Abstract
Breast cancer is the most prevalent malignant neoplasm in females. Genetic variations in the xenobiotic metabolising cytochrome enzymes; Family 1 Subfamily A Member 1 (CYP1A1) and Family 1 Subfamily B Member 1 (CYP1B1) might play a role in the individual susceptibility to breast cancer and its prognosis. The goal of this study is to estimate the incidence of single nucleotide polymorphisms (SNPs) in CYP1A1 (rs1048943, Ile462VaI, and rs4646903/MSP1) and in CYP1B1 (rs1056836, Leu432Val) genes in patients with breast cancer. This case-control study included 180 female patients with breast cancer and 180 healthy control subjects from Kirkuk/Iraq. Genomic DNA was extracted from venous blood samples and tested for SNPs by the direct DNA sequencing technique. A statistical analysis was done to identify if there is any association between SNPs and the increasing odd of breast cancer and its stage, grade and molecular subtype at diagnosis. The common (reference) genotype of CYP1A1 gene rs1048943 is AA. The AG and GG variant genotypes were significantly more common in the breast cancer patients and conferred an increased odd of breast cancer and its later stages (stages III and IV) and poor differentiation (P < .01) but not with the molecular subtypes. The common genotype of CYP1A1 rs4646903 is TT. The variant genotypes TC and CC are not associated either with increased risk of breast cancer (P > .05) or with its stage, grade or molecular subtypes (P > .05). The GG genotype of CYP1B1 rs1056836 was the common genotype. The CG and CC variant genotypes were not associated with the increased risks of breast cancer (P > .05) or its stage, grade or molecular subtypes (P > .05). In conclusion, variants genotypes of CYP1A1 rs1048943 might play a role in breast cancer pathogenesis and prognosis and can have a place in cancer screening and tailored medicine in the future in the Iraqi population. Future larger scale studies including other genes might help to better understand the role of the SNP in breast risk and its prognosis.
Introduction
Worldwide, breast cancer ranks first among other malignant diseases in women with more than two (2.3) million people suffering annually and about a third of this number die. Unfortunately, the incidence is increasing. 1 It contributed to 6.9% of all deaths in 2020. 1 Geography markedly influences the incidence of breast cancer; it is most common in the Western World at > 80 new cases per 100,000 females and lowest in Central America, Eastern and Middle Africa, and South Central Asia (less than one half of the Western World). 1 Breast cancer ranks first of all cancer in Iraq in both sexes. 2 Its incidence has been increasing sharply. According to the Iraqi cancer registry, it peaked over a decade from 21.75 to 34.6/100,000 female population in 2010 and 2019, respectively, which represent 32.34% and 34% of all female cancer at the same order. 2 Breast cancer is on the top of the list of cancer deaths. 2 It was responsible for 21% and 23% of all cancer deaths in Iraq in 2010 and 2019, respectively. 2
Breast cancer, akin to all other cancers, is a genetic disorder that results from the interplay of different factors that disrupt the fine and delicate genetic make-up of cells that ultimately result in cancer. 3 These elements are hereditary, familial and environmental. Almost three quarters of breast cancers in the United States are attributed to the last two elements. 4 Exposure to xenobiotics is the major environmental player. 5
The leap advances in molecular technologies opened the door wide for personalised medicine with regard to breast cancer risk assessment, diagnosis, prediction and prognosis and ‘one size fits all’ management strategy is no longer valid. 6 For instance, the fact that the earlier breast cancer is detected, the better the patient’s prognosis is, has promoted the researcher to develop risk assessment models or tools, such as Breast and Ovarian Analysis of Disease Incidence and Carrier Estimation Algorithm (BOADICEA). 7 This model depends on traditional risk factors such as imaging and mammography in addition to genetic profiling of BRACA1 and BRACA2 genes. 7 For therapeutic and prognostic purposes, breast cancer was divided, molecularly, into four types: Luminal A (oestrogen receptor [ER] positive, Her2 negative AND Ki-67 low < 14%, OR Ki-67 intermediate 14–19%, progesterone receptor [PR] high > 20%); Luminal B [ER positive, Her2 negative, AND Ki-67 intermediate 14–19%, and PR low/negative OR Ki-67 high > 20%, OR Her2+] and Her2 overexpression [ER negative, PR negative and Her2 positive]; and triple negative/basal-like [ER negative, PR negative and Her2 negative].6,8
A seminal review article shed a light on the role of Cytochromes P450 (CYPs) in breast cancer and discussed their future promising role in its personalised medicine. 9 Cytochromes P450 (CYPs) are a superfamily of enzymes that function as monooxygenases. 10 They play an important role in the oxidation of steroids, fatty acids, xenobiotics and synthesis and clearance of hormones. 10 Of the important cytochromes, are those involved in xenobiotic metabolising genes such as CYP1A1 and CYP1B1.11-13 CYP1A1 and CYP1B1 are involved in breast carcinogenesis by various mechanisms such as metabolic activation of polyaromatic hydrocarbons (PAHs) and hormonal carcinogenesis. 14
Both enzymes biotransform PAHs to carcinogens that cause DNA damage through formation of DNA adducts with subsequent mutations that are pillars in carcinogenesis.15-19 Interestingly, PAH can induce expression of the enzymes which creates a vicious circus of PAHs activation and enzyme expression. 20
Regarding hormonal carcinogenesis, CYP1A1 can perform 2-hydroxylation of the 17β-estradiol (E2) at a C2 position into 2-hydroxyestradiol (2-OH-E2) while CYP1B1 can hydroxylate 17β-estradiol at a C4 position to 4-hydroxyestradiol (4OH E2). 21 In all, 2-hydroxyestradiol and 4-hydroxyestradiol can be further oxidised to form quinones (estradiol-2,3-quinone and etradiol-3,4-quinone, respectively) that react to DNA to cause depurinated nucleotides adducts as intermediate mutation stimulator with subsequent tumour initiation. 22 Quinines and their precursor can be detoxified by phase II xenobiotic metabolising enzymes such as catechol-o-methyl transferase (COMT) and glutathione s-transferases (GSTs).15-18,22
Minor genetic changes at nucleotides level known as single nucleotide polymorphism (SNP) that are found in phase I xenobiotic metabolising genes, such as CYP1A1 and CYP1B1, can alter the metabolism of the xenobiotics and hormones and consequently increase the susceptibility to various cancers including breast.11-13,23,24 In addition, meta-analysis studies showed that there is a significant difference in the risk of breast cancer between different populations who have the same SNP.12,25
Regarding the tailored medicine, the expression of CYP1B1 gene in hormone-dependent breast tumours plays a vital role in the control of the tumour progression, metabolism, treatment and resistance and toxicity to drugs. 26 In a recent articles review, overexpression of CYP1B1 was associated with the resistance to treatment and higher stage and poor differentiation. 9 Expression of CYP1A1 has been found to be high in breast tumour cells with a positive correlation to tumour grade and menopausal status in newly diagnosed patients with adenocarcinoma of the breast. 27 In addition, it has been found that CYP1A1 is overexpressed in breast cancers that are resistant to anti-oestrogen treatment. 28 Several preclinical studies were performed to target the AhR, CYP1A1 and CYP1B1 expression with promising outcome awaiting future clinical translation. 9
Kirkuk governorate is known for its heavy oil industry that is associated with high concentration of heavy metals and PAH in the air and soil.29-32 Many studies have found an association between petroleum industry and breast cancer through different chemicals such as polycyclic aromatic hydrocarbons (PAHs).33-36 Thus, it would be of value to examine if there is a correlation between genetic variation in the form of SNPs in genes coding cytochrome enzymes that are metabolising xenobiotics; CYP1A1(rs1048943, rs4646903) and CYP1B1(rs1056836) in breast cancer patients in Kirkuk governorate/Iraq applying case control study design.
Subjects, Materials and Methods
Subjects
This retrospective case control study encompassed 180 female patients who have been diagnosed with breast cancer by histopathology over the last 5 years who attend Kirkuk Centre for Oncology (KCO) for treatment, follow-up or consultation during the period from 1 June 2020 till 1 October 2020. An equal number of healthy women (180) of matched age from Kirkuk/Iraq were considered as the study controls subjects. The study is in line with the Declaration of Helsinki for ethical principles of medical research and approved for by Kirkuk Medical College Ethics Committee (No: 15/2020). The risk and aims of the study were explained to the participant and their acceptance for the enrolment in the study was obtained via verbal or written consents. We excluded cases with known family history of cancer specially breast and ovarian ones.
We formulated a questionnaire to include all the demographic and clinic-pathological characteristics of the study population including age, menarche, marital status, parity, educational level, history of chronic disease, family history of cancer, histopathological type, stage and grade and molecular subtypes of the breast cancer at diagnosis. We used TNM staging that is adopted by the Union for International Cancer Control (UICC) and the American Joint Commission on Cancer Staging and End Results Reporting (AJCC). 37 Nottingham Modification of Bloom-Richardson Grading System was used for tumour grading. 38
Blood samples, DNA extraction and polymerase chain reaction
We extracted genomic DNA from peripheral venous blood samples collected from anti-cubital veins and placed it in EDTA containing tube. Genomic DNA was purified using the DNeasy Blood & Tissue Kit (250)® (Cat. No. 69506, Qiagen, GmbH) according to the manufacturer’s instructions. The purity and the concentration of the extracted DNA was assessed using NanoVue™ Plus Spectrophotometer (GE Health Care, USA) and used for polymerase chain reactions (PCRs).
PCRs had a total volume of 25 µL consisting of 20 ng of DNA, 10 µL of PCR master mix (Taq PCR Master Mix Kit, Cat. No. 201445, Qiagen Medical Ltd, GmBH), and 10 picomol (1µl) of each primer. The final volume was reached with molecular grade water. Rotor-Gene Q® thermocycler machine (Qiagen Medical Ltd, GmBH) was used according to the cycling programme consisting of one holding of 94°C for 5 minutes (min), followed by 45 cycles of [(94°C for 20 second (s)/(62°C for 30 s)]/ (72°C for 30 s). The same primers were used for PCRs and DNA direct sequencing of SNPs of CYP1A1 (rs1048943 and rs4646903) and CYP1B1 (rs1056836) cytochrome genes. They were synthesised by the Genetic and Molecular Research Unit of Koya University, Kurdistan, Iraq. The sequences of primers and their sources are shown in Table 1.
Details of the tested genes, reference sequence (rs) of the SNP, their location. The sequence of the primers used for the PCR and their sources are also displayed.

PCR, polymerase chain reactions; SNP, single nucleotide polymorphisms.
The specificity of the PCR products was verified by gel electrophoresis using 2% agarose gel at 5 volt/cm2 and 1X TBE buffer for 1 hour using DNA ladder to confirm size and visualised under UV light (Vilber lourmat®, France). All the PCR samples were purified using AccuPrep® PCR purification Kit (BIONEER Corp, Korea) according to the supplier guidelines and were sent for direct sequencing to Macrogen® Company (Seoul, Korea). Chromas 2.6.6 software (Technelysium Pty Ltd, Australia) was used to view and analyse the DNA direct sequencing chromatograms. The Human Genome Variation Society (HGVS) guidelines were applied to describe the Sequence Variants: 2016 Update, throughout this work and to minimise confusion and overlap in nomenclature. 38
Molecular subtyping
We extracted ER, PR and HER2 immunohistochemistry (IHC) data from patients’ medical records. We divided breast cancer according to their expression into simply four molecular subtypes: Luminal A (ER + and or /PR +/HER2-), Luminal B (ER + and or PR +/HER +), Human Epidermal Receptor 2 (HER2) overexpressing (ER-/PR-/HER +) and basal-like or triple negative (ER-/PR-/HER2-) as described before. 41 The authors have, ethically approved, access to the IHC slides of the patients found at Azadi Teaching Hospital-Kirkuk-Iraq.
Statistical analysis
The data were analysed statistically using GraphPad Prism 8® software (San Diego, CA, USA). The chi square was used to calculate the OR (odds ratio) and 95% CIs (confidence intervals) and to evaluate the association between polymorphisms and the risk of breast cancer, its stage, grade and molecular subtypes. The significance of the association was calculated by the Fisher’s exact test. Quantitative (numerical) parameters were analysed by unpaired T test (Student’s T-test). P value < .05 was recognised as statistically significant.
Results
Demographic and clinico-pathological characteristic of the study population
The age range of the breast cancer patients was 25 to 73 years, while it was 30 to 67 years for the control subjects. The difference between the patients and the controls regarding the mean age, parity, menarche age, educational level was statistically not significant (P < 0.05) as depicted in Table 2. Almost two third (65%) of the cases were in the age group 40 to 60 years. Details of the age at diagnosis of breast cancer are shown in Table 2.
The demographic and clinico-pathological characteristics of the breast cancer patients (n = 180) and control subjects (n = 180).

Most of the breast cancer cases had an invasive ductal carcinoma (86%) of mainly stages II and III (~76%) and moderately differentiated (52%), as shown in Table 2. The frequency of other histopathological types and grades of tumour are detailed in Table 2. The expression of ER, PR and HER2 showed that most of the cases were Luminal A (122, 67.8%), followed by Luminal B (22, 12.2%) then comes the triple negative (20, 11.1%), and HER over expressing (16, 8.9%) as shown in Table 2.
Associations of SNP genotype variants with breast cancer
The study population was examined for the genotyping of the SNP in the genes of interest using DNA direct sequencing as the gold standard method for genotyping as described earlier. Examples of SNP direct sequencing are shown in Figure 1 (CYP1A1 rs1048943), Figure 2 (CYP1A1 rs4646903) and Figure 3 (CYP1B1 rs1056836).

Determination of CYP1A1 rs1048943 genotypes by direct sequencing. (A) Gel electrophoresis for CYP1A1 rs1048943 which shows a single specific amplicon of the expected size of 212 bp. (B) Direct sequencing chromatogram showing a homozygous AA (arrow) genotype which codes for isoleucine. (C) The heterozygous GA (arrow) chromatogram genotype that codes for the amino acids; isoleucine/valine. (D) The black arrow points to one signal for the homozygous GG genotype that codes for valine.

Direct sequencing of SNPs in CYP1A1 rs4646903. (A) PCR was done and then run on 2% gel to confirm specificity before sending to direct sequencing, the electrophoresis showed a single band of 342 bp. (B) Heterozygous CT genotype is seen in the direct sequencing chromatogram (black arrow) (C). Homozygous variant (CC) of the SNP is shown in this chromatogram. (D) Single signal of homozygous thymine TT (black arrow).

Direct DNA sequencing result of CYP1B1 rs1056836. (A) Gel electrophoresis of PCR products of 12 samples of CYP1B1 rs1056836 which show a single specific product of the 135 bp. (B) The black arrow points to the CC genotype which codes for leucine. (C) The heterozygous CG genotype (arrow) is shown which has two signals for cytosine and guanine which codes for leucine/valine, respectively (D) The single signal for the homozygous GG genotype that codes for valine is indicated by the arrow.
CYP1A1 rs1048943 SNP genotyping shows that the AA genotype is the common genotype (reference genotype) in the control group (72%) and the breast cancer group (50%). The AG variant genotype is more common in the cancer group (39%) than in the control one (21%) and associated with an increased odd of breast cancer (OR: 2.7, 95% CI [1.6-4.2]). The GG variant of CYP1A1 gene rs1048943 SNP increased the cancer risk by more than two folds (OR: 2.4, 95% CI [1.3-5.3]). Details of the reference genotype and variants frequencies in the study population and their correlations are clarified in numerical details in Table 3.
Genotypes frequencies of CYP1A1 rs1048943, CYP1A1 rs4646903 and CYP1B1 rs1056836 in both the control (n = 180) and patient subjects (n = 180) and their association with breast cancer risk. The number and percentage are within the same study group.
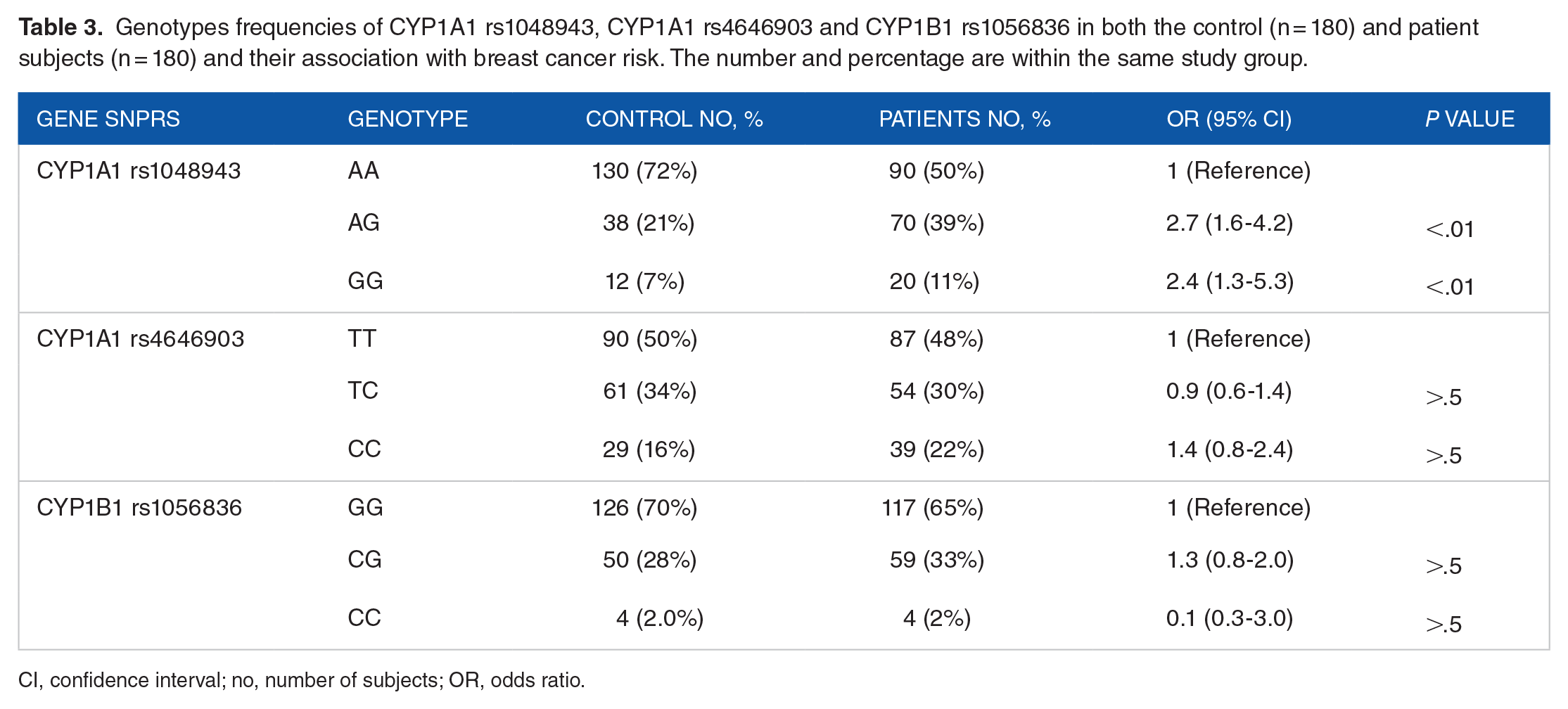
CI, confidence interval; no, number of subjects; OR, odds ratio.
The TT is the common genotype (reference) of SNP rs4646903 of CYP1A1 in both the (50%[ control and ]48%) cancer subjects. The TC variant genotype was the second most prevalent (control, 24% and patients, 30%) that does not confer any significant increases in the risk of cancer (P > 0.05). The CC genotype of the SNP rs4646903 has the lowest prevalence and conferred no significant association with breast cancer (P > 0.05). All details of the SNP number, odds ratio and P values are shown in Table 3.
The frequency of genotypes of CYP1B1 gene (rs1056836) among the 180 patients was CC (65.0%), CG (33%) and GG (2%), while in the control group it was among CC (70%), CG (28%) and GG (2%). As can be seen in Table 3, the CG and GG genotypes do not elevate the odd of breast cancer as detailed in Table 3. There was no association between the above genotypes and age at breast cancer onset. The details of the data are not shown.
Variant genotypes association with breast cancer stage
The stage of a tumour is a pillar in the treatment and prognosis workup. We assigned stages I and II (with all their sub-stages) as early stages while stages III and IV (with all their sub-stages) as late stages. Variant genotypes of CYP1A1 rs1048943; AG and GG were significantly associated (OR: 2.7, 95% CI [1.4-4.9], P < .01) and (OR: 8.0, 95% CI [2.5-23.4], P < .01), respectively, with late stages of breast cancer (III and IV) relative to the AA reference genotype.
The genotypes of CYP1A1 rs4646903 (TC and CC) and CYP1B1 rs1056836 (CG and CC) do not confer any significant association with the stages of the breast cancer (P > .05). Details of the statistical analysis of all the SNPs and their genotypes are shown in Table 4.
Association of the genotype variants of CYP1A1rs1048943, CYP1A1rs4646903 and CYP1B1 rs1056836 with the tumours stage in 180 breast cancer patients. The shown percentages are for the same genotype.
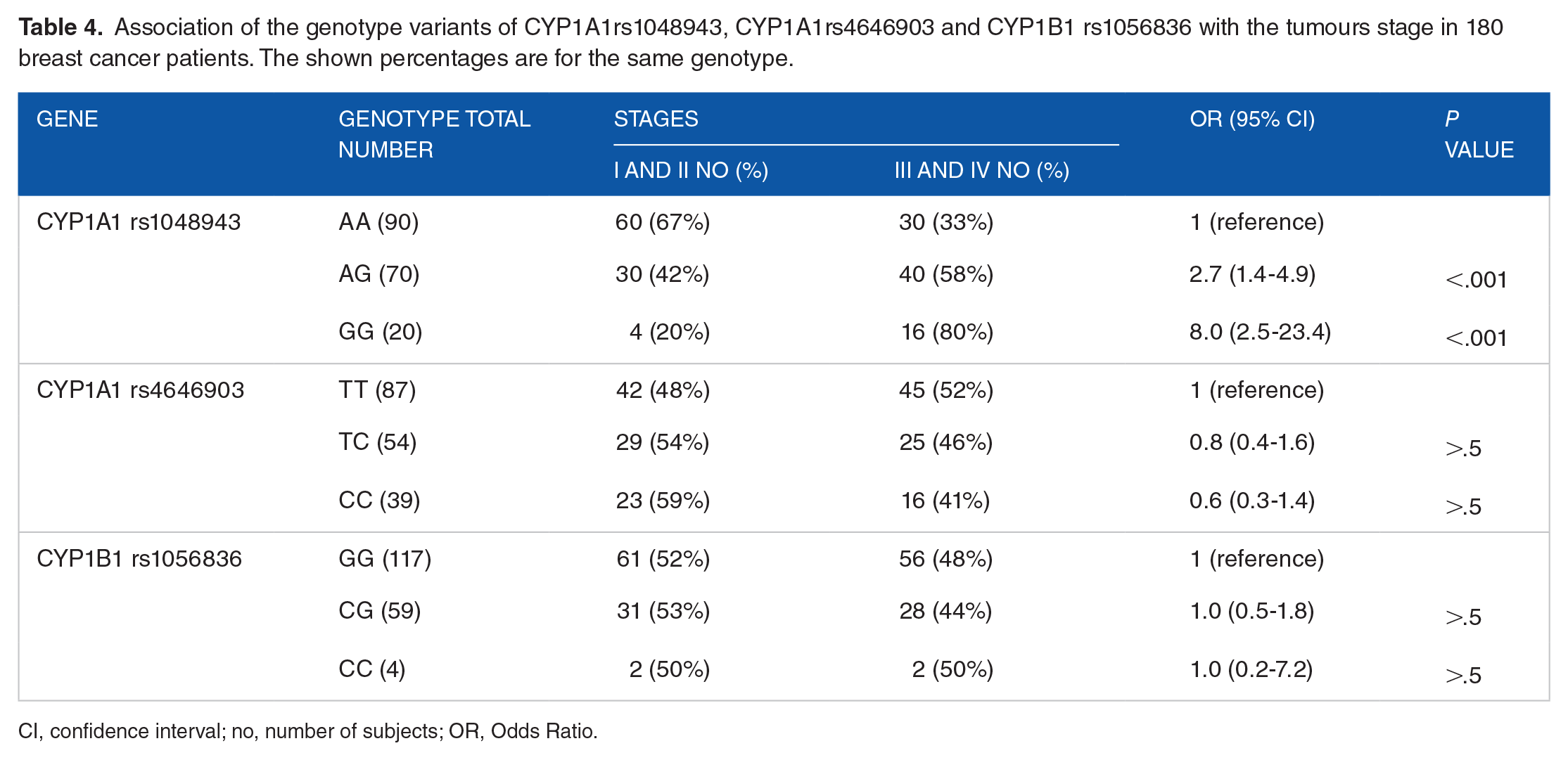
CI, confidence interval; no, number of subjects; OR, Odds Ratio.
The association of genotype variants with breast cancer grade
The degree of cell differentiation (grade) is another accepted prognostic factor. Grades I and II were considered one category, while grade III was considered poorly differentiated. The genotype variants; AG and GG of CYP1A1 rs1048943, had strong associations (OR: 4.0, 95% CI [2.0-7.6], P < .0001) and (OR: 4.5 [1.64-12.5], P < .01) respectively, with a poor differentiation of grade III. All the variant genotypes of SNP of CYP1A1 rs4646903 and CYP1B1 rs1056836 revealed no associations with the grade of the breast tumour. Table 5 contains details of the percentages of the genotypes grade and the degree of association as measured by OR with 95% confidence interval and P value.
Association of genotype variants of CYP1A1rs1048943, CYP1A1rs4646903 and CYP1B1 rs1056836 in) with tumours grade in 180 breast cancer patients.

CI, confidence interval; n, number of subjects, OR, Odds Ratio.
Associations of SNP genotype variants with breast cancer molecular subtypes
The majority of breast cancer cases had the Luminal A expression pattern (122, 67.8%) that is followed by Luminal B (22, 12.2%) and the triple negative (20,11.1%) and HER2 over expressing pattern was the least common (16, 8.9%). No association was found between the cytochrome genotypes; CYP1A1rs1048943, CYP1A1rs4646903 and CYP1Brs1056836 and molecular subtypes. Table 6 contains details of the percentages of the genotypes and molecular subtypes and the significance of association (difference) as measured by P value.
Association of genotype variants of CYP1A1rs1048943, CYP1A1rs4646903 and CYP1B1 rs1056836 in with tumours molecular subtypes in 180 breast cancer patients.

no, number of subjects.
CC genotype was not included in the statistical due to presence of zero value in one of the molecular subtypes.
Examples of IHC patterns are shown in Figure 4. Pictures A, B and C show positive expression of ER, PR and HER2, respectively, in a patient who was assigned as Luminal B. The remaining pictures; D, E and F did not show any expression of ER, PR and HER2 sequentially and been molecularly labelled triple negative.

Expression of ER, PR and HER2 by IHC of two patients. (A) The nuclear expression of ER is visible under low power field (LPF) microscopy. High power field (HPF) view is shown at the right lower corner of the picture. (B) PR is seen under LPF as brown DAB nuclear staining. HPF view is shown in the right lower corner. (C) HER2 has plasma membrane expression, HPF view staining is shown in the right lower corner which is seen as plasma membrane brown staining sparing the nucleus. Pictures A, B and C collectively indicate Luminal B molecular subtype. Pictures D, E and F show no nuclear or plasma membrane staining of ER, PR and HER2 indicating Triple negative molecular subtype of breast cancer.
Discussion
The high incidence of breast cancer has made it a hot spot for research. Breast cancer nowadays is in the heart of personalised medicine, and this work is within this context. In this work, an attempt was made to identify genetic variants that make people more susceptible to breast cancer, especially those in the cytochrome genes that play a role in xenobiotic metabolism namely, CYP1A1 (rs1048943 and rs4646903) and CYP1B1 (rs1056836). 14 CYP1A1 (rs1048943) is a hot spot for genetic polymorphism. The common genotype is homozygous AA which codes isoleucine. Its transition to AG and GG results in coding for isoleucine/valine and valine/valine, respectively, that in this work are associated with increased risks of breast cancer. This finding is justifiable, since these changes are associated with increased expressions and activities of this Phase I enzyme that lead to potential carcinogen activation.42-45 This causes an increased free radical generation that culminates in DNA damage.42-45 In addition, these amino acid changes influence polychlorinated biphenyls metabolism and increase endogenous production of steroid hormones (mainly estrogens).42-45
This association is consistent with other studies conducted in Iraq. It was associated with increased risk of prostate cancer, 46 cervical cancer 47 and lung cancer. 48 Two seminal meta-analysis reviews that examined the association between CYP1A (rs1048943) and breast cancer found conflicting results.23,49 One Japanese study revealed that AG genotype was associated with reduced risks (protective effect). 23 While there was a consistent positive association between the variant and increased occurrence of breast cancer in Indian population, 50 there was no association in the African-American and white women. 51 However, the included studies in the meta-analysis reviews showed similar patterns of distribution of the genotypes of the above SNP similar to our observations.23,49 The controversy in the relation can be attributed to the fact that occurrence of cancer is not a simple cause and effect relation. There is large number of players in the field of carcinogenesis such as the genome as a whole and environmental factors.
It was interesting to find, in the current work, that the above variant genotypes are associated with poor prognosis since higher tumour stages and poor differentiations were more common in the patients harbouring the variants when compared to common genotype. The mechanism by which these variants influence the stage and grade is yet to be identified. However, this relation seems to be racial and cancer type modified since a Polish research found that Ile462Val is not associated with stage or grade of cervical cancer. 52 Similarly, an Iranian breast cancer study showed no associations of the variants with stage but with breast cancer grade. 53 Another study found that these variants are associated with better drug response in breast cancer. 54 Some studies studied CYP1A1 mRNA expression in breast cancer cell lines and its inhibition was associated with impaired proliferation and increase apoptosis. 55 In our work, we did not do gene expression work, which is useful to find any association between breast tissue CYP1A1 expression and breast cancer occurrence and its stage and grade. From clinical point of view, having history of the drugs given to these patients and knowing their response to treatment would add a merit of drug predictive value to this work. Knowing the relation between the gene polymorphism and expression and breast cancer characteristics (stage, grade and drug response) will pave the way, in the future, for CYP1A1 dependent precision medicine regarding risk stratification, diagnosis, drug response prediction and prognosis. It might be even part of predictive or prognostic indices.
The variant genotypes CYP1A1 rs4646903 SNP (MspI) (TC, CC) conferred no elevated odd of cancer when compared to the common genotype (TT). In a multicentre study, in Korea, there was almost a two-fold increase in breast cancer risks in people carrying the variant genotypes. This study contradicts a Chinese study which observed reduced breast cancer risk in those harbouring these genotype. 24 Despite being located in 3 un-translated region of CYP1A1, some studies speculated that such polymorphism might be a reflection of linkage disequilibrium, or the resulting RNA influence the activity of other genes with regard to their expression or stability of the proteins they code. 56
CYP1B1 cytchrome was selected since it codes enzyme that is involved in the metabolism of PAH, androgens and oestrogen substrates and is able to catalyse 4-hydroxyl-estrogens, which is pillar in hormonal carcinogenesis model of breast cancer. 14 SNP rs1056836 G/C transversion results in leucine being replaced by valine at codon 432 which is located at heme-binding domain. The valine amino acid increases the activity of the CY1B1 enzyme for 4-hydroxylation of estradiol. The ideal theoretical response to this extra-hydroxylation is a greater risk of breast cancer through the hormonal carcinogenesis model.
However, this scenario is not consistent throughout all the studies that were conducted in different geographical and ethnic settings that prompted several meta-analysis studies.12,25 It can be positive correlations as seen in Nigerian population where CYP1B1 Val432Leu variant increases the risk of breast cancer. 57 On the other hand, this variant had inverse correlations with breast cancer risk, in other words, it is protective in population who are of mixed African origin. Similar to our observations, population from Asia did not show any correlation between breast cancer risk and the CYP1B1 genotypes.12,25 In addition, in this work, no statistically significant association was identified between CYP1B1 genotypes neither with its stage nor with grade. As explained earlier, the outcome of genetic variation is not a simple direct cause and effect relation.
In this work, all the genotypes of the studied genes showed no significant correlation with ER-, PR- and HER2-dependent molecular subtypes. It is generally accepted that triple negative molecular subtype is associated with late stage and poor differentiation58,59 and, in our work, the minor genotypes of CYP1A1 rs1048943 have similar associations. Accordingly, it is logical to think that some sort of positive correlation would exist between triple negative breast cancer and genotypes of CYP1A1 rs1048943. We could not find other studies that look at the association to enable comparison or extract explanation. Larger study size might give a better insight about the relation.
Conclusion and Recommendations
Knowing that CYP1A1 rs1048943 increases the risk of breast cancer and its stage and aggressiveness might make it a future player in the field breast cancer prediction and prognosis, at least, in Iraq where the study was conducted. The advances in molecular biology have brought innovative technologies that have made genotyping relatively accessible. This gives the clinicians the chance to design appropriate management plan. It would have been better to have a larger sample size sample that enables us to statistically correlate the genotypes with subgroups of age and histopathological types. Breast tissue expression of the enzymes using immunohistochemistry in combination with tissue measurement of the level of some xenobiotics will help us to see the actual effect of the genetic variation on the level of xenobiotics. We did not have drug history of the patients or their survival data. All these missing data are vital to have a better insight into the role of CYP1A1 in breast cancer. SNPs in CYP1A1 rs4646903 and CYP1B1 rs1056836 need studying in a larger sample to prove or disapprove their role in breast cancer.
Footnotes
Acknowledgements
We are indebted to the Kirkuk Oncology Centre for facilitating our study; we greatly thankful to the oncologist and the staff of the centre. We extend our thanks and gratitude to the Department of Histopathology, Azadi Teaching Hospital, Kirkuk for the generous help.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work is fully funded by the researchers with no financial support from any party.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Salih Q Ibrahem: Design of the study, laboratory work, writing up. Hussien Q Ahmed: Collection of the cases and their data and data, analysis of the results and writing up. Khalida M Amin: Collection of the cases, data analysis.
