Abstract
Background:
Triple-negative breast cancer (TNBC) lacks effective targeted therapies, underscoring the need for novel molecular targets. Gonadotropin-releasing hormone receptor (GnRHR) has been shown to suppress TNBC proliferation and metastasis. G protein–coupled receptor 173 (GPR173), known to regulate GnRHR in neuroendocrine cells, has an undefined role in TNBC. This study aimed to determine whether GPR173 modulates TNBC progression through GnRHR-mediated signaling.
Methods:
GPR173 and GnRHR expression levels were analyzed in TNBC tissues and correlated with patient prognosis. In vitro, TNBC cell lines were modified to knock down or overexpress GPR173 and GnRHR. Cell proliferation, migration, invasion, and expression of dual specificity phosphatase 1 (DUSP1), phosphorylated/total protein kinase B (AKT), phosphorylated/total extracellular signal–regulated kinase (ERK), and matrix metallopeptidase 2 (MMP2) were evaluated.
Results:
GPR173 and GnRHR expression was significantly reduced in TNBC tumors compared to normal breast tissues. Low expression of either protein correlated with poorer overall survival and increased lymph node metastasis. In vitro, GPR173 knockdown promoted TNBC cell proliferation, migration, and invasion, and reduced GnRHR expression. These changes were accompanied by increased phosphorylation of AKT and ERK, and elevated MMP2 expression. Notably, the pro-proliferative, pro-migratory, and pro-invasive effects of GPR173 knockdown were reversed by rescue overexpression of GnRHR. This GnRHR overexpression was accompanied by upregulation of DUSP1, dephosphorylation of AKT and ERK, and decreased MMP2 levels.
Conclusions:
Based on these in vitro data, GPR173 likely constrains the pro-proliferative, pro-migratory, and pro-invasive phenotypes of TNBC cells by enhancing GnRHR signaling. These findings highlight GnRHR and GPR173 as potential therapeutic targets for TNBC.
Keywords
Introduction
Breast cancer is the most common malignant tumor among women globally, with more than 2.3 million new cases reported in 2022. 1 Triple-negative breast cancer (TNBC), a particularly aggressive subtype, is defined by its lack of estrogen receptor, progesterone receptor, and human epidermal growth factor receptor-2 expression. Patients with TNBC typically face a higher risk of recurrence and metastasis, with limited targeted treatment options available compared with other breast cancer subtypes.2,3 Consequently, there is an urgent need to identify new therapeutic targets and elucidate the underlying molecular mechanisms that drive TNBC progression.
The gonadotropin-releasing hormone receptor (GnRHR) is a G protein–coupled receptor (GPCR) that is predominantly expressed in the pituitary gland, where it plays a central role in reproductive function. However, its expression has also been documented in a variety of nonpituitary tissues and cancer cells, including those of the breast, ovary, and prostate. 4 In these contexts, the activation of GnRHR has been shown to inhibit cancer cell proliferation and metastasis, often through autocrine or paracrine signaling loops. 5 Our previous work has specifically demonstrated that GnRHR can suppress proliferation and metastasis in TNBC. 6 Furthermore, a number of preclinical and clinical studies have highlighted the potential of targeting the GnRHR pathway as a viable therapeutic strategy for TNBC.7,8
Another GPCR, G protein–coupled receptor 173 (GPR173), is a member of a receptor subfamily primarily expressed in the central nervous system. GPR173 has been implicated in a diverse range of physiological processes, including reproduction, mood regulation, and appetite control.9-11 Crucially, an established regulatory relationship exists between these 2 proteins: in vitro studies have demonstrated that GPR173 plays a key role in upregulating GnRHR expression in pituitary cells. 12 Recent studies have also begun to uncover its role in cancer, with evidence suggesting it can inhibit cell proliferation in glioblastoma. 13 Despite this, the function of GPR173 and its interplay with GnRHR in breast cancer remain largely unexplored. This leaves a significant gap in our understanding of how this receptor might influence tumor biology.
Therefore, this study aimed to investigate the precise role of GPR173 in TNBC. Specifically, we sought to systematically evaluate the expression characteristics and prognostic significance of GPR173 and GnRHR in TNBC tissues, and to explore the impact of the GPR173–GnRHR signaling axis on tumor cell proliferation and metastatic potential and its molecular mechanisms, thereby providing new potential approaches for targeted therapy of TNBC.
Methods
Bioinformatics
Gene mRNA expression data were sourced from the University of ALabama at Birmingham CANcer data analysis portal (UALCAN) (https://ualcan.path.uab.edu/). 14 Survival data were sourced from the Kaplan-Meier plotter website (https://kmplot.com/analysis/). 15 Gene expression data for breast cancer cell lines were retrieved from the cancer cell line encyclopedia (CCLE) (https://sites.broadinstitute.org/ccle/), specifically including the TNBC cell lines BT-20, BT-483, MDA-MB-231, and MDA-MB-468.16,17
Tissue microarray immunofluorescence
A human TNBC tissue microarray (product name: Triple-Negative Breast Cancer with Survival Data; Catalog No. BRC1601), containing 160 samples (Supplemental Table S1), was purchased from Guilin Fanpu Biotech (Guilin, Guangxi, China). A specific lot/batch number for this product was not available. Demographic information including age and sex for the de-identified samples was provided by the supplier (Supplemental Table S1). Ethnicity data were not provided by the supplier. Deparaffinization, rehydration, microwave-based antigen retrieval (EDTA, pH 8.0), and blocking were performed using standard protocols. Slides were incubated overnight at 4°C with primary antibodies against GPR173 and GnRHR (Immunoway, YT1998 and YT1941), followed by HRP-conjugated secondary antibodies and tyramide signal amplification (CY3/FITC). DAPI was used for nuclear counterstaining, and slides were imaged under a fluorescence microscope. Patients were stratified into “low” and “high” expression groups based on the median optical density of immunofluorescence staining for GPR173 or GnRHR in tumor tissues as the cut-off value. Of the 80 patient cases included in the tissue microarray, a total of 42 were excluded from analysis due to technical failures during staining or imaging (eg, weak fluorescence signals, high background, or tissue detachment). Importantly, each patient contributed 2 paired tissue samples: tumor and adjacent normal tissue. For paired analysis, both samples needed to exhibit interpretable fluorescence signals. If either the tumor or the corresponding normal tissue failed quality control, the entire case was excluded from the final dataset. This exclusion criterion was not pre-established but was essential to ensure valid and statistically comparable paired analyses.
Cell culture
The human TNBC cell line MDA-MB-231 was obtained from Wuhan Pricella Biotechnology Co., Ltd. (Catalog No. CL-0150; RRID: CVCL_0062). This cell line was originally derived from the pleural effusion of a patient diagnosed with metastatic breast adenocarcinoma. According to the manufacturer, the identity of the cell line was authenticated through short tandem repeat profiling, and the cells were confirmed to be free of mycoplasma contamination. The culture medium was MEM (Wuhan Pricella Biotechnology Co., Ltd., PM150410P) supplemented with 10% fetal bovine serum (Gibco) and 1% penicillin-streptomycin (Wuhan Pricella Biotechnology Co., Ltd., PB180120). Cells were cultured at 37°C with 5% CO2 in a humidified environment.
Transfection
Both the GPR173 siRNA and the GnRHR-overexpression plasmid constructs were designed and purchased from General Biologicals (Chuzhou, Anhui, China). The siRNA sequences targeting GPR173 were 5′-GGACUGAUUAUGUGCGUGATT-3′ (sense) and 5′-UCACGCACAUAAUCAGUCCTT-3′ (antisense). The coding sequence for human GnRHR (GenBank Accession No. NM_000406.2) was cloned into the pCDNA3.1 mammalian expression vector. The integrity and correct sequence of the inserted GnRHR cDNA were confirmed by supplier through PCR amplification and Sanger sequencing. For GPR173 knockdown experiments, a nontargeting scrambled siRNA sequence (same supplier) was used as a negative control (NC group). An empty pCDNA3.1 vector (same supplier), lacking the GnRHR insert, served as the negative control for overexpression experiments. In experiments involving combined GPR173 knockdown and GnRHR overexpression (si-GPR173 + oe-GnRHR), the control group was co-transfected with both the scrambled nontargeting siRNA and the empty vector plasmid. A total of 2 × 105 cells per well were seeded in a 6-well plate and cultured overnight at 37°C. Two 1.5-mL centrifuge tubes were prepared. In one tube, 250 μL of Opti-MEM and 5 μL of lipofectamine 3000 transfection reagent (Invitrogen, L3000-015) were added, whereas in the other tube, 250 μL of Opti-MEM and 2 μL of P3000 reagent were added. The final siRNA concentration was 50 nM, and the plasmid amount per well was 1 μg. The contents of the 2 tubes were mixed thoroughly and left to stand at room temperature for 15 minutes. The mixture was then added to each well of the plate, which was incubated for 72 hours to continue culture.
Cell counting kit-8 assay
Cells were seeded in a 96-well plate at a density of 1 × 104 cells per well, with 3 replicates per group. Next, 10 μL of cell counting kit-8 (CCK-8) solution (Beyotime, C0038) was added to each well. The samples were incubated for 3 hours, after which absorbance was measured at 450 nm with a microplate reader to produce the growth curve. All experiments were independently repeated 3 times.
Colony formation assay
Cells were seeded in 12-well plates at a density of 400 cells per well and cultured in an incubator for 2 weeks. Then, 4% paraformaldehyde (Sinopharm Chemical Reagent Co., Ltd.) was added to fix the cells for 15 minutes. After discarding the paraformaldehyde, 1 mL of crystal violet staining solution (Genview, DC079-25G) was added to stain the cells for 15 minutes. The cells were dried, photographed, and counted. The experiment was independently performed 3 times.
Wound healing assay
Cells were seeded in 12-well plates at a density of 3 × 105 cells per well, with 2 replicate wells per group. When the cell density reached confluence, a micropipette tip was used to make a scratch along the central axis of each well. Detached cells were removed with phosphate-buffered saline washing, and fresh culture medium was added. At 0, 24, and 48 hours, the 12-well plate was imaged at 3 distinct positions per well to observe cell migration distances. Migration rate at N h = (scratch width at 0 h—scratch width at N h) / scratch width at 0 h. Each experiment was independently repeated 3 times.
Transwell
A total of 1 × 105 cells were seeded into Matrigel-coated invasion chambers. Next, 600 µL of medium containing 10% fetal bovine serum was added to the lower chamber. The chamber was incubated for 48 hours. Noninvading cells on the upper membrane surface were removed with a cotton swab, fixed in paraformaldehyde (Sinopharm Chemical Reagent Co., Ltd.), and stained with crystal violet solution (Genview, DC079-25G). Images were captured, and cells were counted. The assay was performed in 3 independent biological replicates.
Western blotting
Cells were lysed on ice using lysis buffer containing protease and phosphatase inhibitors. Total protein concentration was determined using a Bradford assay (Beyotime, P0006). Equal amounts of protein were mixed with 5× loading buffer, denatured at 95 °C for 10 minutes, separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis, and transferred to polyvinylidene fluoride membranes. After blocking, membranes were incubated overnight at 4 °C with primary antibodies and then with HRP-conjugated secondary antibodies (Supplemental Table S2) for 1 hour at room temperature. Protein bands were visualized using an ECL kit (PROTEINTECH, PK10001) and quantified using ImageJ software. Phosphorylated proteins (eg, p-AKT, p-ERK) were normalized to their total protein levels. All protein levels, including MMP2 and total proteins, were further normalized to GAPDH. Western blotting (WB) experiments were independently repeated 3 times to ensure reproducibility.
Statistical analysis
Data were analyzed using SPSS 22.0 statistical software. Measurement data following a normal distribution were analyzed with an independent 2-tailed t-test or paired samples t-test. Statistical significance was set at a P value of less than .05. The Benjamini-Hochberg procedure was applied to adjust P values from all hypothesis tests, yielding a corrected α of .042.
Reporting standards
This study was conducted in accordance with the ethical and reporting recommendations set forth by the International Committee of Medical Journal Editors (ICMJE). The Materials Design Analysis Reporting (MDAR) checklist was completed and submitted alongside this manuscript to ensure transparency and reproducibility in the experimental design and data reporting. 18 No additional reporting guidelines, such as CONSORT, PRISMA, or ARRIVE, were applicable to this study.
Results
GPR173 and GnRHR expression levels were reduced in TNBC compared with normal breast tissue, with lower levels associated with poorer prognosis
The expression levels of GPR173 and GnRHR were obtained from the UALCAN database, comparing 116 TNBC tumor samples with 114 normal breast tissue samples. Results indicated that GPR173 and GnRHR expression levels were significantly lower in TNBC than in normal breast tissues (Figure 1A). The association between GPR173 and GnRHR expression levels and overall survival (OS) among patients with TNBC was analyzed using the Kaplan-Meier plotter database. Results indicated that patients with low expression of GPR173 or GnRHR had poorer OS (Figure 1B).

The expression of GPR173 and GnRHR mRNA is reduced in TNBC tissues and correlates with poorer overall survival. (A) Comparative expression levels of GPR173 (left panel) and GnRHR (right panel) mRNA in TNBC tissues (n = 116) versus normal breast tissues (n = 114). Data were obtained from the TCGA breast cancer dataset and analyzed using the UALCAN portal (http://ualcan.path.uab.edu/). Expression levels are presented as TPM. Box plots illustrate the median (indicated by a line within the box), the interquartile range, and whiskers extending to 1.5 times the interquartile range. P values indicating statistical significance between groups were obtained directly from the UALCAN portal; however, the specific statistical tests employed for these comparisons were not explicitly stated in the output. (B) Kaplan-Meier OS curves for TNBC patients stratified by GPR173 (left panel) and GnRHR (right panel) mRNA expression levels. Analyses were conducted using the Kaplan-Meier Plotter database (https://kmplot.com/analysis/). Patients were categorized into high- and low-expression groups utilizing the “auto select best cutoff” (percentile) feature of the platform. For GPR173, the cut-off value applied in the analysis was −0.31 (expression range of probe: −3 to 3); low expression (n = 62) was associated with poorer OS compared with high expression (n = 64) (HR = 0.28, 95% CI: 0.11-0.69; log-rank P = 0.0034). For GnRHR, the cut-off value used in the analysis was −0.7 (expression range of probe: −3 to 1); low expression (n = 35) was linked to poorer OS compared with high expression (n = 91) (HR = 0.42, 95% CI: 0.19-0.94; log-rank P = 0.029). CI, confidence interval; GnRHR, gonadotropin-releasing hormone receptor; GPR173, G protein–coupled receptor 173; HR, hazard ratio; OS, overall survival; TPM, transcripts per million; TCGA, the cancer genome Atlas; TNBC, triple-negative breast cancer.
Low expression of GPR173 and GnRHR was associated with lymph node metastasis
To investigate correlations of GPR173 and GnRHR expression with the clinicopathological characteristics of TNBC, we used immunofluorescence to examine a TNBC tissue microarray composed of paraffin-embedded cancer tissues and their corresponding adjacent normal tissues from 80 patients with TNBC. However, due to technical issues, only 38 patients were included in the analysis (Figure 2A; Tables 1 and 2). Samples were excluded not only for technical reasons, such as signal loss or tissue detachment, but also due to the necessity of having complete tumor-normal pairs. Cases that lacked valid immunofluorescence data from either the tumor or adjacent tissue were not included in the paired analysis. Protein expression levels of both GPR173 and GnRHR were lower in cancer tissues than in the normal tissues (Figure 2B). Based on the expression levels of GPR173 and GnRHR, patients were stratified into high- and low-expression groups for each protein. Both GPR173 and GnRHR were associated with lymph node status, with a higher incidence of N3 observed in the GPR173/GnRHR low-expression group compared to the high-expression group (Table 3), suggesting potential inhibitory effects on lymph node metastasis. GPR173 and GnRHR expressions were not correlated with age, histological grade, or T stage (Table 3).

Reduced protein expression of GPR173 and GnRHR in human TNBC tissues compared to paired adjacent normal breast tissues, as observed in a tissue microarray. (A) Representative immunofluorescence images from the TNBC tissue microarray. Each spot on the microarray corresponds to an individual specimen, either TNBC or normal tissue, with specific locations detailed in Supplemental Table S1. The GPR173 protein is depicted in red (CY3), the GnRHR protein in green (FITC), and the nuclei are counterstained with DAPI (blue). Scale bar = 5000 µm. (B) Quantitative analysis of GPR173 and GnRHR protein expression levels, measured as OD from the tissue microarray. Due to technical issues during processing, complete paired data for OD measurements were available for 38 TNBC tissues and their corresponding adjacent normal tissues. The data are presented as mean ± SD. Statistical significance was determined using a paired t-test. For GPR173, P = .005; for GnRHR, P < .001 when comparing TNBC tissues to normal tissues. DAPI, 4′,6-diamidino-2-phenylindole; GnRHR, gonadotropin-releasing hormone receptor; GPR173, G protein–coupled receptor 173; OD, optical density; SD, standard deviation; TNBC, triple-negative breast cancer.
Distribution of clinical and pathological characteristics of patients with TNBC.

Abbreviation: TNBC, triple-negative breast cancer.
Optical density of GPR173 and GnRHR in normal and TNBC tumor tissues.

Abbreviations: GPR173, G protein–coupled receptor 173; GnRHR, gonadotropin-releasing hormone receptor; TNBC, triple-negative breast cancer.
The relationship between GPR173/GnRHR and clinical pathological characteristics of TNBC.

Abbreviations: GnRHR, gonadotropin-releasing hormone receptor; GPR173, G protein–coupled receptor 173; TNBC, triple-negative breast cancer.
Patients were stratified into “low” and “high” expression groups based on the median optical density of immunofluorescence staining for GPR173 or GnRHR in tumor tissues as the cut-off value.
Bold values indicate statistical significance (P ≤ 0.05).
Expression of GPR173 and GnRHR in TNBC cell lines
The expression levels of GPR173 and GnRHR in the TNBC cell lines BT-20, BT-483, MDA-MB-231, and MDA-MB-468 were obtained from the CCLE website. The results indicated that both genes exhibited higher expression in BT-483 cells and lower expression in BT-20 cells, with intermediate expression levels observed in MDA-MB-231 cells (Figure 3).

Basal mRNA expression levels of GPR173 and GnRHR in various TNBC cell lines. (A) mRNA expression levels of GPR173 in the TNBC cell lines BT-20, BT-483, MDA-MB-231, and MDA-MB-468. (B) mRNA expression levels of GnRHR in the TNBC cell lines BT-20, BT-483, MDA-MB-231, and MDA-MB-468. CCLE, cancer cell line encyclopedia; GnRHR, gonadotropin-releasing hormone receptor; GPR173, G protein–coupled receptor 173; TNBC, triple-negative breast cancer; TPM, transcripts per million.
GPR173 inhibition enhances proliferative, migratory, and invasive phenotypes in TNBC cells
MDA-MB-231 cells were divided into 2 groups in vitro: (1) NC group, transfected with a nonspecific RNA sequence, and (2) si-GPR173 group, transfected with GPR173 siRNA. Western blotting analysis confirmed that GPR173 expression was reduced in the si-GPR173 group, indicating effective interference by GPR173 siRNA (Figure 4A). CCK-8 assays demonstrated increased cell viability following GPR173 interference (Figure 4B). Colony formation assays showed that cell clone numbers increased after GPR173 knockdown (Figure 4C). Scratch assay results indicated enhanced cell migration following GPR173 interference (Figure 4D), whereas Transwell assays revealed an increase in cell invasion capability (Figure 4E).

GPR173 inhibition enhances proliferative, migratory, and invasive phenotypes in TNBC cells. (A) GPR173 protein was reduced in si-GPR173 versus NC by WB. Right: densitometric quantification of GPR173 normalized to GAPDH (GPR173/GAPDH, a.u.). (B) Cell viability was significantly increased following the knockdown of GPR173, as assessed by the CCK-8 assay at 0, 24, 48, and 72 hours post-transfection. (C) The colony formation assay demonstrated that the number of colonies was significantly higher in the si-GPR173 group after 2 weeks of culture. (D) Wound healing assay demonstrating enhanced migration in the si-GPR173 group. Representative images at 0, 24, and 48 hours after scratching are shown. (E) Transwell invasion assay demonstrating increased invasion following GPR173 knockdown. Cells were seeded in Matrigel-coated chambers and fixed/stained after 48 hours; representative micrographs are shown. a.u., arbitrary units; CCK-8, Cell Counting Kit-8; GPR173, G protein–coupled receptor 173; SD, standard deviation; TNBC, triple-negative breast cancer; WB, Western blotting.
Overexpression of GnRHR mitigates pro-proliferative, pro-migratory, and pro-invasive effects of GPR173 inhibition
Cells were divided into 4 in vitro experimental groups: (1) NC group, transfected with a nonspecific RNA sequence and an empty plasmid; (2) si-GPR173 group, transfected with GPR173 siRNA and an empty plasmid; (3) oe-GnRHR group, transfected with a nonspecific RNA sequence and a GnRHR plasmid; and (4) si-GPR173 + oe-GnRHR group, transfected with GPR173 siRNA and a GnRHR plasmid (Supplemental Table S3). Western blotting analysis showed that GPR173 interference led to decreased GnRHR expression, while GnRHR plasmids effectively induced GnRHR overexpression (Figure 5A). CCK-8 assay results indicated that GPR173 interference increased cell viability, whereas combined GPR173 interference and GnRHR overexpression significantly reduced cell viability (Figure 5B). Colony formation, scratch, and Transwell assays demonstrated that GnRHR overexpression reversed the effects of GPR173 interference on cell proliferation, migration, and invasion (Figure 5C to E).
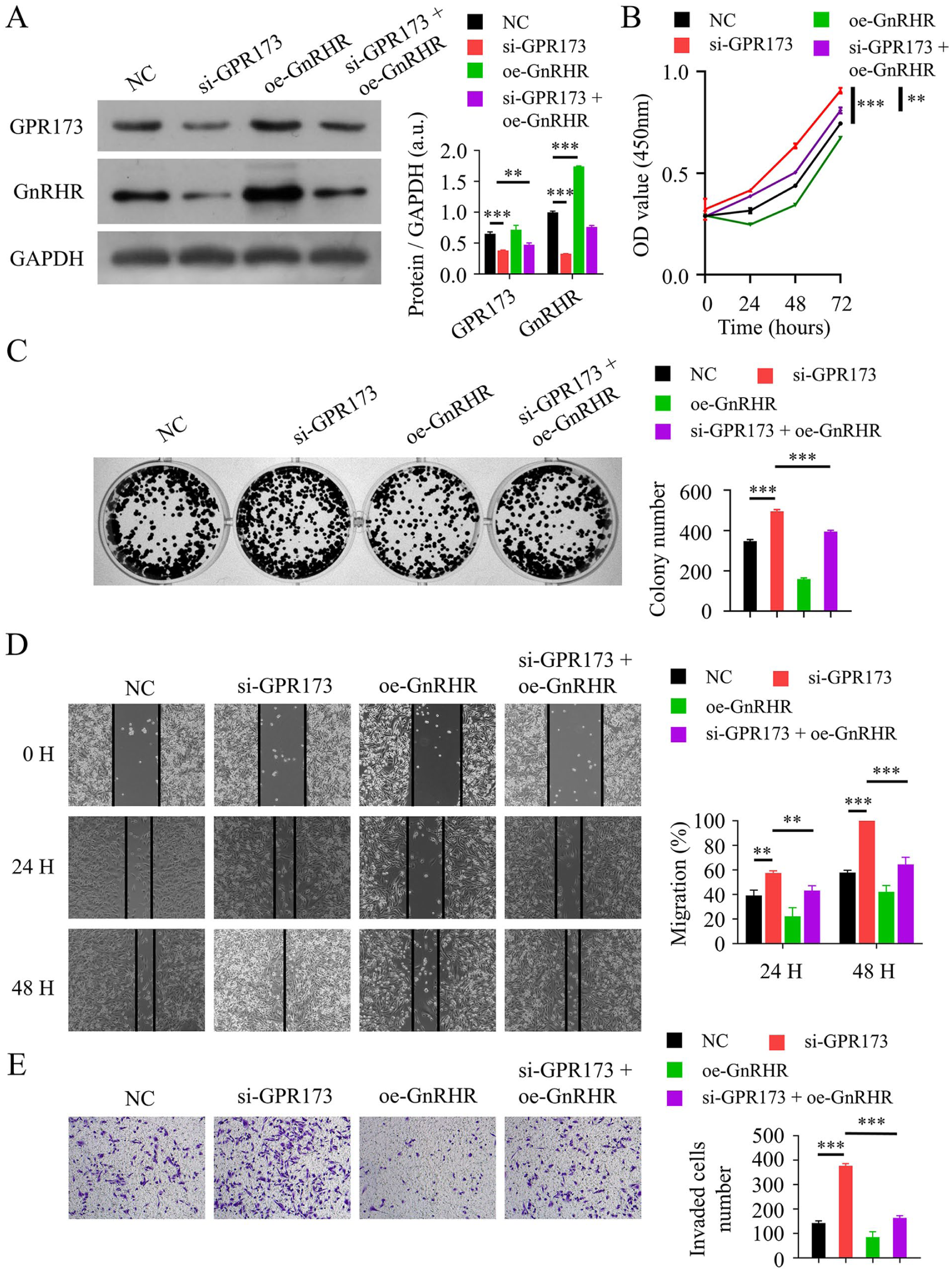
Overexpression of GnRHR mitigates pro-proliferative, pro-migratory, and pro-invasive phenotypes induced by GPR173 inhibition in TNBC cells. (A) Western blot shows reduced GPR173 after si-GPR173 and restoration of GnRHR by GnRHR plasmid in MDA-MB-231 cells. Right: densitometric quantification of protein levels normalized to GAPDH (Protein/GAPDH, a.u.); bars represent GPR173 or GnRHR as indicated in the legend. For statistical testing, group comparisons were performed separately within each protein. (B) CCK-8 assays at 0, 24, 48, and 72 hours indicate that the increased viability following GPR173 knockdown was attenuated by GnRHR overexpression. (C) Colony formation increased after GPR173 knockdown and was attenuated by GnRHR overexpression (colonies ⩾50 cells per colony were counted per well; 14 days). (D) Wound healing assays showed enhanced migration with si-GPR173, which was reduced by GnRHR overexpression (quantified as % wound closure at 24 and 48 hours). (E) Transwell invasion assays (Matrigel-coated inserts, 48 hours) demonstrated increased invasion with si-GPR173 and attenuation by GnRHR overexpression (cells counted across ⩾ 4 random fields per insert). a.u., arbitrary units; CCK-8, Cell Counting Kit-8; GnRHR, gonadotropin-releasing hormone receptor; GPR173, G protein–coupled receptor 173; SD, standard deviation; TNBC, triple-negative breast cancer; WB, Western blotting.
GnRHR overexpression is associated with increased DUSP1 expression and reduced p-AKT/p-ERK and MMP2
Cells were divided into 2 groups in vitro: (1) NC, transfected with an empty plasmid, and (2) oe-GnRHR, transfected with a GnRHR plasmid. RNA sequencing was used to identify differential mRNA expression profiles between the 2 groups (Figure 6A). The data indicated that DUSP1 transcripts were increased in the oe-GnRHR group (Figure 6B).

GnRHR upregulates DUSP1 (RNA-seq) and decreases p-AKT/AKT, p-ERK/ERK, and MMP2 in TNBC cells. (A) Venn diagram illustrating the overlap of differentially expressed mRNAs identified through RNA sequencing in MDA-MB-231 cells transfected with a GnRHR-overexpression plasmid (oe-GnRHR) compared with cells transfected with an empty vector (NC). (B) RNA sequencing indicated increased DUSP1 transcripts in oe-GnRHR compared with NC. (C) Western blot analysis of p-AKT, AKT, p-ERK, ERK, and MMP2 across NC, si-GPR173, oe-GnRHR, and si-GPR173 + oe-GnRHR. Right/below: densitometric quantification of p-AKT/AKT and p-ERK/ERK ratios, and MMP2 normalized to GAPDH (a.u.). AKT, protein kinase B; a.u., arbitrary units; DUSP1, dual specificity phosphatase 1; ERK, extracellular signal–regulated kinase; GnRHR, gonadotropin-releasing hormone receptor; MMP2, matrix metallopeptidase 2; NC, negative control; SD, standard deviation; WB, Western blotting.
In a 4-group design as described above, Western blotting was performed to assess p-AKT, AKT, p-ERK, ERK, and MMP2. Interference with GPR173 increased p-AKT, p-ERK, and MMP2 relative to NC, whereas GnRHR overexpression in the si-GPR173 background attenuated these changes (Figure 6C; Supplemental Table S4). These findings indicate that the phenotypic rescue by GnRHR is accompanied by DUSP1 upregulation and suppression of AKT/ERK phosphorylation and MMP2.
Discussion
Triple-negative breast cancer is an aggressive subtype of breast cancer with limited actionable targets. Chemotherapy remains the mainstay, yet responses are modest and toxicities are substantial. Identifying novel proteins involved in TNBC proliferation and metastasis would deepen our understanding of TNBC’s biological behavior and aid in discovering potential therapeutic targets. Our previous study showed that GnRHR inhibits TNBC cell proliferation and metastasis. 6 This study examines proteins in the upstream and downstream pathways of GnRHR.
In seeking to identify upstream regulators of GnRHR, we focused on GPR173, a putative upstream regulator. However, no studies have reported on GPR173 regulation in TNBC. Bioinformatics analysis showed lower GPR173 expression in TNBC than in normal breast tissue, and patients with higher GPR173 expression had better prognoses than those with lower expression. These findings suggest that GPR173 may play a role in inhibiting TNBC progression.
To select a suitable cell line for this study, we first examined GPR173 and GnRHR expression levels in various TNBC cell lines using the CCLE database. Expression levels of both genes were moderate in MDA-MB-231 cells, suggesting that these cells are appropriate for further research. We transfected MDA-MB-231 cells with siRNA targeting GPR173, and WB analysis confirmed decreased GPR173 expression, indicating successful GPR173 knockdown. Additionally, GnRHR expression decreased following GPR173 knockdown, consistent with a positive regulatory relationship between GPR173 and GnRHR in vitro. This finding aligns with previous studies in hypothalamic cells, where GPR173’s regulatory mechanism may involve activation of the cAMP-PKA pathway. 19 Inhibition of GPR173 expression also led to increased cell proliferation and metastatic potential, aligning with clinical observations that high GPR173 expression correlates with better patient prognosis.
Since GnRHR expression decreased following GPR173 interference, we conducted a rescue experiment by overexpressing GnRHR to investigate the regulatory relationship between GnRHR and GPR173. We found that GPR173 inhibition increased cell proliferation as well as migratory and invasive phenotypes, which were subsequently attenuated by GnRHR overexpression. These observations suggest that GnRHR may act downstream of GPR173 to partially counter the in vitro pro-proliferative, pro-migratory, and pro-invasive changes observed upon GPR173 inhibition. This mechanism has not been previously reported.
To investigate GnRHR downstream pathways, we performed RNA sequencing after GnRHR overexpression, which indicated that DUSP1 transcripts were increased in the GnRHR-overexpression group. Previous reports indicate that DUSP1 expression also increased upon GnRHR activation in cultured mouse pituitary cells,20,21 consistent with our findings. Prior studies have shown that DUSP1 can dephosphorylate and inactivate AKT and ERK and downregulate MMP2, thereby inhibiting cell proliferation and metastasis.22-24 Accordingly, we measured levels of p-AKT, AKT, p-ERK, ERK, and MMP2, and observed that GPR173 interference increased p-AKT/AKT, p-ERK/ERK, and MMP2 levels. GnRHR overexpression attenuated these changes. Given that GnRHR overexpression was also associated with DUSP1 upregulation, and DUSP1 is known to dephosphorylate AKT and ERK, it is plausible that GPR173 may modulate p-AKT, p-ERK, and MMP2 levels, at least in part, potentially via a GnRHR-DUSP1 axis; however, a causal role for DUSP1 in mediating these effects remains to be established. However, further research is required to definitively establish DUSP1 as a direct functional intermediary in this specific context. Kurowska et al found that PNX-14 inhibited MAP3/1 (ERK1/2) phosphorylation in vitro, consistent with our results. They also observed a transient increase in AKT phosphorylation at 5 and 15 minutes after PNX-14 treatment, followed by a subsequent decrease. In our study, we monitored AKT phosphorylation over a longer period, which produced consistent outcomes. 25
A limitation of this study is that all experiments were conducted in vitro using the MDA-MB-231 cell line only. Future studies are necessary to validate these findings across additional TNBC cell lines and in vivo models. The precise regulatory relationship between GPR173 and GnRHR also remains incompletely defined and warrants further investigation.
While DUSP1 upregulation accompanied the GnRHR-mediated rescue, a causal role for DUSP1 in mediating these effects remains to be established. Future studies will test this directly by combining GPR173 knockdown and GnRHR overexpression with DUSP1 silencing (si-DUSP1) and by validating DUSP1 changes at both mRNA and protein levels. In addition, in vivo models (eg, orthotopic implantation or tail-vein colonization assays) across multiple TNBC cell lines will be employed to evaluate metastatic dissemination and to substantiate the translational relevance of the GPR173-GnRHR-DUSP1 axis.
Conclusions
Our findings suggest that GPR173 may constrain the pro-proliferative, pro-migratory, and pro-invasive phenotypes of MDA-MB-231 TNBC cells by upregulating GnRHR. These in vitro effects were accompanied by reduced AKT/ERK phosphorylation and lower MMP2 levels. Further studies, including in vivo validation and targeted loss-of-function approaches, are required to establish causality and to evaluate the translational relevance of the GPR173-GnRHR axis.
Supplemental Material
sj-docx-1-onc-10.1177_11795549251380919 – Supplemental material for Mechanism of GPR173-Mediated Suppression of TNBC Proliferation and Metastatic Potential via GnRHR Upregulation
Supplemental material, sj-docx-1-onc-10.1177_11795549251380919 for Mechanism of GPR173-Mediated Suppression of TNBC Proliferation and Metastatic Potential via GnRHR Upregulation by Dan Xing, Caiping Chen, Chao Han, Li Xue and Xiang Lu in Clinical Medicine Insights: Oncology
Supplemental Material
sj-docx-2-onc-10.1177_11795549251380919 – Supplemental material for Mechanism of GPR173-Mediated Suppression of TNBC Proliferation and Metastatic Potential via GnRHR Upregulation
Supplemental material, sj-docx-2-onc-10.1177_11795549251380919 for Mechanism of GPR173-Mediated Suppression of TNBC Proliferation and Metastatic Potential via GnRHR Upregulation by Dan Xing, Caiping Chen, Chao Han, Li Xue and Xiang Lu in Clinical Medicine Insights: Oncology
Supplemental Material
sj-docx-3-onc-10.1177_11795549251380919 – Supplemental material for Mechanism of GPR173-Mediated Suppression of TNBC Proliferation and Metastatic Potential via GnRHR Upregulation
Supplemental material, sj-docx-3-onc-10.1177_11795549251380919 for Mechanism of GPR173-Mediated Suppression of TNBC Proliferation and Metastatic Potential via GnRHR Upregulation by Dan Xing, Caiping Chen, Chao Han, Li Xue and Xiang Lu in Clinical Medicine Insights: Oncology
Supplemental Material
sj-docx-4-onc-10.1177_11795549251380919 – Supplemental material for Mechanism of GPR173-Mediated Suppression of TNBC Proliferation and Metastatic Potential via GnRHR Upregulation
Supplemental material, sj-docx-4-onc-10.1177_11795549251380919 for Mechanism of GPR173-Mediated Suppression of TNBC Proliferation and Metastatic Potential via GnRHR Upregulation by Dan Xing, Caiping Chen, Chao Han, Li Xue and Xiang Lu in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
Not applicable.
Ethical Considerations and Consent to Participate
The human TNBC tissue microarray (Catalog No. BRC1601) used in this study was purchased from Guilin Fanpu Biotech (Guilin, Guangxi, China). Ethical approval for the collection and research use of these patient samples was obtained by the Guilin Fanpu Biotech Ethics Committee (Approval No. Fanpu [2018] No. 23; Approval Date: October 25, 2018). The supplier confirmed that all samples were collected with appropriate informed consent from the participants and fully anonymized prior to distribution. The use of these commercially obtained, anonymized samples in this study was additionally reviewed by the Medical Ethics Committee of the First Hospital of Jiaxing (Jiaxing, Zhejiang, China), which confirmed that no further institutional ethical approval was required, given the existing external ethical clearance and complete anonymization of patient data.
Consent for Publication
Not applicable. The study did not involve any individual person’s data in any form (including individual details, images, or videos). All human tissue samples were fully anonymized and commercially sourced with prior ethical approval and informed consent obtained by the supplier.
Author Contributions
All authors contributed to the study conception and design. Data curation was performed by Xiang Lu and Dan Xing. Funding acquisition was performed by Xiang Lu and Caiping Chen. Writing—original draft was performed by Dan Xing. Methodology was performed by Dan Xing and Chao Han. Investigation was performed by Li Xue. Validation was performed by Caiping Chen. Formal analysis was performed by Li Xue. Visualization was performed by Chao Han. Conceptualization, project administration, resources, software, writing—review & editing was performed by Xiang Lu. All authors reviewed the manuscript and approved the final submitted version.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The Jiaxing Key Discipline of Medicine (Mastropathy, Innovation Subject, 2023-FC-001); The Jiaxing Science and Technology Plan Project (2022AY30019); The Breast Cancer Precision Diagnosis and Treatment Center of the First Hospital of Jiaxing (2021-ZZZX-06); The Jiaxing Research Hospital Association Research Fund (2021JYHA003); The Jiaxing First Hospital hospital-level project (2020YJZD016).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availabilty statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
