Abstract
Background:
Evaluation of biological changes at the molecular level has important clinical implications for improving the survival rate of esophageal squamous cell carcinoma (ESCC). Therefore, we plan to analyze and elucidate the expression of microRNA-133b (miR-133b), M2 pyruvate kinase (PKM2), and signal transducer and activator of transcription 3 (STAT3) in ESCC and their associated clinicopathological significance.
Methods:
The 72 patients with ESCC were selected as the experimental study group. Normal adjacent tissues (NAT) were matched as the control group. In this study, in situ hybridization was used to detect the expression of miR-133b in ESCC, and tissue expressions of PKM2 and STAT3 were detected by immunohistochemistry, and literature review was conducted.
Results:
Studies had shown that the positive expression of miR-133b in NAT was significantly higher than that in ESCC (χ2 = 9.007, P = .003). PKM2 and STAT3 in ESCC had a significantly higher positive expression levels than those of NAT (χ2 = 56.523, P = .000; χ2 = 72.939, P = .000). From correlation analysis, there was a negative correlation between miR-133b and PKM2(r = −0.515, P < .001), a negative correlation between miR-133b and STAT3(r = −0.314, P = .007), and a positive correlation between PKM2 and STAT3(r = 0.771, P < .001).
Conclusions:
In ESCC, our study demonstrated that downregulation of miR-133b and upregulation of PKM2 and STAT3. We predict that miR-133b may inhibit the STAT3 pathway by downregulating PKM2.
Introduction
In the world, esophageal cancer is one of the ten most common malignant tumors. 1 In China, the incidence of esophageal cancer ranks sixth and the death rate ranks fourth. 2 Histological type, the most common is esophageal squamous cell carcinoma (ESCC). 3 In terms of 5-year survival rates, esophageal cancer is reported to be less than 20%. 4 Cancer cell invasion of blood vessels and distant metastasis are the main causes of death of esophageal cancer. 5 Therefore, evaluation of biological changes at the molecular level has important clinical implications for improving the survival rate of patients with esophageal cancer.
miRNAs are closely related to cell proliferation, apoptosis, differentiation, and growth. 6 The miR-133b is abnormally expressed in cancers. 7 miR-133b promotes the phosphorylation of SMAD3 by activating the TGF-β/Smad3 signaling pathway, affects the expression of target genes, and mediates the occurrence of EMT in tumor cells. 8 miR-133b can also directly target fibroblast growth factor receptor 1 or inhibit Sirt1 expression through Wnt/catenin signaling pathway to reduce tumor cell vitality, and inhibit osteosarcoma cell growth, invasion, and migration. 9 We had analyzed the microarray hybridization results of ESCC and normal adjacent tissues (NAT), and found that miR-133b was significantly lower in ESCC than in NAT.
In tumor cells, M2 pyruvate kinase (PKM2) is a well-known the rate-limiting enzyme in the final step of aerobic glycolysis. PKM2 is highly and specifically in a variety of tumor cell lines and cancer tissues, and plays a major role in tumor metabolism and malignant proliferation.10,11 Our team studied ESCC and found that PKM2 is highly expressed. 12 The over activation of signal transduction and transcription activating factor 3 (STAT3) can induce the abnormal expression of multiple target genes, promotes cell growth and proliferation, malignant transformation, and hinders cell death in a variety of ways. 13 STAT3 can be activated by phosphorylation of EGFR, JAK, Src, and other classical tyrosine kinases, and it is the junction of multiple carcinogenic tyrosine kinase signal channels. 14 In our previous work, we found that high PKM2 activity transformed JAK-STAT3 pathway into PKM2-STAT3 pathway, resulting in glucose metabolism disorder and tumor invasion and metastasis of ESCC.15,16
In ESCC, whether miR-133b targets PKM2-STAT3 expression to inhibit invasion and metastasis has not been reported. We are the first to report the expression of miR-133b, PKM2, and STAT3 in ESCC, and further analyse the relationship with clinicopathological characteristics.
Materials and Methods
Patients
72 cases of ESCC were selected as the study group, and 72 cases of NAT were selected as the control group from January 2020 to December 2021 in the Affiliated Hospital of Southwest Medical University. Inclusion criteria are given as follows: (1) ESCC was diagnosed by histopathology. (2) The patients had not received radiation or chemotherapy prior to surgery. (3) Complete clinical and pathological data are available. Exclusion criteria are given as follows: (1) Patients with other malignant tumors. (2) Patients with a history of blood transfusion and immunotherapy. There are 64 males and 8 females in the study. The age is between 43 and 90, with an average age of 63. The tumor was 0.7 to 7.0 cm in diameter. The NAT was more than 5 cm from the tumor margin, and there was no tumor cell infiltration confirmed by pathology. This study protocol was reviewed and approved under the guidance of the Institutional Ethics Committee of the Affiliated Hospital of Southwest Medical University (No. KY2022274), with the informed consent of patients or their families.
In situ hybridization
The miR-133b ISH kit (No. MK10754) and supporting reagents were purchased from BOSTER. The sequence of has-miR-133b probe was follow: 5′-TAGCTGGTTGAA GGGGACCAAA-3′. The experiment was conducted in strict accordance with the instructions. All slides were hybridized overnight in 42° incubator, and DAB staining was strictly observed under the microscope by skilled pathologist before termination. The positive tissue section proved by the pre-experiment is used as the positive control, and the pre-hybridization solution replaces hybridization solution as negative control.
Immunohistochemistry
The expression of PKM2 and STAT3 in 72 cases of ESCC and NATs was detected by EnVision method. Rabbit anti human monoclonal antibody PKM2 (Catalog number: 4053S; 1:800 dilution) and STAT3 (Catalog number: 9145S; 1:100 dilution) were purchased from Cell Signaling Technology (CST) in the United States. The secondary antibody (Catalog number: #KIT-9921) was purchased from MXB Biotechnology and DAB staining kit (Catalog number: #ZLI-9018) was purchased from Zhongshan Jinqiao Biotechnology. It is very important to do a good job in antigen repair. The positive tissue section confirmed by the experiment was used as the positive control, and the first antibody detected by IHC was replaced by phosphate buffer solution (PBS) as the negative control.
Result determination
The classical immune response score (IRS) system was used to evaluate immunostaining semi-quantitatively. 17 The score was based on staining intensity and cell positive percentage. Staining strength definition: 0 was negative; 1 was weak; 2 was mild; 3 was strong. Percentage of positive cells: 0 was negative; 1 was less than 10% positive cells; 2 were 10% to 50% positive cells; 3 were 51% to 80% positive cells; 4 were more than 80% positive cells. The optimal cores defining the expression levels of miR-133b, PKM2, and STAT3 were determined by staining intensity multiplying positive percentage of cells. Those whose IRS values above the 4 score was high expression; whereas below the 4 score were low expression. The results were carefully evaluated by two experienced pathologists (LXM and LQ).
Observations
We further investigated the relationship between the expression levels of miR-133b, PKM2, and STAT3 in ESCC and the clinicopathological characteristics of patients, including sex, age, tumor diameter, pathological grade, depth of invasion, vascular metastasis, neural metastasis, and lymph node metastasis, and conducted statistical analysis.
Statistical analyses
The experimental data were analyzed and processed by SPSS 16.0 statistical softwar.
When the data were normal distribution, the two-tailed independent sample T-test was used to analyze the statistical difference between the two groups. The correlation between clinicopathological parameters and protein expression was analyzed by χ2 test. When there were less than 5 categorical variables in the contingency table, Fisher exact test was used. Pearson correlation was used for correlation analysis. A P value < .05 was considered statistically significant.
Results
Expression of miR-133b, PKM2, and STAT3 in ESCC
Brown particles with positive expression of miR-133b and STAT3 were found in the nucleus of esophageal squamous epithelial cells, and brown particles were found in the cytoplasm of esophageal squamous epithelial cells with positive PKM2 (Figure 1). The results showed that the expression level of miR-133b in NAT (63.9%, 46/72) was significantly higher than that in ESCC (38.9%, 28/72), and the difference between the ESCC and NAT was statistically significant (χ2 = 9.007, P = .003). The expression of PKM2 in ESCC (84.7%, 61/72) was significantly higher than that in NAT (22.2%, 16/72) (χ2 = 56.523, P = .000). The expression of STAT3 in ESCC (80.6%, 58/72) was significantly higher than that in NAT (9.7%, 7/72) (χ2 = 72.939, P = .000) (Table 1).

Detection of miR-133b, PKM2, STAT3 expression in normal adjacent tissues (A, C, E), esophageal squamous cell carcinoma tissues (B, D, F). (A) miR-133b was highly expressed. (B) The expression of miR-133b was low. (C) PKM2 low expression. (D) High expression of PKM2. (E) STAT3 expression was low. (F) STAT3 was highly expressed. Magnification was 200-fold, scale bar represents 50 μm correspondingly.
Expression of miR-133b, PKM2, and STAT3 in ESCC and NAT.

Abbreviations: ESCC, esophageal squamous cell carcinoma; miR-133b, microRNA-133b; NAT, normal adjacent tissues; PKM2, M2 pyruvate kinase; STAT3, signal transducer and activator of transcription 3.
Expression of miR-133b, PKM2, and STAT3 with clinicopathological characteristics
Clinicopathological characteristics data was classified and organized in ESCC, including sex, age, tumor diameter, pathological grade, depth of invasion, vascular metastasis, and nerve metastasis, meanwhile, the high and low expression of miR-133b, PKM2 and STAT3 had no significant differences with them (Supplementary Table 1).
Correlation expression of miR-133b, PKM2, and STAT3 in ESCC
The correlation expression of miR-133b, PKM2 and STAT3 in ESCC was further analyzed (Supplementary Tables 2, 3, and 4). From correlation analysis, there was a negative correlation between miR-133b and PKM2 (r = −0.515, P < .001) (Figure 2), a negative correlation between miR-133b and STAT3 (r = −0.314, P = .007) (Figure 3), and a positive correlation between PKM2 and STAT3(r = 0.771, P < .001) (Figure 4).

Correlation between miR-133b and PKM2.

Correlation between miR-133b and STAT3.

Correlation between PKM2 and STAT3.
Discussion
Early detection of regulatory targets related to the malignant phenotype of esophageal cancer invasion and metastasis, thereby inhibiting the invasion and metastasis of cancer cells, has important clinical significance for improving the survival rate of patients. Our study demonstrated that the expression level of miR-133b in NAT was significantly higher than that in ESCC, while PKM2 and STAT3 expression in ESCC were significantly higher than those in NAT. Further analysis showed that there was a negative correlation between miR-133b and PKM2, a negative correlation between miR-133b and STAT3, and a positive correlation between PKM2 and STAT3. The expression of miR-133b, PKM2, and STAT3 in ESCC did not differ significantly in clinicopathological characteristics, which may be related to the number and representativeness of patients, and this speculation needs to be verified by a large sample and multicenter study. Comprehensive analysis showed downregulated expression of miR-133b and upregulated expression of PKM2 and STAT3 in ESCC. In ESCC, miR-133b was a tumor suppressor gene, PKM2 and STAT3 were oncogenes. Based on our previous experimental results,15,16 we speculate that PKM2 may be a target gene for miR-133b, which negatively regulates the expression of PKM2 and STAT3, promoting the invasion and metastasis of ESCC, however, further validation is needed.
Studies shows that most miRNAs are highly conservative, sequential, and tissue-specific, and have great potential to become new biomarkers.18,19 At present, an important challenge of miRNA research is how to quickly predict, search and determine the target gene of miRNA. It makes sense to understand the function of miRNA and the biological processes involved in miRNA and the occurrence of diseases, and finally to use miRNA in clinical practice. The miR-133b showed low expression in ESCC,20 -22 which was consistent with our result.
We further reviewed the literature about miR-133b (Figure 5). There are seven reports about miR-133b regulating PKM2 expression to inhibit tumor invasion and metastasis.23 -29 Overexpression of miR-133b inhibits the expression of target gene PKM2 and Warburg effect, which can be helpful to improve the diagnosis and treatment of ovarian cancer. 23 Overexpression of miR-133b can inhibit the growth and proliferation of lung cancer stem cells by downregulating the expression of PKM2. 24 It has been proved that overexpression of miR-133b makes radio-resistant lung cancer cells desensitized by inhibiting PKM2-mediated glycolysis. 25 miR-133b can inhibit the proliferation of tongue squamous cell carcinoma by downregulating PKM2 expression. 26 Downregulation of miR-133b will further increase PKM2 expression and promote tumor progression. 27 miR-133b exerted a tumor inhibitory effect by negatively regulating the Warburg effect of gastric cancer cells. 28 miR-133b downregulates PTBP1 and induces the transformation of PKM subtype from PKM2 to PKM1, resulting in the growth inhibition of rhabdomyosarcoma. 29
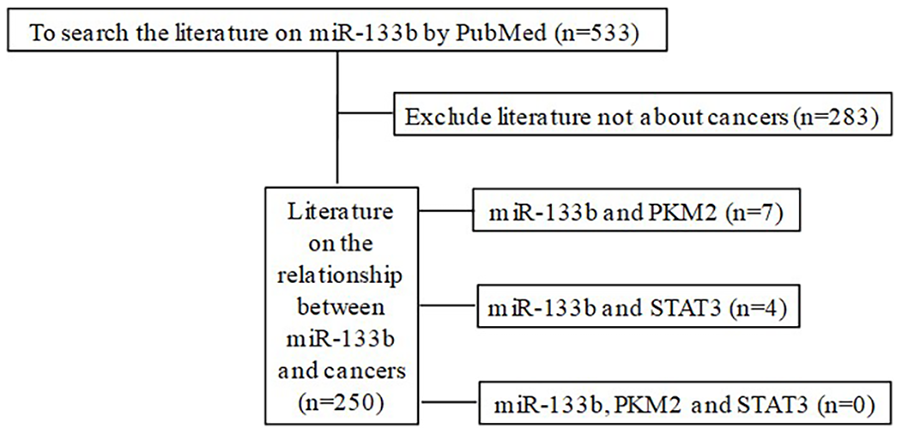
Diagram of miR-133b search.
The scientists are very concerned about the important role of STAT3 in ESCC.30 -33 At present, there are only three studies on miR-133b and STAT3.34 -36 The low expression of miR-133b was negatively associated with EGFR and increased p-STAT3 levels. 34 miR-133b was downregulated in wild-type and mutant cell renal cell carcinoma and induced apoptosis in vitro through signal cascade involving JAK2, STAT3 and Bcl-2. 35 miR-133b altered the regulation of the STAT3 signaling pathway and the expression of downstream proteins in nasopharyngeal carcinoma cells. 36 We further analyzed the relationship between PKM2 and STAT3. Research suggested that overexpression of PKM2 promotes colorectal cancer cell migration and adhesion by regulating STAT3-associated signaling. 37 Gao et al 38 showed that the energy metabolism of gastric cancer is controlled through the mTOR/PKM2 and STAT3/ c-Myc signaling pathway, thereby improving the malignant biological behavior of gastric cancer. At present, only our team has reported on the relationship between PKM2 and STAT3 in vivo and in vitro of ESCC, PKM2 positively regulates STAT3 expression and promotes malignant progression.15,16,39
Conclusions
Esophageal cancer still has a poor prognosis, and the mechanism of its invasion and metastasis is still unclear. We are the first to report the association of miR-133b, PKM2 and STAT3 in esophageal cancer. miR-133b was downregulated in ESCC, suggesting that it has the function of tumor suppressor gene. Based on our previous findings, we found that high PKM2 activity makes JAK-STAT3 pathway become PKM2-STAT3 pathway. Regarding future research directions, we plan to reveal the mechanism by which miR-133b regulates PKM2 expression and blocks the PKM2-STAT3 signaling pathway to inhibit invasion and metastasis of esophageal cancer at the cellular and animal levels. The research of this project will provide a new perspective for controlling the growth, invasion, and metastasis of esophageal cancer cells, thereby providing scientific basis for molecular prediction and targeted treatment of esophageal cancer invasion and metastasis.
Supplemental Material
sj-docx-1-onc-10.1177_11795549231219502 – Supplemental material for miR-133b Promotes Esophageal Squamous Cell Carcinoma Metastasis
Supplemental material, sj-docx-1-onc-10.1177_11795549231219502 for miR-133b Promotes Esophageal Squamous Cell Carcinoma Metastasis by Cong-Gai Huang, Qing Liu, Shu-Tao Zheng, Tao Liu, Yi-Yi Tan, Tian-Yuan Peng, Jiao Chen and Xiao-Mei Lu in Clinical Medicine Insights: Oncology
Footnotes
Author Contributions
HCG made major contributions to the data analysis and manuscript writing. LQ and ZST participated in the manuscript writing. LT and TYY collected the data. PTY and CJ analyzed the data. LXM had the main primary idea and participated in the manuscript revision. All authors discussed, carefully read, and approved the final manuscript.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was partly supported by Natural Science Foundation of China (81960527 and 82260568), Leading talent project of scientific and technological innovation in Tianshan talents training plan of Xinjiang Uygur Autonomous Region (2022TSYCLJ0031), the Natural Science Foundation of Xinjiang Uygur Autonomous Region(2022D01D69), Key Research and Development Project of the Xinjiang Uygur Autonomous Region (2020B03003-1),Xinjiang Medical University National Innovation Research Group Cultivation Project(xyd2021C001),and Project from State Key Laboratory of Pathogenesis, Prevention, Treatment of High Incidence Diseases in Central Asia (SKL-HIDCA-2022-14).
Statement of Ethics
This study protocol was reviewed and approved by the guidelines of the Institutional Ethics Committee of the Affiliated Hospital of Southwest Medical University (No. KY2022274; approval date, October 14, 2022). The patient agreed to participate in this study.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
