Abstract
Multidrug resistance and invasiveness of non-typhoidal Salmonella (NTS) serovars have in recent times brought to the fore the public health risk associated with salmonellosis. This study was aimed at profiling NTS serovars isolated from food animals and humans for their susceptibility to antibiotics and plasmid replicon types. Forty seven NTS serovars were profiled for their susceptibility to antibiotics using the disk diffusion method. Polymerase chain reaction based replicon typing assay was used for profiling plasmid replicon types detected in Salmonella isolates. High rate of resistance were found for amoxicillin/clavulanic acid (40/47; 85.1%), cefuroxime (38/47; 80.9%) and ceftazidime (30/47; 63.8%). Thirty one (65.9%) and 33 (70.2%) showed intermediate resistance to ofloxacin and ciprofloxacin respectively. Plasmids of sizes ranging from 14.3 to 16.7 kb were detected in 24 (51.1%) of Salmonella isolates with some serovars harbouring multiple plasmids. FIA, FIB, Frep and W plasmid replicon types were detected in 11, 4, 2 and 1 of the Salmonella isolates respectively. Three of the isolates harboured both FIA and FIB replicon types. The high rate of resistance to β-lactams observed in Salmonella serovars harbouring different plasmid replicon types in this study highlight potential public health threat and the need for prudent use of antibiotics in human and veterinary medicine.
Introduction
The continuous circulation of non-typhoidal Salmonella serovars (NTS) within the food chain poses a major public health risk.1,2 The emergence and dissemination of invasive multidrug resistant strains mainly in sub-Saharan Africa portends a serious danger as well. 2 The World Health Organization (WHO) prioritized fluoroquinolone resistant Salmonella species as pathogens for which there is high need for research and development of new antibiotics. 3 Several factors including efflux pump activity, biofilm formation, mobile genetic elements among others have been reported to mediate antimicrobial resistance (AMR) in Salmonella.4 -6 Of these factors mobile genetic elements principally plasmids, facilitate transmission and dissemination of AMR among Salmonellae through horizontal gene transfer (HGT).4,7 Plasmids are wide spread among Salmonella serovars where they mediate not only virulence but also resistance to antibiotics. 8 Han et al 9 in a previous study reported multidrug resistance plasmids that carried at least 8 antimicrobial resistance genes in Salmonella Heidelberg isolated from poultry. Incompatibility groups (Inc) of plasmids found in Salmonella serovars are said to be associated with serovars, animal sources and differ in their replication mechanism and accessory gene content. 10 Emond-Rheault et al 8 reported the large distribution of IncFII and IncI1 plasmid replicon types among many Salmonella serovars but serovar restricted occurrence of IncHI2A/IncHI2 and IncA/C2 replicon types. In Nigeria several studies have reported the occurrence of plasmids in Salmonella serovars but there is paucity of data on the classes of plasmids that are borne by serovars circulating the food chain. Therefore in this study we profiled Salmonella serovars isolated from different sources for their plasmids, plasmid replicon types and antibiotic susceptibility.
Materials and Methods
Bacterial isolates
Forty seven NTS serovars isolated previously form food animals and apparently healthy humans 1 were used in this study. Thirty one of the Salmonella serovars were isolated from chicken, 2, 5 and 9 were isolated from cattle, human and sheep respectively.
Susceptibility testing
Susceptibility of Salmonella isolates to 10 antibiotics (ceftazidine [30 μg], cefuroxime [30 μg], gentamicin [10 μg], cefixime [5 μg], ofloxacin [5 μg], amoxicillin-clavulanic acid [30 μg], ciprofloxacin [5 μg], nitrofurantoin [200 μg], ciprofloxacin [5 μg] and amikacin [100 μg]) was done by the Kirby-Bauer disc diffusion method according to the guidelines of the European Committee on Antimicrobial Susceptibility Testing. 11 The susceptibility testing was done in duplicates with Escherichia coli ATCC 25922 used as quality control organism. The isolates were inoculated into brain heart infusion broth (Oxoid, Basingstoke, UK) and incubated at 37°C for 18 hours. Bacteria broth culture was streaked on nutrient agar (Oxoid, Basingstoke, UK) and incubated for 24 hours at 37°C. Cell suspension was made by emulsifying single colonies bacterial cells 5 mL of sterile normal saline and adjusted to 0.5 McFarland standard. Cell suspension was then applied onto Muller-Hinton agar using sterile swab sticks (Oxoid, Basingstoke, UK) and antibiotic discs were dispensed and incubated for 24 hours at 37°C. Zone of inhibition was measured and recorded as sensitive, intermediate and resistant. 6
Plasmid DNA extraction
Norgen Plasmid MiniPrep extraction kit (Norgen Biotek Corp, ON, Canada) was used for plasmid extraction and the extraction was done according to the manufacturer’s instructions. Isolates were inoculated into brain heart infusion broth (Oxoid, Basingstoke, UK) and incubated at 37°C for 24 hours after which cells were harvested by centrifuging at 13 000 rpm for 5 minutes. Supernatant was discarded and 1 mL of sterile water was added to cell pellet vortexed and centrifuged at 13,000 rpm for 5 minutes. Two hundred microlitres of Resuspension Solution AZ (containing RNase) was added to washed bacterial cells and vortexed then incubated at room temperature for 5 minutes. Two hundred and fifty microlitre of lysis buffer was added to the cell suspension and gently inverted several times until the suspension become clear and viscous. Then 350 µL of buffer TN was added and immediately mixed by inverting the tube several times and centrifuged for 10 minutes. Lysate was transferred into spin column and centrifuged for 1 minute then separated from the collection tube while the flow through was discarded. Six hundred microlitre of wash solution E was added to the column and centrifuged for 1 minute after which the flow through was discarded. To dry the column, it was spun for 2 minutes. Fifty microlitres of elution buffer K was used in eluting plasmid DNA and plasmid was stored at −20°C until use. Separation of plasmid on agarose gel was done and molecular weights of plasmids were estimated In-silico as previously described by Ajayi et al. 4
PCR based replicon typing of plasmid
Plasmid replicon types were determined by simplex and multiplex polymerase chain reaction (PCR) using pairs of primers listed in Table 1. A 20 μL PCR reaction mix containing 10.8 μL nuclease free water, 0.6 μL forward primer, 0.6 μL reverse primer, 4 μL DNA template and 4 μL of 5X PCR Master Mix. The PCR programming parameters are as follows: Initial denaturation at 95°C for 5 minutes, followed by 30 cycles of denaturation at 95°C for 30 seconds, annealing temperature at 60°C for 40 seconds for primer targets except FrepB, K/B and B/O that had annealing at 52°C. Extension was programmed at 72°C for 2 minutes, followed by a final extension at 72°C for 10 minutes. The PCR products were electrophoresed at 100 V for 1 hour in 1.5% agarose gel stained with ethidium bromide and visualized under ultraviolet trans-illuminator (Clever Scientific UK).
List of primers used for plasmid replicon typing in both simplex and multiplex PCR.
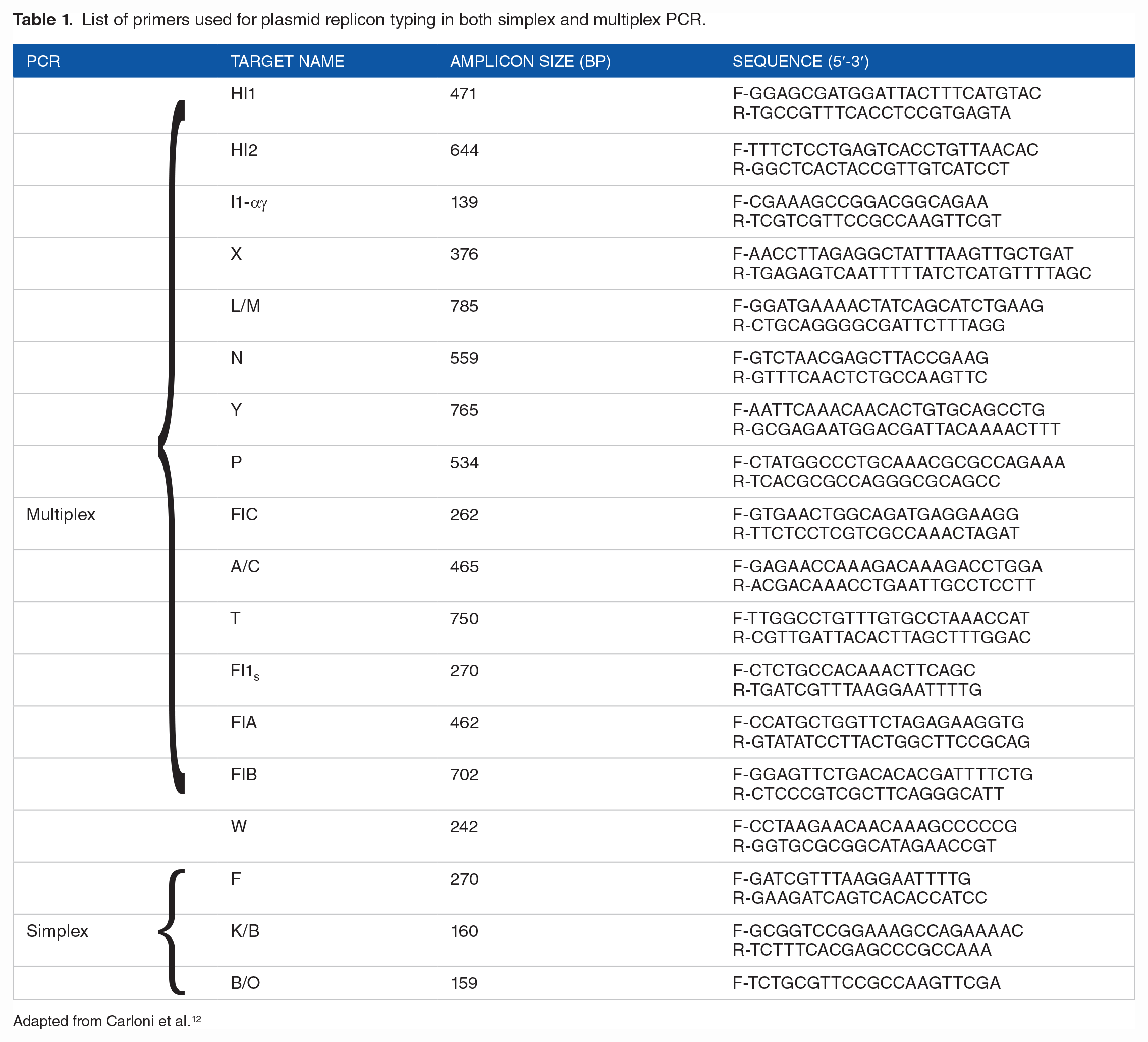
Adapted from Carloni et al. 12
Data analysis
Microsoft Excel (Microsoft Cooperation, 2013 USA) and GraphPad Prism version 8.0.2 (GraphPad Software Inc. USA) were used for data analysis. Counts and percentages were used to describe categorical data. Chi square test was used to determine if the possession of plasmid was associated with multidrug resistance. Confidence interval was pegged at 95% and P-value <.05 was considered statistically significance.
Results
Antimicrobial susceptibility profile
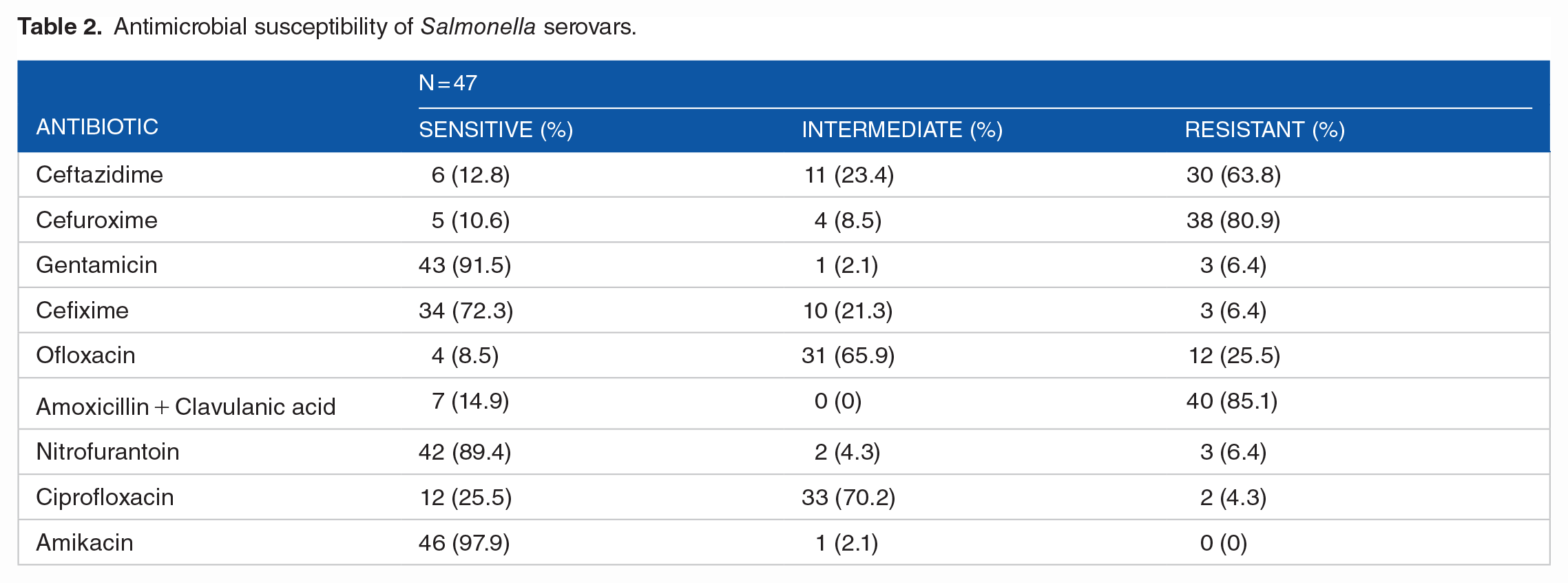
Salmonella isolates showed varied susceptibility to test antibiotics as shown in Table 2. Forty six (97.9%) of the isolates were sensitive to amikacin, 1 (2.1%) showed intermediate susceptibility, while none was resistant to amikacin. Isolates were also largely sensitive to gentamicin 43 (91.5%) and nitrofurantoin 42 (89.4%). High rate of resistance to amoxicillin + clavulanic acid 40 (85.1%) and cefuroxime 38 (80.9%) was recorded. Thirty (63.8%) of the Salmonella isolates were also resistant ceftazidime. Majority of the isolates 31 (65.9%) and 33 (70.2%) displayed intermediate susceptibility to ofloxacin and ciprofloxacin respectively. None of the isolates were sensitive to all test antibiotics. In terms of classes of antibiotics to which the isolates were resistant, 45% of them were resistant to β-lactams, while resistance to quinolones and aminoglycosides were both 22% and resistance to nitrofurantoin was 11% as shown in Figure 1.
Antimicrobial susceptibility of Salmonella serovars.
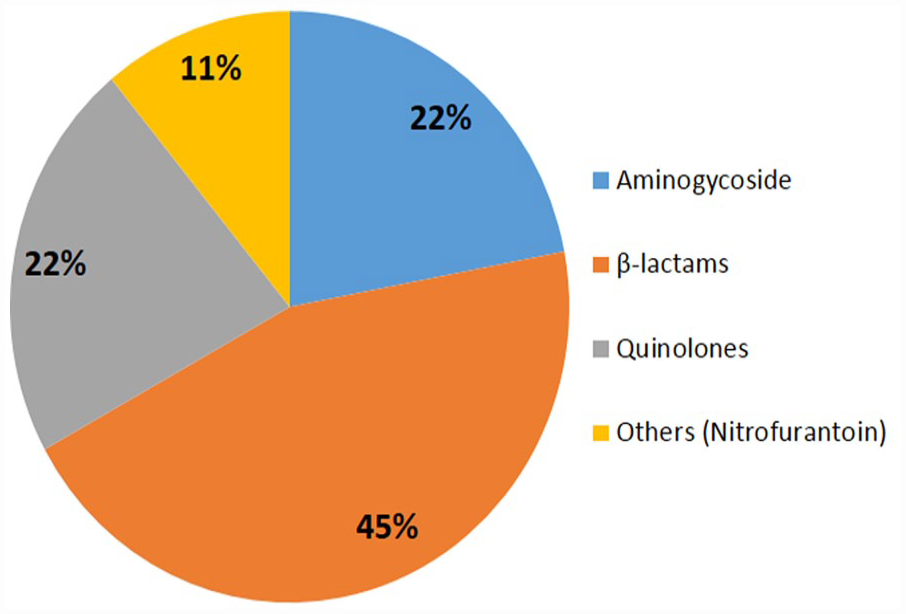
Percentage resistance of Salmonella to different classes of antibiotics.
Plasmid profile of Salmonella serovars and replicon types
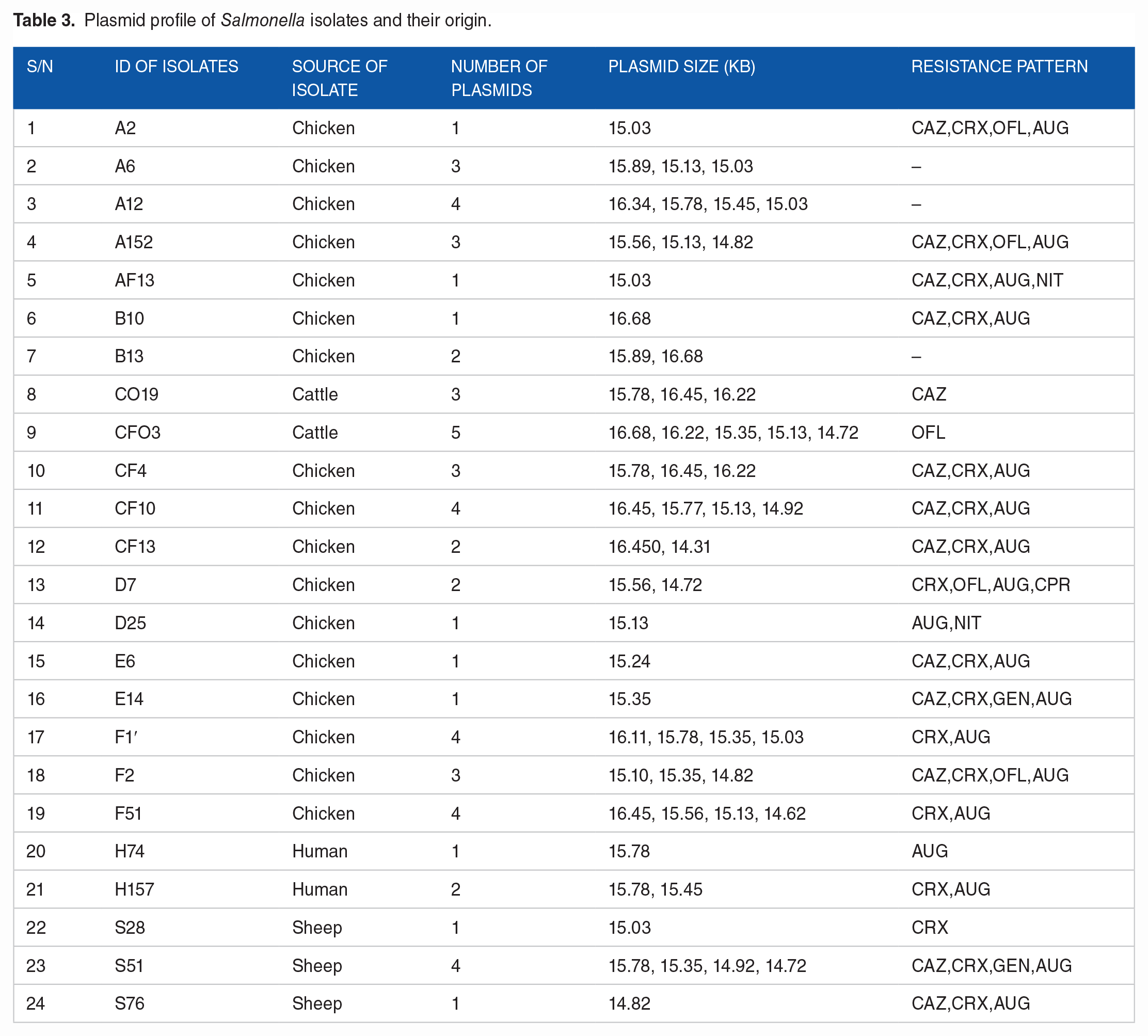
Plasmids of sizes ranging from 14.3 to 16.7 kb were detected in 24 (51.06%) of the Salmonella isolates. Some of the isolates harboured multiple plasmids with isolate CFO3 having 5 plasmids. Isolates S51, F51, F2, CF10 and A12 possessed 4 plasmids each, while isolates A6, A152, CO19, CF4 and F2 harboured 3 plasmid each as shown in Table 3 and Figure 2. Isolates A6, A12 and B13 harboured 3, 4 and 2 plasmids respectively but did not display any phenotypic antibiotic resistance pattern. On the other hand Isolates A2, AF13, E14 and B10, E6, S76 possessed one plasmid each and were resistant to 4 and 3 antibiotics respectively. Furthermore, isolates CO19 and CFO3 from cattle possessed 3 and 5 plasmids respectively but were resistant to only one antibiotic (Table 3). The association of multidrug resistance and the presence of plasmid was found not to be statistically significant (Table 4).
Plasmid profile of Salmonella isolates and their origin.
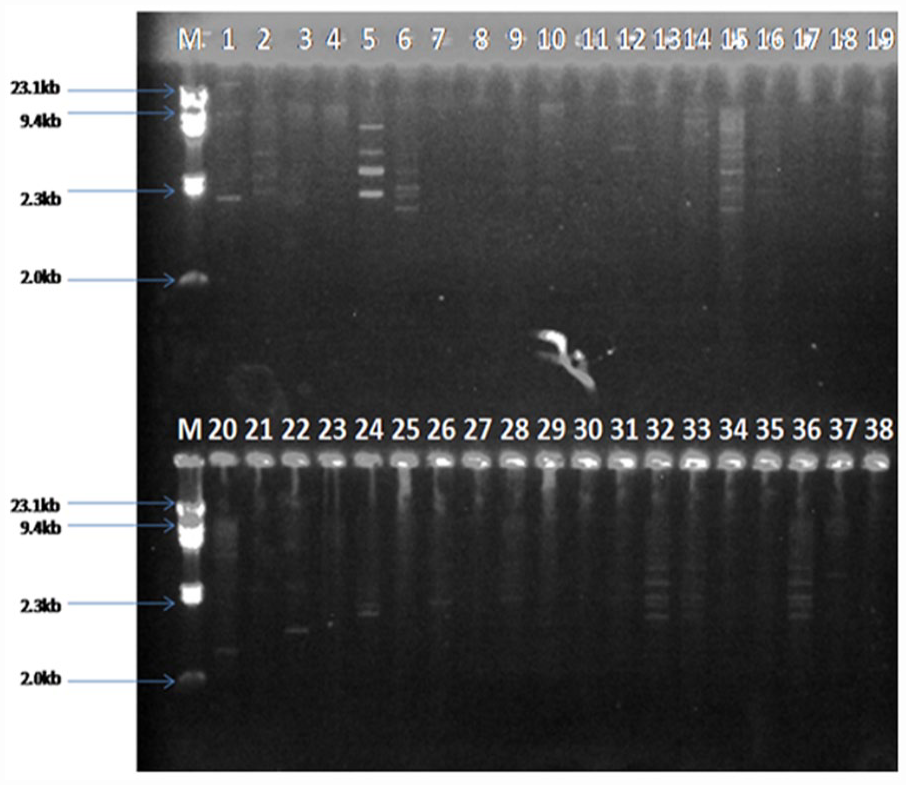
Gel image showing plasmid profile of Salmonella isolates. Lane M: Lamda/Hind III digest DNA ladder. Lane 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 12, 14, 15,16, 19, 20, 22, 24, 26, 28, 32, 33, 36 and 37 are positive for single and multiple plasmids.
Association between possession of plasmids and MDR among Salmonella serovars.
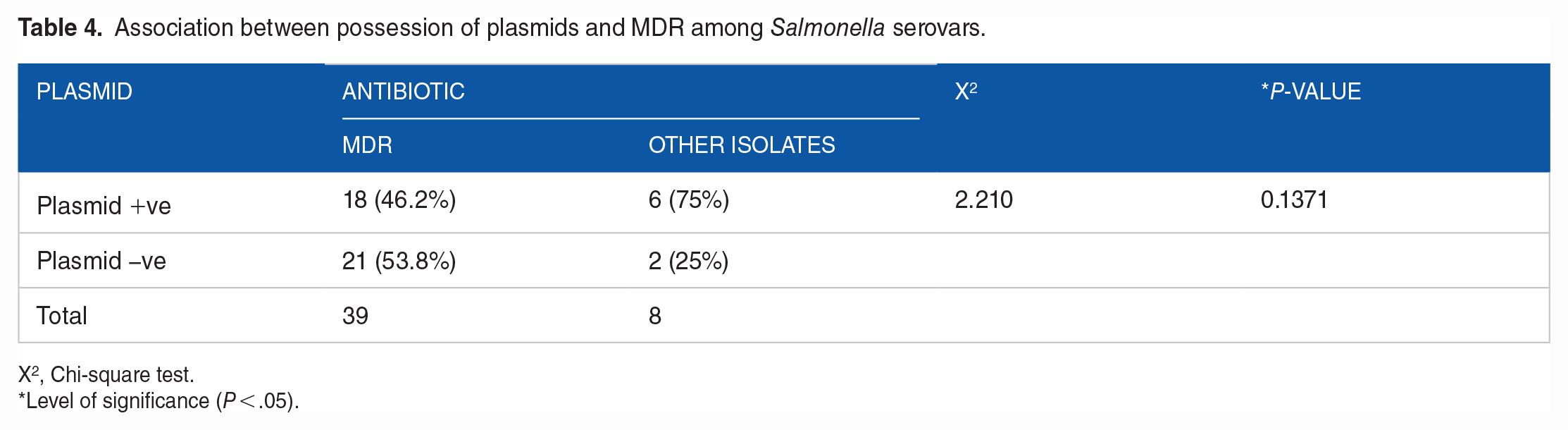

X2, Chi-square test.
Level of significance (P < .05).
Nine (37.5%) of the Salmonella isolates possessed plasmids that could not be typed. However, 11 (45.8%) of the isolates harboured FIA plasmid replicon type. Replicon type FIB was present in 4 (16.7%) isolates. Two of the isolates possessed plasmid replicon type Frep, while replicon type W was detected in only one isolate. Three Isolates possessed both replicon type FIA and FIB (Figure 3).
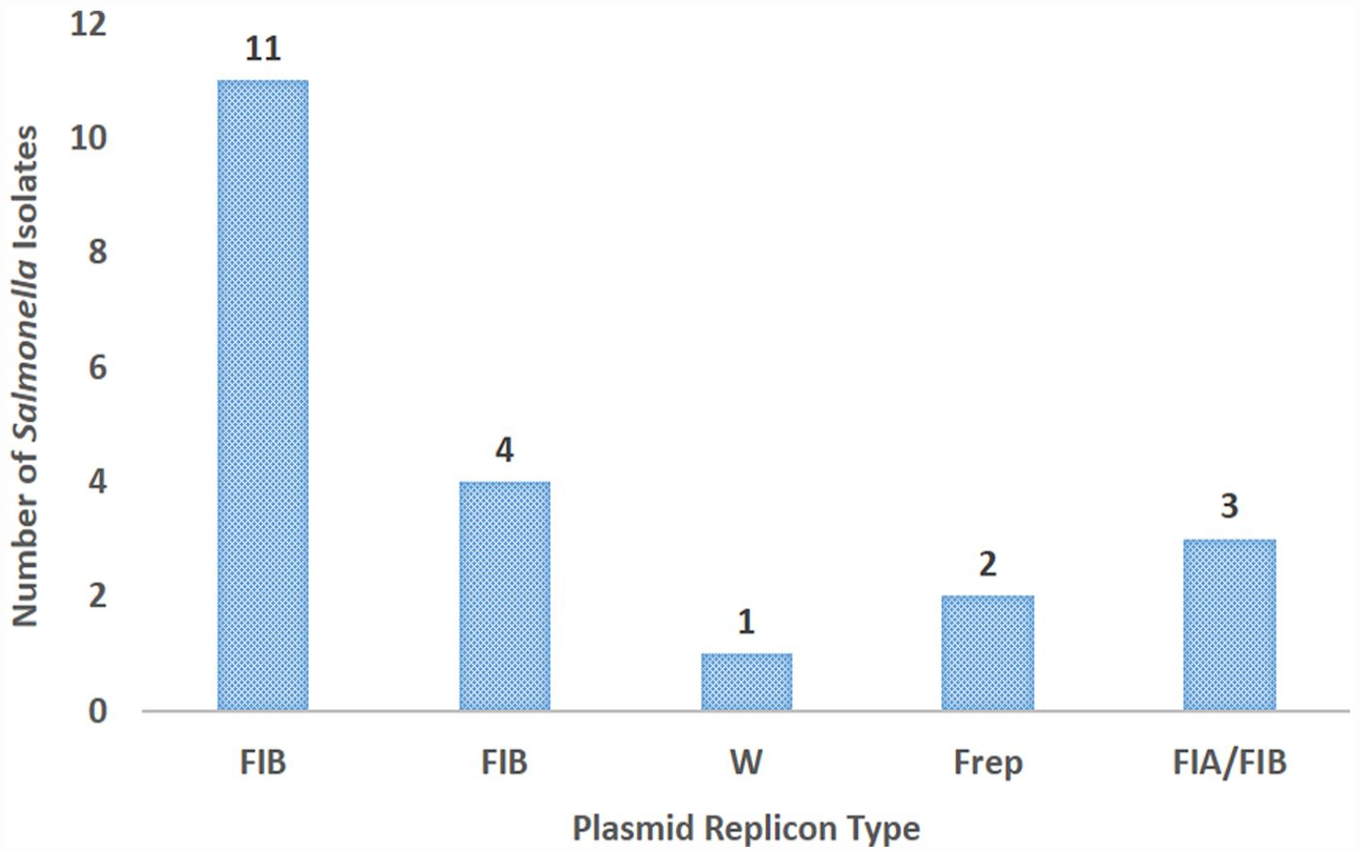
Plasmid replicon types detected in Salmonella isolates.
Discussion
Salmonellosis is a major food borne zoonoses of global proportion that is becoming difficult to treat as a result of antimicrobial resistance (AMR). 13 In this study high resistance to amoxicillin + clavulanic acid, cefuroxime and ceftazidime was observed among Salmonella serovars. Rodrigues et al 14 in a similar study reported high resistance of Salmonella serovars isolated from chicken to amoxicillin + clavulanic acid and cefotaxime in Minnesota, United States of America. In a study in Lagos Nigeria, Akinyemi et al, 15 reported amoxicillin + clavulanic acid, cefuroxime and ceftazidime resistance in Salmonella species from febrile patients. Majority of Salmonella isolates in this study were resistant to β-lactam class of antibiotics followed by aminoglycoside. This is in contrast to the report of Pławińska-Czarnak et al, 16 who reported a higher resistance to aminoglycoside than β-lactams in Salmonella species isolated from raw meat in Poland. Although increasing resistance of Salmonella to fluoroquinolones have been reported, 3 in this study majority of the isolates displayed intermediate susceptibility to ofloxacin and ciprofloxacin. The apparent difference in level susceptibility may be due the source of the isolates, since clinical isolates are known to be more resistant to fluoroquinones. 17
Plasmids are wide spread among Salmonella serovars where they mediate both virulence and resistance to antibiotics. 8 Plasmids are major players in the horizontal transfer of AMR determinants in Salmonella. 18 In this study 51.06% of the Salmonella isolates harboured plasmids of different sizes. Plasmids were detected in isolates from chicken, cattle, human and sheep. In a previous study Ogunleye et al 19 reported antimicrobial resistance determinants that were plasmid borne in Salmonella serovars isolated from poultry, cattle and lizard in Oyo and Ogun states Nigeria. Similarly, Jibril et al 20 reported plasmids mediated antibiotic resistance in Salmonella serovars that were isolated from poultry in North-west Nigeria. In another study in Lagos, Akinyemi et al 21 reported the occurrence of plasmids ranging from 3.2 to 30.2 kb in Salmonella species isolated from different sources. Elsewhere in Uganda, Bosco et al 22 reported 68.7% of Salmonella species of human and food animal origin that harboured plasmids.
Some of the isolates possessed plasmids but did not display any phenotypic antibiotic resistance pattern. The plasmids they harboured could be plasmids that mediate virulence factors. Ajayi et al 4 in their study reported plasmids of Salmonella that mediated ability to bind and invade host intestinal epithelium thereby establishing infection. On the other hand multiple plasmids harboured by the isolates as observed in this study could also mediate both virulence and antimicrobial resistance. Hsu et al 23 in a similar study in the United States of America reported multiple plasmids that mediated both virulence and AMR in Salmonella Dublin that was isolated from cattle, retail beef and humans.
Although IncHI2 plasmids are said to be predominant in antibiotic resistant Salmonella isolates, 24 in this study FIA and FIB associated with the IncF group were the predominant plasmid replicon types among Salmonella isolates with 1 and 2 other isolates having W and Frep replicon types. Furthermore, no association of MDR with possession of plasmid was established here. This is in contrast to the study of Mutai et al 7 who reported IncHI1as the most prevalent plasmid replicon type detected in MDR S. Typhi strains isolated from patients in Kenya. Jibril et al 20 identified multiple antimicrobial resistance genes located on IncN and IncQ1 plasmids of S. Schwarzengrund and S. Muenster in an In Silico prediction analysis using whole genome sequences of Salmonella serovars isolated from poultry in Nigeria. The replicon type disparity observed among serovars in these studies including ours might be the fact that some replicon types are serovar restricted as indicated by Emond-Rheault et al. 8 However, this does not negate the fact that plasmid replicon types most especially FIA and FIB are ubiquitous among Salmonella serovars and E. coli strains with the possibility of exchanges among both genera. 24
Conclusions
This study highlights the occurrence of multidrug resistant NTS serovars with dominant FIA and FIB plasmid types circulating the food chain in Lagos State. This warrants further extensive studies to profile and attribute plasmid replicon types to serovars with their concomitant antibiotic resistance profile.
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
FLA, AA, UUE contributed to data collection, FLA, AA, UUE contributed to data analysis and preparation of draft, AIA, SIS and AA, contributed to conceptualization of study, AIA and SIS supervision and correction of draft. All authors read and approved final draft.
