Abstract
Background:
SH3-domain binding protein-1 (SH3BP1), which specifically inactivates Rac1 and its target protein Wave2, has been shown to be an important regulator of cancer metastasis. However, the effects of SH3BP1 in melanoma progression remain unclear. The current study aimed to explore the function of SH3BP1 in melanoma and its possible molecular mechanism.
Methods:
TCGA database was used to analyze the expression of SH3BP1 in melanoma. Then, reverse transcription–quantitative polymerase chain reaction was performed to detect the expression of SH3BP1 in melanoma tissues and cells. Next, genes related to SH3BP1 were analyzed by LinkedOmics database, and protein interactions were analyzed by STRING database. These genes were further subjected to Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analysis. In addition, the signaling pathway of SH3BP1 action was screened by bioinformatics analysis. Finally, the function of SH3BP1 and its mediated signaling pathway in melanoma progression were investigated in vitro and in vivo.
Results:
SH3BP1 was significantly upregulated in melanoma tissues and cells. The pathways regulated by SH3BP1 are closely related to the occurrence and development of tumors. And we found that overexpression of SH3BP1 promoted the proliferation, migration, and invasion of melanoma cells by increasing Rac1 activity and Wave2 protein levels in vitro. Similarly, overexpression of SH3BP1 facilitated melanoma progression by upregulating Wave2 protein expression in vivo.
Conclusion:
In summary, this study revealed for the first time that SH3BP1 promoted melanoma progression through Rac1/Wave2 signaling pathway, providing a new therapeutic target for melanoma.
Introduction
Melanoma is a malignant tumor of pigment-producing melanocytes in the skin, which is characterized by a high degree of invasion and distant organ metastasis. 1 In recent years, it has become one of the tumors with the fastest growing incidence with an annual growth rate of approximately 3%, and the age of onset is showing a younger trend. 2 Although significant progress has been made in the diagnosis and treatment of melanoma, the prognosis of melanoma patients is still disappointing. 3 At present, the survival rate of early melanoma is reaching 90%, but the 5-year survival rate of advanced melanoma is less than 5% due to the limited treatment options. 4 Therefore, understanding the molecular mechanism of invasion and metastasis is the key to improve the prognosis of melanoma. The exploration of new potential biomarkers and effective therapeutic targets for melanoma provides a basis for improving the prognosis of patients with melanoma, which has important clinical significance.
The process of melanoma metastasis and recurrence is multi-step, complex, biological, and pathological, which involves that tumor cells break away from the tumor body, enter the systemic circulation or lymphatic system, recolonize, and grow at the distal end. 5 It has previously been reported that Kruppel-like factor 10 (KLF10) overexpression significantly inhibited the proliferation, invasion, and migration of melanoma cells and promoted their apoptosis via the PI3K/Akt signaling pathway. 6
RIG-I promotes apoptosis and inhibits proliferation by G1 phase cell cycle arrest in the melanoma cell lines via induced the phosphorylation of p38 mitogen-activated protein kinase (MAPK) and MAPK kinase 3 and MAPK kinase 4. 7 The lamellipodia is an important structure for cell migration and plays an important role in the invasion and metastasis of cancer cells. 8 Rac1 is a member of the Rho family of Guanine Trinucleotide Phosphate (GTPases), which produces sheet-like protrusions called lamellipodia. 9 Previous studies have found that Rac1 is dramatically upregulated in prostate cancer, 8 breast cancer, 10 glioma, 11 hepatocellular carcinoma, 12 and osteosarcoma, 13 and promotes the migration and invasion of tumor cells by activating the Wave2 signaling pathway complex. The research also finds that Rac/Wave2 also plays an important role in the invasion and metastasis of mouse melanoma. 14 Nevertheless, the molecular mechanism of Rac1/Wave2 migration and invasion still needs to be further elucidated.
SH3-domain binding protein-1 (SH3BP1) belongs to the RhoGAP family and is essential for cell movement because it can specifically target Rac1 GAP. 15 Recently, the role of SH3BP1 and SH3BP1-Rac1 pathway in cancer metastasis has attracted more attention. For example, in prostate cancer, overexpression of SH3BP1 can enhance the migration ability of tumor cells. 16 In hepatocellular carcinoma, through Rac1-WAVE2 signaling, SH3BP1 promotes tumor invasion and microvessel formation contributing to hepatocellular carcinoma metastasis and recurrence. 17 However, it is still not clear whether SH3BP1 participates in the regulation of the melanoma progression by activating the Rac1/Wave2 signaling pathway.
In this study, we aimed to investigate whether SH3BP1/Rac1/Wave2 axis regulated melanoma progression through bioinformatics analysis, in vitro and in vivo experiments.
Materials and Methods
Clinical sample collection
The 29 cases of melanoma tissues and 21 cases of congenital mole tissues were obtained from patients who visited the Middle Military Command General Hospital of PLA (Wuhan, China) between May 10, 2021, and December 30, 2021. All patients in this study gave informed consent and were approved by the Middle Military Command General Hospital of PLA (Wuhan, China) (approval number: [2021]016-11).
Cell lines and culture conditions
All cell lines were purchased from the Cell Bank of Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). Human epidermal melanocyte (HEMa-LP) and human melanoma cell lines (A375, A875 and SK-MEL-1) were all cultured in Dulbecco’s Modified Eagle Medium (DMEM; Gibco, USA). All media were supplemented with 10% fetal bovine serum (FBS; Gibco, USA) and 1% penicillin/streptomycin. All cells were cultured in a humidified incubator at 37 °C with 5% CO2.
Cell transfection
Lipofectamine 2000 Transfection reagent (Invitrogen, Carlsbad, CA, USA) was used to transfect the A375 and A875 cell lines according to the manufacture’s protocol. Cells were cultured in 6-well plates with 5% CO2 at 37 °C until they reached a confluence of 70% to 80% and then were transfected with si-SH3BP1 or PCMV-SH3BP1.
Bioinformatics data analysis
LinkedOmics 18 is an openly accessible database with multi-omics and clinical data on 32 cancer types and a total of 11 158 patients from The Cancer Genome Atlas (TCGA) project, offering a unique platform for biologists and clinicians to access, analyze, and compare within and across tumor types. In this study, we used LinkedOmics database to identify SH3BP1-associated genes and carry out gene expression correlation analyses of SH3BP1. Next, the STRING database was used for the protein-protein interactions of target genes, which were imported into Cytoscape. We obtained the top 10 hub genes according to the degree and calculated the enrichment modules through the Mcode in Cytoscape. Then, we conducted Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway function enrichment analyses of target genes in FunRich.
Reverse transcription–quantitative polymerase chain reaction (RT-qPCR)
RT-qPCR was performed to determine the messenger RNA (mRNA) level. 5 According to the manufacturer’s instructions, total RNAs were extracted from cells and tissues using Trizol reagent (Thermo, USA). Next, 1 µg of total RNA was reversely transcribed into cDNA using TaqMan Reverse Transcription Kit (Applied Biosystems, USA). Then, RT-qPCR was performed to detect the mRNA level by SYBR Green (Toyobo, Japan). The data were measured using the 2-ΔΔCT method and normalized by glyceraldehyde-3-phosphate dehydrogenase (GAPDH). The primers used were as follows: SH3BP1: forward 5’-GGGGTACCCCCAAGATGATGAAGAGGCAG-3’, reverse 5’-CCGCTCGAGCGGACTATCATTTGGGCAGCT-3’; GAPDH: forward 5’-GCTCTCTGCTCCTCCTGTTC-3’, reverse 5’-CGACCAAATCCGTTGACTCC-3’.
Western blot analysis
The protein expression of cells or tissues was detected by western blot analysis as previously described. 19 In brief, cells or tissue samples were collected and homogenized using radioimmunoprecipitation assay buffer. Equal total proteins (30 µg/lane) were separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (Bio-Rad, USA) and transferred onto the polyvinylidene difluoride membrane (Millpore, USA). Next, the membranes were blocked by 5% bovine serum albumin (BSA) for 2 hours at room temperature, followed by incubation with primary antibodies against Wave2 (1:500, ABclonal, China) and GAPDH (1:1000, ABclonal, China) at 4 °C overnight. After washing with 1× phosphate-buffered saline (PBS) for 3 times, the membranes were incubated with HRP-conjugated secondary antibodies (1:5000, ABclonal, China) for 2 hours at room temperature. The bands were then developed using enhanced chemiluminescence chromogenic substrate (GE Healthcare, UK) and analyzed by the Image J software. GAPDH was used as a control in this study.
Glutathione (GST) pull-down analysis
According to the previous description, GST pull-down experiment was used to measure the level of GTP-bound small GTPase when NSC23766 (Rac1 inhibitor, 100 µg/mL) was added or not. 20 Cell lysates were ice lysed for 30 minutes at 4 °C, 12 000 × g, centrifuged for 5 minutes. Then these cell lysates were transferred to a new EP tube, glutation-Sepharose GST-fusion protein was added, and incubated at 4 °C for 1 hour. After that, the cells were washed with GST-Fish buffer for 3 times. After centrifugation at 2000 × g at 4 °C for 3 minutes, GST-Fish buffer was resuspended and 4× protein loading buffer was added for western blot analysis.
Cell counting kit-8 (CCK-8) analysis
The rate of the proliferation and inhibition of melanoma cells was measured by CCK-8. In brief, the cells were seeded on 96-well plates (5000 cells/well) and subjected to different treatments. Next, the cells were treated with 10 μL of CCK-8 solution (Beyotime, China) at 37 °C for 1 hour. The optical density (OD value) of each well was measured by a microplate reader at 450 nm.
Wound healing assay
The migration ability of melanoma cells was assessed using the transwell system (Costar, USA) and scratch wound healing assay as previously described with some modifications. 21 Briefly describing the transwell migration experiment, after different treatments, 1 × 105 cells were seeded onto the upper chamber and incubated with the serum-free medium. The medium containing 10% FBS was added to the lower chambers. After 48 hours, the migrated cells were fixed with 4% paraformaldehyde, stained with 10% crystal violet at room temperature, and observed under a light microscope in 5 random fields (100×). Briefly describing the scratch experiment, the cells were plated into 6-well culture plates and cultured until 100% confluence. A wound was produced with a sterile plastic tip (0.5 mm), and then the cells were washed and cultured in the fresh medium. The level of migration in each group at 0, 24, and 48 hours was evaluated using photographs (CKX41, Olympus).
Transwell assay
The melanoma cells were collected, digested with trypsin, and then resuspended in a serum-free medium to adjust the concentration to 2 × 104 cells/mL. After adding the DMEM culture medium to dilute Matrigel at a ratio of 1:5, we spread it on the upper chamber of the transwell chamber, dried it at room temperature, and followed the procedure of the cell migration experiment. Finally, the number of cells which stained with crystal violet under the microscope was the number of invaded cells.
Transient transfection assay
The SH3BP1 small interfering RNA (siRNA) and overexpression plasmid (PCMV-SH3BP1) all came from GenePharma (Shanghai, China). Melanoma cells were transfected with them using Lipofectamine 2000 (Thermofisher, USA) according to the manufacturer’s instructions.
Establishment of tumorigenicity analysis in mice
Animal care and euthanasia were carried out with the approval of the Laboratory Animal Management and Use Committee of Hubei Center for Disease Control and Prevention on October 30, 2020 (Wuhan, China) (approval number: 20201030). Melanoma cells were transfected with a SH3BP1 PCMV plasmid and then injected subcutaneously into the right back flank of 4- to 5-week-old female nude mice (Nanjing Biomedical Research Institute of Nanjing University) (5×106 cells/mouse, 4 mice/group). Mice were killed by cranio-cervical dislocation.
Immunohistochemical analysis
Immunohistochemical (IHC) analysis was performed as previously described. 22 The tumor tissues were collected and perfused with 10% formaldehyde solution for 24 hours at 4 °C, followed by embedment in paraffin, which was then cut into sections with a thickness of 5 μm. The sections were deparaffinized, rehydrated, heated, and blocked the nonspecific antigen with 5% BSA for 1 hour. Then the sections were incubated with primary antibodies for ki67 (1:100, ABclonal, China) and MMP9 (1:100, ABclonal, China) at 37 °C overnight, followed by visualized using the DAKO REAL EnVision Inspection System (DAKO, Denmark). All images were observed under a light microscope (Nikon, Japan).
Statistical analysis
The data were expressed as mean ± SE. Student’s t test was performed to determine the significance between 2 groups. One-way or 2-way analysis of variance with Bonferroni post hoc tests were for multiple groups.
Results
Expression of SH3BP1 in melanoma tissues and cells
As shown in Figure 1A, we first searched the mRNA expression of SH3BP1 in melanoma tissues and normal tissues through the TCGA database. The results showed that the mRNA expression of SH3BP1 in melanoma tissues was higher than that in normal tissues. Then, to verify the role of SH3BP1 in melanoma, we further evaluated the expression of SH3BP1 in clinical tissue samples. We detected the mRNA expression levels of SH3BP1 in 21 cases of congenital mole tissues and 29 cases of melanoma tissues by RT-qPCR. The results showed that the expression level of SH3BP1 in the melanoma group tissue was significantly upregulated (Figure 1B) compared with the normal control group. Finally, SH3BP1 expression was also detected in melanoma cells, which included A375, A875, and SK-MEL-1. As shown in Figure 1C, compared with the normal melanocyte group, the expressions of SH3BP1 in A375, A875, and SK-MEL-1 were significantly upregulated. Moreover, the expression level of SH3BP1 in A375 and A875 was upregulated more significantly. Thus, we chose A375 and A875 for subsequent cell experiments.

Expression of SH3BP1 in melanoma tissues and cells. (A) TCGA database analyzed the mRNA expression of SH3BP1 in cutaneous melanoma and normal tissues. (B) RT-qPCR was used to detect the expression of SH3BP1 in congenital mole (n = 21) and melanoma (n = 29). (C) RT-qPCR was used to detect the expression of SH3BP1 in melanocytes (HEMa-LP) and melanoma cells (A375, A875 and SK-MEL-1).
Bioinformatics data analysis of SH3BP1-related genes and enriched pathways
In this study, we first used LinkedOmics database to identify SH3BP1-associated genes and carry out gene expression correlation analyses of SH3BP1. As shown in Figure 2A, we performed gene expression correlation analyses on genes related to SH3BP1 and drew a Volcano Plot. Figure 2B and C shows the Heatmaps of the genes associated with SH3BP1. Figure 2B shows that the genes were positively correlated with SH3BP1, while Figure 2C shows that they were negatively correlated. The correlation coefficient|R|> 0.55 genes which positively or negatively related to SH3BP1 were analyzed for protein-protein interaction analysis. Then, these 285 genes were introduced into STRING, among which the protein interactions network was observed through cytoscape, which was then used for mapping to obtain the protein interactions between 100 genes (Figure 3A). With the hubba plug-in in Cytoscape, 10 hub genes were obtained by degrees method. The Mcode plug-in in Cytoscape was used for module analysis with the condition of Degree Cutoff. Among these interacting proteins, we also found that SH3BP1 regulated the Rac1 signaling pathway, which was consistent with previous reports (Figure 3B and C). Finally, we further performed GO and KEGG pathway enrichment analysis for these genes (Figure 3D). As shown in Table 1, these genes were mainly enriched in signal pathways which related to cancer, such as MAPK and Ras, and were closely related to DNA replication and cell apoptosis.

Genes differentially expressed in association with SH3BP1 in melanoma according to LinkedOmics. (A) The correlation between SH3BP1 and genes differentially expressed in melanoma was evaluated by Pearson test and presented by volcano plot. (B) Heatmap of differentially expressed genes showed the top 50 significant genes positively and (C) negatively correlated with SH3BP1 in melanoma.
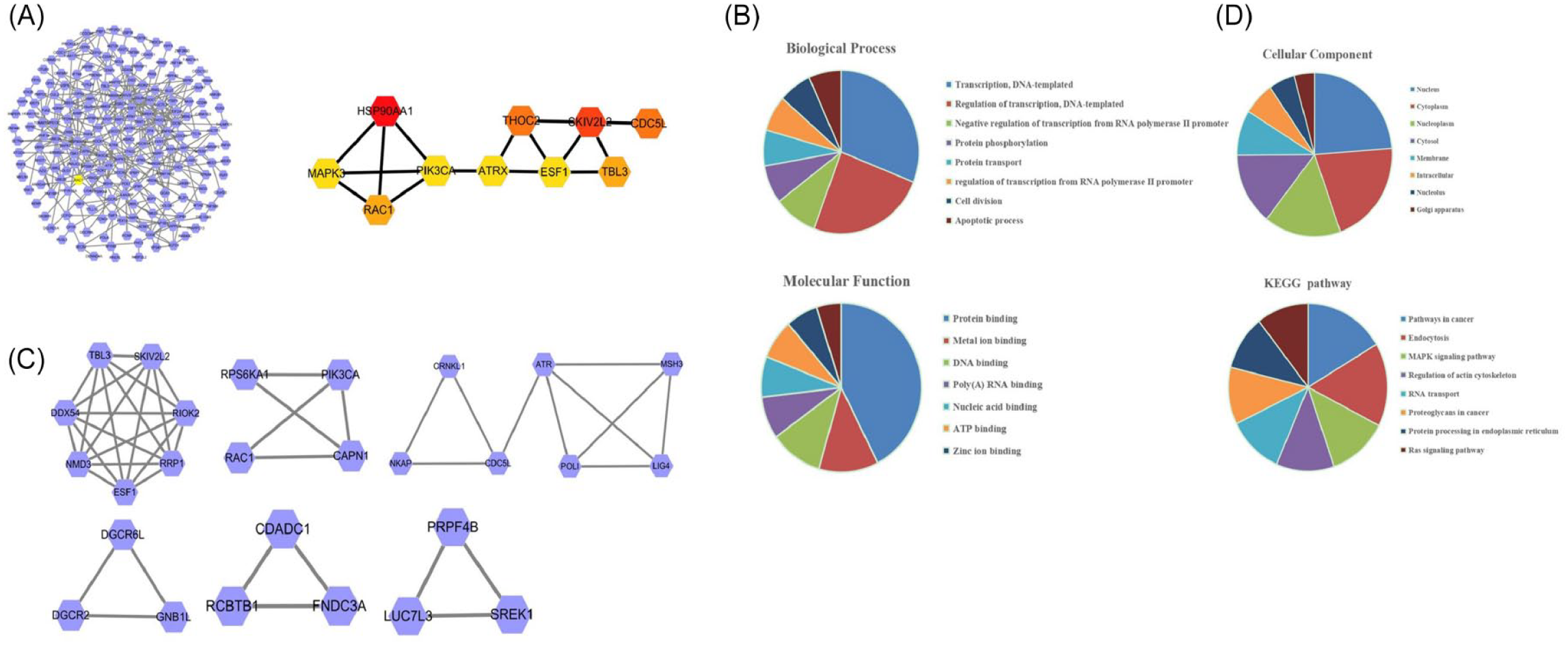
Bioinformatics data analysis of SH3BP1 enriched pathways. (A) STRING software constructed the PPI network. (B, C) Genes involved in 3 Mcodes as analyzed by Cytoscape. (D) The significantly enriched GO and KEGG pathways of SH3BP1 co-expression genes in melanoma were analyzed.
Enrichment analysis of SH3BP1-associated genes.

Abbreviations: ATP, adenosine triphosphate; MAPK, mitogen-activated protein kinase; SH3BP1, SH3-domain binding protein-1.
The effects of SH3BP1 on proliferation, migration, and invasion of melanoma cells
To explore the role of SH3BP1 in melanoma metastasis, we evaluated the function of SH3BP1 in the proliferation, migration, and invasion of melanoma cells. The PCMV-SH3BP1 or si-SH3BP1 was separately employed to upregulate or downregulate SH3BP1 in A375 and A875 cells. Its knockdown or overexpressed efficiency was evaluated by RT-qPCR assay, separately. The interference and overexpression efficiency of SH3BP1 in A375 and A875 cells were significant (P < .05). As shown in Figure 4B, compared with the control group (si-NC), the proliferation ability of melanoma cells (A375 and A875) in the SH3BP1 knockdown group (si-SH3BP1) was significantly downregulated. In contrast, the proliferation ability of melanoma cells in the SH3BP1 overexpression group (PCMV-SH3BP1) was significantly upregulated. Then, we evaluated the migration ability of melanoma cells through the scratch wound healing assays and transwell invasion assays. As shown in Figure 4C and D, compared with the si-NC group, the migration ability of melanoma cells in the si-SH3BP1 group was dramatically downregulated. Nevertheless, the migration ability of melanoma cells in the PCMV-SH3BP1 group was substantially upregulated. Finally, we also evaluated the invasion ability of melanoma cells through the transwell invasion experiment. As shown in Figure 4D, the invasion ability of melanoma cells in the si-SH3BP1 group was significantly downregulated compared with the si-NC group. However, the migration ability of melanoma cells in the PCMV-SH3BP1 group was markedly upregulated.
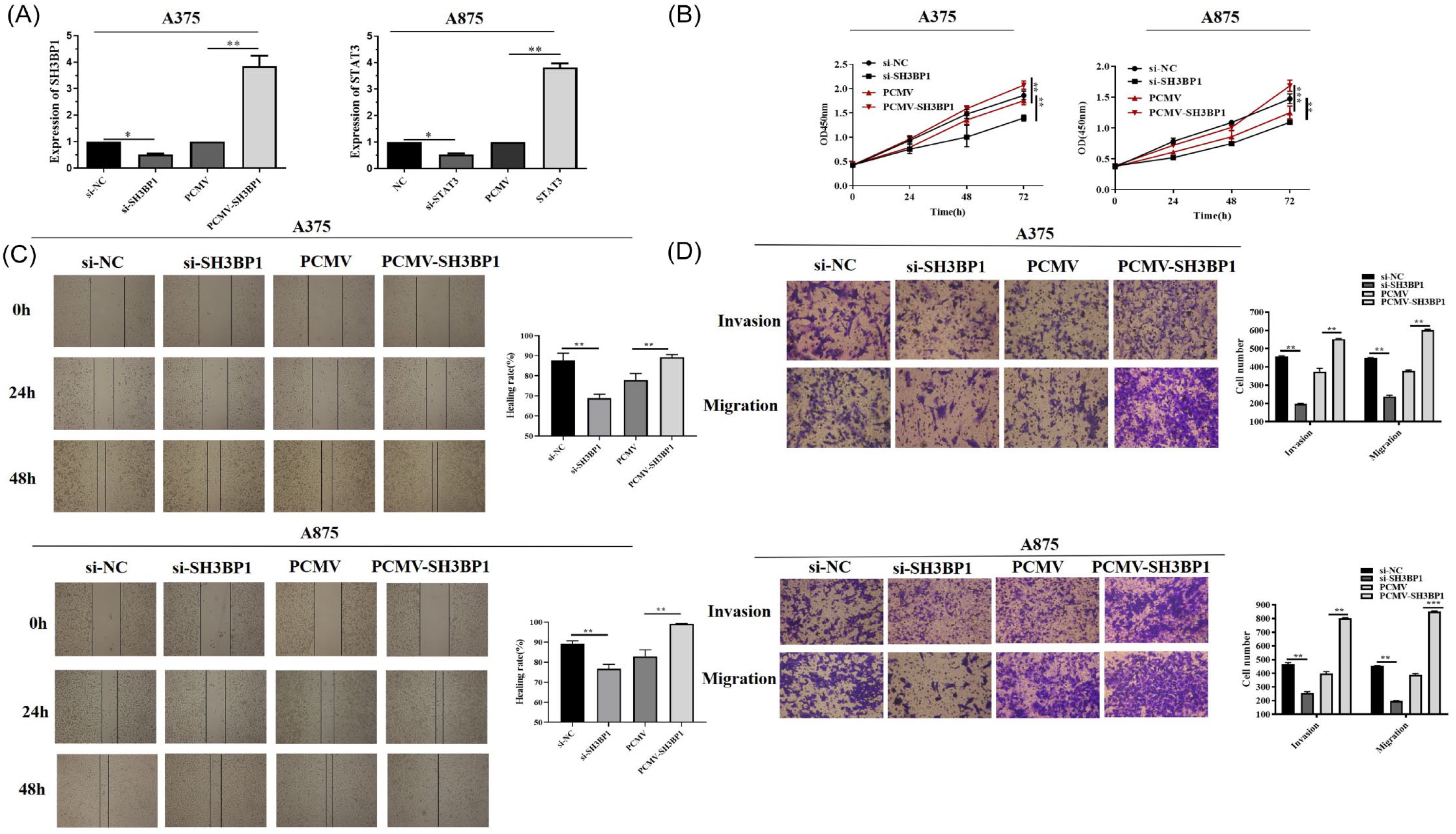
The effects of SH3BP1 on the proliferation, migration, and invasion of melanoma cells. (A) Cell transfection efficiency with si-SH3BP1 and PCMV-SH3BP1 was evaluated by RT-qPCR assay. (B) Cell viability of A375 and A875 was measured by CCK-8. A375 and A875 were transfected with control siRNA (si-NC), SH3BP1 siRNA (si-SH3BP1), control plasmid (PCMV) and PCMV-SH3BP1 plasmid (PCMV-SH3BP1). (C) The transverse migration of cells transfected with si-NC, si-SH3BP1, PCMV, and PCMV-SH3BP1 as determined by scratch wound healing assays in A375 and A875 at a magnification of 40×. (D) The invasion and vertical migration of cells transfected with si-NC, si-SH3BP1, PCMV, and PCMV-SH3BP1 as determined by transwell assays in A375 and A875.
SH3BP1 activated Rac1 to advance the expression of Wave2 in melanoma
As we all know, SH3BP1 affects cell behavior and maintains cell motility by regulating the Rac1 signaling pathway. 15 Therefore, we first verified whether Rac1 was involved in the invasion and migration of melanoma through GST pull-down analysis. As shown in Figure 5, the activity of Rac1 was prominently improved by SH3BP1, whereas dramatically reduced by NSC23766. The promotion of Rac1 activity by SH3BP1 could be partially inhibited by NSC23766 in both A375 and A875. Furthermore, we also tested the protein expression level of Wave2. The results showed that the expression level of Wave2 protein in the SH3BP1 overexpression group was significantly upregulated compared with the control group, whereas memorably reduced by NSC23766. The addition of NSC23766 treatment partially inhibited the promotion of SH3BP1 on the expression of Wave2 protein. These results were consistent with the changing trend of Rac1 expression.

SH3BP1 activated Rac1 to advance the expression of Wave2 in melanoma. GST pull-down analysis was used to determine the activity of Rac1 in A375 (A) and A875 (B) in response to co-processing SH3BP1 overexpression and Rac1 inhibitor NSC23766 (100 μg/mL). Western blot analysis was used to determine the protein level of Wave2 in A375 and A875 in response to co-processing SH3BP1 overexpression and Rac1 inhibitor NSC23766 (100 μg/mL).
SH3BP1 participated in the regulation of the proliferation, migration, and invasion of melanoma cells by regulating the Rac1/Wave2 signaling pathway
We have clarified that SH3BP1 promoted the proliferation, migration, and invasion of melanoma cells and clarified its activation of the Rac1/Wave2 signaling pathway. Therefore, we further explored the possibility of Rac1/Wave2 signaling pathway participating in SH3BP1 regulation of the proliferation, migration, and invasion of melanoma cells. As shown in Figure 6A, compared with the control group (PCMV + PBS), the proliferation ability of melanoma cells (A375 and A875) in the SH3BP1 overexpression group (PCMV-SH3BP1 + PBS) was significantly upregulated, whereas dramatically reduced by NSC23766. The promotion of proliferation ability by SH3BP1 could be partially inhibited by NSC23766 in both A375 and A875. Then, we evaluated the migration ability of melanoma cells through the scratch wound healing assays and transwell invasion assays. As shown in Figure 6B and C, compared with PCMV + PBS group, the migration ability of melanoma cells in the PCMV-SH3BP1 + PBS group was dramatically upregulated, whereas dramatically reduced by NSC23766; the promotion of migration ability by SH3BP1 could be partially inhibited by NSC23766 in melanoma cells. Finally, we also evaluated the invasion ability of melanoma cells through the transwell invasion experiment. As shown in Figure 6C, the invasion ability of melanoma cells in the PCMV-SH3BP1 + PBS group was significantly upregulated compared with the PCMV + PBS group, whereas dramatically reduced by NSC23766. The promotion of invasion ability by SH3BP1 could be partially inhibited by NSC23766 in melanoma cells.

SH3BP1 participated in the regulation of the proliferation, migration, and invasion of melanoma cells by regulating the Rac1/Wave2 signaling pathway. (A) Cell viability of A375 and A875 in response to co-processing SH3BP1 overexpression and Rac1 inhibitor NSC23766 (100 μg/mL) was measured by CCK-8. (B) The transverse migration of cells in response to co-processing SH3BP1 overexpression and Rac1 inhibitor NSC23766 (100 μg/mL) as determined by scratch wound healing assays in A375 and A875 at a magnification of 40×. (C) The invasion and vertical migration of cells in response to co-processing SH3BP1 overexpression and Rac1 inhibitor NSC23766 (100 μg/mL) as determined by transwell assays in A375 and A875.
SH3BP1 affected tumor growth of melanoma in vivo
We have previously found that SH3BP1 inhibited the proliferation, migration, and invasion of cells in vitro. Therefore, we further evaluated whether SH3BP1 could affect tumor growth of melanoma in vivo. Melanoma cells were transfected with SH3BP1 plasmid and implanted subcutaneously into severe combined immunodeficiency mice to allow tumor formation. Tumor sizes were measured every 7 days, and the mice were killed 5 weeks later. As shown in Figure 7A to C, the xenograft tumors were significantly bigger in the SH3BP1-overexpression mice group compared with the control group. Moreover, we also detected the expression of Wave2 protein in tumor tissues by western blot analysis. The results showed that overexpression of SH3BP1 significantly increased the expression level of Wave2 in vivo (Figure 7D). Furthermore, we also detected the expression of ki67, an indicator of cell proliferation and MMP9 and an indicator of migration/invasion through IHC. The results showed that overexpression of SH3BP1 markedly increased the ability of migration and invasion in vivo (Figure 7E).

SH3BP1 promoted the growth of melanoma by promoting the expression of Wave2. (A) Optical images of tumor tissues dissected from A375 tumor-bearing mice after 16 days. (B) Relative tumor growth curves. (C) Tumor weight. (D) Western blot analysis was used to detect the expression of Wave2 protein in tumor tissues from A375 tumor-bearing mice. (E) IHC of tumor tissue sections dissected from A375 tumor-bearing mice.
Discussion
In this article, we confirmed that SH3BP1 was significantly upregulated in melanoma tissues and melanoma cell lines, which was consistent with the predicted results in the TCGA database. The LinkedOmics database was used to analyze the positively/negatively related genes of SH3BP1, and the selected genes were imported into the STRING database for protein interaction analysis. The results showed that SH3BP1 regulated the Rac1 signaling pathway. Then, we clarified that SH3BP1 promoted the proliferation, migration, and invasion of melanoma cells by activating Rac1 and promoting the protein expression of Wave2. Finally, we clarified that SH3BP1 regulated tumor growth by promoting the protein expression of Wave2 in vivo.
SH3BP1, which was originally identified by SH3 domain screening, is a GAP protein. 23 As a key effector of Sema3E-Plexind1 signaling, SH3BP1 is involved in signal integration, cytoskeleton regulation, and cell behavior regulation. 24 A large number of studies have shown that tumor metastasis can be attributed to the migration of cancer cells to a large extent and the change in cell movement determines the ability of cell migration. 25 Melanoma is the most malignant tumor of the skin and remains the leading cause of skin cancer–related death due to its highly metastatic nature. 26 Studies have shown that the proliferation, migration, invasion, and angiogenesis of tumor cells are closely related to melanoma metastasis.5,27 More recently, the overexpression of SH3BP1 in cancer has been widely reported and is closely related to the occurrence and development of tumors. For example, SH3BP1 is highly expressed in gastric cancer and is considered to be one of the independent prognostic factors for the poor prognosis of gastric cancer patients. 28 In this study, we also found that the expression of SH3BP1 in melanoma tissues was significantly upregulated by TCGA database analysis. The expression level of SH3BP1 was further detected in tissues and cells and found to be consistent with previous studies.
Due to its important role in cell movement, SH3BP1 overexpression is usually associated with tumor metastasis. Previous studies have shown that SH3BP1 participates in the regulation of breast cancer metastasis by regulating Epithelial-Mesenchymal Transition. 29 SH3BP1 in tumor microenvironment promotes liver cancer metastasis through Wave2 pathway. 17 In this study, we demonstrated that overexpression of SH3BP1 increased the proliferation, migration, and invasion ability of melanoma cells. On the contrary, knocking down SH3BP1 reduced the proliferation, migration, and invasion ability of melanoma cells.
Next, we explored that how SH3BP1 was involved in the regulation of melanoma. Previous studies have indicated that SH3BP1 promotes the secretion of vascular endothelial growth factor by regulating the Rac1/Wave2 signaling pathway, thereby enhancing the microvascular formation and infiltration of liver cancer cells. 17 In cervical cancer, SH3BP1 participates in the regulation of cell migration, invasion, and cisplatin resistance by activating the Rac/Wave2 pathway. 20 In the present study, we also found protein interaction between SH3BP1 and Rac1 pathway through bioinformatics data analysis of LinkedOmics database and STRING database. Further through GST pull-down experiment and western blot analysis, we clarified that SH3BP1 activated Rac1 and promoted the protein expression of Wave2. However, the activation of SH3BP1 can be partially reversed by NSC23766, which moreover also partially reversed the effects of SH3BP1 on the proliferation, migration, and invasion of melanoma cells, revealing that the Rac1/Wave2 signaling pathway was involved in the regulation of SH3BP1 on the invasion of melanoma. Finally, we measured the effect of SH3BP1 on the growth of melanoma in vivo.
Conclusion
In summary, the present study first described the mechanism of SH3BP1 in regulating the melanoma progression via Rac1/Wave2 pathway. This provided a new perspective on the treatment and prognosis of melanoma.
Footnotes
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Author Contributions
Conceptualization: Jie Pu, Ting Sun, and Zhiguo Yu.
Data curation: Wenxian Tong.
Formal analysis: Jie Pu.
Investigation: Ting Sun.
Methodology: Zhiguo Yu.
Literature research: Wenxian Tong.
Supervision: Zhengchun Kang.
Writing—original draft: Jie Pu, Ting Sun, and Zhiguo Yu.
Writing—review & editing: Zhengchun Kang.
Ethics Statement
All protocols of patient samples were approved by the Middle Military Command General Hospital of PLA (Wuhan, China).
Data Availability
Publicly available data sets were analyzed in this study.
