Abstract
Background:
The purpose of this study was to compare different invasive methods for Helicobacter pylori (H. pylori) detection, namely PCR for H. pylori specific ureC gene, Rapid urease test (RUT), and histopathological examination by modified Giemsa staining.
Methodology:
Endoscopic gastroduodenal biopsy materials were collected from dyspeptic patients who underwent endoscopic examination upon fulfilling the inclusion criteria. Three to four samples were collected from each patient after taking informed consent and proper clinical history. A rapid urease test (RUT) was done on spot with in-house RUT media from 1 specimen. One to two specimens were preserved in 10% formaldehyde for histopathology and PCR for ureC gene was done from 1 specimen. Collected biopsy specimens from gastric and duodenal mucosa of 142 patients were categorized as H. pylori-positive cases and H. pylori-negative cases based on the case definition used in the study upon positivity of 3 diagnostic tests.
Results:
Among 142 biopsy specimens, 34.5% were categorized as H. pylori-positive cases, 35.2% as H. pylori-negative cases, and finally 30.2% as doubtful or indeterminate cases. Rapid urease test was the most sensitive method, closely followed by ureC gene PCR and histopathology, with a sensitivity of 94.2%, 83.0%, and 76.5%, respectively. Whereas histology was the most specific, having 98.0% specificity followed by 83.0% in PCR. RUT was the least specific, with 55.5% specificity.
Conclusion:
While histopathology could detect H. pylori infection with the highest specificity, for definitive diagnosis combination of any 2 methods should be used, if available.
Introduction
Helicobacter pylori colonize gastric epithelium causing a spectrum of site-specific diseases that is, peptic ulcer disease, gastric adenocarcinoma, and mucosa-associated lymphoid tissue (MALT) lymphoma.1,2 For prevention of serious chronic and life-threatening consequences; It is essential to identify and treat H. pylori infection at an early stage when patients usually attend gastroenterology centers with the complaint of dyspepsia. 3
There are several invasive and noninvasive avenues for detecting H. pylori infection in the affected population. Biopsy specimens collected during endoscopic procedures are often the preferred sample for isolating H. pylori 4 From biopsy tissue organisms can be directly detected by histological stain or microscopy following culture, by PCR, or indirectly by RUT.2,3
In other cases, serological tests, antigen detection from stool, and Urea Breathe Test (UBT) can be performed successfully to diagnose H. pylori infection among patients.1,4,5
Choice of the test depends on availability, cost-effectiveness, clinical situation, previous treatment with antibiotics and proton pump inhibitors (PPI), the population prevalence of infection, etc. 6 For a country like Bangladesh where the prevalence of H. pylori infection is thought to be widespread, and with the lack of modern diagnostic facilities and financial resources, choosing the right method for early and definitive infection detection could be crucial, and may be lifesaving in many instances. Hence, in this study we compared 3 available invasive diagnostic methods for the detection of H. pylori in Bangladesh including RUT, ureC gene PCR, and histopathological examinations of biopsy specimens.
Materials and Methods
This cross-sectional study was carried out at the Microbiology and Immunology department of Bangabandhu Sheikh Mujib Medical University (BSMMU), from September 2018 to July 2019.
Patient selection & sample collection
Patients aged 18 to 65 years who underwent upper gastrointestinal tract (GIT) endoscopy at the gastroenterology department of Dhaka Medical College and Hospital, due to dyspepsia were primarily selected for this study. 5 Individuals already receiving triple therapy for H. pylori eradication, therapy in the previous 2 months, had taken anti-ulcerant drugs including proton pump inhibitors (PPI), painkillers such as nonsteroidal anti-inflammatory drugs (NSAID), or any antibiotic 1 month before enrollment were ultimately excluded from this study.7-9
Case definition
According to the established case definition, participants were identified as H.pylori positive if they tested positive in at least 2 of those 3 testing methods (RUT, H.pylori histology, and PCR for ureC gene). Alternatively, participants were identified as H.pylori negative if they tested negative in all 3 tests. Study participants testing positive in one of 3 testing methods were identified as indeterminate/doubtful cases.
Laboratory procedures
Four to five gastric tissue specimens were surgically removed from the lesion. From those specimens, one collected from the antrum was used to detect H. pylori infection with the help of rapid urease test media. Another 2 specimens were taken in a 1.5 ml microcentrifuge tube containing 1 ml phosphate buffer solution for ureC gene PCR and stored at −20°C. One to two specimens from the margins of ulcers were collected and then put into a bottle containing 10% formalin and sent to the pathology department for histopathological examinations.
Histological examination
For H. pylori detection, a modified Giemsa stain was used to stain and detect H. pylori (Figure 1). An independent pathologist without prior knowledge of the patient’s condition and clinical history performed the histological examinations.
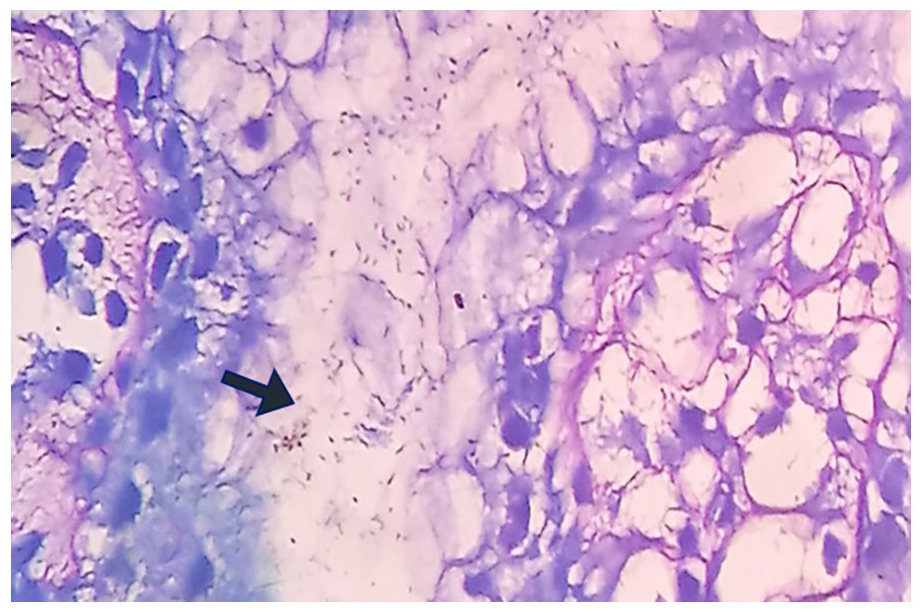
Histological section of gastric mucosa showing H. pylori over the surface mucosa (Modified Giemsa stain × 100).
DNA extraction and ureC gene PCR:
QIAamp DNA Mini Kit (Hilden, Germany) was used to isolate DNA from specimens. PCR was used to identify the ureC gene using the following forward and reverse primers and thermocycling conditions as described previously— AAGCTTTTAGGGGTGTTAGGGGTTT (forward); AAGCTTACTTTCTAACACTAACGC (reverse).
10
The final PCR product was run in 2% agarose gel and documented using a gel doc system (Figure 2).
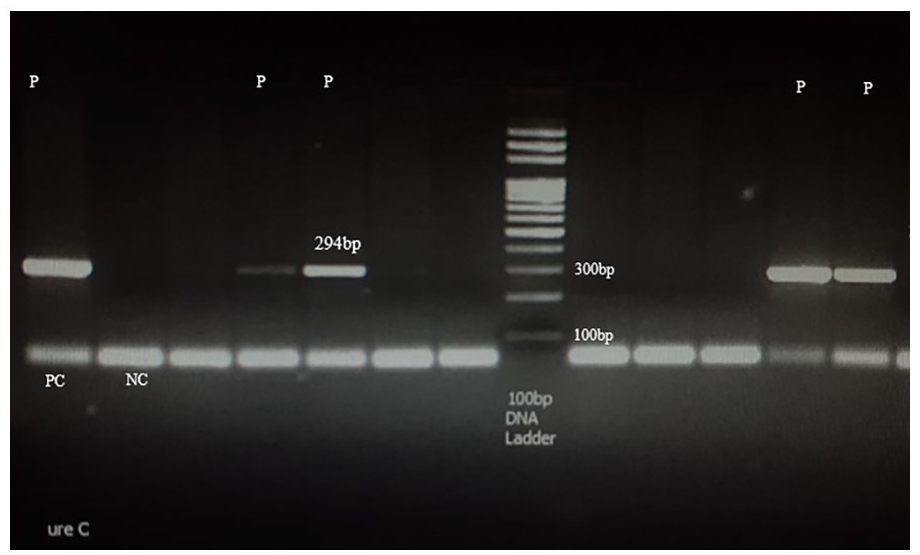
Agarose gel electrophoresis analysis showed amplified DNA product of ureC (294 bp). Lane P, positive results. Positive control (PC): ureC positive H. pylori, negative control (NC): amplified product of PCR without DNA).
Statistical analysis
Data were stored in a password-protected computer and all statistical analyses were performed using the SPSS software package (Version 23.0). The sensitivity and specificity of different detection methods were calculated considering the case definition of H. pylori-positive cases as the gold standard. The Chi-square test was used to seek association between variables. For all statistical tests, P value less than .05 was considered significant.
Results
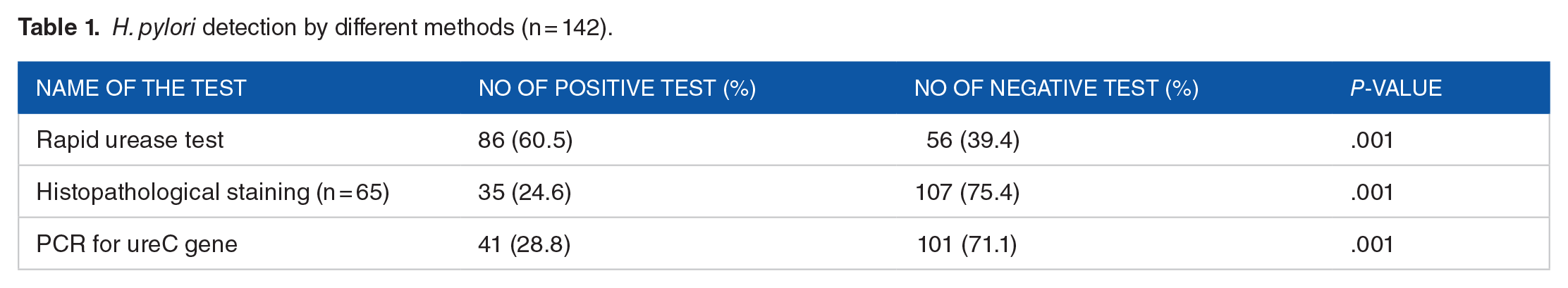
Table 1 shows the results from 3 different detection methods for H. pylori.
H. pylori detection by different methods (n = 142).
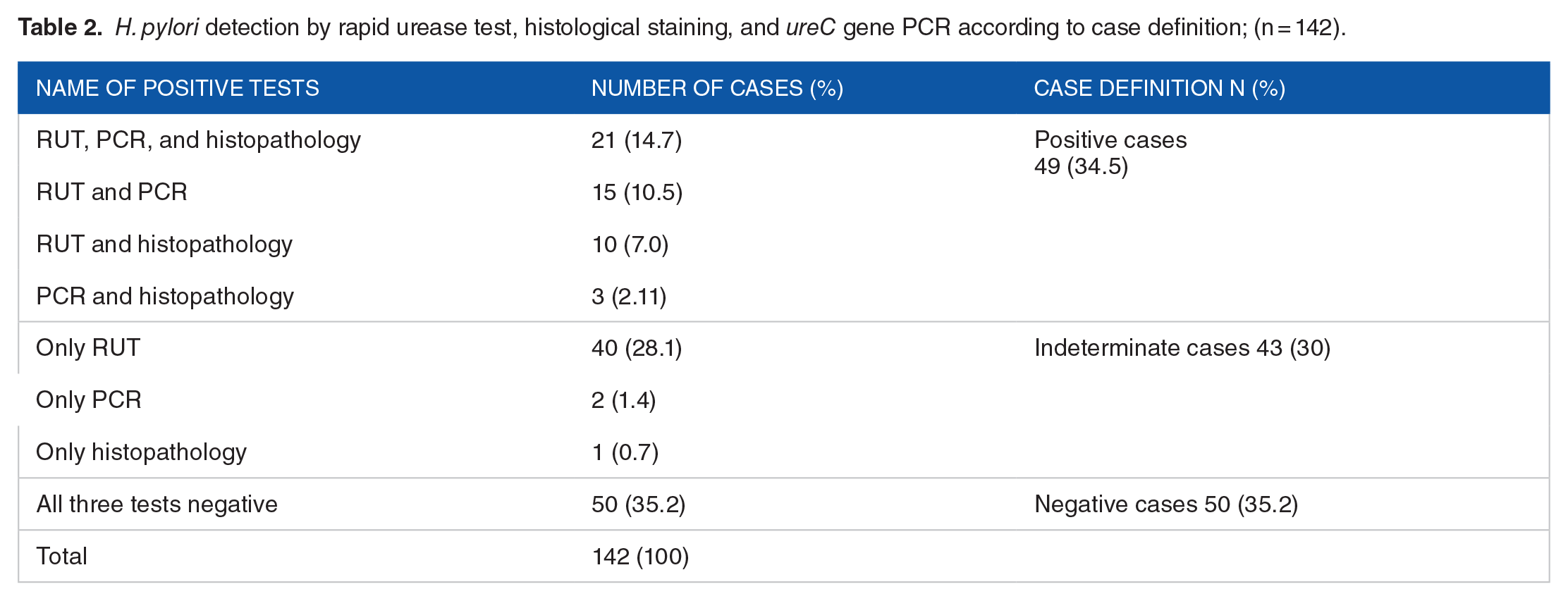
A combination of RUT and UreC gene PCR could detect 15 (10.5%) cases as positive; RUT, histology (modified Giemsa staining) could detect 10(7%) cases, and PCR and Histopathology could detect 3 (2.11%) cases as positive. Forty-three (30.2%) cases were identified as doubtful or indeterminate as they were found positive in only one test (Table 2).
H. pylori detection by rapid urease test, histological staining, and ureC gene PCR according to case definition; (n = 142).
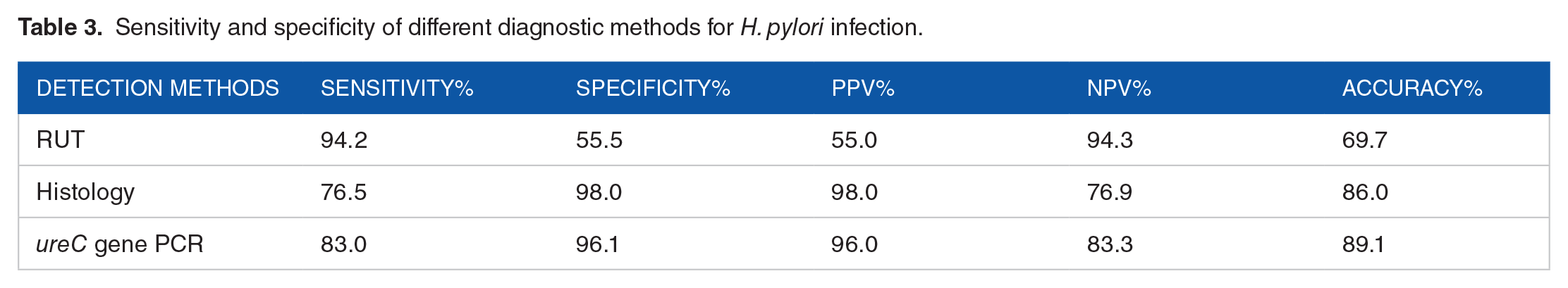
Sensitivity and specificity of all 3 tests for detecting H. pylori infection were calculated, considering the case definition of positive cases as the gold standard (Table 3).
Sensitivity and specificity of different diagnostic methods for H. pylori infection.
Discussion
Currently, there is no established gold standard for the diagnosis of H. pylori infection. Some researchers have suggested histology, some immunohistochemistry while others urea breath test as the gold standard for the detection of H. pylori.11-13 Choice of diagnostic test should depend on clinical indication, local availability, and costs of the different tests, as well as patient preferences. We, therefore, tried to evaluate 3 other diagnostic tests as the best available alternative for diagnosing a patient’s actual H. pylori status suitable for our country.
In our study, among 142 cases, RUT was positive in 86 (60.5%) cases. On the other hand, in previous studies conducted in Bangladesh, RUT could detect the bacteria in 54.05% and 53.3% of cases respectively.14,15 In the present study, the bacteria were detected in 35 (24.6%) samples by histological methods. Similar findings were reported by other Bangladeshi studies that found 34.4% and 25.71% H. pylori-positive cases respectively by histology in the gastric biopsy specimens.15,16 ureC gene was identified by PCR in 41(28.9%) cases in our study. Similar findings were observed in another study where 53% of H. pylori cases were detected using ureC gene PCR. 17
In our study, out of 142 total cases, 49 (34.5%) were identified as H. pylori-positive, and 50 (35.2%) were defined as H. pylori-negative. Forty-three (30%) cases were found to be positive only by one test. Similar findings were reported by one Iranian and another Bangladeshi study.18,19 In contrast to our study, H. pylori detection rates were identified to be 76.7% and 60.2% in other studies, which is much higher than what we reported here.15,20
There are several reasons behind such discrepancies in findings. For example, H. pylori infection rates vary across different geographic areas, with several factors such as socioeconomic conditions of patients, detection methods used, and sampling error during gastric biopsy-based testing playing important roles. 21 In our study, a small number of biopsy tissue specimens (1-2 in most cases) were used to detect the presence of infection. Besides, the uneven distribution of pathogens within gastric mucosa could also contribute to the lower detection rate of the organism in our study. Also, with the improved hygienic and socioeconomic conditions in this region, there is a gradual decline in H. pylori infection detection, as reported by some Asian and Middle Eastern studies.22-24
Regarding sensitivity and specificity of 3 invasive tests, RUT was found to be the most sensitive (94.2%), which is close to findings from other studies 95% and 96%.25,26 In contrast, specificity was lowest, at 55.5%, compared to 78.5% and 96.1% in other studies.26,27 More prolonged reading time, in absence of inhibitors, causing bacterial overgrowth of in-house RUT might have resulted in 40 (28.1%) cases being assigned to the indeterminate group. This is because RUT increases sensitivity at the expense of specificity with longer incubation time. 28
If we compare our results with other noninvasive methods serological assay was found to be 54% sensitive and 91.4% specific by a Bangladeshi study. 18 When compared with other noninvasive methods serological assay showed 85% sensitivity and 79% specificity. Stool antigen detection by ELISA showed 93.3% sensitivity; 93.2% specificity and UBT showed 96% sensitivity and 93% specificity. 29
In this study, 3 (6.1%) cases were found to be PCR and histology positive but RUT negative. Formalin contamination of biopsy forceps might have resulted in false-negative results. 30 A small gastric biopsy containing a small number of organisms might also be the contributing factor of negative RUT, as approximately 105 copies of bacteria should ideally be present in the sample for a test to return as positive. 4
PCR of ureC gene showed 83.0% sensitivity and 96.1% specificity in our study, similar to an Iranian study where sensitivity and specificity were 93.5% and 95.6%, respectively. 25 Accuracy of PCR was highest, at 81.1%, as molecular methods are known to be rapid and sample transport conditions are said to have little influence on the accuracy of the test. In this study, 10 (20.4%) cases were found to be PCR negative for ureC gene, whereas positive by the other 2 tests as the colonization of H. pylori in the gastric mucosa occurs in a patchy manner so obtaining a biopsy sample without any bacteria or fewer than 50 bacteria may produce misleading results in PCR. 31
Among the 3 invasive tests used in our study, histology was most specific (98.0%) with a sensitivity of 76.5%, compared to 77.8% specificity and 95.6% sensitivity in another study. 25 The sensitivity and reliability of histology depend on several factors including the number and the site of biopsy specimen collection. Ideally, a total of 4 (2 from the antrum of the gastric mucosa and another 2 from the corpus area) specimens should be collected. 4 In this study we used only 1 or 2 samples for histological examination, which might have had a detrimental effect on diagnostic accuracy. In addition, several previous studies display significant variation in results across observers, suggesting that the pathologist’s skills are essential when it comes to H. pylori detection.32,33
Conclusion
Although noninvasive methods like urea breath test can be more sensitive and less cumbersome for patients at times, invasive methods can be a cheaper alternative in patients already needing endoscopy. In conclusion, the diagnostic accuracy of the tests for H. pylori can be arranged in the following order: PCR > histology > RUT. Although RUT was the most sensitive method it was the least specific. Histology is the more specific one, and readily available and cost-effective than PCR in our country. So histopathology is the most reasonable choice in our country as it can also diagnose the disease. With the emergence of the recent COVID-19 pandemic, PCR machines have become an essential part of diagnostic laboratories. In the near future when the need for RT PCR for SARS-COV2 will decline these machines can be repurposed to utilize readily available kits to provide rapid detection of H.pylori.
Footnotes
Acknowledgements
We are grateful to the department of gastroenterology of Dhaka Medical College Hospital for their support.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Partial funding for this study was received from Bangladesh Medical Research Council (BMRC).
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Conceptualization: SK, SA, CKR, SHAH
Methodology: SK, SA, MMR, SHAH, CKR
Laboratory work and data analysis: SK, NNT, RA, MMR
Writing first draft: SK
Writing- editing and revision: MMAM
Supervision: SA, CKR, SHAH
Data availability statement
Data would be made available by the corresponding author for anyone interested upon request.
Ethical statement
This study received ethical clearance from Institutional Review Board (IRB) at Bangabandhu Sheikh Mujib Medical University.
