Abstract
In China, esophageal cancer (EC) is one of the most prevalent malignant tumors of the digestive system. EC has a high incidence and mortality rate, of which esophageal squamous cell carcinoma (ESCC) accounts for more than 90%. Due to a lack of effective prevention and treatment methods, the 5 year survival rate is less than 30%. In recent years, microecology has become a hot spot in cancer research, and dysbiosis may play an important role in the etiology of EC. Presently, research on the relationship between the microbiome and ESCC remains in its early stages. This narrative review examines the relationship between the oral and esophageal microbiota and ESCC. A better understanding of this relationship may facilitate early detection and the optimization of treatment strategies.
Keywords
Introduction
Esophageal cancer (EC) is ranked seventh globally in cancer incidence and sixth in mortality in the Global Cancer Statistics 2020 report. 1 China has a high incidence of EC, accounting for about 50% of global cases and deaths, 2 with esophageal squamous cell carcinoma (ESCC) representing about 90% of cases. 3 Due to late-onset symptoms and a lack of accurate early diagnosis and screening methods, the majority of patients reach the middle or late stages of treatment, and overall survival rates range from 15% to 25% after 5 years of treatment. 4 Several risk factors contribute to the development of EC, including smoking, alcohol consumption, diet, environmental factors, genetic factors, and socioeconomic status. 5 However, the specific etiology remains unclear. Therefore, it is necessary to explore the mechanism underlying the occurrence and development of EC to find new methods of diagnosis and treatment.
Owing to the development of high-throughput DNA sequencing technology, scientists have gained a deeper understanding of the complexity of microorganisms. In humans, microorganisms typically coexist with their hosts and are crucial in digestion, immune function, and metabolism. 6 However, some microorganisms are also vital to the occurrence and development of cancer.7,8 Studies have shown that the human gut microbiota can influence the initiation and progression of gastrointestinal tumors by damaging DNA, activating oncogenic signaling pathways, generating tumor-promoting metabolites, and suppressing anti-tumor immune responses. 9 In recent years, epidemiological evidence has suggested that poor oral health, periodontal disease, and tooth loss are associated with EC risk,10-12 and this association may involve alterations in the diverse community of microbes present in the oral cavity. As the entrance to the digestive tract, the mouth is one of the most complex parts of the human body. 13 As the esophagus is adjacent to the oral cavity, it is generally believed that esophageal microorganisms originate from the oral cavity and stomach. Recently, colonizing bacteria have been found in the esophageal mucosa. 14
While numerous studies have confirmed the relationship between the intestinal microbiota and tumors, research on the oral and esophageal microbiota, especially in the context of ESCC, is limited. As a narrative review, this article systematically examines the current evidence regarding the potential links between ESCC and microbial dysbiosis in the oral cavity and esophagus. This work improves our understanding of the pathogenesis of ESCC and may inform the search for new therapeutic targets.
A systematic search of PubMed was conducted to identify relevant publications, with priority given to recent studies (2015-2025). The search strategy incorporated the following key terms: oral microbiome, esophageal microbiome, salivary/esophageal microbiota, Porphyromonas gingivalis, P. gingivalis, Fusobacterium nucleatum, F. nucleatum, periodontal disease, periodontitis, oral hygiene, tooth loss, brushing, esophageal squamous cell carcinoma, ESCC, esophageal cancer, orodigestive cancer, and cancer. We conducted backward citation tracking by systematically screening reference lists of all included articles, followed by manual verification of potentially relevant publications.
Oral Microbiome in Healthy People
The human oral microbiome is a complex ecological environment with over 700 known microbial species. 13 Normal oral microorganisms mainly include 6 phyla—Bacteroidetes, Firmicutes, Proteobacteria, Actinobacteria, Fusobacteria, and Spirochaetes—which account for 96% of species. 15 Nearing et al 16 examined the salivary microbiota of 1049 healthy Canadian individuals and found that oral microorganisms are similar among individuals at the genus level, with almost 99% of the samples containing 11 genera, predominantly Veillonella (21.49%), Streptococcus (11.86%), Neisseria (13.04%) and Prevotella_7 (11.55%). The study showed that oral microbes are abundant and similar among individuals in the same population.
The human oral cavity contains complex ecological niches that include saliva, the soft tissue surfaces of the oral mucosa and tongue, and the hard tissue surface of the teeth. Each of these components has its own microbial composition, influenced by pH level, oxygen concentration, nutrition, and oral hygiene practices. 17 Dong et al 18 collected clinical samples from 3 oral niches (saliva, tongue dorsum, and supragingival plaque) in 27 healthy individuals, and identified distinct niche-specific bacterial signatures. The class Bacteroides showed higher abundance in saliva, while Clostridia preferentially colonized the tongue dorsum. In contrast, Flavobacteria were more likely to be enriched in supragingival plaque.
Oral microbial communities can prevent the invasion of exogenous pathogens and participate in the maturation of the host’s innate and adaptive immune systems. Some oral bacteria have immunomodulatory roles in maintaining appropriate host immune responses and host-microbe homeostasis. 19 Oral microorganisms and the oral environment together constitute a relatively stable oral microecology, which plays an important role in both oral and overall health.
Oral Microbiome and ESCC
In a healthy state, microorganisms and the host jointly maintain the homeostasis and dynamic balance of the body. When the microbial balance is disturbed—that is, when an ecological imbalance occurs—there can be significant metabolic and immune effects on the host, potentially leading to a variety of diseases, including cancer. 20
Saliva represents the overall oral microbiota. The salivary microbiota mainly consists of microorganisms isolated from various ecological niches in the mouth. Due to its easy sampling, high genetic stability, and representativeness, saliva is commonly used to study the oral microbiota, and certain trends in salivary microbes have been reported to correlate with ESCC. 21 For instance, Chen et al 22 found that the alpha diversity of patients with ESCC is significantly lower (P < 0.001) than that of healthy controls or those with esophageal squamous dysplasia (ESD), and the relative abundance of Prevotella ( 42.4% vs 36.1%), Streptococcus (21.9% vs 16.1%), and Porphyromonas (8.9% vs 6.5%) is significantly increased among ESCC patients. In contrast, another study found that the alpha diversity of patients with ESCC is significantly higher than that of healthy controls (P < 0.001) and that there is no difference in the composition of the microbiota. 23 It was discovered that the saliva of patients with ESCC contains a higher abundance of Leptotrichia, Porphyromonas, and Streptococcus and a lower abundance of Haemophilus, Alloprevotella, and Prevotella, and these results were verified by Meng et al. 24 In addition, the combined use of Porphyromonas/Prevotella and Porphyromonas/Alloprevotella ratios increases the ESCC diagnostic efficiency (AUC = 0.826). Some studies found no significant differences between patients with ESCC and healthy controls in terms of alpha diversity, but beta diversity analyses suggested that ESCC patients and control individuals have a statistically significant difference in community composition.25-27 The percentage of Firmicutes and Bacteroidetes in ESCC patients is reportedly higher than that in control participants, whereas the percentage of Proteobacteria is lower. At the genus level, the abundance of Prevotella_7 and Streptococcus increases, while that of Neisseria decreases. 25
The tongue coating, which covers the rich surface of the tongue papillae, tongue cleft, and recess, acts as a reservoir for oral pathogens, for which it provides a suitable low oxygen environment and nutrients. 28 Jiang et al 29 used sterile throat swabs to collect oral microorganisms from the tongue coatings of 56 patients with ESCC and 53 control participants. It was found that in patients with ESCC, the microbial richness and diversity are slightly higher and the abundance of Fusobacteriales, Fusobacteria, Fusobacteriota, Leptotrichiaceae, and Leptotrichia is increased.
Dental plaque is a biofilm formed on the surface of teeth and is mainly classified as supragingival plaque and subgingival plaque. The bacterial community composition of the 2 categories of plaque is relatively similar, and plaque bacteria are more abundant than saliva and tongue bacteria. 30 Oral bacteria present in dental plaque can potentially be used to identify patients with EC. Chen et al 31 analyzed the microbiome in the dental plaque at the gingival margin of 34 patients with of ESCC and 18 healthy participants to identify bacterial biomarkers associated with ESCC. Significant differences were observed in the alpha and beta diversities between the ESCC and control groups. Specifically, Streptococcus species, Veillonella parvula and P. gingivalis are more prevalent in ESCC than in healthy individuals. It was further demonstrated that P. gingivalis promotes ESCC tumor progression in vitro and in vivo. Furthermore, Kawasaki et al 32 calculated the copy number of 7 pre-selected bacteria by real-time polymerase chain reaction (PCR) in 61 patients with of EC (58 ESCC cases and 3 esophageal adenocarcinomas cases) and 62 age-matched controls. Aggregatibacter actinomycetemcomitans, P. gingivalis, Streptococcus anginosus, Treponema denticola, and Tannerella forsythia were more abundant in subgingival plaque in the EC group. In addition, T. forsythia (OR = 7.92, 95% CI: 2.20-28.50) and S. anginosus (OR = 32.80, 95% CI: 3.25-331.00) infection in the subgingival plaque was associated with a high risk of EC.
The above studies indicate that different ecological niches (saliva, the tongue coating, and dental plaque) in the oral cavity of ESCC patients have their own microbial characteristics, and changes in oral microbiota composition may be related to ESCC. However, due to the limitations of these case-control studies, a causal relationship between oral microorganism characteristics and ESCC cannot be inferred. Two other prospective case-control studies have revealed a causal link between oral microbiome changes and the etiology of ESCC. Peters et al 33 evaluated oral bacteria in 25 patients with ESCC and 50 matched controls. It was found that Treponema vincentii, Neisseria weaveri, and Prevotella nanceiensis increase the risk of ESCC (P < 0.05) and that P. gingivalis is marginally associated with the risk of ESCC (P = 0.09). In addition, Liu et al 34 collected oral swab samples from 84 patients with severe ESD and esophageal lesions (including severe ESD, carcinoma in situ, and ESCC) and 168 controls. Eleven species were found to predict the risk of malignant esophageal lesions, including Actinomyces odontolyticus, Dialister invisus, Actinomyces viscosus, Leptotrichia hofstadii, Fusobacterium mortiferum, Prevotella shahii, F. nucleatum, Prevotella baroniae, Prevotella melaninogenica, Rothia dentocariosa, and Lachnoanaerobaculum umeaense.
Although only a handful of studies, with small sample sizes, have been conducted, significant differences in the oral microbiota between patients with ESCC and healthy individuals have been revealed (Table 1). The findings may have substantial implications for predicting ESCC development, and the role of the identified bacterial species warrant further investigation. The aforementioned studies showed that although alpha and beta diversity differ between ESCC patients and healthy individuals, ESCC patients exhibit more changes in the abundance of different microbial species. Therefore, oral dysbiosis may be associated with EC. The oral microbiota is rich and diverse, and sampling is convenient, noninvasive, and low cost. Therefore, the microbiota is expected to become a tumor screening marker. In high-risk populations especially, key oral pathogens associated with ESCC are potential microbial biomarkers that could be used during screening and surveillance to facilitate early detection and preventive interventions.
Oral microbiome and ESCC.
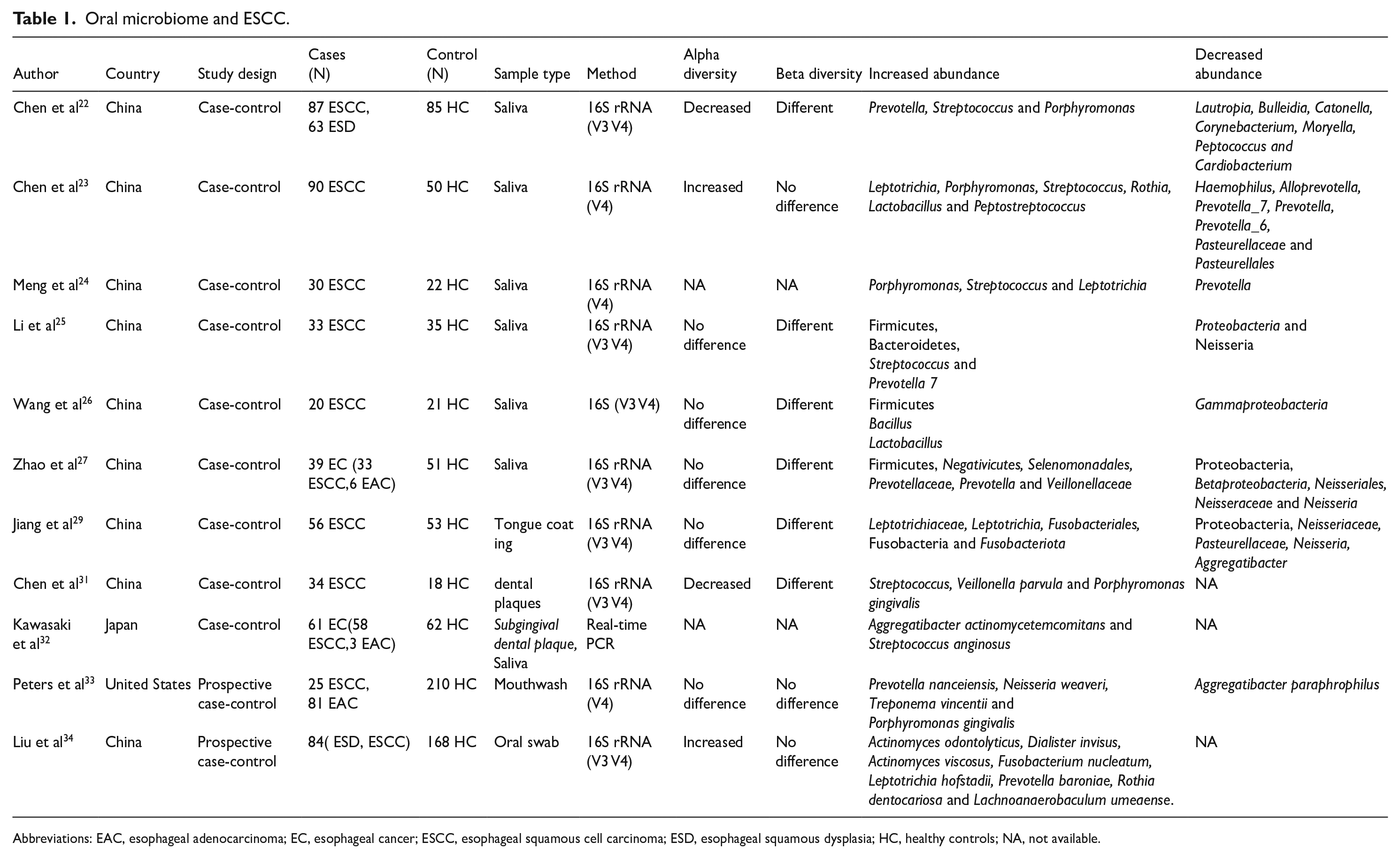
Abbreviations: EAC, esophageal adenocarcinoma; EC, esophageal cancer; ESCC, esophageal squamous cell carcinoma; ESD, esophageal squamous dysplasia; HC, healthy controls; NA, not available.
Microbiota of the Normal Esophagus
Initially, it was generally believed that the esophagus had no colonized microbiota and that its microbiota was transient and similar to the oropharyngeal microbiota. In the early 1980s, traditional isolation culture methods confirmed for the first time that the esophagus was not a completely sterile environment.35,36 The esophagus has its own unique microbiota, dominated by Streptococcus.37-39 However, due to the limitations of specimen sources, culture methods, and detection techniques, the complexity of the esophageal microbiota has been greatly underestimated.
Since its development, 16S rRNA gene sequencing technology has been widely used to study the human microbiota and has provided a fundamental breakthrough in the study of the esophageal microbiota. In 2004, Pei et al 14 used 16S rDNA PCR technology for the first time to analyze esophageal mucosal biopsies from 4 adults without esophageal diseases undergoing upper gastrointestinal endoscopy. It was determined that the distal normal esophagus harbors colonized bacteria, primarily comprising 6 phyla: Bacteroides, Firmicutes, Actinobacteria, Fusobacteria, Proteobacteria, and TM7. There are 3 major genera: Streptococcus, Prevotella, and Veillonella, which account for 39%, 17%, and 14% of the species, respectively. In 2020, Yin et al 40 analyzed the microbiota in esophageal brush specimens from 27 healthy participants, and the results were consistent with those of Pei et al. 14 It was also found that the microbial composition of the upper, middle, and lower esophagus is very similar.
In summary, despite limited research on the normal esophageal microbiota, small sample sizes, and variations in specimen sources and detection techniques, a resident microbiota is generally believed to exist in the normal esophagus, primarily comprising gram-positive bacteria (Table 2). Firmicutes (mainly Streptococcus) is the most common type and is vital in maintaining a normal esophageal environment in the esophagus.
Microbiota of the normal esophagus.

Esophageal Microbiota and Esophageal Disease
Studies have confirmed a relationship between chronic inflammation and EC. Therefore, the esophageal microbiota may play an important role in the progression of EC. An imbalance in the esophageal microbiota is associated not only with esophagitis but also with ESD.
In 2009, Yang et al 41 performed a microbiome analysis using distal esophageal biopsy samples from 34 participants, including 12 healthy individuals, 12 patients with esophagitis, and 10 patients with Barrett’s esophagus (BE). In total, 166 species belonging to 9 phyla and 70 genera were identified. In general, 2 types of esophageal microbiomes can be distinguished. The type I microbiome is strongly associated with a normal esophagus, chiefly comprising Streptococcus (78.8%). The type II microbiome is primarily associated with esophageal abnormalities and mainly comprises Veillonella, Prevotella, Haemophilus, Neisseria, Porphyromonas, and Fusobacterium. Liu et al 42 also found that the esophageal bacteria composition of healthy individuals, patients with reflux esophagitis, and BE patients is different, suggesting that different bacterial communities might be involved in the occurrence of esophageal diseases.
Yu et al 43 analyzed the upper digestive tract microbiome of 142 ESD patients and 191 individuals without ESD and found that a decrease in the abundance of species in the esophageal microbiome is associated with ESD. Therefore, ESD may be caused by a change in the upper digestive tract microbiome, and this change directly correlates with the EC development. These studies suggest that changes in the esophageal microbiome may have a relationship with esophageal diseases.
Esophageal Microbiome and ESCC
Due to the difficulty of esophageal tissue sampling, studies on esophageal microecology are limited, and research on the correlation between esophageal microbiome changes and ESCC remains in the initial stage. Studies have shown that when ESCC occurs, esophageal microbial diversity changes significantly.44-46 The abundance of Streptococcus decreases,47-49 and in EC, that of gram-negative anaerobic bacteria or microaerophiles such as Fusobacterium, Prevotella, Porphyromonas, and Neisseria increases.46-51 Fusobacterium is more abundant (3.2% vs 1.3%), whereas Streptococcus is less abundant (12.0% vs 30.2%). 47
In a study by Li et al, 49 subjects were divided into 5 groups: 70 healthy individuals, 70 patients with of esophagitis, 70 patients with low-grade intraepithelial neoplasia, 19 patients with high-grade intraepithelial neoplasia, and 7 patients with ESCC. Significant differences in Neisseria, Streptococcus, Haemophilus and Porphyromonas were observed among the groups. The Streptococcus abundance decreased during ESCC progression, whereas the abundance of the other genera increased. Finally, the combination of Streptococcus and Neisseria proved useful, as a predictor for ESCC and its precancerous lesions (AUC = 0.738). Yang et al 46 estimated the microbial dysbiosis index by combining 19 related taxa of patients with ESCC and people with a physiologically normal esophagus, including Prevotella, Fusobacterium, and Selenomonas. It was found that the microbial dysbiosis index can effectively distinguish ESCC from normal esophageal conditions.
Compared with the amplification and sequencing of partial 16S regions, full-length 16S amplicon sequencing significantly improves annotation accuracy and resolution and enables species-level annotation. Shen et al 52 conducted full-length 16S rDNA sequencing of 19 ESCC cases and found that the relative abundance of Labrys ginsengisoli, Peptoanaerobacter stomatis, Selenomonas sputigena, Streptococcus constellatus, and Fusobacterium periodonticum is increased in ESCC tissues, whereas that of Lactobacillus murinus and Escherichia coli is decreased.
The esophageal microbiota not only affect the occurrence of ESCC but also correlate with its prognosis.50,53,54 According to Liu et al, 53 the diversity of the esophageal microbiota is related to the stage of ESCC. In comparison with patients with lymph nodes-negative, those with lymph nodes-positive exhibit a higher abundance of Bacteroidetes, Spirochaetes and Firmicutes and a lower abundance of Proteobacteria. In the lymph node-positive group, the abundance of Prevotella and Treponema is higher. Compared with patients with ESCC at stages T1-2, patients with ESCC at stages T3-4 have a significantly higher abundance of Streptococcus. After adjusting for clinicopathological characteristics, the Streptococcus and Prevotella combination is associated with a poor prognosis, indicating its utility as an independent prognostic marker. This study showed that changes in bacterial abundance are linked to the tumor stage, and some microorganisms may be involved in ESCC development.
In studies of esophageal microbiota, most tissue specimens are obtained from endoscopic biopsies or surgical samples. Liu et al 55 investigated the esophageal microbiota in ESCC patients by comparing the differences in bacterial communities between surgical and endoscopic esophageal mucosal tissues. They found that the microbial composition of surgical esophageal tissues differed from that of mucosal tissues. Compared with mucosal samples, surgical tissues exhibited increased microbial richness and improved evenness, suggesting that surgical tissues may be more suitable for studying EC microbiota. Therefore, the source of samples should be considered in research on esophageal microbiota.
The above studies demonstrated that the esophageal microbiota of ESCC patients differs significantly from the normal esophageal microbiota (Table 3). However, due to the difficulty of obtaining biopsy samples, the study sample sizes were small, and most studies were cross-sectional case-control studies. Larger prospective cohort studies are needed for more reliable data.
Esophageal microbiome and ESCC.
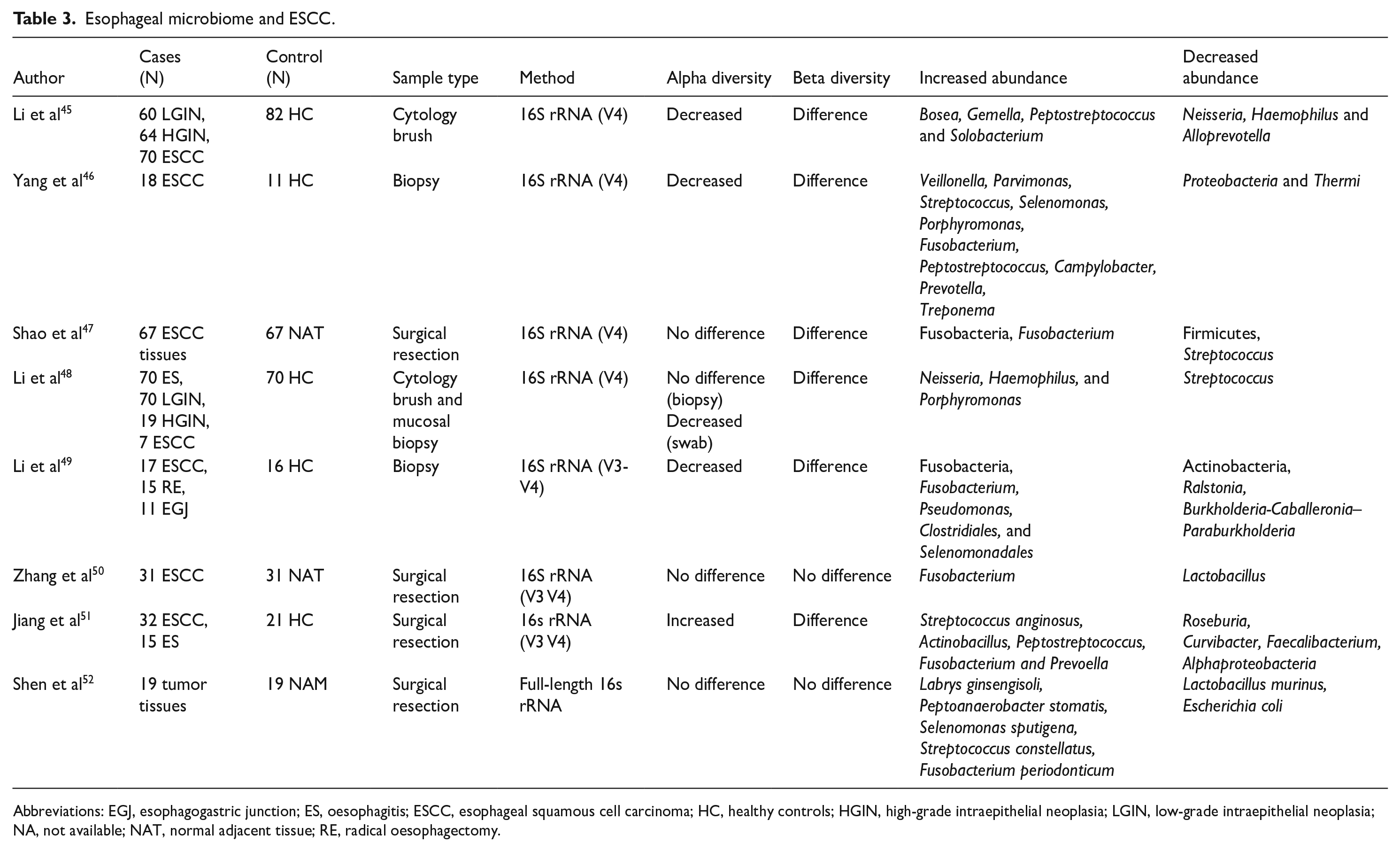
Abbreviations: EGJ, esophagogastric junction; ES, oesophagitis; ESCC, esophageal squamous cell carcinoma; HC, healthy controls; HGIN, high-grade intraepithelial neoplasia; LGIN, low-grade intraepithelial neoplasia; NA, not available; NAT, normal adjacent tissue; RE, radical oesophagectomy.
Certain Bacteria and ESCC
The synergistic action of multiple bacteria is widely believed to be related to carcinogenesis. Nevertheless, specific microbial infections have been confirmed to induce microecological disturbances and promote tumor progression. For instance, P. gingivalis, a pathogen with an important role in periodontitis, has been observed to influence the development of systemic diseases.
Gao et al 56 confirmed that P. gingivalis colonizes ESCC tissues and adjacent tissues with infection rates of 61% and 12%, respectively, and is not found in normal esophageal mucosa. In addition, P. gingivalis is associated with various clinicopathological characteristics, including degree of differentiation, lymph node metastasis, and clinical stage. Patients positive for P. gingivalis demonstrate a poor prognosis. In addition, the serum levels of P. gingivalis immunoglobulin G and immunoglobulin A antibodies are significantly greater in patients with ESCC than in those with esophagitis or normal esophageal conditions. Patients with ESCC that exhibit high serum levels of either immunoglobulin G or immunoglobulin A antibodies have poor prognoses, and high levels of both antibodies are associated with the worst prognoses. 57 P. gingivalis can promote the upregulation of GARP leading to TGF-β activation, induce the epithelial-mesenchymal transition, and subsequently promote ESCC invasion and metastasis. 58
The role of another well-known periodontal pathogen, F. nucleatum, has also been noted in EC. More F. nucleatum DNA is detected in ESCC tissues than in normal esophageal mucosa. F. nucleatum DNA quantity is positively correlated with tumor stage, and the overall survival of F. nucleatum–positive patients is shortened.59,60 Researchers believe that F. nucleatum abundance could be combined with other indicators to predict the metastasis and prognosis of ESCC. 60 Furthermore, in patients with ESCC, a higher abundance of intratumoral F. nucleatum is associated with a poorer neoadjuvant chemotherapy response and a higher tumor recurrence rate.61,62 F. nucleatum invades ESCC cells and induces the NF-κB pathway through the NOD1/RIPK2 pathway, promoting tumor progression. 63 In addition, KEGG enrichment analysis of DNA microarray data has demonstrated that F. nucleatum may promote aggressive tumor behavior by activating chemokines such as CCL20. 59 Furthermore, Li et al 64 discovered that the virulence factor F. nucleatum-Dps, secreted by F. nucleatum, binds to the transcription factor ATF3 and upregulates PD-L1 expression on the surface of ESCC cells. This mechanism enables tumor cells to evade T-cell-mediated attack while protecting intracellular F. nucleatum from immune clearance. Furthermore, F. nucleatum can infiltrate T cells and ultimately induce T-cell death, thereby impairing their tumor-killing activity.
These results have revealed that P. gingivalis and F. nucleatum may encourage ESCC occurrence and development. These are potential prognostic and predictive biomarkers of ESCC, as well as potential targets to improve the treatment response to antibiotic intervention. In addition, the interactions between distinct microbial populations influence the pathogenesis and progression of host diseases. 65 Targeting keystone symbiotic bacteria, designing synthetic microbial consortia, and developing phage or quorum sensing inhibitors to precisely modulate pathogenic bacteria will emerge as potential microbiome-based therapeutics for EC.
Factors Influencing ESCC Microbiota
The oral and esophageal microbiota composition is affected by factors like alcohol consumption, smoking, dietary habits, and oral hygiene. For instance, a 2018 cross-sectional study of 1044 US adults found that heavy drinking alters the oral microbiome composition. Heavy and nonheavy drinkers differ in terms of oral microbiota diversity and overall bacterial profiles. The abundance of Lactobacilli decreases with increasing alcohol consumption, while the abundance of Actinomyces, Leptotrichia, Cardiobacterium, and Neisseria increases. Neisseria, in particular, promotes ethanol synthesis of acetaldehyde, a carcinogen associated with EC. 66 Rao et al 67 assessed 120 ESCC cases, including 60 drinkers and 60 nondrinkers, and confirmed that drinking is related to esophageal microbial diversity in ESCC patients. The microbial diversity of drinkers is significantly lower than that of nondrinkers, and drinkers display a higher abundance of Pasteurellales, especially Pasteurellaceae. In contrast, Clostridiales, Clostridium, Lachnospiraceae, Clostridiaceae, Helicobacteraceae, Helicobacter, and Catonella are more abundant in nondrinkers.
Several other studies have examined the effects of smoking on the microbiota. Wu et al 68 evaluated the relationship between smoking and the oral microbiome in 1204 US adults and found that the oral microbiome composition differs between former smokers and nonsmokers (P < 0.001). Former smokers display a higher abundance of Atopobium and Streptococcus than never-smokers, while smokers exhibit a lower abundance of Capnocytophaga, Peptostreptococcus and Leptotrichia. Smoking may benefit the growth of oral anaerobic bacteria by suppressing the aerobic metabolic pathway and enhancing the oxygen-independent pathway. Vogtmann et al 69 examined 278 Chinese men who underwent EC screening. According to their findings, compared with never-smokers, smokers display increased microbial diversity, and Dialister invisus and Megasphaera microuciformis are more abundant. Li et al 70 verified that the composition of the esophagus and salivary microbiota of drinkers and smokers differs from that of nondrinkers and nonsmokers and that the abundance of Neisseria, Porphyromonas, Prevotella, and Fusobacterium in the saliva of smokers and drinkers is higher.
In terms of diet, Nobel et al 71 obtained esophageal samples from 47 outpatients who underwent an endoscopy and discovered that dietary fiber intake is positively correlated with Firmicutes abundance and negatively correlated with Neisseria, Prevotella, and Eikenella abundance. In terms of oral hygiene, Chen et al 12 found that fewer daily brushing sessions and a greater number of lost teeth increase ESCC risk. One explanation is that oral hygiene affects bacterial diversity, and therefore, maintaining good oral hygiene is crucial for oral and esophageal health.
Summary and Outlook
Research on the relationship between human oral and esophageal microecology and EC is still in its early stages, compared with research on intestinal microecology. Although studies have demonstrated that changes in the oral and esophageal microbiota may be closely related to the development and progression of ESCC, these studies were limited by their cross-sectional design, sample size, and detection techniques. In cross-sectional studies, the microbiota profiles obtained from EC and control groups have limited reliability and accuracy. Moreover, a cross-sectional study can only suggest an association between changes in the microbiome and the occurrence of disease and cannot prove a causal relationship. In addition, many of the studies had very limited sample sizes; most studies included less than 100 cases. Small sample sizes preclude the detection of truly significant relationships between variables, and some findings may be spurious and unable to be replicated. Therefore, to clarify the specific role of certain bacteria in EC, prospective cohort studies with large samples are needed. Longitudinal tracking of microbiota changes is essential to understand the role of the oral and esophageal microbiota in ESCC etiology.
The most commonly used microbiome detection technology is 16S rRNA gene sequencing in the V3-V4 region, but this method is ineffective for the identification of microorganisms in tumor tissues, possibly due to interference caused by the high proportion of host DNA. 5R 16S rRNA gene sequencing can greatly improve the coverage and resolution of bacterial species detection, especially in microbial samples with low biomass. 72 Additional metagenomic sequencing methods can help to reveal species-level classification in more detail and assess the function of the microbiome.
Conclusions
In conclusion, changes in the oral and esophageal microbiota are closely correlated with EC. However, metagenomic analyses and experimental verification via large-scale prospective cohort studies are required to establish a causal relationship, identify specific pathogenic bacteria, and understand underlying mechanisms, as well as to discover microecological targets for precise ESCC treatment. Future detection and quantification of specific bacterial biomarkers will enable comprehensive assessment of disease risk, pathological progression, and prognostic outcomes in patients, thereby facilitating optimized clinical decision-making for improved therapeutic efficacy in EC management.
Footnotes
Ethical Considerations
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Author Contributions
JK wrote the initial manuscript and performed revisions. SG conceptualized the study and critically revised the manuscript. JW, YL, and RL participated in partial revision and editing. All authors read and approved the final manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the General Science and Technology Project of the Open Joint Fund of the National Clinical Key Specialty Construction of Oncology, The First Affiliated Hospital of Henan University of Science and Technology (KJ, grant number ZLKFJJ20230504; LY, grant number ZLKFJJ20230513), and the National Natural Science Foundation of China (GS; grant number 81972571) and the Science and Technology Projects of Luoyang (WJ; grant number 2302010Y).
Declaration Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Not applicable.
