Abstract
The emergence and spread of antibiotic resistance (ABR) have been a public health challenge globally. The burden is even higher in low-income countries where there is a lack of appropriate healthcare systems, and inappropriate antibiotic disposal practices and utilization. Due to poor solid waste disposal practices in developing nations, municipal solid waste dumpsite (MSWDS) can be a reservoir for ABR bacteria. However, only a few studies demonstrated the prevalence of ABR in non-clinical environments such as MSWDS. This study assessed the prevalence of ABR bacteria at Bahir Dar City MSWDS, to understand the public health risks related to poor solid waste disposal systems. Nine soil samples were collected from the dumpsite. Bacteria were isolated, identified and tested for ABR. Seventy-one distinct colonies were isolated from all samples and identified into 10 bacterial genera based on morphological features and biochemical tests. For ABR tests, gentamicin (GN, 10 μg), streptomycin (ST, 30 μg), tetracycline (TE, 30 μg), ciprofloxacin (CIP, 5 μg), nalidixic acid (NAA, 30 μg), sulfonamide (SA, 250 μg), chloramphenicol (C, 30 μg), erythromycin (E, 15 μg), vancomycin (V, 30 μg), and amoxicillin (AMX, 25 μg) were used. The most frequently isolated bacteria were
Introduction
Antimicrobial resistance is one of the top 10 global public health and development threats causing 1.27 million deaths directly and contributing to 4.95 million deaths indirectly worldwide.1 -3 Bacterial pathogens are among the leading pathogenic microorganisms and have been posing serious public health problems by developing antibiotic resistance (ABR).1 -3 The problem of antimicrobial resistance is aggravated by the lack of interest by biotech companies in developing new antibiotics, as the pathogen will develop resistance to the new antibiotic sooner or later. 4 Continued efforts in the understanding of the ecology and resistance patterns of bacteria are important in tackling both the emergence and spread of antimicrobial resistance as well as associated disease burden at the national and global levels. 5 Most studies regarding the prevalence of ABR give attention only to samples from health institutions, in clinical environments. 6 Furthermore, studies on the natural resistome or from non-clinical environments are considered to be equally important.7 -9 There are thus few studies conducted regarding the prevalence of ABR from non-clinical sources, such as soil from any environment and waste dumpsite soils. There is growing interest in the understanding of natural antibiotic resistome in non-clinical sources,1,10 which can serve as a reservoir for resistance genes. 10
In general, ABR develops through natural processes that lead to genetic changes in pathogens. The emergence and spread of ABR are accelerated by human practices, mainly inappropriate use of antimicrobials in humans, animals and plants, and environmental contamination by improper disposal of unused and expired antimicrobials and public wastes, animal farms and pharmaceutical industries.2,11,12
Several studies reported the presence of ABR in domestic wastes.13 -16 These studies emphasized that continuous studies are needed to get deep insights into the real impacts of ABR on the public health system in non-clinical environments. Municipal solid waste dumpsites (MSWDS) are among the potential sources for the emergence of antibiotic-resistant microorganisms.14,17 -20 Mismanaged solid wastes may contain several hazardous chemicals, pharmaceutical wastes, residual antimicrobial agents and pathogenic microorganisms.21 -23 More importantly, untreated pharmaceutical wastes discharged into the dumpsite can kill susceptible microorganisms. 24 This selection pressure favors the resistant bacteria to emerge, which often results in mutations in the bacterial population.24 -26 This leads to a mass increase in antibiotic-resistant bacteria, as the dumped pollutants will wipe out the competing susceptible normal flora. 27 Horizontal gene transfer through various mechanisms can also accelerate the spread of resistance to susceptible strains of bacteria. 28 All of these processes can lead to the emergence and dissemination of new antimicrobial resistance variants with novel resistance mechanisms against various classes of commonly used antimicrobial agents. 24
Recent studies have reported a high prevalence of pathogenic bacteria from MSWDS.29 -33 Several ABR enteric bacteria were identified from dumpsites by Mwaikono et al. 32 Furthermore, a study reported a large diversity of ABR from waste dumpsite soil with both intrinsic and acquired resistance mechanisms. 33 A huge volume of solid waste is being generated in municipalities of populated countries like Ethiopia. However, limited research is available regarding the prevalence of antimicrobial-resistant bacteria from such solid waste dumpsites. The rates and mechanisms of resistance are influenced by geographic variations. 34 The type of determinant factors for the emergence of ABR within a locality vary greatly. The local abiotic and biotic factors and the types of waste generated vary from place to place. Microbial proliferation thus depends on several factors, such as local environmental conditions and types and the amount of nutrients available at the site. Therefore, the risks to public health from different wastes in municipal dumpsites vary greatly. 35 The prevalence of ABR bacteria varies from location to location, and can also change rapidly with time. As a result, ABR needs to be monitored and managed in every potential site because of their public health implications. Bahir Dar City MSWDS is among poorly managed solid waste dumpsites in Ethiopia. The area could be a hot spot for the emergence and subsequent spread of ABR traits to the environment. This study thus aimed to investigate the prevalence of ABR in Bahir Dar city MSWDS, with the view of generating scientific information that can trigger immediate preventive actions by the city administration and other concerned bodies. The results of the emergence and potential spread of ABR to humans from environmental sources can inform policymakers to implement intervention strategies.
Methods
Study area description
This study was conducted in Bahir Dar City (northwest Ethiopia), which is some 565 km away from Addis Ababa (the capital of Ethiopia). The study site, the main solid waste disposal site in Bahir Dar city, is located between 11° 32′ 23″ to 11° 32′ 37″ N latitude and 37° 23′ 12″ to 37° 23′ 24″ E longitude (Figure 1). It is situated at about 7 km outskirts of the city at an elevation of 1790 m above sea level and covers an area of about 22 hectares. The area has a warm temperature with a mean annual temperature of 13.5 to 27.7°C and a mean annual rainfall of about 1500 mm of which 54% of the fall is in July and August when the rainfall can reach 250 to 300 mm per month. 36
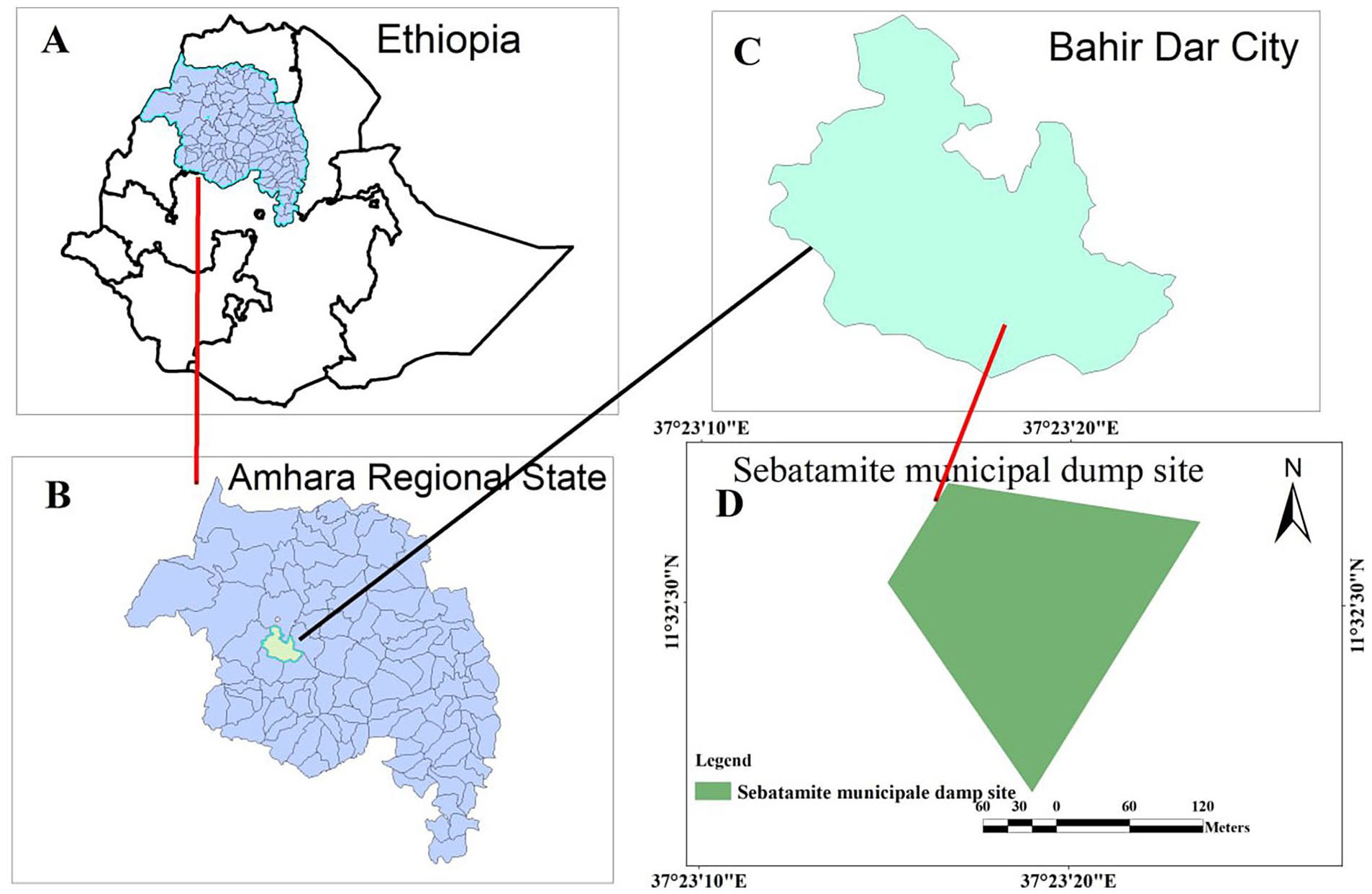
Map of the study area, Bahir Dar City Municipal Solid Waste Dumpsite.
A study by Tassie 37 demonstrated that, daily, over 98 tons of solid waste are being generated in Bahir Dar city. 37 The author added that the wastes proportionally constitute residential (54%), commercial (24.2%), institutional (17%), and street sweeping (3.56%) Generally, there is an unrestricted dumping process in Bahir Dar City and due to that pharmaceutical drugs can be observed scattered at the dumping.
Soil sample collection
As shown in Figure 1 above, the solid waste dump site has roughly a triangular shape (a triangle cut at its tip). To get representative samples, 3 sampling sites were systematically selected along a symmetrical line of the height of the site within Bahir Dar city MSWDS. After removing all surface debris, the site was dug into 4 to 6 cm and soil samples (50 g from each site) were collected with a sterile spatula and placed into sterilized zip-lock polythene bags. The collected soil samples were transported to Bahir Dar University, Microbiology Laboratory in an icebox, and the culturing was begun immediately. A total of 9 soil samples were collected from a 3-round sampling period every 2 months for 6 months (in November 2020, February and May 2021).
Isolation and identification of bacteria
Isolation and purification of bacteria from soil samples of the dumpsite were conducted using standard microbiological methods. 38 A serial dilution technique was used to isolate the bacteria from the 9 soil samples. About 5 g of soil sample from each site was pooled to obtain 1 homogeneous sample. From the mix, 1 g of homogenized soil sample was mixed with 9 ml of sterile normal saline solution (0.85% NaCl) and vortexed for 1 minute to make a homogenous suspension. From the suspension, tenfold serial dilution from the 10−5 to 10−9 was made in 20 ml test tubes each containing 9 ml diluent and 0.1 ml of each dilution was spread on nutrient agar (HiMedia, India). For each dilution, triplicate plates were used and incubated at 28°C for 24 to 48 hours in an inverted position. Morphologically distinct bacterial colonies were collected and purified on the same medium by the streak plate method. The pure isolates were maintained in nutrient broth (HiMedia, India) at 4°C for identification and antibiotic susceptibility test.
The isolates were first categorized based on Gram staining (and cell shape) and their colony characteristics (color, shape, size, margin, opacity, elevation, surface, texture), and further identified using several biochemical tests as described in Bergey’s Manual of Determinative Bacteriology.39,40 The biochemical tests used include catalase, endospore, IMRiV (Indole, Methyl red, Vogas-Proskaur, Citrate), starch Hydrolysis, urease, H2S, and gas production, fermentation of glucose, lactose, sucrose or mannitol.
Various selective and/or differential media were also used to distinguish among Gram-negative bacterial isolates.39,40 Isolates plated on Eosin–Methylene Blue Agar plate and incubated for 24 hours at 37°C, and showed golden greenish shiny color were taken as evidence for the presence of
Antimicrobial susceptibility test on bacterial isolates
After identifying the bacterial isolates, the standard Kirby-Bauer et al’s disk diffusion method was carried out to determine their antimicrobial resistance profiles following standard procedures.41,42 Bacterial inoculum was prepared by suspending 4 to 5 morphologically identical colonies from each isolate in 5 ml nutrient broth (HiMedia, India) and incubated for 4 hours at 37°C. The Clinical and Laboratory Standards Institute (CLSI) recommends using bacteria with McFarland 0.5 turbidity to prepare bacterial suspensions to a specified turbidity for antimicrobial testing. The bacterial suspension was compared with 0.5 McFarland turbidity standards, as it presents the number of bacteria within a given range to standardize the testing. A McFarland standard of 0.5 should have an OD600 between about 0.08 and 0.1, which is comparable to a bacterial suspension of 1.5 × 108 CFU/ml. After adjusting the turbidity, the surface of the prepared Mueller Hinton Agar (MHA) medium (containing (g/L): casein acid hydrolyzate (17.5), beef extracts powder (2), starch (1.5), and agar (17)) (Accumix, India) was evenly inoculated 3 times using a sterile cotton swab while rotating the plate with the culture. The plates were left for 15 to 20 minutes at room temperature to let it dry.
The standard commercially available antibiotic disks used in this study were gentamicin (GN, 10 μg), streptomycin (ST, 30 μg), tetracycline (TE, 30 μg), ciprofloxacin (CIP, 5 μg), nalidixic acid (NAA, 30 μg), sulfonamide (SA, 250 μg), chloramphenicol (C, 30 μg), erythromycin (E, 15 μg), vancomycin (V, 30 μg), and amoxicillin (AMX, 25 μg) (Becton, Dickinson, and Company, Sparks, Maryland, USA). The disks were aseptically laid on the surface of the inoculated agar plates with proper spacing using sterile forceps and incubated at 37°C for 18 to 24 hours. The diameter of the inhibition zone around the disks was measured to the nearest millimeter.
The breakpoints of enterobacteriales are recommended and used for other members in this family such as
Data analysis and interpretation
Descriptive statistics, such as means and standard deviation are used to present the data. The antimicrobial susceptibility profiles of the bacterial isolates are reported as susceptible or sensitive (S), intermediary resistant (I), or resistant (R) according to the annually published microbiological breakpoints by the CLSI. 43 The prevalence of multidrug resistance (MDR) isolates is also determined. MDR is when antimicrobial resistance is shown by a species of microorganism to at least 1 antimicrobial drug in 3 or more antimicrobial categories. 44
Results
Isolation and identification of bacteria
In this study, 71 distinct bacterial colonies were isolated and identified based on colony morphology, Gram test and a series of other biochemical tests (Additional File 4). Ten isolates related to
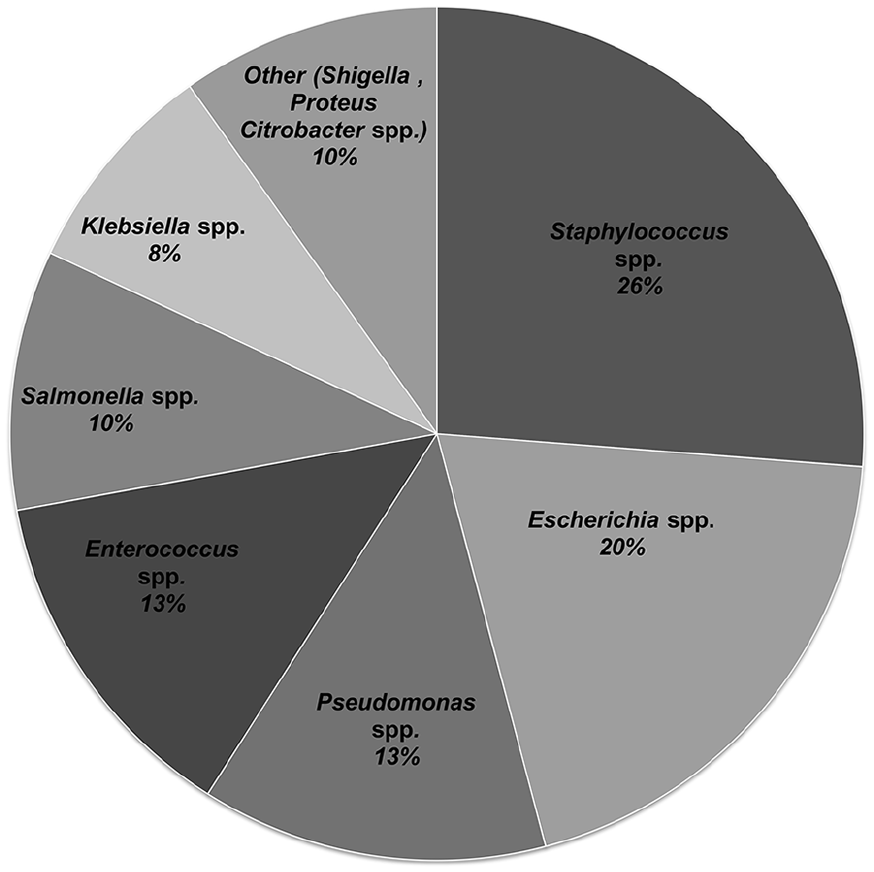
The relative proportion of bacteria isolated from Bahir Dar city MSWDS, Ethiopia.
Antimicrobial susceptibility profiles of the isolates
As shown in Table 1 and Additional File 2, all the tested isolates were resistant to amoxicillin followed by nalidixic acid (75.5%), vancomycin (75%), tetracycline (55.35%), streptomycin (54.5%), and sulfonamide (50%) in that order. On the other hand, large proportions of the isolates were sensitive to ciprofloxacin (83.1%), chloramphenicol (73.7%), and gentamicin (63.5%) (Table 1).
Antimicrobial susceptibility profiles of bacterial isolates recovered from Bahir Dar city SWDS, Ethiopia.
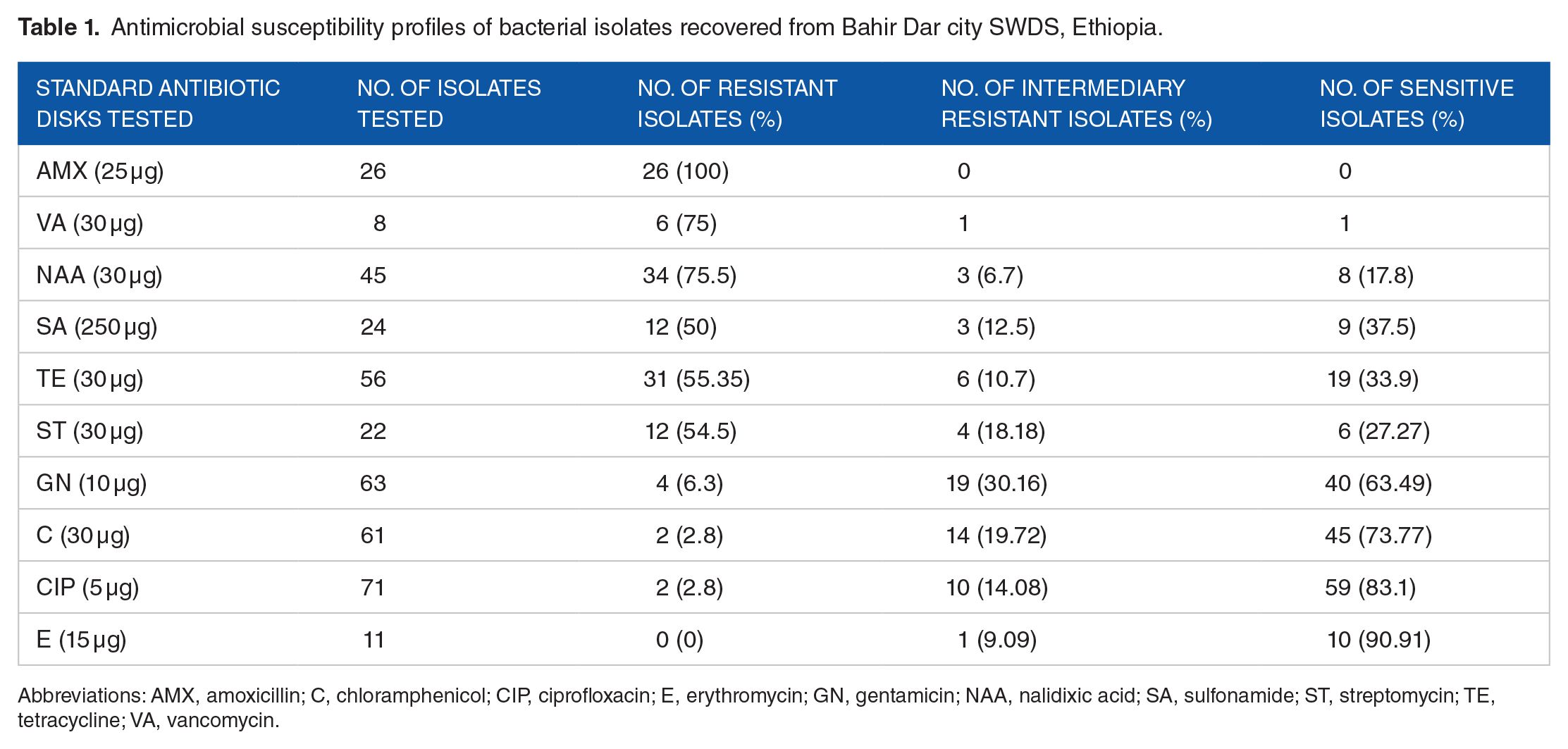
Abbreviations: AMX, amoxicillin; C, chloramphenicol; CIP, ciprofloxacin; E, erythromycin; GN, gentamicin; NAA, nalidixic acid; SA, sulfonamide; ST, streptomycin; TE, tetracycline; VA, vancomycin.
In this study, the overall antibiotic resistance rate was 95.08%. All isolates were resistant to at least 1 antibiotic, except 2 isolates related to
The proportion of multidrug resistant bacterial isolates recovered from Bahir Dar city MSWDS, Ethiopia.
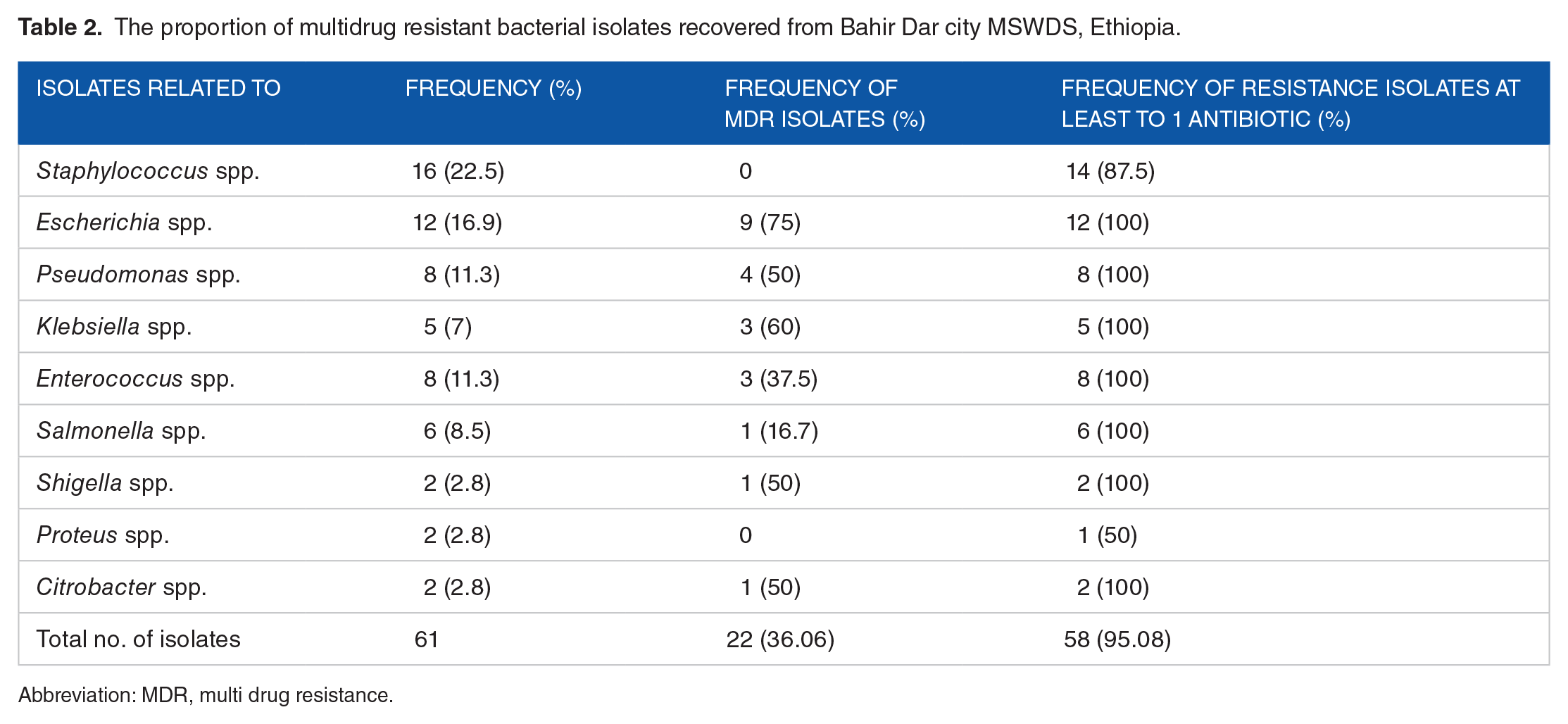
Abbreviation: MDR, multi drug resistance.
Isolates related to
Discussion
The development of antibiotic resistance (ABR) has been among the top concerns of the global healthcare system.2,45 Particularly, this problem is severe in low-income settings as the result of poor waste management systems, inadequate hygienic and sanitation practices and low healthcare system.22,46 A comprehensive study by Mwaikono et al 47 revealed that municipal waste dumpsites harbor large diversity and complex communities of bacteria that have triggered further research on the public health implications of wastes in such man-made environments. As a result, researchers have started to focus on the ecology of ABR in non-clinical and natural environments, particularly on the factors aggravating the emergence of ABR bacteria. With this view, this study was initiated to contribute scientific information on the prevalence of ABR bacteria isolated from a solid waste dumpsite in Bahir Dar City.
In the present study, bacteria related to
Several other bacteria, which were not detected in this study, including
Even though conclusive evidence can be obtained from strain level and serotype identification, the bacteria isolated in the present study are more likely pathogenic, which may be attributed to the disposal of complex solid wastes in the municipal waste dumpsite that originated from multitudes of sources in Bahir Dar City.
The prevalence of ABR has been studied more in clinical settings compared to natural or man-made environments. High prevalent rates of antibiotic-resistant bacteria have been reported in clinical samples, including
In the present study, 10 antibiotics from 8 antibiotic categories (Additional File 3), were tested on the isolated bacteria. The choice for the antibiotics and the corresponding test bacteria species was done based on recommended use as explained in CLSI. A high degree of ABR rate was recorded from Bahir Dar city MSWDS. The overall ABR rate recorded in the current study is the highest (95.08%) compared to that of previous reports from non-clinical samples. Almost all of the isolated bacteria in the current study were resistant to at least 1 antibiotic drug showing that ABR is distributed among the microbial communities in the area. Such an elevated ABR rate may also be due to intrinsic resistance. However, it is recently speculated that intrinsic resistance may serve as the origin of acquired resistance.56,57 The likely reasons for such a high level of antibiotic and multidrug-resistant isolates may thus be due to the presence of bacteria that could be carrying resistance determinants and traces of leftover antibiotics and expired drugs that are improperly discarded with wastes.
Most isolates were resistant to amoxicillin (100%), nalidixic acid (75.5%), and vancomycin (75%), and sizable proportions of the isolates were also resistant to sulfonamide, tetracycline and streptomycin. Some previous studies have also reported a high prevalence of ABR from non-clinical environments. For example, Oviasogie and Agbonlahor
48
reported similar observations that most of the bacterial isolates were intermediary resistant or resistant to tetracycline, nalidixic acid and sulfonamide. Contrary to the results of the current study, a lower resistance rate to amoxicillin (33%-69%) was reported by previous studies conducted in Ethiopia.
6
Similarly, most of
On the other hand, most isolates were susceptible to ciprofloxacin, chloramphenicol and gentamicin. It is to be noted that gentamicin is effective against a wide range of bacterial infections, mostly Gram-negative bacteria including
In this study, a substantial prevalence of multidrug resistance (MDR) bacterial isolates was recorded from soil samples of Bahir Dar City MSWDS. The overall MDR rate was 36.06%, where most isolates demonstrated multidrug resistance features. A comprehensive review by Muhie
61
on antibiotic use and resistance patterns in Ethiopia found a 59.7% pooled prevalence of MDR. However, almost all of the studies included in this review were clinical or hospital-based. Thus, our study documented the highest prevalence of MDR from non-clinical samples in Ethiopia. High prevalence rates of MDR were recorded among isolates related to
It should be noted that the prevalence of ABR bacteria observed in the current study refers only to those bacteria that are cultivable. Better insights into the prevalence of ABR in the study area could be obtained through the metagenomics approach, which incorporates uncultivable bacteria that could also have ABR traits.
Limitations of the Study
We understand that identification of the isolates to species level would be very important. Moreover, soil samples from the nearby environment where the soil is not affected by the dumped wastes should have been included to verify whether the source of the isolated bacteria is municipal solid waste or not.
Conclusions
In this study, almost all isolated bacteria from the Bahir Dar City municipal solid waste dumpsite were antibiotic-resistant (ABR), creating a high public health concern. Especially, isolates related to
Supplemental Material
sj-docx-1-ehi-10.1177_11786302241260508 – Supplemental material for High Prevalence of Antibiotic Resistance Bacteria Isolated From Bahir Dar City Municipal Solid Waste Dumpsite, North West Ethiopia
Supplemental material, sj-docx-1-ehi-10.1177_11786302241260508 for High Prevalence of Antibiotic Resistance Bacteria Isolated From Bahir Dar City Municipal Solid Waste Dumpsite, North West Ethiopia by Baye Sitotaw, Fikremariam Ayalew, Abayeneh Girma, Kindu Geta, Beselam Tadesse and Alemayehu Godana Birhanu in Environmental Health Insights
Supplemental Material
sj-docx-2-ehi-10.1177_11786302241260508 – Supplemental material for High Prevalence of Antibiotic Resistance Bacteria Isolated From Bahir Dar City Municipal Solid Waste Dumpsite, North West Ethiopia
Supplemental material, sj-docx-2-ehi-10.1177_11786302241260508 for High Prevalence of Antibiotic Resistance Bacteria Isolated From Bahir Dar City Municipal Solid Waste Dumpsite, North West Ethiopia by Baye Sitotaw, Fikremariam Ayalew, Abayeneh Girma, Kindu Geta, Beselam Tadesse and Alemayehu Godana Birhanu in Environmental Health Insights
Supplemental Material
sj-docx-3-ehi-10.1177_11786302241260508 – Supplemental material for High Prevalence of Antibiotic Resistance Bacteria Isolated From Bahir Dar City Municipal Solid Waste Dumpsite, North West Ethiopia
Supplemental material, sj-docx-3-ehi-10.1177_11786302241260508 for High Prevalence of Antibiotic Resistance Bacteria Isolated From Bahir Dar City Municipal Solid Waste Dumpsite, North West Ethiopia by Baye Sitotaw, Fikremariam Ayalew, Abayeneh Girma, Kindu Geta, Beselam Tadesse and Alemayehu Godana Birhanu in Environmental Health Insights
Supplemental Material
sj-xlsx-4-ehi-10.1177_11786302241260508 – Supplemental material for High Prevalence of Antibiotic Resistance Bacteria Isolated From Bahir Dar City Municipal Solid Waste Dumpsite, North West Ethiopia
Supplemental material, sj-xlsx-4-ehi-10.1177_11786302241260508 for High Prevalence of Antibiotic Resistance Bacteria Isolated From Bahir Dar City Municipal Solid Waste Dumpsite, North West Ethiopia by Baye Sitotaw, Fikremariam Ayalew, Abayeneh Girma, Kindu Geta, Beselam Tadesse and Alemayehu Godana Birhanu in Environmental Health Insights
Footnotes
Acknowledgements
Bahir Dar University is highly acknowledged for hosting the study. We also acknowledge Amhara Public Health Institute for providing the test bacterial strains.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by Bahir Dar University.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
BS, FA, AG, KG, and AGB conceived the study and worked on the experiments and formal analysis. BS drafted the manuscript. AGB and BT participated in the validation, writing and editing of the manuscript. All authors read and approved the final manuscript.
Ethics Approval and Consent to Participate
The conducted research is not related to either human or animal use.
Consent for Publication
Not applicable.
Availability of Data and Materials
All data generated or analyzed during this study are included in this published article and the additional files.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
