Abstract
Introduction:
Glioblastoma multiforme (GBM) is one of the deadliest cranial tumors occurring in adults. Various biomarkers have been tested for their significance in diagnosis, prognosis, and treatment of GBM. Some well-studied markers in GBM are Isocitrate dehydrogenase 1 (IDH1), Murine double minute 2 (MDM2), Epidermal Growth Factor Receptor (EGFR), and p53. The aim of this study was to investigate the protein expression of these markers in GBM patients of Pakistan.
Methods:
A total of 102 surgically resected formalin-fixed paraffin-embedded specimens from patients diagnosed and treated at Aga Khan University Hospital were included in this study. Immunohistochemistry (IHC) for IDH1, MDM2, EGFR, and p53 was performed using Dako EnVision System and respective monoclonal antibodies. Survival analysis was performed to check association of markers protein expression with prognosis in GBM patients.
Results:
There were 73 males and 29 females in this study, with a median age of 49 years at the time diagnosis. Overexpression of molecular markers was as follows: 52% for EGFR, 26% for p53, 72% for IDH1, and 83% for MDM2. We did observe that EGFR was significantly associated with increased age of our patients and with worse survival. Age > 40 years was a predictor for worse prognosis as well.
Conclusion:
EGFR overexpression and advanced age were worse prognostic indicators.
Introduction
Glioma is the most common and aggressive brain tumor in adults, originating from glial cells. It represents around 30% of central nervous system tumors, 45% of malignant, and 80% of primary malignant central nervous system tumors. There are different types of gliomas based on the type of cells involved, such as astrocytomas, ependymomas, and oligodendrogliomas, which are further graded as I, II, III, and IV according to the World Health Organization classification system. The most common form of glioma is Glioblastoma Multiforme (GBM), classified as grade IV, and it accounts for 70% of astrocytomas and 15% of all other intracranial neoplasms. 1 GBM is a rare tumor with a global incidence of approximately 10 per 100 000 people. Despite the advances of current therapies for GBM, it is still a lethal disease with a poor prognosis. Because of the aggressive nature of GBM, patients have a bleak median survival of only about 14 to 15 months following diagnosis. 2
GBM is categorized as an aggressive malignancy and is thought to arise from genetic aberrations and deregulation of signaling pathways. 3 The existing treatment strategy for GBM is surgery, chemotherapy, and radiotherapy. Most recent treatments include the use of monoclonal and polyclonal antibodies as immunotherapy. The function of such therapies is to target specific cancer pathways and these are less toxic to normal cells as compared to other types of treatments, such as chemotherapy and radiotherapy. 4 For diagnostic or therapeutic purposes, many important biological markers have been identified, such as epidermal growth factor receptor (EGFR), p53, murine double protein 2 (MDM2), and isocitrate dehydrogenase1 (IDH1). Therefore, it is necessary to discover effective therapeutic targets and agents for the treatment of GBM patients.
EGFR is a transmembrane glycoprotein receptor which plays a vital role in cell growth and the development of tumor-like complex signaling cascades that modulate growth, signaling, differentiation, adhesion, migration, and survival of cancer cells. It is involved in the progression of many cancers including breast, lung, esophageal, and head and neck cancers. 5 In astrocytic tumors, overexpression of EGFR is reported to be 90% and approximately 40% to 50% of primary glioblastomas harbor mutated EGFR. 1
The P53 gene, located on human chromosome 17p13 (p53), is a vital tumor suppressor protein that acts as a sensor of cellular stress and influences many signaling pathways, such as cell cycle arrest, apoptosis, senescence, and differentiation. 6 The expression of p53 gene was shown to be upregulated in many cancers, such as endometrial cancer, colorectal cancer, head and neck squamous cell carcinoma, and breast cancer.7-10 Likewise, the network of p53 target genes also acts as a regulator of aging and cancer prevention. 11 Existing studies have also reported that p53 plays a significant role in regulating glioma development and progression. 12
The Murine Double Minute 2 (MDM2) gene is an oncogene that is amplified in a variety of human cancers. 13 The p53 protein regulates the expression of MDM2, and MDM2 is essential for the negative regulation of p53 as well. It is an E3 ubiquitin ligase, localized in the nucleus, located on chromosome 12 (12p14.3-q15). MDM2 protein seizes p53 expression via direct binding with its E3 ubiquitin ligase activity. The correlation between p53 and MDM2 is particularly important for the proliferation of tumor growth. 14 Modifications in MDM2 gene may be found in both primary and secondary GBM. In primary GBMs, it is amplified in about 10% of the cases and overexpressed in 50%, while in secondary GBMs, it is overexpressed in 10% of tumors. 3
The IDH genes encode 2 metabolic enzymes, namely IDH 1 and IDH 2, present in the cytoplasm and mitochondria and located on the long arm of Chromosome 2 (2q33.3). They carry out decarboxylation of isocitrate to α-ketoglutarate by reducing NADP in the cytosol. These metabolic enzymes are essential for cellular respiration. IDH1 mutation is commonly found in low-grade glioma and about 90% of high-grade glioma (GBM) contains wild-type IDH1 and the remaining are mutant IDH1. The mutation in IDH1 is thought to be a prognostic marker for glioblastoma patient survival. 15 The most common IDH1 mutation is R132H, a novel point mutation in exon 4 codon 132 which replaces amino acid arginine with histidine. Another lesser common mutation is R172 which occurs in IDH2. IDH1/2 mutations may be loss of function or gain of function. It is theorized that once mutated, the IDH1/2 enzyme loses its affinity to substrate IDH for conversion to α-ketoglutarate and instead gains a neomorphic enzymatic affinity for conversion of α-ketoglutarate to oncometabolite D-2-hydroxyglutarate which in turn causes several genome-wide epigenetic modifications and progression of glioma. 16
The existing data on the genetics of Pakistani GBM patients is very limited. Understanding the genetics of a patient population is crucial for an accurate and comprehensive diagnosis, such as using molecular classification for gliomas, and also for applying latest treatment options, such as targeted therapy. Keeping in mind the current need of therapeutic targets for GBM patients and low survival rate, the objective of our study was to investigate the protein expression of different prognostic markers p53, EGFR, IDH1, and MDM2 and to correlate their protein expression with patient characteristics and overall survival of Pakistani GBM patients.
Materials and Methods
Study subjects and samples
Medical records from a total of 102 patients admitted to Aga Khan University Hospital Karachi, Pakistan were reviewed. Patients who were admitted between January 2015 and December 2016 with biopsy-proven GBM were selected. The samples and medical records were accessed from November 2017 to October 2019. Inclusion criteria were complete demographic, clinicopathologic data, adequate follow-up, and availability of sufficient paraffin-embedded tumor material. Patients and surgical samples that did not fulfill all the listed criteria were excluded from the study. All the samples diagnosed or treated in our institute, including these specimens, were stored according to the institutional policies for storing specimens. Our institute is accredited by Joint Commission International (JCI) and College of American Pathologists (CAP), and the appropriate storage practices according to their guidelines are to store specimens for 10 years. The specimens were used and returned to the institutional storage.
Ethical considerations
This study was approved by the Ethical Review Committee of Aga Khan University (ERC#2018-0511-808). All patients provided written informed consent for participation in the study.
Immunohistochemistry
Tissue sections (3-5 μm thick) were obtained from formalin-fixed paraffin-embedded tissue blocks for immunohistochemical staining and mounted on precoated FLEX IHC Microscope Slides (Dako, Denmark). The sections were dried at 55°C for 30 minutes. These sections were deparaffinized in xylene and rehydrated through serial-graded (100%, 90%, 80%, and 70%) water–ethanol solutions and then rinsed in distilled water. The Envision FLEX + Visualization kit, Mouse, High pH (K800221, Dako, Denmark) was used for performing immunohistochemistry (IHC) for all antibodies. To unmask the antigen, the slides were treated with Target Retrieval Solution, High pH (50x Tris/EDTA buffer, pH 9) (Dako, Denmark) in a preheated (90°C-95°C) water bath for 20 minutes, while proteinase K was added for 30 mins for EGFR only. The slides were then allowed to cool for 15 to 20 minutes at room temperature. After antigen retrieval, endogenous peroxidase was blocked with Peroxidase-Blocking Reagent (Dako, Denmark) for 10 minutes. The slides were rinsed well with Tris Buffer Saline with Tween 20 (TBST) (Dako, Denmark). The sections were then incubated for 30 minutes with Primary Antibodies against IDH1, EGFR, MDM2, and p53 at room temperature and rinsed with TBST buffer. Anti-human primary antibodies against the following antigens were used: mouse monoclonal p53 (Dako, clone DO-7; 1:50 dilution), mouse monoclonal EGFR (Dako, clone H11; 1:200 dilution), rabbit polyclonal IDH1 (Abcam, ab81653; 1:200 dilution), and mouse monoclonal MDM2 (Abcam, clone 2A10, ab16895; 1:100 dilution). Afterwards, these sections were treated with secondary antibody EnVision FLEX + Mouse (LINKER) (Dako, Denmark), and bound antibody was identified using the polymer technology EnVision/HRP, DAB + Chromogen (Dako, Denmark). Slides were then counterstained with hematoxylin for 30 seconds. After counterstaining, the specimens were dehydrated in a graded water–ethanol series and mounted with Dako Toluene-Free Mounting Medium (Dako, Denmark) and observed under light microscope.
Evaluation of slides
The stained sections were observed microscopically by 2 pathologists, blinded to each other and the case history. At least 200 tumor cells from 5 to 10 different fields each were observed. This ensured that thousands of cells from multiple areas of the tissue were reviewed before assigning a score. Results were based on a combination of intensity of staining (mild, moderate, or strong) and the percentage of cells stained. The following criteria were used for scoring the slides: 0 / negative (when no staining was observed), 1 (< 5% cells are stained), 2 (5%-50% cells stained), and 3 (> 50% of cells stained).17,18
Statistical analysis
SPSS version 22 (SPSS Inc, Chicago, IL, USA) was used for analysis. Descriptive analysis was reported as mean ± SD or median, depending on the normality of the population. Differences between categorical variables (biomarker + and biomarker −) were assessed using Pearson’s Chi-Square test or Fisher’s Exact test, where appropriate. For overall survival analysis, only those patients with known status (dead or alive) and follow-up data were included. Follow-up was taken in months from the date of surgery until last follow-up, or death, whichever occurred first. Patients who were alive at last contact were considered censored observation for survival analysis. Kaplan–Meier survival curves and log-rank tests were used to compare survival between 2 groups. A P-value of less than .05 was considered significant for all tests.
Results
A total of 102 GBM patients (73 males, 29 females) were included, with a mean age of 47.35 ± 16.129 and a median age of 49 years at the time of diagnosis. The minimum and maximum patient ages were 20 and 81 years, respectively. The site of lesion was categorized as frontal (n = 11), temporal (n = 27), occipital (n = 2), parietal (n = 14), frontoparietal (n = 6), frontotemporal (n = 3), temporoparietal (n = 6), and parieto-occipital (n = 3). For 30 patients, the site of lesion was unspecified or other than the ones listed above.
Figure 1 shows strong positive expression of all markers observed in GBM biopsies as brown coloration. EGFR positivity displayed as membranous and cytoplasmic staining, p53 positivity was exclusively nuclear, while IDH1 and MDM2 positivity was cytoplasmic, and in some cases, nuclear positivity was also observed for MDM2.
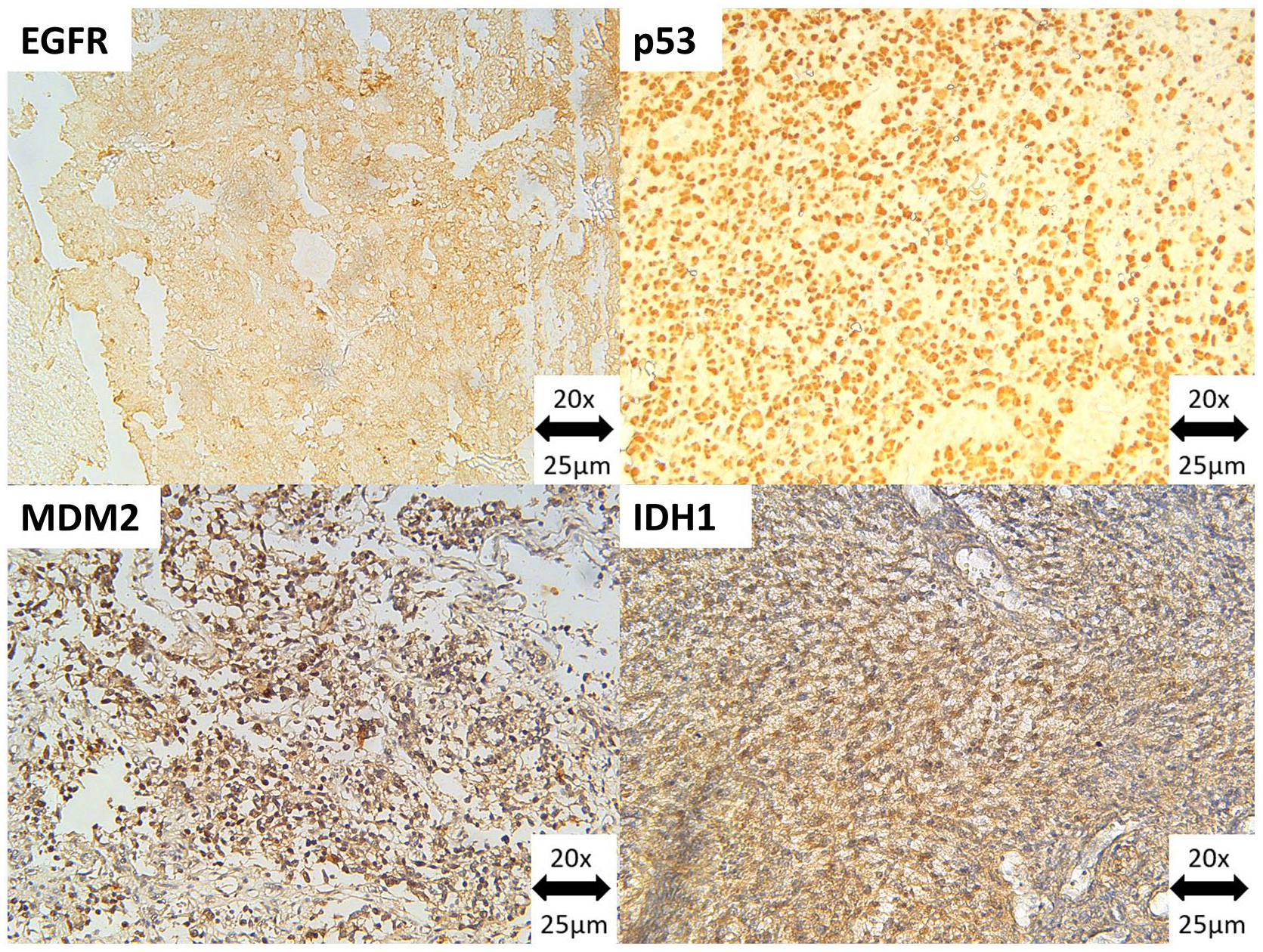
Strong immunohistochemical positivity for (A) EGFR in GBM, observed as membranous and cytoplasmic brown coloration, (B) p53 in GBM, observed as nuclear brown coloration, (C) IDH1 in GBM, observed as cytoplasmic brown coloration, (D) MDM2 in GBM, observed as cytoplasmic (and in some cases nuclear) brown coloration at 20X magnification.
The positive overexpression of molecular markers was found to be 53/102 (52%) for EGFR, 27/102 (26%) for p53, 73/102 (72%) for IDH1, and 85/102 (83%) for MDM2. Staining intensity was noted as follows: for EGFR, there were 13 mild, 23 moderate, and 17 strong positives; for p53, there were 14 mild, 17 moderate, and 10 strong positives; for IDH1, there were 21 mild, 36 moderate, and 16 strong positives; and for MDM2, there were 19 mild, 38 moderate, and 28 strong positives. For p53 statistical analysis, the mild positives were merged with negative results. This was done so that any cells expressing p53 normally/wild type (which shows as weak positivity in a few scattered cells) would not be counted among those tissues that are overexpressing p53, as previously performed. 18 Correlations among markers and patient characteristics have been listed in Tables 1 to 4. The only significant association found was between age and EGFR status (P < .001) which showed that EGFR was overexpressed with increasing age in our patients.
Patient characteristics in relation to p53 status.

Abbreviations: CI, confidence interval; EGFR, epidermal growth factor receptor; IDH, isocitrate dehydrogenase; MDM, murine double minute; OR, odds ratio.
P-value calculated using Pearson’s Chi-Square test or Fisher’s Exact test, where appropriate.
P-value considered significant at < .05.
Patient characteristics in relation to EGFR status.

Abbreviations: CI, confidence interval; EGFR, epidermal growth factor receptor; IDH, isocitrate dehydrogenase; MDM, murine double minute; OR, odds ratio.
P-value calculated using Pearson’s Chi-Square test or Fisher’s Exact test, where appropriate.
P-value considered significant at < .05.
Patient characteristics in relation to IDH1 status.

Abbreviations: CI, confidence interval; EGFR, epidermal growth factor receptor; IDH, isocitrate dehydrogenase; MDM, murine double minute; OR, odds ratio.
P-value calculated using Pearson’s Chi-Square test or Fisher’s Exact test, where appropriate.
P-value considered significant at < .05.
Patient characteristics in relation to MDM2 status.
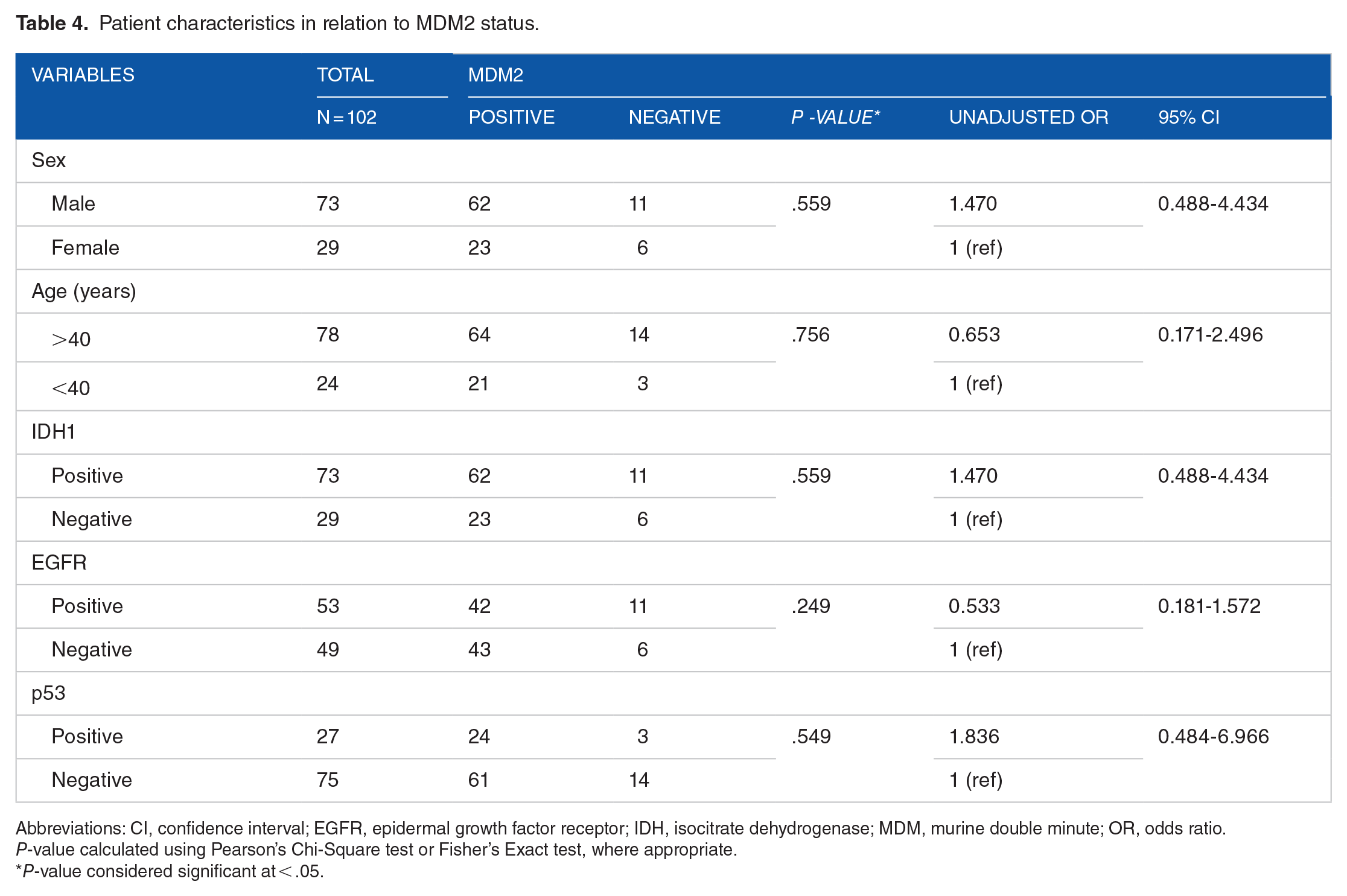
Abbreviations: CI, confidence interval; EGFR, epidermal growth factor receptor; IDH, isocitrate dehydrogenase; MDM, murine double minute; OR, odds ratio.
P-value calculated using Pearson’s Chi-Square test or Fisher’s Exact test, where appropriate.
P-value considered significant at < .05.
Complete follow-up data and last status (whether dead or alive) were known for 90 out of 102 patients, and these patients were used for survival analysis. Figure 2 shows the Kaplan–Meier survival curves for GBM patients in relation to markers status. EGFR-overexpressed patients were seen to have significantly lower survival as compared with their negative counterparts (P = .009). Moreover, increasing age was also correlated with a worse prognosis as patients > 40 years of age had median survival of 5 months, whereas patients younger than 40 years had median survival of 19 months (P = .002). Correlations among survival and markers status and patient characteristics are shown in Table 5.
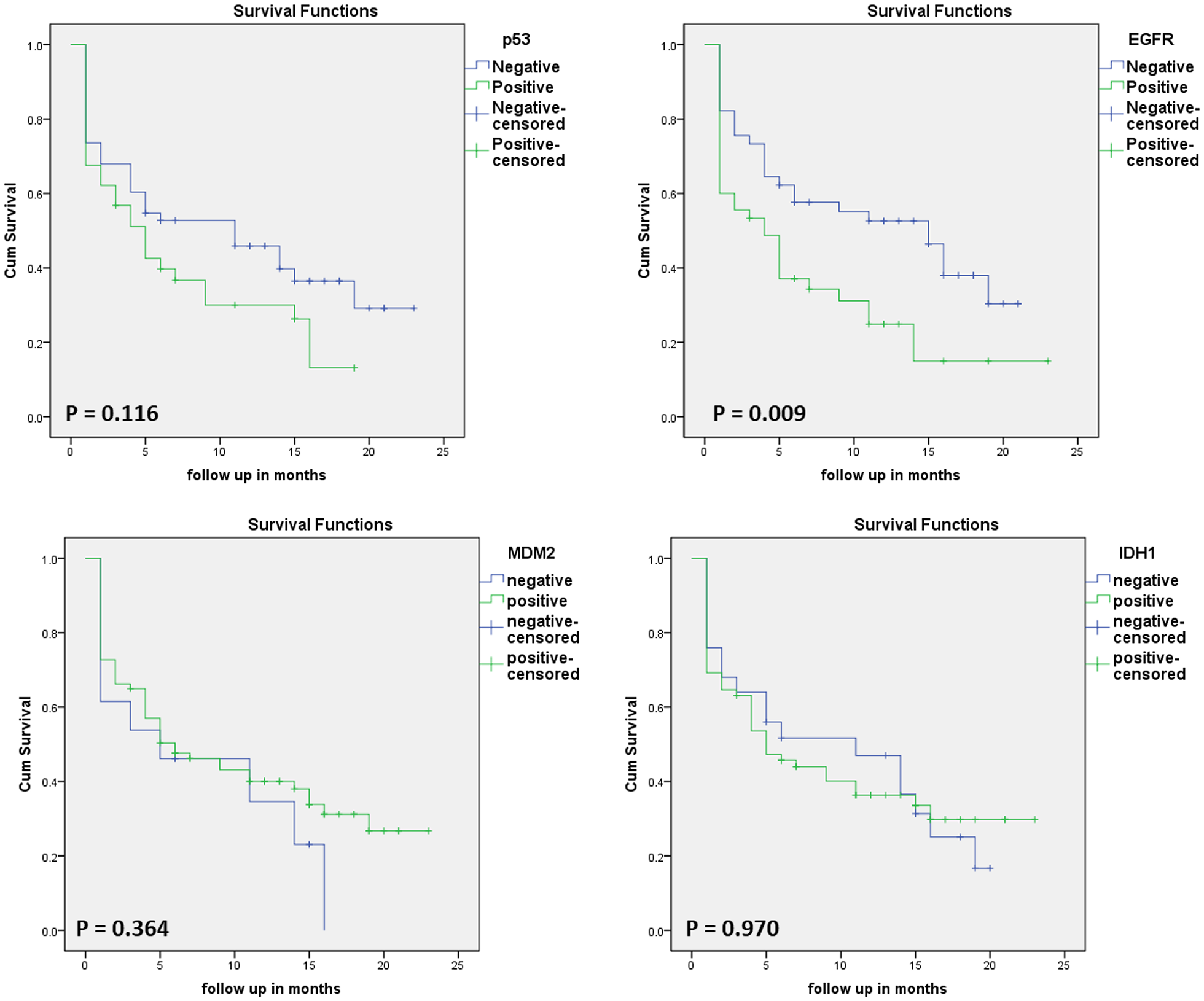
Kaplan–Meier survival curves in 66 GBM patients selected according to (A) p53 status, (B) EGFR status, (C) MDM2 status, and (D) IDH1 status.
Correlation of patient characteristics with overall survival.

Abbreviations: CI, confidence interval; EGFR, epidermal growth factor receptor; IDH, isocitrate dehydrogenase; MDM, murine double minute.
P-value calculated using log-rank test and considered significant at < .05.
Discussion
It has been observed that unusual expression of EGFR, p53, IDH1, and MDM2 proteins in GBM patients has an impact on overall survival, although some studies have cited an improvement, while others have affirmed a worse effect on prognosis. The aim of our research was to study the correlation between overall survival of GBM patients and immunohistochemical expression of EGFR, p53, IDH1, and MDM2 proteins. We found that EGFR, IDH1, and MDM2 proteins were overexpressed in > 50% of our patients, while p53 was overexpressed in about 26% of patients. On correlating the overexpression of these protein markers with overall survival, we found that only EGFR was significant for patient survival. Although another noteworthy observation for EGFR was that increasing age was significantly associated with higher EGFR expression (P < .001). Moreover, EGFR overexpression and increasing age were worse prognostic factors for our population.
Considering the demographics of our population, we had a higher number of males as compared to females in our data which corresponds with previous observations about the gender prevalence of GBM. 2 The mean age of our patients at about 47 years is slightly lower than the peak incidence estimated to be around 55 to 60 years of age 19 in Western countries; however, it is in line with the findings of Ahsan et al 20 which was a study based on central nervous system tumors in the Pakistani population. Age above 40 years was a worse prognostic indicator for our patients as determined on Kaplan–Meier survival analysis.
Montgomery et al previously reported positivity percentages for EGFR, IDH1, MDM2, and p53 for a cohort of 36 patients and found 100% positivity for EGFR, MDM2, and p53, while only 1 patient was positive for the mutant form of IDH1. In comparing these results with our cohort, we found lesser positivity percentages for EGFR, MDM2, and p53. The differences may be due to the differing populations under study (Brazilian and Pakistani), and the differences in cohort sizes (36 vs 100). Also, the authors defined positivity as starting from 0% to 25% which also includes any cases that had zero positive cells in their total count, whereas we considered zero cells stained as negative. The higher scoring criteria of our study were established to rule out any artifacts and normally expressed positive signals and to only include true overexpression of all respective protein markers. Moreover, in the case of IDH1, a comparison is not wholly applicable because Montgomery et al 1 detected only the mutant type of IDH1 in their study, while the antibody we tested in our cohort did not specify between the mutant and wild-type forms of IDH1, only noting the immunohistochemical expression of both of the 2 IDH1 states.
Our study used the H11 clone of EGFR which detects both the wild type and the deletion mutant form of EGFR (EGFRvIII). The same clone was used by Lopez-Gines et al to detect EGFR overexpression in 40 cases of glioblastoma, and they found EGFR overexpression in 50% of cases, which corroborates our findings. In addition, a close correlation between the levels of EGFR protein expression and EGFR gene-copy number alteration was also reported. This suggests that high EGFR protein levels, such as those observed in our study, are in fact due to changes at the genetic and transcriptional level and that IHC is a valuable detection technique in this regard. 21 Another study assessing the expression of EGFR in formalin-fixed paraffin-embedded sections of GBM found 43.5% EGFR-positive cases in 76 patients. In the same cohort, p53 positivity was also observed in 64% of these patients which is a higher number than our own observation. 22 Considering the effect of these markers on overall survival, it has been noted that EGFR-positive patients have longer survival and a better overall response to therapy as compared to EGFR-negative patients. These findings were noted for a cohort of 52 patients in which 30 were positive for EGFR. However, we had no such observations for our patients. Our EGFR-positive patients had a median survival of 4 months and EGFR-negative patients had a median survival of 15 months (P = .009). 23 In another detailed meta-analysis of 17 studies based on IHC conducted in the year 2017, hazard ratios were pooled, and in a cumulative of 1458 patients, the pooled HR was 1.72, meaning that a high EGFR IHC expression was a significantly poor prognosis factor for GBM patients, similar to our findings. 24
Moreover, we observed a significant correlation between age and EGFR status which leads us to suggest that more aged patients harbor EGFR mutations and may be better recipients of EGFR-targeted therapies, such as those using tyrosine kinases and monoclonal antibodies. However, this can only be determined by further molecular analysis of the type and frequency of EGFR mutations occurring in these patients to determine their suitability for such a therapy as these therapies target only specific mutations in the EGFR gene. The association of EGFR and advanced patient age may be due to the fact that primary GBM is more prevalent in patients aged 50 years and above, and EGFR mutations are more frequent in primary GBM tumors (about 35%) as compared with secondary GBM where it occurs rarely. 25 In addition to EGFR mutations, EGFR amplification is also a commonly occurring event in primary advanced-age GBM, along with other genetic events, such as MDM2 amplification and expression. 26 Furthermore, the EGFR-positive patients of this cohort suffered from statistically worse prognosis as compared with EGFR-negative patients. This ties in with the fact that advanced-age primary tumors are characterized by increased EGFR expression and these patients may have developed primary tumors at late stages of their life, leading to worse prognosis.
Regarding IDH1, a 2012 study by Sipayya et al 27 found that GBM patients had 9.7% (7/72) IDH1-positive specimens in a total of 195 gliomas. However, another study correlating the expression of IDH1 mutant protein using IHC detected no positive samples in GBM patients of India. 28 Contrarily, we observed high IDH1 positivity (72%) in our patients and this deserves further molecular investigations to check whether IDH1 gene mutation corresponds to the observed protein overexpression. To investigate the concordance of IDH1 IHC positivity and molecular analysis, a meta-analysis was performed by Pyo et al 29 on 1360 cases of glioblastoma from 16 studies. The meta-analysis calculated that the overall concordance rate was 0.947, reflecting that IDH1 IHC is a suitable and reliable method for testing IDH1 mutations. In a literature review assessing the importance of IDH1 in GBM, it was concluded that IDH1-mutant patients had significantly longer progression-free survival as compared with wild-type IDH1 patients and that IDH1 status was an independent prognostic factor for survival in GBM patients. 30 This, however, could not be reflected in our data as we observed no significant difference in the survival times of IDH1-positive and IDH1-negative patients which were 5 months and 11 months, respectively (P = .97).
It is generally understood that IDH1/2-mutant tumors have a better prognosis as compared with IDH1/2 wild-type tumors. A study hypothesized that this may be due to neovascularization and tumor angiogenesis as GBM is a highly vascularized tumor. 15 However, no significant differences in hypoxia and angiogenesis between IDH1 wild-type and IDH1-mutant tumors was seen. Another study has attributed the increased survival advantage of IDH1-mutant tumors to the fact that tumor-related immune response is severely downregulated in IDH1-mutant tumors, including leukocyte chemotaxis. 31 This results in much lower infiltration of immune cells, such as macrophages, microglia, monocytes, and neutrophils in glioma, while the presence of these infiltrative cells in other tumors is linked to lower survival.
We did not observe any significant difference in survival of IDH1-positive and IDH1-negative patients. This may be due to the difference of treatments that our patients received in comparison to other clinical practices. Due to the lack of health insurance and other financial difficulties, many patients failed to follow-up with their complete treatment protocol which further complicated survival. Furthermore, the prognostic effect of IDH1 is apparent for patients with IDH1 wild-type versus IDH1-mutant patients. 32 However, we analyzed the expression level of IDH1 protein only and further molecular investigations regarding the status of IDH1 (whether mutated or not) may help highlight the prognostic differences further. Similar to our findings, another study group has also concluded no survival differences based on IDH1 mutation for a glioblastoma cohort with long-term follow-up. Out of 172 patients, only 2 had mutated IDH1 R132H which was confirmed using IHC and sequencing. Hence, there may be factors other than IDH1 mutation affecting the survival of our GBM patients. 33
The IHC expression of p53, as detected by the clone DO-7, may be a marker for inactivating p53 mutations as it detects total p53. Weakly staining scattered cells are considered normal/wild type, while strong nuclear staining is considered overexpression due to inactivating point mutation of TP53. Additionally, a complete lack of expression may be due to a null mutation/deletion of TP53. To discriminate between these types of expression, the p53 mild positives of our study were considered negative in statistical analysis, so that any normal/wild-type expression of p53 was not included in the p53-overexpressing cells and does not influence the correlations. This is in line with our previous study on p53 expression in oral cancer. 18
To assess the role of mutant p53 in malignant gliomas, a 2014 study by Wang et al investigated mutant p53 in patients being treated with temozolomide. It was found that the expression of mutant p53 was significantly inversely proportional to survival and a fivefold increase in chemosensitivity to temozolomide was seen when knockdown of mutant p53 occurred. 34 It is noted that p53 pathway is deregulated in 84% of glioblastomas and TP53 mutations lead to high expression of oncogenic variants of p53 protein. 35 Birner et al investigated 114 cases of primary GBM and found 25.4% positive patients. They also observed that p53-positive patients were significantly younger and had markedly better overall survival. Although we observed no correlation of p53 with age (P = .384) or survival (P = .116), we did observe the same 26% positivity of p53 in our patients. Since Birner et al considered only > 50% immunostaining as positivity, we may base the similarity on positivity threshold. Another study conducted on the Korean population had a higher positivity of 48.4% of p53 evaluated immunohistochemically in GBM. 36
To study the differing expression patterns of MDM2 in primary vs secondary glioblastomas, an immunohistochemical and molecular examination was undertaken on 29 patients of primary GBM and 27 patients of secondary GBM. MDM2 immunoreactivity was seen in 52% of primary GBM and 11% of secondary GBM. 37 We had a significantly higher 83% MDM2 positivity in our patients. Although other studies have concluded MDM2 to be linked to poor overall survival in GBM patients, 38 we found no significant difference in our patient set.
Statistically significant correlations have also been found between p53 and MDM2, and EGFR and MDM2 by other authors. In addition, a strong negative correlation has also been found for p53 and survival with a higher percentage of p53 positivity leading to a lower survival in months. 1 While there are other studies that have denied such correlations and concluded an effect of MDM2 apart from p53 as well, with one such study citing that only 1 out of 15 primary GBMs that overexpressed MDM2 also contained a p53 mutation. 37 Similarly, we found no association of p53 and MDM2 in our patients (P = .549) even though an inverse relationship between MDM2 and p53 has also been suggested due to the fact that MDM2 targets p53.
The patient population in this study is heterogenous and poorly studied. Comparisons between our study and those from genetically and geographically diverse regions might not draw fair conclusions. Further molecular and immunohistochemical approaches are needed on a broader set of patients to determine the impact of these markers on GBM patients. From this study, the role of these markers remains in doubt, although EGFR presented as a worse prognostic indicator for the Pakistani GBM population. Differences in analytical techniques, antibodies used, and even among criteria used to assess the same technique may account for the varying results. Systematic reviews and meta-analysis researching only immunohistochemical studies and their results will yield a more comprehensive view of these markers and their eventual utility in a clinical setting.
Conclusions
The role of p53, EGFR, MDM2, and IDH1 in initiating, maintaining, and progressing glioblastoma has been documented and we found increased expression of these markers in most of our GBM patients. However, we only concluded the adverse effect of EGFR overexpression on survival, out of all markers tested. Increasing age was also correlated with higher EGFR expression and poor survival.
Footnotes
Acknowledgements
None.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Seed Money Grant, Aga Khan University Hospital, Karachi, Pakistan awarded to Muhammad Shahzad Shamim (PF 94/1216).
Author Contributions
SMAA conceived and designed the study, performed literature search and writing. SAE and MSS performed patient data collection, data interpretation, and project supervision and provided financial support. YA and ZA performed statistical analysis, data interpretation, and writing. HAF performed literature search and assisted in data entry and data interpretation. All authors reviewed and edited the final draft of the article and took responsibility for its content.
Ethical approval
This study was approved from ethical review committee of AKUH (ERC#2018-0511-808).
