Abstract
Coriaria myrtifolia occurs as natural flora of warm temperate climates of northern Algeria which commonly found in hedges, forest and ravine edges. This actinorhizal species was known to establish a mutualistic symbiosis with members of phylogenetic cluster 2 (including strains associated to Coriaria spp., Ceanothus, Datiscaceae, and Dryadoideae) within the genus Frankia. Attempts to isolate C. myrtifolia microsymbionts from native plants growing in 4 locations in Algeria permitted to only recover asymbiotic Frankia strains (unable to reestablish nodulation and to fix nitrogen) from phylogenetic cluster 4 and several non-Frankia actinobacteria including members of Micrococcus, Micromonospora, Nocardia, Plantactinospora, and Streptomyces genera. The biodiversity of Frankia microsymbionts of C. myrtifolia root nodules was assessed using PCR-amplification followed by partial nucleotide sequencing of glnA1 (glutamine synthetase type 1) gene. On the 12 different glnA1 gene sequences obtained in this study, 9 were detected for the first time, and were mainly closelyrelated to Mediterranean genotypes previously described in the Grand Maghreb countries (Morocco and Tunisia) and in Europe (France) but without clear separations from other cluster 2 genotypes.
Introduction
Coriaria myrtifolia is endemic to western Mediterranean area including Northern Africa of the Grand Maghreb.1,2 It is mainly found in hedges, forest, and ravine edges under warm temperate climates of the Rif in Morocco and coastal to tellian atlas sub-sectors of Algeria. As an actinorhizal plant, C. myrtifolia forms a symbiotic association with the actinobacterial genus Frankia, which results in the development of nitrogen-fixing root nodules.3-6 The Frankia strains 7 associated with C. myrtifolia form a distinct lineage 8 which has been grouped in cluster 2 together with other microsymbionts whithin Coriaria spp., Ceanothus, Datiscaceae, and Dryadoideae root nodules.9-12
Members of Frankia cluster 2 have long been considered as uncultivable actinobacteria. However, several Frankia strains which are unable to fulfill Koch’s postulates (non-nodulating and/or non nitrogen-fixing) have been established from Coriaria, Datisca, and Ceanothus species.13-15 These asymbiotic isolates have been affiliated to phylogenetic cluster 4 of the genus Frankia.9-12
In our previous studies we used a dual approach combining comparative genomics and physiological bioassays to successfully establish axenic cultures of 2 strains from Frankia cluster 2 for the first time.16,17 Here we report results of cultivation attempts of cluster 2 strains from root nodules collected from native C. myrtifolia in Algeria and we provided an overview of genetic diversity of the uncultured Frankia cluster 2 populations directly in root nodules using GlnA1 gene sequence.
Materials and Methods
Root nodules sampling: A total of 44 root nodules were collected from C. myrtifolia growing in 4 locations in Algeria (Figure 1) and consisted of 3 nodules from one tree in site GKB: Les Gorges de Kharrata, Bejaia (N 36°31′9.379″, E 5°16′45.616″, 314 m), 11 nodules from 2 trees in site CDB: Les Cascades de Kefrida, Derguina, Bejaïa (N36°34′14.634″, E5°17′24.175″,550 m), 12 nodules from 3 trees in site PGM: La Plage de la Grotte Merveilleuse Jijel (N36°41′4946″ E5°31′45.8322″,15 m) and 18 nodules from 3 trees in site PR: La Plage Rouge, Ziama mansoria, Jijel (N36°39′24.289″, E5°26′5.866″,3 m).
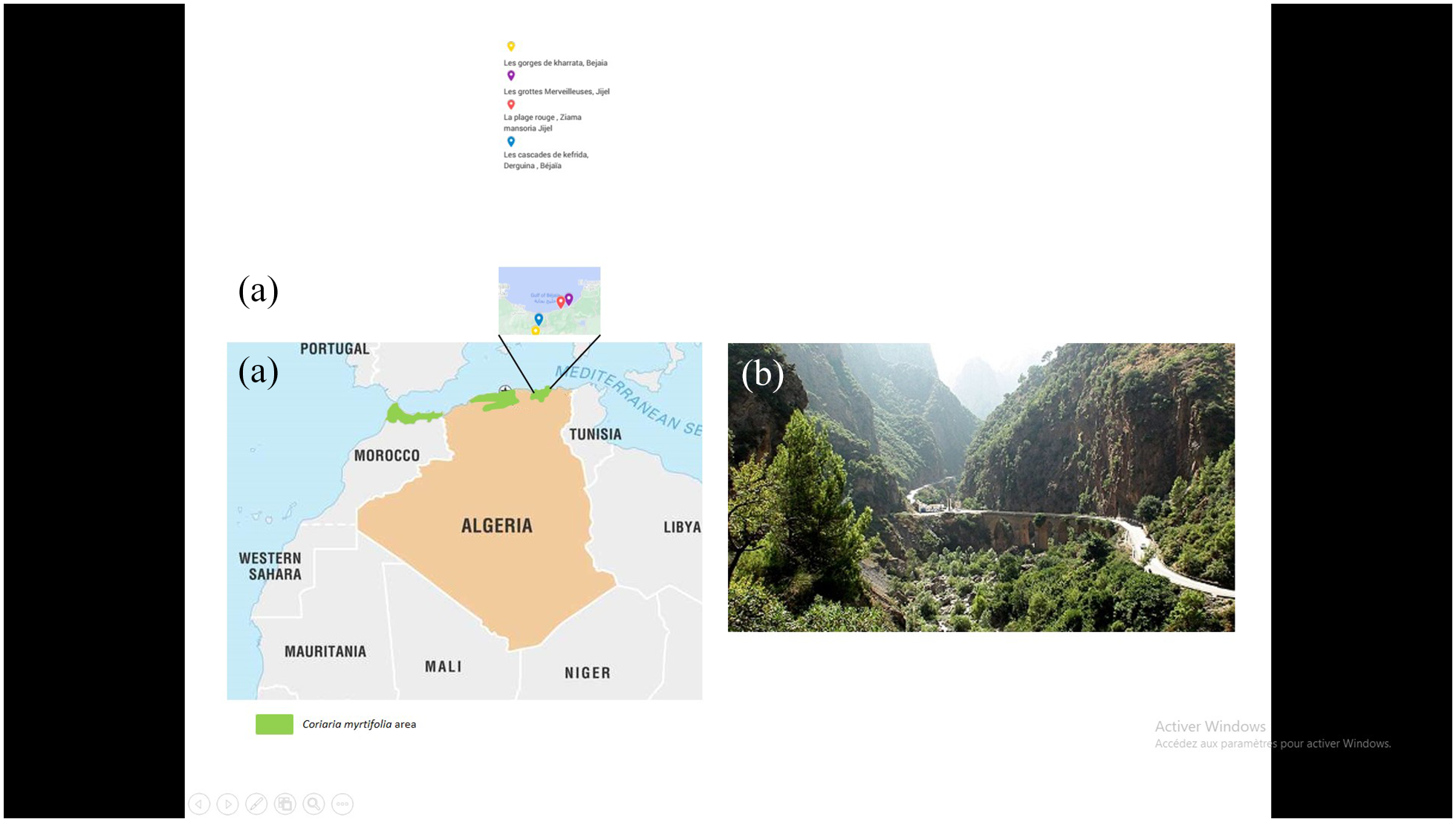
Distribution area of Coriaria myrtifolia in Algeria (A), Coriaria myrtifolia strand in Les gorges de kharrata, Bejaia, Algeria.
Microsymbiont isolation: Harvested root nodules were processed as described by Ghodhbane-Gtari et al 18 . After cleaning thoroughly with sterile tap water individual lobes (n = 833) were surface sterilized by shaking in 30% (v/v) H2O2 for 30 minutes and aseptically rinsed several times with sterile distilled water. To check the efficiency of nodule disinfection, an aliquot of 300 μl of the last washing solution was inoculated on nutrient agar plates and incubation for several days. Nodules resulting showing contamination-free washing liquid were further considered for nodule microsymbiont isolation by incubation of aseptically crushed individual lobe in BAP medium 19 buffered to range of pH (pH 6.8-pH 9) and supplemented with several organic acids as carbon source including: acetic acid, aspartic acid, butyric acid, citric acid, glutamic acid, malic acid, propionic acid, pyruvic acid, succinic acid, ribose uridine and thymidine and at a final concentration of 10 mM. 16
DNA extraction, PCR amplification, and sequencing
DNA extraction from root nodules was performed as described by Gtari et al 20 . Individual lobe from the 44 root nodules were surface sterilized as described for microsymbiont isolation protocol, ground in nitrogen liquid and taken up in the extracting buffer (100 mmol l−1 Tris-HCl [pH 8]; 20 mmol l−1 EDTA [pH8.2]; 1.4 M NaCl, and 2% w/v cetyl trimethyl ammonium bromide; CTAB). For cultivated strains, DNA was extracted from 1 month-old liquid cultures for Frankia strains and for non-Frankia actinobacteria from 3 to 5 well-grown colonies on solid Luedemann medium. After washing 3 times in sterile saline buffer, bacterial cells were taken up in the same extracting buffer used for nodules and forced several time through a 0.7 ×30 mm sterile needle to homogenize the mycelium. The resulting pellets, from grinded nodules or bacterial cells collected through centrifugation (3000 rpm for 30 minutes), were used for further DNA extraction steps. After incubation for 30 minutes in extracting buffer, DNA was chloroform extracted and ethanol precipitated through centrifugation (12 000 rpm, 30 minutes at 4°C), the DNA pellet was dissolved in 50 μl TE (10 mmol l−1 Tris-HCl [pH 8]; 20 mmol l−1 EDTA [pH 8.2]). PCR amplifications were performed using universal primers S-D-Bact0008-a-S- 20 (5′- AGAGTTTGATCCTGGCTCAG-3′) and S-DBact-1495-a-A-20 (5′-CTACGGCTACCTTGTTACGA-3′) 21 for almost full 16S rRNA gene and primers DB41 (5′-TTCTTCATCCACGACCCG-3′) and DB44 (5′-GG CTTCGGCATGAAGGT-3′) 22 for 477 bp fragment of glnA1 gene, in 50 µl final reaction volume containing 10 ng genomic DNA, 1x Taq polymerase buffer (Promega, Madison, WI, USA), 1.5 mmol l–1 MgCl2, 0.1 µM each dNTP, 0.2 µM each primers and 2 U Taq DNA polymerase. The thermal program consisted of 3 minutes at 95°C followed by 35 cycles of 94°C for 30 seconds, 55°C for 30 seconds, and 72°C for 45 seconds. PCR-products were purified using the QIAquick Wizard PCR purification Kit (Promega, Madison, VVI, USA) and then cycle-sequenced in both directions using an ABI cycle sequencing kit (Applied Biosystem 3130) as described by Ghodhbane-Gtari et al 18 . The nucleotide sequences of glnA1 and 16S rRNA genes were aligned using ClustalW 23 and compared to corresponding homologous sequences retrieved from GenBank. Phylogenetic trees were constructed using MEGA version 11. 24 Bootstrap values were determined from 1000 replicates. 25
C. myrtifolia inoculation test
To fulfill Koch’s postulates, plant inoculation experiments were performed as previously described. 16 Seeds of C. myrtifolia, locally collected from Algeria were surface sterilized by shaking in 30% (v/v) H2O2 for 30 minutes and aseptically rinsed several times with sterile distilled water, germinated on moistened filter paper under daylight illumination at 28°C and then transferred to sterilized moist sand. Collected cells from 4 weeks old Frankia cultures were washed several times with sterile distilled, water crushed in sterile glass homogenizer and used to inoculate 3 week-old C. myrtifolia seedlings. Seedlings were weekly checked for nodulation for 3 months. Seedlings inoculated with Frankia coriaria strain BMG5.1 and sterile water were used as positive and negative controls respectively.
Results
Isolation of Frankia from C. myrtifolia root nodules
On the 833 lobes derived from the 44 sampled root nodules and used for isolating Frankia micosymbionts, only 3 isolates have been obtained in BAP medium from La Plage de la Grotte Merveilleuse Jijel site. These 3 isolates showed 2 of the 3 typical morphological structures of the genus Frankia; hyphae and sporangia but never produce vesicles (Figure 2). The strains were unable to grow on nitrogen free BAP medium and to reinduce root nodules on C. myrtifolia seedlings. Based on partial glnA1 gene and 16S rRNA gene sequences the 3 isolates represent the almost the same strain which clustered within phylogenetic cluster 4 (Figure 3).
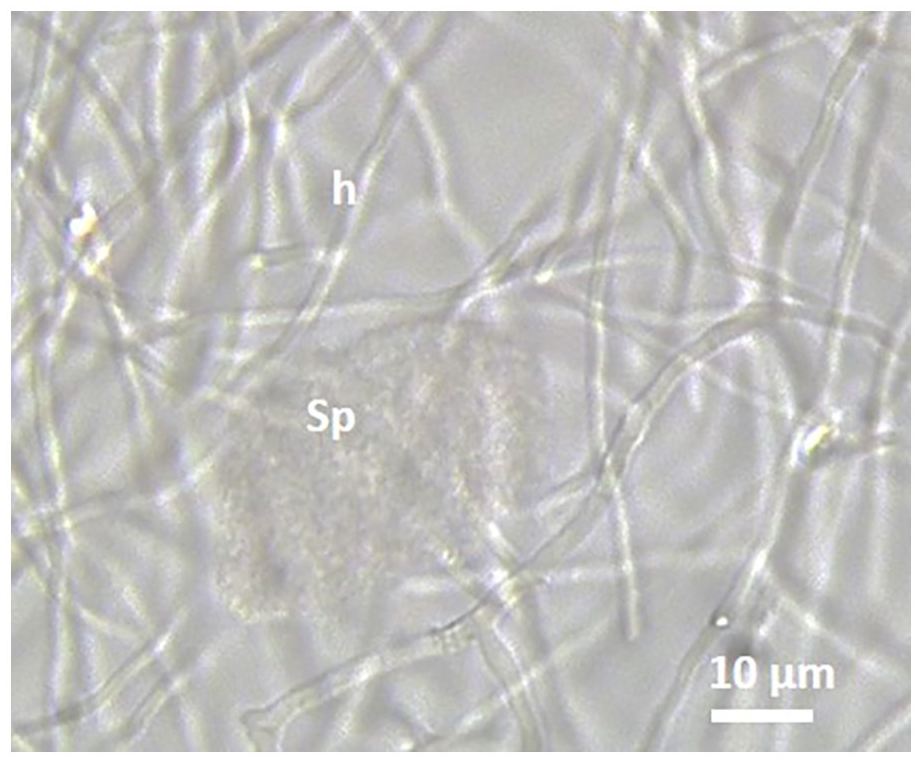
Phase-contrast microscopy of strain BMG5.36 grown for 4 weeks at 28°C on liquid BAP medium showing the 2 morphological structures; hyphae (h) and sporangia (sp) but never produces vesicle.
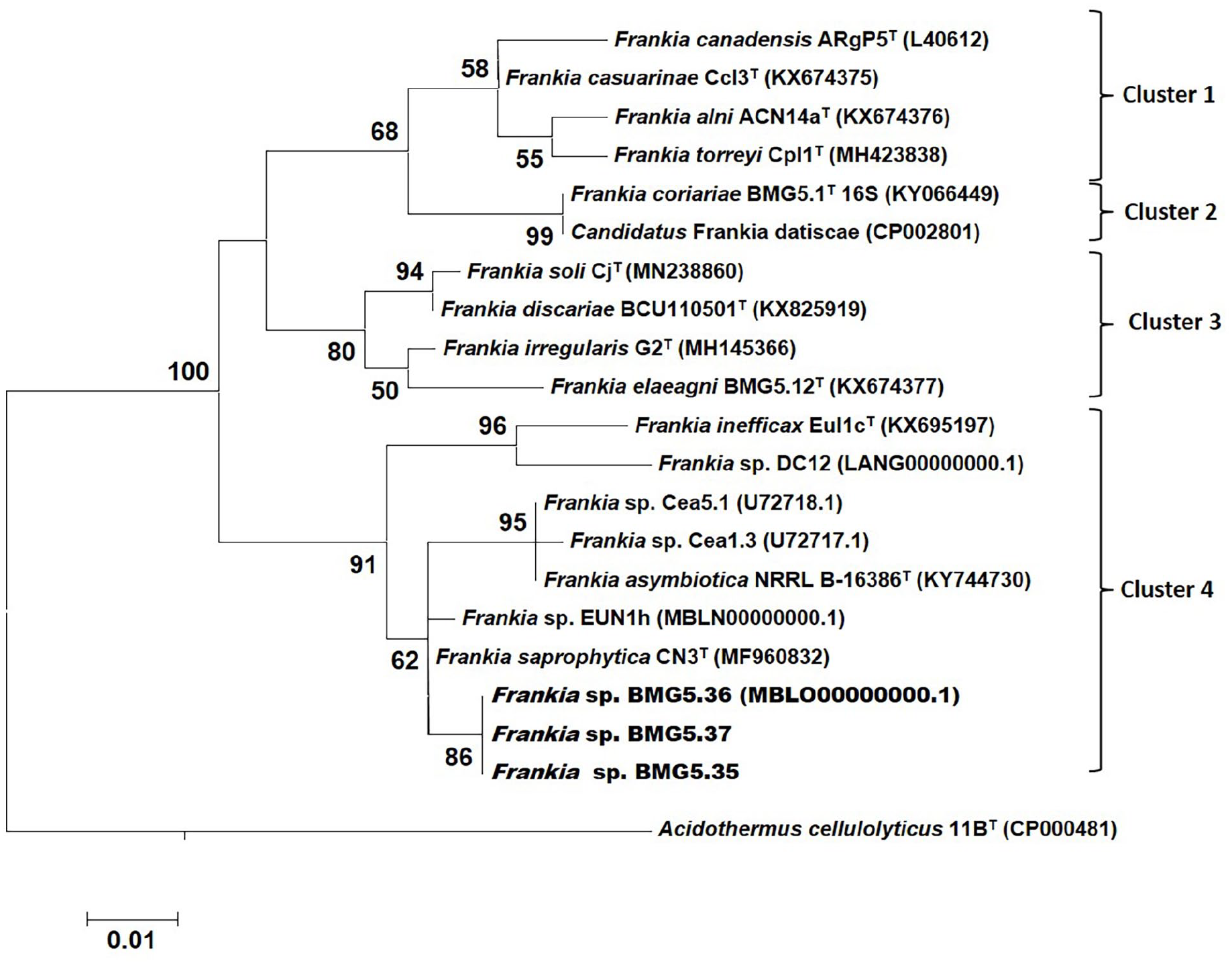
Maximum-likelihood tree based on partial 16S rRNA gene showing the phylogenetic position of Frankia isolated from C. myrtifolia root nodules sampled in Algeria (in bold). Acidothermus cellulolyticus strain 11BT (CP000481) was used as out-group. Bootstrap values (in %) determined from 1000 replicates. Only values above 50% were shown.
Isolation of non-Frankia actinobacteria from C. myrtifolia root nodules
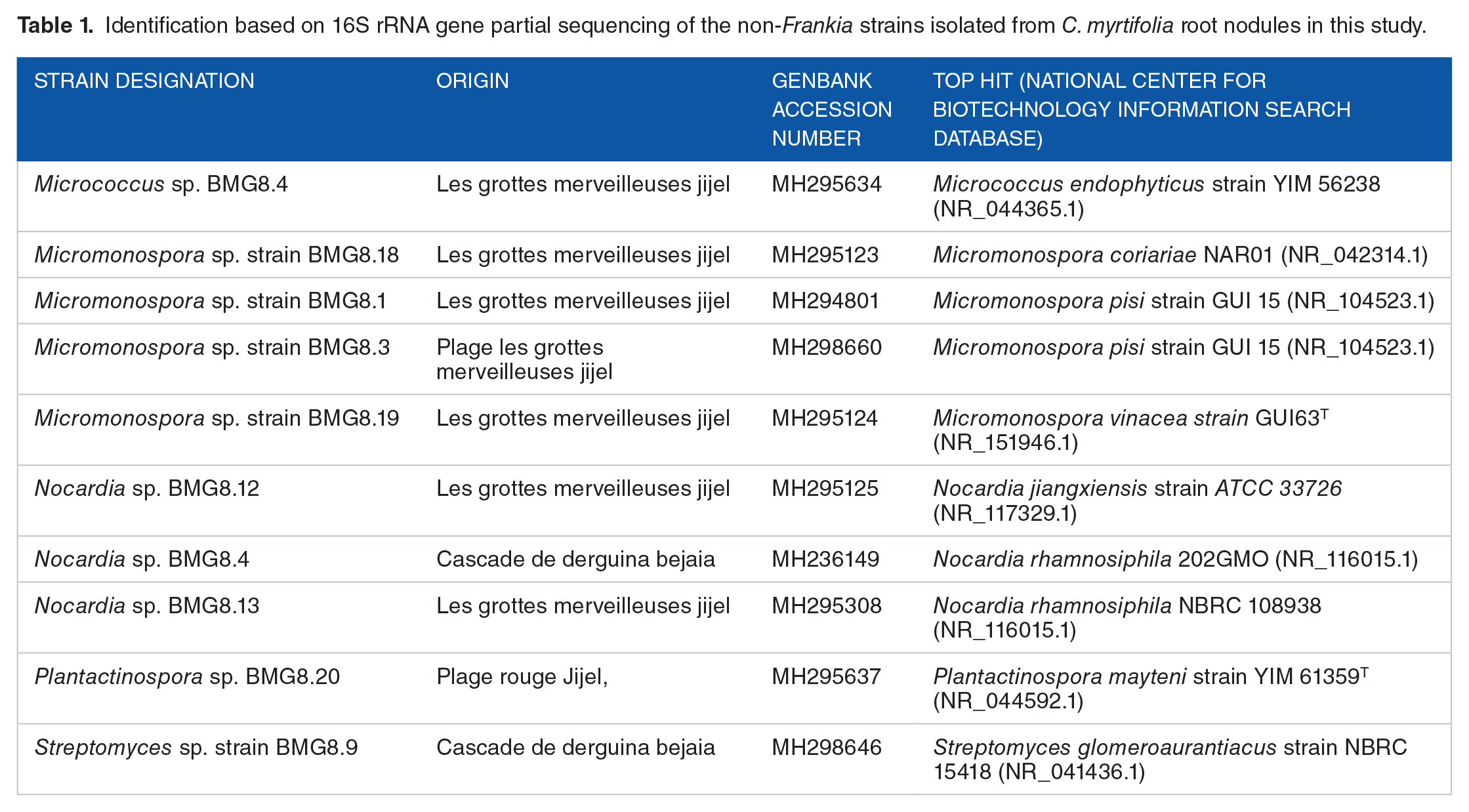
After incubation of 833 lobes in isolating BAP medium, 160 showed contaminations of fast growing fungi and bacteria within 2 to 5 days of incubation that have been discarded. These fast growing contaminants may have resisted the sterilization procedure or have happened during the rest of manipulation. Other vials showed slowly outgrowing mycelia which shown obvious substantial improvement of growth rate after transfer on solid BAP medium containing nitrogen source. Based on 16S rRNA gene sequence analysis, these isolates were assigned to members of Micrococcus, Micromonospora, Nocardia, Plantactinospora, and Streptomyces genera (Table 1).
Identification based on 16S rRNA gene partial sequencing of the non-Frankia strains isolated from C. myrtifolia root nodules in this study.
Overview of genetic diversity of cluster 2 microsymbionts
PCR-sequencing of the partial glnA1 gene was performed on DNA directly extracted from single lobes representing each C. myrtifolia tree growing in the 4 locations in Algeria. BLAST (Basic local alignment search tool) analysis 26 indicated that all obtained 24 sequences have the best BLAST hit with glnA1 sequences of Frankia from phylogenetic cluster 2 and none of them was affiliated to the asymbiotic phylogenetic cluster 4. Further multiple alignment of the total sequences showed that there are 11 different glnA1 gene sequences from Algeria. Nine of them differed from all previously described among C. mytiolia micosymbionts. Three subclusters may be defined (Figure 4) occurring in 4 sampling locations. A large subcluster (62.5% of obtained glnA1 gene sequences) includes C. mytifolia micosymbionts in 3 sites (Les Cascades de Kefrida, Derguina, Béjaïa, La Plage de la Grotte Merveilleuse Jijel and La Plage Rouge, Ziama mansoria, Jijel) which were homologous to those detected in Morocco 11 and in Tunisia 27 and France 11 together with microosymbionts detected in Datisca glomerata, Dryadoideae (Purshia tridentata, Cercocarpus betuloides, Cercocarpus ledifolius, C. foliolosa) and Ceanothus ( C. cuneatus, C. prostratus and C. integerrimus) root nodules sampled from North American Midwest in California, USA. 28 A second subcluster representing 16.6% of obtained glnA1 gene sequences in 2 sites (La Plage de la Grotte Merveilleuse Jijel and La Plage Rouge, Ziama mansoria, Jijel), together with microsymbionts from C. myrtifolia from France and Morocco, C. micophylla in Mexico and C. nepalensis in Pakistan 11 together with microsymbionts of Cercocarpus betuloides and Datisca glomerata in California. 28 A third subcluster representing 16.6% of analyzed sequences from 2 sites (Les Gorges de Kharrata, Bejaia and Les Cascades de Kefrida, Derguina, Béjaïa) were associated with C. myrtifolia microsymbionts from France and Morocco, C. japonica micosymbionts from Japan, and C. microphylla micosymbionts from Mexico. 11

Phylogenetic tree based on nearly 477-bp of glnA1 and inferred by neighbor-joining treeing method. GlnA1 sequences obtained in this study are in bold. Acronym of each sequence indicates site of collection; Cascade de kefrida, Derguina, Bejaïa (CDB), Plage Rouge, Ziama Mansoria, Jijel (PR), Plage de la Grotte Merveilleuse Jijel (PGM), and Les Gorges de Kharrata, Bejaia (GKB) and the plant number (1, 2, and 3) and the range of nodule (-, A, and B). Frankia saprophytica strain CN3 (NZ_KI912267.1) as out-group. Bootstrap values (in %) determined from 1000 replicates. Only values above 50% were shown.
Discussion
GlnA1 sequencing has been chosen to assess diversity of C. myrtifolia microsymbionts because it has been shown to be more variable than other genetic markers including 16S rRNA and nifH genes. 22 Our study of the native Frankia microsymbionts from this actinorhizal species using PCR-amplification and sequencing of glnA1 gene sequence showed a heterogeneity among the 4 sampling provenances in Algeria confirming previous reports for cluster 2 microsymbionts. By analyzing a large collection of cultivated strains and nodules, Vanden Heuvel et al 28 reported a low genetic diversity among Frankia microsymbionts in sympatric populations of actinorhizal species of Dryadoideae, Ceanothus, and Datisca glomerata in North American Midwest. Nouioui et al 11 also showed a low diversity of micosymbionts from 5 disjunct Coriaria species: C. myrtifolia, C. arborea, C. nepalensis, C. japonica, and C. microphylla, growing in the Mediterranean area (Morocco and France), New Zealand, Pakistan, Japan, and Mexico. These authors also reported the absence of cospeciation between the microsymbionts and their respective Coriaria species. Here we confirmed that the biogeography of sampling sites in Algeria does not affect the microsymbiont genotypes. Moreover, glnA1 gene sequences detected in this study were homologous to those retrieved from C. myrtifolia nodules collected in neighbor countries (Tunisia and Morocco) or in Europe (France) and even in other Coriaria species; C. nepalensis (Pakistan), C. japonica (Japan), and C. microphylla (Mexico) 11 and Dryadoideae, Ceanothus and Datisca glomerata in North American Midwest. 28 However, C. arborea micosymbionts still form a distinct sublineage of phylogenetic cluster 2. Nguyen et al 29 indicated that even at a global scale and for almost associate host species, the glnA1-based phylogeny did not result in an unambiguous separation of African, Eurasian and North American Cluster 2 strains.
Despite the utilization of the most recent protocol for cultivating cluster 2 microsymbionts which previously permitted the isolation of 2 symbiotic strains,16,17 no Frankia isolate able to fulfill Koch’s postulates was obtained in this study. While, 3 asymbiotic Frankia strains have been successfully isolated and cultivated in axenic conditions. Except few cases,30-32 almost all cluster 4 members have been established from cluster 2 host plant species; Coriariaceae, Datiscaceae, Dryadoideae, and Ceanothus.13-15 It is worth noting that no of these asymbiotic strains have been detected using the PCR-amplification and sequencing of glnA1 gene on DNA directly extracted from root nodules. This is coherent with several other studies that never reported asymbiotic Frankia strains based on similar molecular methods directly applied on root nodules of cluster 2 host species.8,11,18,22,28,33 However, Ramírez-Saad et al 34 were the first to report that beside symbiotic Frankia within root nodules of Ceanothus, asymbiotic stains were also present in the outer layers of the nodules. Recently other studies have confirmed this nodule surface emplacement of Casuarina incompatible Frankia strains.35,36
Alongside with the asymbiotic Frankia strains, the protocol used in the present study permitted the isolation of several other actinobacteria which were affiliated to Micrococcus, Micromonospora, Nocardia, Plantactinospora, and Streptomyces genera. Numerous are the studies which reported the isolation of non-Frankia actinobacteria from actinorhizal root nodules18,20,34,37-45 as it was shown before for legume nodules which contain both rhizobial and non-rhizobial bacteria.46,47 Three type strains; Micromonospora coriariae 42 ; Nocardia casuarinae 44 and Nocardia alni 48 have been isolated respectively from Coriaria; Casuarina and Alnus root nodules. Interestingly, a Micromonospora strain isolated in the present study is very closely related to M. coriariae species suggesting a widespread occurrence of members of this species within C. myrtifolia root nodules. Plant growth promoting proprieties have been attributed to several of these non-Frankia actinobacteria such as nitrogen fixation,20,40,41,49 phosphorous solubilization18,50,51 hormone production,18,44,51 siderophores, 18 or antagonisms against pathogenic fungal and insect predators.18,50 Stimulation of actinorhizal plant nodulation in co-inoculation condition with Frankia was also reported.41,52 More interesting 2 Nocardia strains, which have been isolated from Casuarina glauca nodules, were reported to use the Frankia infection pathway resulting in the induction of nodule-like structures in original host species and the promotion of its growth. 45
In conclusion native C. myrtifolia in Algeria harbors Frankia microsymbionts which are closely related to other microsymbionts of disjunct Coriaria species within phylogenetic cluster 2. These Frankia strains have resisted isolation and cultivation in axenic condition even with the most recent protocol we previously reported. 16 Instead we obtained strains that failed to reinduce root nodules on C. mlyrtifolia seedlings and to fix nitrogen, which have been assigned to phylogenetic cluster 4. The fact that asymbiotic Frankia strains were consistently isolated from cluster 2 host plant species merit further investigation to help clarify their ecological role and their evolutionary radiation in actinorhizal symbiosis. This study provided new non-Frankia actinobacteria that should enhance our understanding of rhizosphere microbiome associated with actinorhizal nodules.
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MG designed the study. AG, ML and HC-S sampled Coriaria myrtifolia nodules. AG, IM and FG-G performed the experiments and analysis. MG wrote the manuscript. All authors approved the submitted version.
