Abstract
Background:
Biomarkers to predict the recurrence risk are required to optimize perioperative treatment. Adjuvant chemotherapy for patients with human epidermal growth factor 2-positive (HER2-positive) early breast cancer is decided by pathological responses of neoadjuvant chemotherapy (NAC). However, whether pathological responses are appropriate biomarkers is unclear. Currently, there are several studies using minimal residual disease (MRD) as a predictor of prognosis in solid tumors. However, there is no standard method for detecting MRD.
Objectives:
This study aimed at prospectively evaluating the relationship between MRD detection and recurrence in Asian patients with HER2-positive early breast cancer.
Design:
Prospective, observational, single-group, and exploratory. This study will include 60 patients from 2 institutions in Japan and the Philippines. The invasive disease-free survival (IDFS) rates of the MRD-positive and MRD-negative groups are compared in patients with HER2-positive early breast cancer who undergo surgery after receiving NAC.
Methods and analysis:
Circulating tumor DNA (ctDNA) levels of patients will be evaluated 6 times: before NAC, after NAC, after surgery, and annually after surgery for 3 years. We will analyze the genetic profile of blood and tissue samples using the Todai OncoPanel (TOP) and the methylation level of DNA. The primary endpoint is IDFS. Secondary endpoints include overall survival (OS) and disease-free survival (DFS). Patient enrollment began in June 2022, and new participants are still being recruited.
Ethics:
This study has been approved by the National Cancer Center Hospital Certified Review Board in March 2022 and has been approved by the Research Ethics Board of the participating center.
Discussion:
Our findings will contribute to determining whether MRD detection using TOP is useful for predicting the recurrence of HER2-positive early breast cancer. If this is proven, MRD detected by TOP could be used in the future as a biomarker to assist in the de-/escalation of treatment strategies in the next interventional trial, thereby avoiding overtreatment in patients at low risk, and in the addition of intensive treatment modalities for those in patients at high risk.
Introduction
Globally, the incidence of breast cancer in 2020 was estimated at 2.3 million cases, with half of these deaths occurring in Asia. 1 The number of breast cancer survivors is increasing because of the rise in the incidence of breast cancer and improvements in treatment strategies for breast cancer.2,3 However, the mortality rates remain high in Asian countries. The 5-year survival rate of patients with breast cancer in Asia is 65% to 89%. 4 Perioperative treatment strategies are important to reduce mortality in breast cancer survivors but biomarkers to determine appropriate adjuvant therapy have not been explored.
Human epidermal growth factor 2-positive (HER2-positive) breast cancer accounts for 15% to 20% of breast cancers and is common in Asia. 5 Neoadjuvant chemotherapy (NAC) with anti-HER2 drugs has dramatically improved pathological complete response (pCR) rate, and prognosis and survivors are increasing. 6 Pathological evaluation of treatment effect is often linked to prognosis, and more intensive treatments, such as adjuvant trastuzumab emtansine (T-DM1), are recommended for patients with HER2-positive breast cancer who cannot achieve pCR after NAC. 7 However, the recurrence rate of invasive disease in patients with HER2-positive breast cancer who achieved pCR after dual HER2 blockade could still be as high as 10%, with higher rates among those with node-positive disease. 8 This is partly because of recurrence, which can be attributed to micrometastatic disease at the initial diagnosis that is not detected by preliminary diagnostics.9,10 Neoadjuvant chemotherapy aims to eradicate these micrometastases, translating to increased pCR and a substantial decrease in relapse risk. 11 Although pCR is often linked to favorable outcomes and low recurrence rates, better diagnostic tools for residual disease are needed. There are some genetic assays for predicting the effect of chemotherapy and prognosis. Oncotype DX is a multigene assay for predicting chemotherapy treatment effects and prognosis in patients with hormonal receptor-positive and HER2-negative breast cancer without NAC.12,13 HER2DX is a test to estimate the risk of recurrence and pCR likelihood in patients with HER2-positive early breast cancer. This test incorporates tumor size, nodal staging, and 4 gene expression signatures tracking immune infiltration, tumor cell proliferation, luminal differentiation, and the expression of HER2 amplicon into a single score. 14 However, these tests do not reflect the status of residual tumor cells after treatment completion.
In recent years, detecting circulating tumor DNA (ctDNA) in patients with early cancer has been used for tracking minimal residual disease (MRD).15,16 Both ctDNA analysis after completion of all treatments and the surveillance ctDNA analysis are associated with recurrence in breast cancer.17 -19 However, there are no standard methods of ctDNA detection, and methods differ depending on the study. 15 Circulating tumor DNA detection in patients with early-stage cancer is challenging due to the low tumor burden.15,16 Therefore, a study on ctDNA detection methods with sufficient sensitivity and specificity for clinical use is needed.
In this study, we plan to detect ctDNA using the Todai OncoPanel (TOP) and analysis of DNA methylation level. We will prospectively evaluate the association between ctDNA detected by this method and prognosis in patients with HER2-positive breast cancer who received NAC in Japan and the Philippines. If ctDNA can be detected correctly, ctDNA-positive patients will have a higher recurrence rate, and ctDNA-negative patients should show a lower recurrence rate.
Methods/design
Aim
This study aims to evaluate the relationship between MRD detection and recurrence in patients with HER2-positive early-stage breast cancer. In addition, we aim to establish a new MRD detection assay for HER2-positive breast cancer.
Study setting
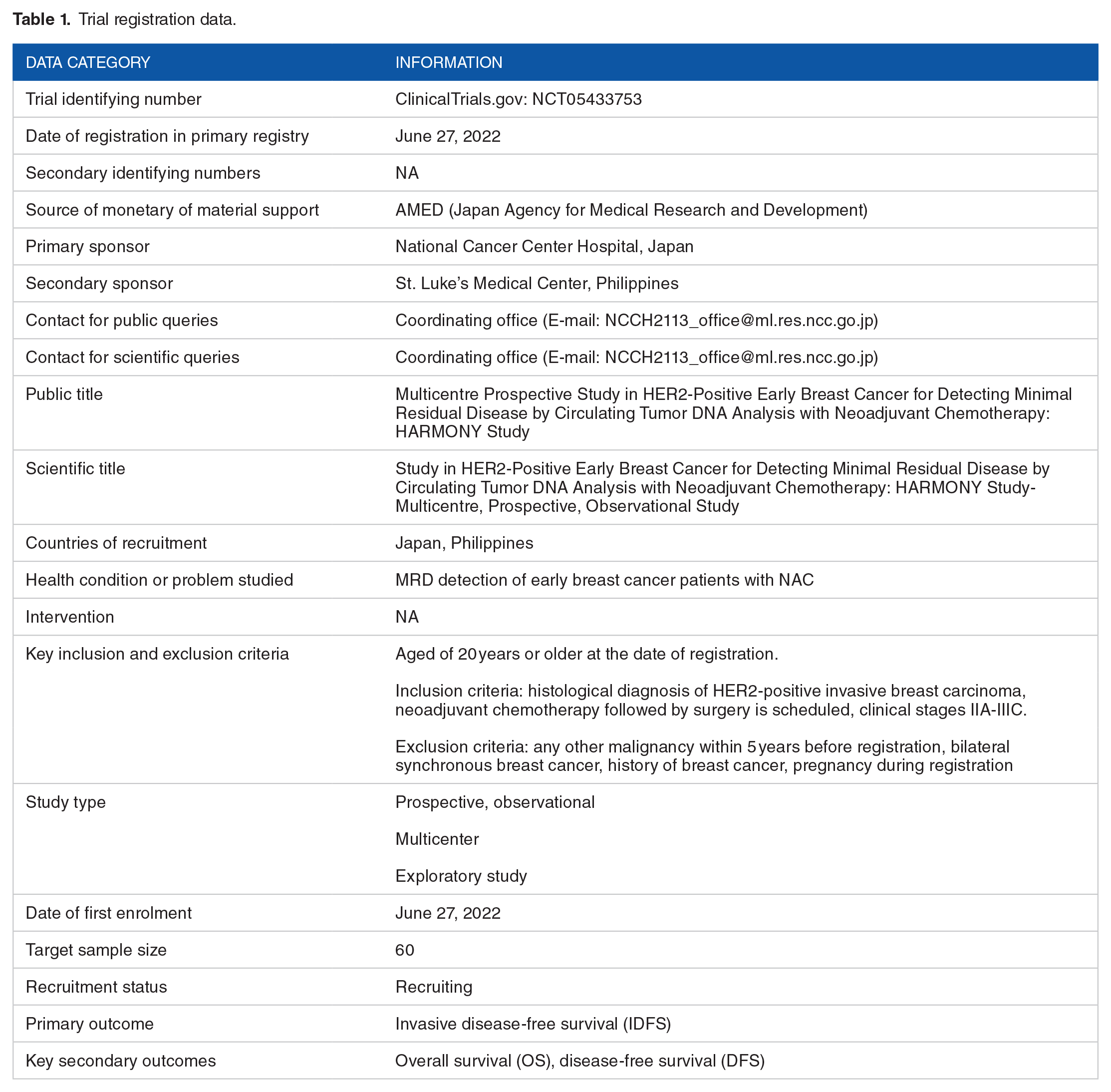
This is a prospective, observational, single-group, exploratory, and multicenter study (National Cancer Center Hospital in Japan and St. Luke’s Medical Center in the Philippines). This study includes cases of HER2-positive breast cancer before NAC followed by surgery, so that the site must be a hospital with breast cancer surgery and a medical oncology department. We chose a participating site with sufficient cases and availability of collection blood and tissue samples with sufficient quality to extract nucleic acid. We summarize study registration data in Table 1 and an overview of this study in Figure 1. Regimens of adjuvant chemotherapy for non-pCR cases are determined by each institution. No financial or nonfinancial incentives will be provided in this study.
Trial registration data.

Flow diagram of the HARMONY protocol.
Endpoints
The primary endpoint is invasive disease-free survival (IDFS). Invasive disease-free survival is defined as the duration from the date of surgery to the date of the judgment of one of the following events, whichever occurs first: ipsilateral and contralateral invasive breast cancer recurrence, regional invasive breast cancer recurrence, distant recurrence, and death attributable to any cause.
The secondary endpoints are overall survival (OS) and disease-free survival (DFS). Overall survival is defined as the duration from the date of surgery to the death attributable to any cause. Disease-free survival is defined as the duration from the date of surgery to the date of the judgment of one of the following events, whichever occurs first: IDFS events, ipsilateral or contralateral ductal carcinoma in situ (DCIS), or second primary nonbreast invasive cancer.
Eligibility criteria
Age: 20 years or older at the date of registration.
Histological diagnosis of HER2-positive invasive breast carcinoma.
Scheduled for neoadjuvant chemotherapy followed by surgery.
Clinical stages IIA-IIIC (AJCC/UICC TNM staging system, seventh edition).
Known hormone receptor status.
Provision of signed informed consent.
Exclusion criteria
Any other malignancy within 5 years before registration, except for adequately treated basal cell or squamous cell skin cancer, or carcinoma in situ of the esophagus, stomach, colon, or cervix.
Bilateral synchronous breast cancer.
History of breast cancer.
Pregnancy during registration.
Sample size calculation and statistical methods
The target enrollment number is 60 patients. Because this is an exploratory study, the sample size is determined by considering feasibility. Each site performs NAC on more than 20 new patients with breast cancer each year, so it will be possible to enroll 60 patients more than a 3-year period. Patient enrollment began in June 2022, and the participant recruitment is currently incomplete.
For all eligible cases, the IDFS, median IDFS, and annual IDFS proportions in the MRD-positive and MRD-negative group are estimated using the Kaplan-Meier method. The 95% confidence intervals of the median IDFS and annual IDFS proportions are estimated using the Brookmeyer and Crowley method and the Greenwood formula, respectively. The same analysis is performed on the pooled population.
The IDFS rates of the MRD-positive and MRD-negative groups are compared using an unstratified log-rank test; however, this result should be interpreted as being exploratory, as the sample size is not based on any statistical tests in this exploratory study. For all eligible cases, the hazard ratio (HR) between the MRD-positive and MRD-negative groups and its 95% confidence interval is estimated using an unstratified Cox proportional hazards model.
The group excluding “ineligible patients” determined by principal investigator or Coordinating Investigators from all registered patients is defined as “all eligible patients.” Among all eligible patients who completed NAC and surgery, the group of patients who submitted the available blood sample collected within 28 days after surgery for MRD analysis is defined as “all eligible examination cases.” This is the primary data set, and we will perform statistical analysis for the following groups; all registered patients, all eligible patients, patients with NAC, patients who completed NAC, patients with surgery, patients who completed surgery, and all examination cases.
Factors planned for subgroup analysis are as follows:
Age, years (<50/⩾50)
Race
Clinical T factor (T1/T2/T3/T4)
Clinical N factor (N0/N1/N2/N3)
Menopausal status (pre/post)
Family history (yes/no)
Smoking history (yes/no)
Hormone receptors (positive/negative) (biopsy tissue sample)
Histological grade, low (grades 1 and 2)/high (grade 3) (biopsy tissue sample)
Anti-HER2 drug (used/unused)
Genetic abnormality before NAC
Pathological response (pCR/non-pCR)
Statistical considerations
This study is designed to evaluate the relationship between recurrence and ctDNA positivity. The dropout rate is estimated at 10%, and the recurrence rate within 3 years is estimated at 20%. Owing to the small sample size, we plan to evaluate the results using point estimates. Individuals in the study group will not be matched to a control group.
Patient enrollment
A trained surgeon or medical oncologist will introduce the study to patients after diagnosis using an informed consent form (ICF). Patients who agree to participate in the study will be registered using the Electronic Data Capture (EDC) system.
Procedures for sample collection
Blood samples are collected 6 times: before NAC, after NAC, after surgery, and annually after surgery for 3 years. When recurrence occurs, a blood sample is collected only at that time, and no further blood samples are collected. The definition of recurrence is IDFS events except “death attributable to any cause.” If the recurrence is DCIS, only an EDC report is required, and no blood sampling is needed. Tissue samples from the biopsy and surgery are collected for genetic analysis. The schedule of enrollment, sample collection, and assessments is shown in Figure 2. Plasma separation is performed at each site by a trained research assistant. Plasma, buffy coat, and tissue samples from each site will be collected at the National Cancer Center, and genomic analysis will be performed by the National Cancer Center Research Institute.

Schedule of the study.
MRD detection methods
In this study, MRD is defined as ctDNA/ctRNA detected in the absence of clinical evidence of tumor after surgery. Cell-free DNA (cfDNA)/cell-free RNA (cfRNA) and buffy-coat DNA will be extracted from blood samples. Buffy-coat DNA will be used for filtering genomic DNA for assessing the clonal hematopoiesis of indeterminate potential (CHIP). Circulating tumor DNA will be detected by TOP and methylation analyses. Cell-free DNA with cancer-related genetic abnormalities will be monitored as ctDNA. We will extract cfRNA and apply TOP to identify ctRNA. As this is the first study of TOP for ctDNA detection in the MRD setting, its sensitivity and specificity for MRD detection will be validated herein. The minimum detection sensitivity of SNV/indel tested by the TOP using commercially available standard tissue DNA is approximately 5%. Furthermore, the minimum detection sensitivity for copy number is 4 copies.
Follow-up
The participants will be evaluated at scheduled visits during screening, treatment, and follow-up. The participants will be followed up every 1 year after the initial surgery. Follow-up is continued until death or a maximum of 3 years after surgery. To promote complete follow-up, the coordinating office will send an e-mail at the follow-up time of each enrolled patient. At any time, patients can withdraw their informed consent regarding the use of biological specimens by requesting the site staff.
Data management
In this study, all patient data are collected by EDC system. User identification code and password are provided from the data manager. Participant data are anonymized using registration number issued by the data manager at the registration. Data cleaning is performed regularly to avoid duplicates, missing data, and inconsistencies.
Monitoring
This study does not require a data monitoring committee (DMC) because it is observational, and there is no need for safety monitoring. Moreover, the sample size is small so an interim analysis has not been planned. Site visit monitoring and auditing will not be undertaken unless critical problems are confirmed.
Protocol amendment
Any protocol amendment or change which may impact on the conduct of the study will be reviewed and approved by the Ethics Committee prior to its activation. The coordinating office will notify important protocol modifications to relevant parties.
Consent or assent
Physicians provide the patient enough time to contemplate and that the patient has understanding on the study participation. If the patient agrees to participate in the study, the physician will obtain the patient’s signature on the ICF reviewed and approved by each site. Informed consent form includes the information about secondary use of patient data and biological samples for future study. Future research will only use data/samples from patients who have consented to secondary use.
Confidentiality
All participant information used during the anonymization process will be stored and managed in a password-protected device. All samples, reports, and data will be anonymized by registration number and stored in an area with limited access. All records containing names or other personal data will be stored in locked file cabinets with limited access.
Access to data
The project principal investigator can access the final cleaned data set, and with the approval of the project principal investigator, other study team members can access anonymized data.
Dissemination policy
Primary publications will be published in English language journals. Conference presentations and articles (review articles) for referral of studies that do not include the results of the analysis of study endpoints, as well as the distribution of patient characteristics and the presentation of safety data after completion of enrollment, can be performed with the approval of the principal investigator.
In principle, the authors of the main published articles on the results of the study (the first publication of the results of the primary endpoint) will be the researchers of each institution, and the following will be finalized by the principal investigator depending on their contributions, including the biostatistician responsible for the analysis and the researchers who have contributed to the study.
All co-authors will review the article content prior to publication and only those who agree to the publication. In principle, the leading presenter of the main conference presentation (the first conference presentation of the results of primary endpoint) will be the principal investigator.
All publications relevant to the study should comply with the ethical standards widely accepted for publication and writing, including the Uniform Requirements for Manuscripts Submitted to Biomedical Journals specified by the International Committee of Medical Journal Editors (Unified Requirements for Manuscripts in Biomedical Journals), the Ethical Considerations in the Conduct and Reporting of Second Section Studies of http://www.icmje.org/index.html#authorship.
A publication containing the results of this study has not yet been published or submitted. In addition, no any parts of this study have yet been presented at a conference.
Storage of sample and information
The study information should be stored for 5 years from submission date of the final analysis report or 3 years from study related publication date, whichever comes later, and is recommended the study site to extend the storage period as possible.
Storage period in the data center shall be semipermanent, in consideration of long-term follow-up and/or secondary research use. After the storage period, samples and information related to this study should be destroyed after anonymization at each institution. Correspondence tables or any other data used while in the anonymization process should be stored and managed in a password-protected device.
Discussion
This study aims to establish a novel MRD detection assay for HER2-positive early breast cancer by evaluating the association between MRD and recurrence in HER2-positive early breast cancer. Minimal residual disease detection using ctDNA is a less invasive and repeatable technique. Minimal residual disease landmark analysis after completion of all treatment is useful for identifying low-risk patients for whom overtreatment can be avoided and high-risk patients requiring intensive therapy. Repeated ctDNA testing in long-term surveillance is useful in predicting recurrence and may allow treatment to be initiated before clinical recurrence.15 -18
Adjuvant chemotherapy of HER2-positive early breast cancer patients received NAC is decided by pathological response. 7 However, the recurrence is experienced even in patients with HER2-positive breast cancer who achieved pCR after dual HER2 blockade. 8 HER2DX is a one of the tools to predict the prognosis of HER2-positive breast cancer. 14 However, a clinically feasible assay to assess the status of the remaining tumor after treatment is lacking.
Multiple studies have shown that detected ctDNA is associated with prognosis.15,16 The I-SPY2 trial in all subtypes of breast cancer suggested that ctDNA levels after NAC, but not pCR, were associated with recurrence. 19 Although the NeoALTTO trial of HER2-positive breast cancer could not detect a significant association between ctDNA and event-free survival, it reported that ctDNA detection at baseline was associated with PAM50 subtypes and lower pCR rate. 20 During the follow-up surveillance, ctDNA is identified before clinical evidence of recurrence. Median lead times for ctDNA are reported at 8.9 months for Coombs and 12.4 months for Lipsyc-Sharf. Circulating tumor DNA surveillance tends to improve sensitivity compared with landmark analysis.17,18 However, it is unclear whether treatment changes based on ctDNA detection will improve outcomes. Trials to evaluate whether ctDNA-guided intervention alters clinical outcomes are underway. 21
There are several approaches for ctDNA analysis of MRD, such as digital polymerase chain reaction (PCR), PCR amplicon-based next-generation sequencing (NGS), hybridization capture-based NGS, whole-genome sequencing, and epigenomic approaches (ie, methylation and fragmentomics). However, a standard method has not yet been established because of the lack of the sensitivity and specificity required for clinical application. 15 In this study, we use the TOP and DNA methylation analysis. Todai OncoPanel is a hybridization capture-based NGS that can analyze a wide range of cancer somatic gene mutations using nucleic acids obtained from tumor tissues with a DNA panel and simultaneously perform fusion gene detection analysis with an RNA panel for a total of more than 900 genes. 22 Currently, we have reported the analysis of ctDNA, cfRNA, circulating tumor cells (CTCs) in advanced cancer using the TOP platform.23,24 In this study, we will evaluate the utility of TOP for MRD detection.
From another perspective, there are 2 different approaches: tumor genotype-informed and genotype-naive methods. Tumor genotype-informed methods have high sensitivity and specificity because they can reduce the influence of genetic aberrations of nontumor origin (ie, CHIP). 25 However, patients who achieve pCR with NAC do not have sufficient tumor volumes for genetic analysis. Genotype-naive methods are useful for detecting genetic aberrations acquired during treatment. 26 Therefore, we will evaluate our detection method to determine whether it can be used as a genotype-naive method. However, we will perform tissue genetic analysis to confirm genotype concordance between the tissue and plasma.
In addition to detection methods, biological features also affect the MRD results. The amount of ctDNA correlates with tumor burden. 27 Moreover, the amount of ctDNA released into blood varies depending on the organ involved. Circulating tumor DNA detection rate in patients with brain tumor is less than 10%. 28 HER2-positive breast cancer is reported to have a higher ctDNA level than the luminal type. 11 In addition, cancer-specific gene mutations (ie, EGFR mutations in lung cancer) are good targets for detecting of MRD. Variants of the BRCA1, BRCA2, and PALB2 gene are reported to be associated with a high risk of breast cancer (odds ratio ranging from 5.0 to 10.6), but their frequency is low. 29 HER2-positive breast cancer has a high frequency of ERBB2 amplification and TP53 mutation. 30 We plan to detect ctDNA/ctRNA using a panel including ERBB2 and TP53.
The limitation of this study is small sample size. Although exploratory, this prospective study will provide the basic data for new MRD assay. Studies on MRD assay are important to enable risk-based individualized therapy.
Conclusions
The findings of this study will provide insights into the utility of a new MRD assay and the relationship between landmark/surveillance MRD and recurrence in patients with early HER2-positive breast cancer.
Declarations
Footnotes
Appendix
Acknowledgements
We thank all the participants for their participation in this study. Editage-Cactus Communications provided editorial support in the form of medical editing, assembling tables, creating high-resolution images based on the authors’ detailed directions, collating author comments, copyediting, fact checking, and referencing.
Declarations
Supplemental material
Supplemental material for this article is available online.
