast, CXCR3-high classical monocytes retained chemotactic activity, and PD-1–enriched T cells reflected immune exhaustion, underscoring the heterogeneity of immune responses in MPN.
This single-cell layer adds substantial value to the article because it demonstrates how transcriptional programming underlies the phenotypic changes observed by flow cytometry, thus bridging biomarker expression with disease biology. For readers, it highlights a mechanistic link between checkpoint activation, cytokine signaling, and fibrosis progression, moving beyond surface marker description to uncover the cellular origins of immune dysregulation. Clinically, these findings strengthen the rationale for targeting immune checkpoints and inflammatory cytokines in MPNs, and they illustrate the power of multiomics integration to refine risk stratification and therapeutic hypotheses.
These results underscore the intersection between molecular lesions and immune dysregulation in MPNs. JAK2 mutations have long been recognized as drivers of proliferative signaling, but emerging data also implicate them in immune checkpoint upregulation through STAT3/STAT5 activation. 43 Our finding that CALR mutations were linked to a more “immune-active” CXCR3-positive profile is intriguing, as CALR-driven disease has been associated with distinct immunogenic properties and responses to cytokines. 44 Transcriptomic enrichment of profibrotic and inflammatory pathways aligns with prior work demonstrating that chronic cytokine stimulation, particularly IL-6 and TGF-β, is linked to the development of marrow fibrosis. 45 Single-cell resolution adds novel insights by revealing heterogeneity within monocyte subsets, a feature not captured by bulk assays. Similar approaches in myelofibrosis have identified inflammatory monocyte clusters with profibrotic signatures. 46 Our data extend these observations to an Egyptian population, highlighting the value of integrated genomic-immune profiling in understanding disease biology and identifying therapeutic targets.
To refine prognostic assessment, we applied LASSO regression incorporating immune checkpoint biomarkers, clinical parameters, and DIPSS scores. The resulting model identified PD-L1/CXCR3 ratios, CXCR3 expression on intermediate monocytes, patient age, and baseline DIPSS as the most informative predictors. The composite biomarker-enhanced score demonstrated superior performance in internal validation compared with DIPSS and MIPSS70 alone, achieving higher concordance indices for overall and progression-free survival (Table 12). Furthermore, it showed promising generalizability in a small, independent external cohort. Kaplan-Meier curves illustrated a clear separation of risk categories, with patients in the high-risk biomarker group experiencing significantly shorter survival (Figure 9). These findings collectively suggest that checkpoint-derived markers could add prognostic precision beyond existing clinical models and represent compelling candidates for biomarker-driven therapy, which should be tested in future interventional studies.
Integrating immune biomarkers into prognostic tools represents an important step toward personalized management of MPNs. Current risk scores, such as DIPSS and MIPSS70, rely primarily on clinical and genetic variables but lack direct incorporation of immune dysfunction. 47 Our findings show that checkpoint profiles, particularly PD-L1/CXCR3 ratios, can meaningfully stratify outcomes when added to traditional models. Clinically, this could be implemented using a tiered workflow: at diagnosis, (1) perform flow cytometry for PD-L1/CXCR3 and CXCR3 on intermediate monocytes alongside routine labs; (2) integrate these results with DIPSS using our LASSO-derived formula to compute a biomarker-enhanced risk score; (3) use the score to guide early intervention (eg, JAK inhibitor initiation, clinical trial enrollment) in high-risk patients; and (4) monitor biomarker trends during therapy to assess response. Similar approaches have been proposed in other hematologic malignancies, where immune-based biomarkers improve the prediction of survival and therapeutic response. Importantly, flow cytometry is widely available and cost-effective, making this strategy feasible even in resource-limited settings like Egypt. 48 Importantly, our study is among the first to provide supporting evidence for a role for such a model in an Egyptian population, addressing regional gaps in prognostic research. The ability to use flow cytometry-based immune ratios, which are cost-effective and widely available, enhances the feasibility of implementing this model in resource-limited health care systems. By embedding immune biology into risk stratification, we not only refine prognostic accuracy but also provide a rationale for therapeutic decisions, such as prioritizing patients for immunomodulatory or antifibrotic interventions. 49
The PROs demonstrated strong associations between symptom burden and immune checkpoint activation. Fatigue, pruritus, and constitutional symptoms were more severe in patients with high PD-L1/CXCR3 ratios, underscoring the symptomatic relevance of immune dysregulation (Table 13). Integrated biomarker and mutational profiling suggested that immune checkpoint inhibitors, JAK-STAT pathway modulators, and epigenetic regulators could be rationally matched to patient subgroups (Table 14). Longitudinal follow-up of patients receiving JAK inhibitors revealed dynamic biomarker modulation: PD-L1 and SIRI decreased during therapy, whereas CXCR3 expression on intermediate monocytes increased in patients with clinical improvement (Table 15). These findings highlight the translational potential of immune profiling for monitoring disease activity and for stratifying patients in future trials of tailored therapeutic strategies.
The alignment between symptom burden and immune checkpoint activation reinforces the clinical relevance of immunologic pathways in MPNs. Prior reports have shown that inflammatory cytokines contribute directly to fatigue and pruritus, 2 hallmark symptoms of myelofibrosis. 50 Our data extend this by linking specific immune checkpoint profiles to PROs in an Egyptian cohort. The in silico drug-gene analysis underscores the therapeutic promise of combining JAK inhibitors with immune modulators. Early clinical studies with PD-1 inhibitors in MPNs demonstrated safety but limited efficacy, suggesting that rational combination approaches may be required. 51 Epigenetic agents such as BET inhibitors, including pelabresib, are also emerging as promising partners by targeting inflammatory transcriptional programs. 52 The longitudinal biomarker dynamics we observed during JAK inhibitor therapy provide real-world evidence that immune signatures are not static but modifiable. This supports their potential role as surrogate markers of treatment response, as previously suggested in smaller cohorts. 53 Together, these findings open avenues for integrating immune checkpoint profiling into both prognostic assessment and therapeutic monitoring, particularly in regions like Egypt, where novel therapies are being increasingly introduced.
CXCR3 expression on intermediate monocytes, PD-L1/CXCR3 ratios, and systemic inflammatory indices (SII and SIRI) defined biologically distinct immune phenotypes that tracked with fibrosis, symptom burden, and survival (Tables 8, 13 to 15). A biomarker-enhanced prognostic score integrating PD-L1/CXCR3, CXCR3/intermediate, age, and DIPSS achieved superior predictive accuracy compared with DIPSS, DIPSS-Plus, and MIPSS70 (AUC 0.85 vs 0.71-0.78) and demonstrated external validation (AUC 0.82). These results support the incorporation of immune biomarkers into risk stratification frameworks, particularly in Egyptian patients who present at younger ages with aggressive features. Longitudinal observations suggest that these immune signatures are modifiable during JAK inhibitor therapy, but randomized studies are required to confirm causality and to establish whether combining JAK inhibition with immune or epigenetic modulation can translate into improved patient outcomes.
To our knowledge, this study is among the first to integrate PD-1/PD-L1-CXCR3 coexpression with systemic inflammatory indices and multiomics profiling in an Egyptian MPN cohort, addressing a gap in regionally tailored prognostic tools. While prior studies in Western populations have highlighted the role of immune checkpoint dysregulation in myelofibrosis and related neoplasms, 54 no data have previously defined how these pathways operate in Middle Eastern or North African cohorts. Our findings reveal distinct immune-inflammatory signatures in Egyptian patients, likely reflecting the influence of genetic background, environmental exposures, and health care disparities. 55
The identification of CXCR3-low intermediate monocytes as predictors of leukemic transformation (AUC 0.876) and the validation of PD-L1/CXCR3 ratios as markers of advanced fibrosis (AUC 0.975) are novel contributions, supported by complementary single-cell and functional analyses. The biomarker-enhanced prognostic score, externally tested in an independent Egyptian cohort (AUC 0.82), further underscores its clinical applicability in a setting where access to advanced molecular diagnostics such as NGS is often limited. Finally, by linking immune checkpoint profiles to PROs, our study introduces a patient-centered dimension that has been largely neglected in MPN research. Together, these advances position our work as a unique contribution to precision medicine in MPNs, with global scientific significance and immediate relevance for underrepresented populations.
Despite its strengths, this study has limitations that should be acknowledged. The sample size of 90 MPN patients, with only 20 included in the scRNA-seq cohort, may limit generalizability, especially for less common subtypes such as PV, which represented just over 11% of cases. Furthermore, the analysis of fibrosis severity was necessarily focused on grades 2-4 due to the single grade 0-1 PMF patient in our cohort; future studies with larger prefibrotic PMF cohorts are needed to define biomarker dynamics in the earliest disease stages. The external validation cohort of 30 patients provided supportive evidence for the prognostic score but remains modest, emphasizing the need for larger, multicenter studies across diverse Egyptian populations to confirm robustness. The functional validation of CXCR3-low intermediate monocytes was performed in vitro using coculture systems and lacks in vivo confirmation. Future studies using animal models or clinical trials will be essential to establish their causal role in driving fibrosis. Similarly, while longitudinal analyses showed biomarker responsiveness to JAK inhibitors, longer follow-up and evaluation under other therapeutic strategies, such as anti-PD-L1 antibodies or epigenetic agents, are needed to assess durability and broader applicability. Another limitation targeted nature of our cytokine analysis, which focused on IL-6 and TGF-β as representative mediators. Expanding into broader proteomic, epigenomic, and metabolomic analyses would provide deeper mechanistic insights into immune and stromal interactions. Furthermore, while our scRNA-seq and in vitro functional assays propose a mechanistic role for CXCR3-low monocytes in fibrosis and inflammation, definitive proof of causality remains to be established through in vivo models or clinical intervention studies targeting this axis. Finally, integration with public datasets for cross-population validation and the design of biomarker-guided interventional trials in Egyptian patients should be prioritized. Addressing these gaps will strengthen the clinical and scientific impact of our proposed biomarkers and prognostic model, advancing their potential for translation into routine practice.
Beyond refining prognostication, our study demonstrates how immune checkpoint-chemokine interactions may directly shape both disease progression and symptom burden, providing a mechanistic basis for biomarker-guided therapy. This dual clinical and biological significance strengthens the translational impact of our findings and positions them for validation in prospective multicenter trials. It is important to note that while several key analyses—particularly those concerning fibrosis progression, leukemic transformation, and survival—were focused on the PMF subgroup, the identified immune checkpoint and CXCR3 dysregulation was observed across all MPN subtypes. The gradient of biomarker abnormality (most pronounced in PMF, intermediate in ET, and mildest in PV) mirrors the known clinical aggressiveness of these diseases. This suggests that the immune phenotypes we describe have broad relevance across the MPN spectrum, though their prognostic cut-offs and therapeutic implications may be subtype-specific. Future studies should validate these biomarkers in larger, subtype-stratified cohorts.
From a translational perspective, integrating PD-L1/CXCR3 and CXCR3-intermediate monocyte biomarkers into practice could follow a clear pathway: prospective validation in multicenter cohorts to standardize cut-offs; development of a streamlined flow-cytometry panel for routine clinical use; and incorporation into dynamic risk models that update with treatment response. This approach would shift MPN risk stratification from a static, genetics-centered model toward a dynamic, immune-informed framework, enabling earlier identification of high-risk patients and personalized surveillance and therapeutic strategies.
Conclusion
Promising immune biomarkers associated with marrow fibrosis, leukemic transformation risk, and inferior survival in Egyptian Philadelphia-negative MPNs include PD-1/PD-L1-CXCR3 coexpression and reduced CXCR3 on intermediate monocytes. Integrating these biomarkers with established clinical scores improved prognostic discrimination and showed promising initial performance in an external cohort; prospective multicenter validation is required to confirm their clinical utility and to evaluate their role as treatment-response markers. Longitudinal associations with JAK2/ASXL1 genotype and modulation during JAK inhibitor therapy highlight biological plausibility and support further mechanistic and interventional studies.
Footnotes
Acknowledgements
None
Data Availability Statement is included at the end of the article.
Ethics Approval
This study was approved by the Medical Research Ethics Committee of Kafr Elsheikh University (Reference No. KFSIRB200-234) on 29 April 2020. The study was conducted in accordance with the ethical principles of the Declaration of Helsinki (2024).
Consent to Participate
All participants received a clear explanation of study aims, procedures, risks, and expected benefits, and provided written informed consent.
Consent for Publication
All authors agree to the terms of the publication consent form.
Author Contributions
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets generated and/or analyzed during the current study are not publicly available due to patient privacy and ethical restrictions, but are available from the corresponding author on reasonable request.
Use of Artificial Intelligence Statement
Generative artificial intelligence (AI) was not used to create or modify any scientific content, data, or conclusions.
