Abstract
Introduction:
Citrobacter spp. is an opportunistic bacteria that have been recognized as significant pathogens in patients with underlying diseases or immunocompromised status. The aim of this study was to identify extended-spectrum β-lactamases in clinical isolates of Citrobacter spp.
Methods:
This cross-sectional study was conducted at Hospital Central “Dr. Ignacio Morones Prieto” in San Luis Potosi, Mexico. Nineteen isolates of Citrobacter spp. were obtained from clinical specimens between April to December 2015. Four isolates were resistant to third-generation cephalosporins. The presence of genes encoding ESBL (blaCTX-M-15, blaTEM-1, blaVEB-1, blaSHV, and blaPER-1) was analyzed by PCR. For this purpose, plasmid DNA was extracted and horizontally transferred to recipient E. coli Top 10.
Results:
blaCTX-M-15 and blaVEB-1 genes were detected in Citrobacter freundii and Citrobacter sedlakii, whereas blaPER-1 gene was identified in 1 isolate of Citrobacter freundii. In contrast, blaSHV gene was not detected in any isolate. One strain carried blaCTX-M-15, blaTEM-1, blaVEB-1, and blaPER-1 genes, most in a 275-kb plasmid.
Conclusion:
This study shows the presence of different types of ESBL in clinical isolates of Citrobacter freundii and Citrobacter sedlakii, which confer resistance to broad-spectrum β-lactams. The plasmid identified in this study harboring ESBL genes could play an important role in the dissemination of antibiotic resistance.
Introduction
Bacteria of the genus Citrobacter belong to the family Enterobacteriaceae and comprise 13 species; Citrobacter are found in a variety of environmental sources, including soil and water, and occasionally are isolated from the gastrointestinal tract of animals and humans.1,2 Despite being considered an unusual nosocomial pathogen, neonates and immunocompromised patients are a frequent target of infections caused by these microorganisms. 3 These conditions include sepsis, urinary tract infections, respiratory and intra-abdominal infections, and central nervous system infections. 4 According a large observational study, Citrobacter species account for 0.8% of all Gram-negative infections in a hospital setting, with a mortality rate in hospitalized patients that ranged from 6.8% to 56%.3-5
As expected, infections caused by multidrug-resistant Citrobacter strains are associated with a higher rate of in-hospital mortality compared to those caused by susceptible strains. These multidrug-resistant strains show high levels of molecular class C (Amp-C) and extended spectrum β-lactamases (ESBL) as well as plasmid-mediated quinolone and carbapenem resistance. 2
ESBL confer resistance to penicillins, first to third-generation cephalosporins, and aztreonam, but not to cephamycins and carbapenems. 6 The production of ESBL has been recognized as a global problem, mainly in the case of E. coli and K. pneumoniae.7-9 However, this condition has been also described in recent years in other species, including the genus Citrobacter.7-9
Few studies have analyzed the presence and distribution of ESBL in clinical isolates of Citrobacter spp. Among them, an outbreak caused by 5 isolates of CTX-M-2 producing C. koseri in hematological patients was reported in 2006. 7 Furthermore, CTX-M-1/3 β-lactamases have been reported in Korea, France and Spain, CTX-M-9/30 in Canada, and CTX-M-14 in China.7,8 In addition, CTX-M-14, TEM, SHV-4, and SHV-12 ESBL have been detected in Japan.7,8
The aim of this study was to detect ESBL in clinical isolates of Citrobacter spp. at the Hospital Central “Dr. Ignacio Morones Prieto” (HCIMP), in the State of San Luis Potosi, located at the center-north of Mexico.
Materials and Methods
Bacterial isolates
This study was conducted after approval (July 23, 2015) by the Research Committee [COFEPRIS 14 CI 24 028 083] and the Research Ethics Committee of the Hospital Central “Dr. Ignacio Morones Prieto” [CONBIOETICA-24-CEI-001-20160427]. The registration number was 48-15. A written informed consent was obtained from all participants or legal guardians/parents for those under the age of 16 years.
The HCIMP has 250 beds and 32 beds in the intensive care unit and provides medical services to mid-and low-income populations from all over the State of San Luis Potosi.
Between July and December 2015, the Microbiology Laboratory of HCIMP identified 19 consecutive and non-repeated isolates of Citrobacter spp. The clinical isolates were identified using Vitek 2C (bioMérieux, Marcy l’Étoile, France) and were transferred to the Section of Medical Genomics from the Research Center of Health Sciences and Biomedicine, UASLP for molecular characterization.
ESBL phenotypic confirmatory test
The phenotypic confirmatory test of ESBL was performed by using the combined disk method on Mueller Hinton Agar, according to the Clinical and Laboratory Standards Institute (CLSI) recommendations, with Cefotaxime (CTX, 30 mg) and Ceftazidime (CAZ, 30 mg) disks, alone and in combination with Clavulanic Acid (CA, 10 mg) (Becton, Dickinson, Sparks, MD). Results were considered positive when the growth-inhibitory zone around either the CTX/CL or CAZ/CL disk was 5 mm or greater. 10
Molecular identification of β-lactamases
DNA amplification of β-lactamase genes was carried out with specific primers (Table S1). In brief, 3 colonies of an overnight culture were suspended in 100 μL of DNase free water and incubated at 94°C for 5 min and at −70ºC for additional 5 min. Then, tubes were centrifuged at 13 000 rpm for 5 min and the supernatant was used as DNA template. 11 The PCR reaction mixture contained 1X buffer, 2.0 mM of each dNTP, 5.0 µM of oligonucleotides for blaTEM, blaCTX-M, blaSHV, blaVEB-1, and blaPER-1 genes, 1.0 U of Taq DNA polymerase, bacterial genomic DNA, and 1.5-3.0 mM MgCl2. PCR conditions were performed at 94°C for 5 min for initial denaturation, followed by 30-40 cycles of 30 s at 94°C, 1 min at 50-60°C, and 1 min at 72°C, followed by a final extension of 5 min at 72°C, using a Multigene thermo-cycler (Labnet International Inc, New Jersey, United States). The amplified products were analyzed by electrophoresis on 2% agarose gels. PCR products were purified using a Wizard DNA Clean-Up system (Promega), according to the manufacturer’s instructions and were sequenced by using the dideoxynucleotide method, in a 3130 Genetic Analyzer device (Applied Biosystems, Foster City, California). BLAST analysis was performed in the NCBI http://www.ncbi.nlm.nih.gov/page.
Plasmid isolation
Plasmid DNA extraction to Citrobacter spp. (R-135 and R-086) was made by using the Plasmid Mini Kit (QIAGEN), which was performed through the following 3 steps: (1) Isolates grown in Luria Bertani medium for 12-16 h to a cell density of approximately 3.0–4.0 × 109 cells/mL were centrifuged at 6000g for 15 min at 4°C. Then, bacterial pellet was resuspended in 0.3 mL of Buffer P1, followed by the sequential addition of P2 and P3 buffers. After an additional centrifugation, the supernatant containing the plasmid was obtained. (2) Supernatant was loaded onto an anion-exchange Genomic tip (QIAGEN 20/G) and incubated for 20 min. (3) Plasmid DNA was eluted from the tip with 0.8 mL of QF buffer. Then, DNA was precipitated with isopropanol, centrifuged at 15 000 rpm for 30 min. Finally, the DNA pellet was washed with 1 mL of 70% ethanol, centrifuged and dissolved in a 10 mM Tris-HCl, pH 8.5. Extracted DNA was visualized on a 1% agarose gel.
Bacterial transformation
In brief, E. coli Top 10 bacteria was permeabilized with 75.0 mM calcium chloride and incubated with plasmid DNA (50-100 ng/µL) and heat-shocked at 42°C for 45 s/ice for 5 min. Afterward, bacteria were resuspended in 250 µL of Luria Bertani (LB) broth and incubated at 37°C under continuous shaking for 1 h. Finally, transformed bacteria were selected by incubating on LB/ampicillin (100 µg/mL) agar plates at 37°C for 24 h. ESBL genes in these bacteria were detected as described above.
Antimicrobial susceptibility testing
Antimicrobial susceptibility testing was carried out by using the broth microdilution method, according to CLSI recommendations. Briefly, antibiotics were serially diluted 2-fold in 50 μL of Mueller-Hinton broth, mixed with 50 μL of bacteria at a density of 106 colony-forming units/mL and incubated for 18 h at 37°C. Quality control was performed using the reference E. coli Top 10 and results were expressed as the minimum inhibitory concentration (MIC).
Results
Bacterial isolates
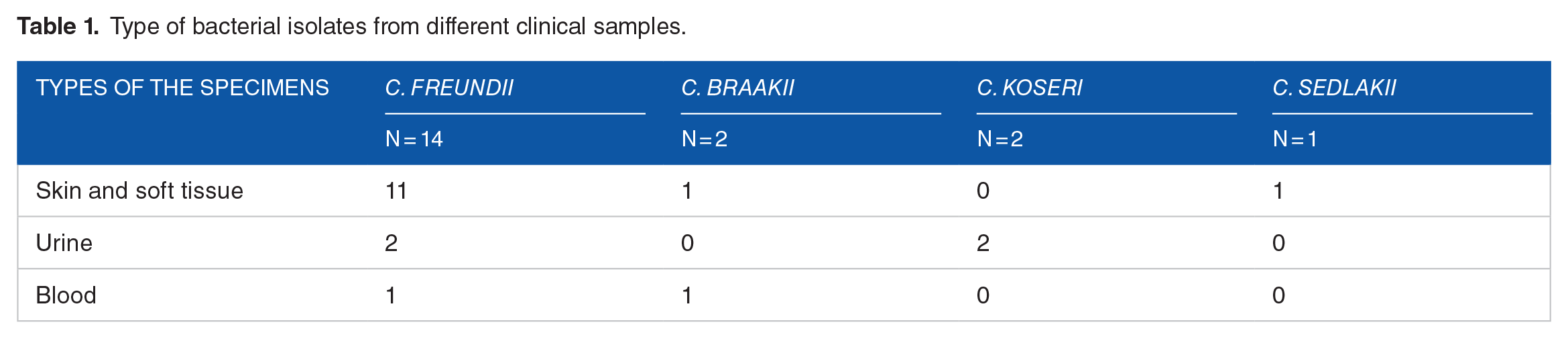
The Microbiology Laboratory identified 14 isolates of C. freundii, 2 isolates of C. koseri, 2 isolates of C. braakii, and 1 isolate of C. sedlakii in the different samples studied (Table 1).
Type of bacterial isolates from different clinical samples.
ESBL phenotypic confirmatory test
Three isolates resistant to third-generation cephalosporins showed an ESBL phenotype: C. freundii R-086, R-135, and C. sedlakii R-099. Moreover, the C. freundii R-134 isolate, which was also resistant to third-generation cephalosporins, did not exhibit a resistance phenotype indicative of the presence of ESBL (Table 2).
Phenotypic and genotypic detection of ESBL for strains studied.
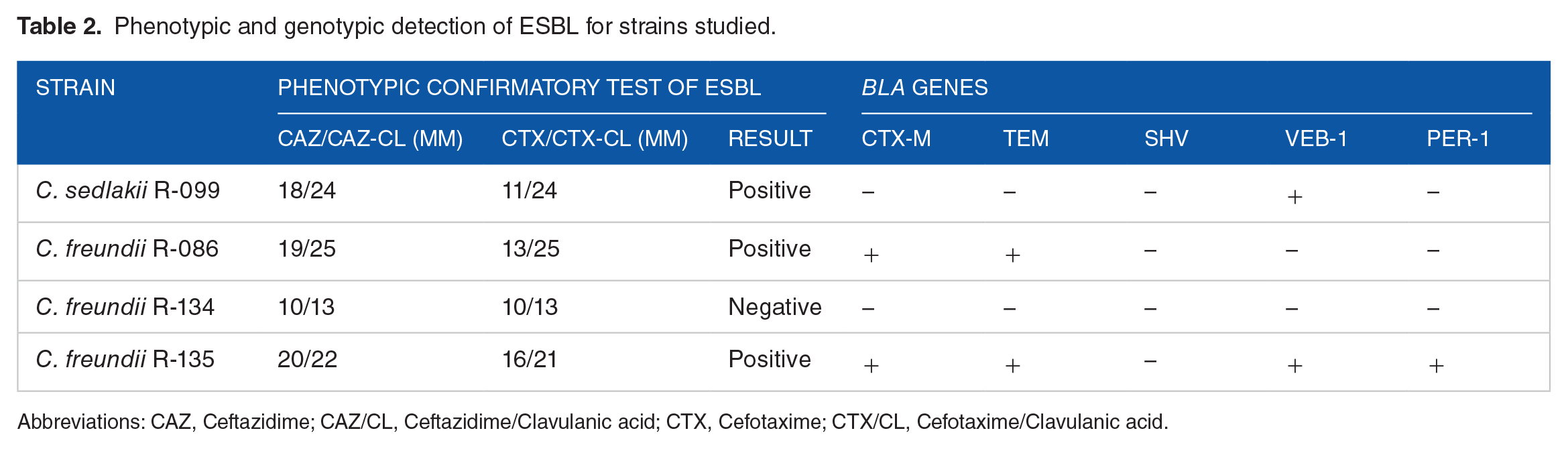
Abbreviations: CAZ, Ceftazidime; CAZ/CL, Ceftazidime/Clavulanic acid; CTX, Cefotaxime; CTX/CL, Cefotaxime/Clavulanic acid.
Molecular identification of β-lactamases
As shown in Figure 1, blaTEM and blaCTX-M genes were identified in C. freundii R-086 and R-135. In 2 additional isolates (C. freundii R-135 and C. sedlakii R-099), blaVEB-1 genes were detected Figure 1c, whereas the blaPER-1 gene was identified in C. freundii R-135 Figure 1d. In contrast, the blaSHV gene was not identified in any isolate (data not shown). Moreover, blaTEM, CTX-M, VEB-1, and PER-1 genes were identified in the C. freundii R-135 isolate (Table 2). Finally, DNA sequence analysis showed that the TEM β-lactamase detected in C. freundii R-086 had 100% homology to TEM-1 (GenBank accession number: ALW82937.1.), whereas the predicted amino acid sequence of the CTX-M β-lactamase showed 99% homology to CTX-M-15 (GenBank accession number: AKO22374.1).
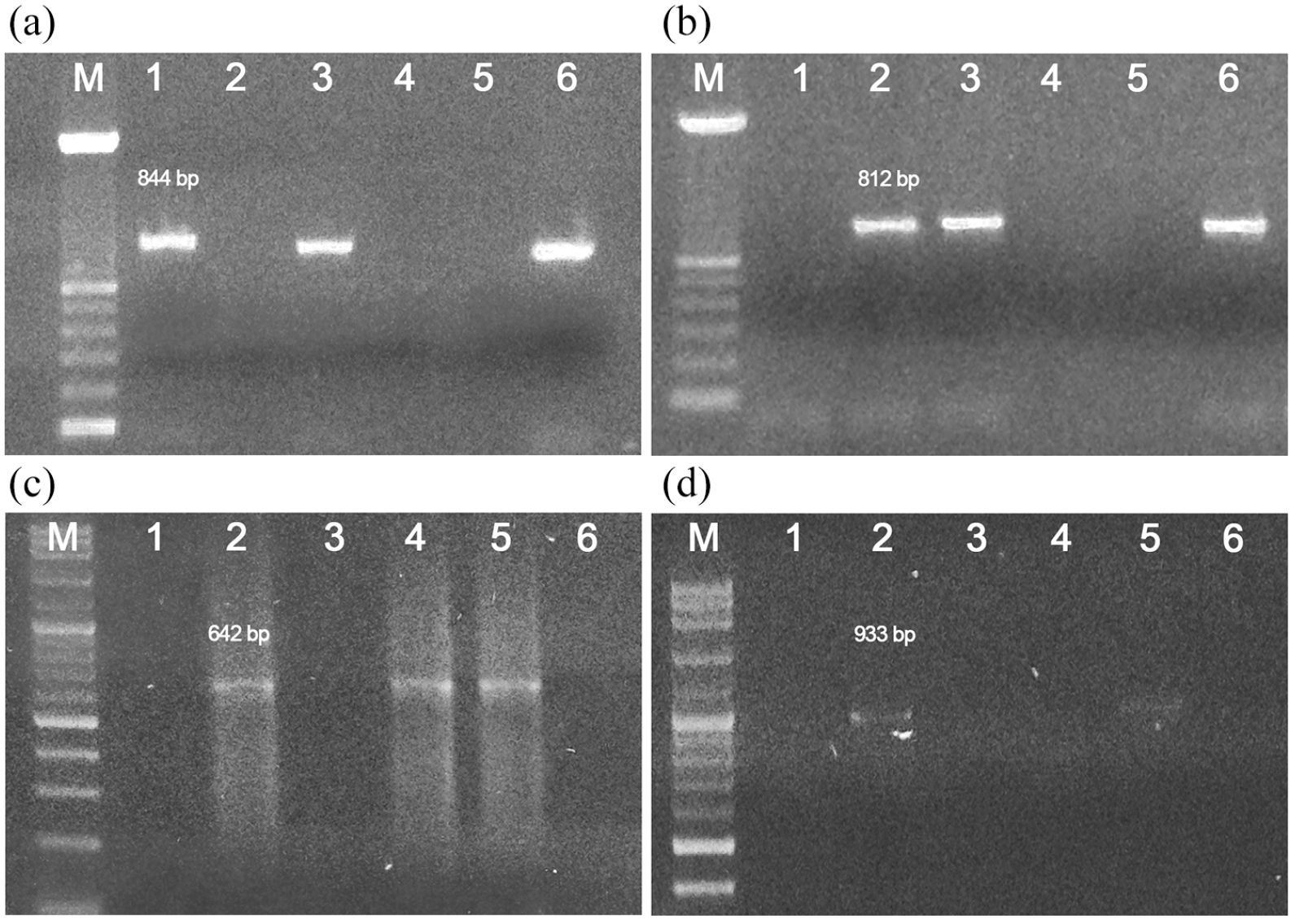
Agarose gel electrophoresis for the PCR products of ESBL: (a) the blaTEM gene. Line M: 100 bp ladder, line 1: positive control, line 2: negative control, lines 3 and 6: R-086 and R-135 strains (blaTEM positive), lines 4 and 5 blaTEM negative strains, (b) the blaCTX-M gene. Line M: 100 bp ladder, line 1: negative control, line 2: positive control, lines 3 and 6: R-086 and R-135 strains (blaCTX-M positive), lines 4 and 5: R-099 and R-134 strains (blaCTX-M negative), (c) the blaVEB-1 gene. Line M: 100 bp ladder, line 1: negative control, line 2: positive control, lines 3 and 6: R-086 and R-134 strains (blaVEB-1 negative), lines 4 and 5: R-099 and R-135 strains (blaVEB-1 positive), and (d) the blaPER-1 gene. Line M: 100 bp ladder, line 1: negative control, line 2: positive control, line 5: R-135 strain (blaPER-1 positive), lines 3, 4, and 6: blaPER-1 negative strains.
Plasmid isolation and bacterial transformation
A 275-kb plasmid DNA was isolated from C. freundii R-086 (R-086p) and R-135 (R-135p) strains (data not shown). On the other hand, In the Top 10-R-135 E. coli transformant we were able to amplify the blaCTX-M-15, PER-1, and TEM-1, genes Figure 2. In contrast, the blaVEB-1 gene was not amplified in R-135p or Top-10-R-135.
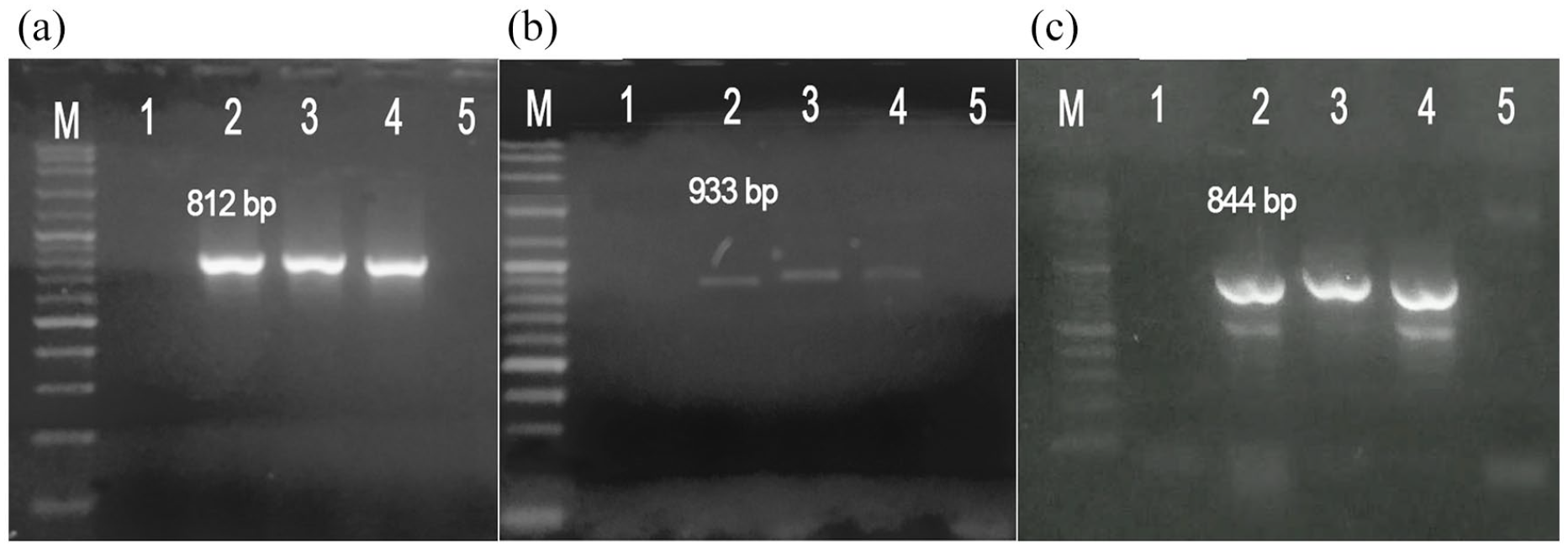

Agarose gel electrophoresis for the PCR products of Top10R-135 strain: (a) the blaCTX-M-15 gene. Line M: 100 bp ladder, line 1: negative control, line 2: R-135 strain (blaCTX-M-15 positive), line 3: R-135p, line 4: Top10R-135 (transformant strain), and 5: E. coli Top-10, (b) the blaPER-1 gene. Line M: 100 bp ladder, line 1: negative control, line 2: R-135 strain (blaPER-1 positive), line 3: R-135p, line 4: Top10R-135 (transformant strain), and 5: E. coli Top-10, and (c) the blaTEM-1 gene. Line M: 100 bp ladder, line 1: negative control, line 2: R-135 strain (blaTEM positive), line 3: R-135p, line 4: Top10R-135 (transformant strain), and 5: E. coli Top-10.
Antimicrobial susceptibility testing
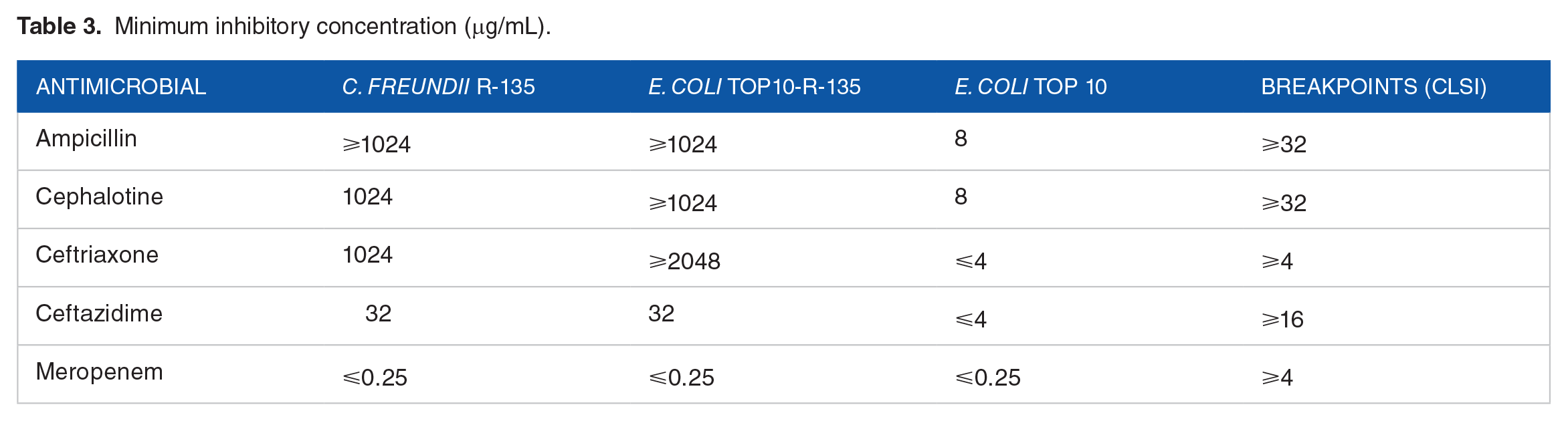
The MIC of antimicrobial agents for C. freundii R-135, E. coli Top 10-R-135, and E. coli Top 10 are shown in Table 3.
Minimum inhibitory concentration (μg/mL).
Discussion
In recent years, infections caused by Citrobacter spp. have become important because they cause nosocomial outbreaks, mainly affecting neonates and immunocompromised patients. 12 Although C. koseri has been most frequently identified in sporadic cases and hospital outbreaks, 6 in our study, the predominant species was C. freundii.
Since its first description in the 1990s, ESBL of the CTX-M family have showed an increased frequency in Enterobacteriaceae, followed by SHV and TEM as well as, to a lesser extent, by the OXA, Tla, GES, VEB, and PER types. However, the frequency of these types is variable in different geographical regions. 6 In our study, we detected that the C. freundii R-086 and R-135 strains synthesized CTX-M-15, an ESBL worldwide distributed and characterized by its efficient ceftazidime hydrolyzing activity.13,14 The first report of CTX-M-15 in Mexico was in 2011, in E. coli isolate, 11 when 2 different groups reported strains of E. coli, Enterobacter cloacae, and K. pneumoniae as carriers of CTX-M-15 ESBL.9,15 Two years later, 58 isolates of E. coli and 16 isolates of K. pneumoniae producers of CTX-M-15 ESBL were detected in a tertiary care hospital in the city of Guadalajara, State of Jalisco, México. 11 The present study is the first report of Citrobacter spp. able to synthesize CTX-M-15 ESBL in Mexico. In the same strain the TEM-1 β-lactamase was identified, which efficiently hydrolyze penicillins and low spectrum cephalosporins and that is not considered an ESBL.
The presence of VEB-1 ESBL in the genus Citrobacter had not been described so far and in the case of PER ESBL, there is only a previous report of C. koseri PER-2 producer. 16 This ESBL has emerged in clinical isolates of E. coli, Pseudomonas aeruginosa, Acinetobacter baumannii, Klebsiella spp., and Proteus mirabilis. 6
The first report on the resistance to β-lactams in C. sedlakii was an isolate resistant to aminopenicillins, carboxypenicillins, low spectrum cephalosporins (but sensitive to broad-spectrum cephalosporins, and carbapenems), which was mediated by a β-lactamase called Sed-1. 12 In our study, the C. sedlakii R-099 isolate was resistant to aminopenicillins, carboxypenicillins, broad-spectrum cephalosporins including cefepime. In this case, the PCR analysis showed the presence of VEB-1 but not of bla TEM, SHV, CTX-M or PER-1. At our best knowledge, this is the first report of a C. sedlakii strain able to produce VEB-1 ESBL.
We identified the blaTEM-1, and blaCTX-M-15 genes in C. freundii R-086, an isolated that also showed resistance to quinolones, aminoglycosides, and inhibitors of the folate pathway (data not shown). Moreover, in the C. freundii R-135 strain, blaTEM-1, blaCTX-M-15, blaVEB-1, and blaPER-1 genes were identified. This phenomenon may be related to the presence of genetic mobile structures such as integrons, transposons, and plasmids that have been previously described in other species of Gram-negative bacilli; In this regard, an analysis of 40 isolates of Citrobacter spp. performed in 2010, the simultaneous presence of blaCTX-M, ampC, SHV, and TEM genes was observed, and in 32.5% of these isolates class 1 integron was identified. In addition, in 48% of CTX-M-15 positive isolates the insertion sequence IS26 was detected. 17 In the same year, a class 1 integron and ISCR1 insertion sequence were identified in a multi-resistant C. freundii isolate, which showed resistance to quinolones and β-lactams by presence of CTX-M-15 ESBL. 18
Previous reports suggest that different plasmids are responsible for the global spread of CTX-M-15 ESBL 19 since blaCTX-M-15 has been identified in plasmids ranging from 40 to more than 200-Kb 20 and belonging to IncFII and IncI1 groups.20,21 In these mobile elements, more resistance determinants have been identified, as blaTEM-1 and blaSHV-12 genes. However, there is only 1 previous report on a plasmid containing the blaCTX-M-15 and blaPER-1 genes, which was detected an Aeromonas caviae strain, which was isolated from a wild-growing Mediterranean mussel. 22 Thus, this is the first report of a plasmid R-135p carrying the blaCTX-M-15, TEM-1, and PER-1 genes in C. freundii. The genetic context of blaCTX-M-15 includes an ISEcp1 sequence, upstream of the gene, which can be an efficient factor for the mobilization and expression of blaCTX-M-15, as it has been observed in previous reports.19,23
We consider that it would be important to investigate the genetic context of β-lactamase genes in both, plasmids and chromosome as well as to determine the structure of the plasmid identified in C. freundii R-135.
In conclusion, our study shows the presence of different ESBL types in clinical isolates of C. freundii and C. sedlakii, which mediate the resistance to broad-spectrum β-lactams. The simultaneous presence of several antibiotic resistance genes seems to be related to genetic mobile elements that may favor their dissemination.
Supplemental Material
sj-docx-1-idr-10.1177_11786337211065750 – Supplemental material for Plasmid Carrying blaCTX-M-15, blaPER-1, and blaTEM-1 Genes in Citrobacter spp. From Regional Hospital in Mexico
Supplemental material, sj-docx-1-idr-10.1177_11786337211065750 for Plasmid Carrying blaCTX-M-15, blaPER-1, and blaTEM-1 Genes in Citrobacter spp. From Regional Hospital in Mexico by Cindy Negrete-González, Edgar Turrubiartes-Martínez, Miriam Briano-Macias, Daniel Noyola, Luis Fernando Pérez-González, Roberto González-Amaro and Perla Niño-Moreno in Infectious Diseases: Research and Treatment
Supplemental Material
sj-docx-2-idr-10.1177_11786337211065750 – Supplemental material for Plasmid Carrying blaCTX-M-15, blaPER-1, and blaTEM-1 Genes in Citrobacter spp. From Regional Hospital in Mexico
Supplemental material, sj-docx-2-idr-10.1177_11786337211065750 for Plasmid Carrying blaCTX-M-15, blaPER-1, and blaTEM-1 Genes in Citrobacter spp. From Regional Hospital in Mexico by Cindy Negrete-González, Edgar Turrubiartes-Martínez, Miriam Briano-Macias, Daniel Noyola, Luis Fernando Pérez-González, Roberto González-Amaro and Perla Niño-Moreno in Infectious Diseases: Research and Treatment
Footnotes
Acknowledgements
We thank to María Anita de Lira Torres, Andrés Flores Santos, and Laura Cerda Ramos, the staff of the Microbiology Laboratory of the HCIMP for their support in the collection of strains. We thank Adriana Martínez Rodríguez for her excellent laboratory assistance and Francisco Rodríguez Velázquez for his valuable technical support on this project.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Grants CONACyT-SALUD-142334, and CONACyT-CB2011-169567, to PNM. The funding agency had no role in the study design, sample collection, data collection and analysis, decision to publish, or preparation of the manuscript.
Author Contributions
CNG acquired clinical data and samples, interpreted results, and drafted the manuscript. CNG and MBM performed the experiments. ETM and PNM co-designed and supervised the study and interpreted the results of experiments. DEN analyzed and interpreted data. RGA and LPG critically revised and edited the manuscript. All authors have read and approved the manuscript.
Ethical Statement
This study was conducted at Hospital Central Dr. Ignacio Morones Prieto (HCIMP) in San Luis Potosi, Mexico after approval by the Research Committee [COFEPRIS 14 CI 24 028 083] and the Research Ethics Committee of the HCIMP [CONBIOETICA-24-CEI-001-20160427]. The registration number was 48-15.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
