Abstract
Automated leaf segmentation pipelines must balance accuracy, scalability, and usability to be readily adopted in plant research. We present an end-to-end deep learning pipeline designed for practical use in plant phenotyping, which we developed and evaluated during a real-world plant growth experiment using Atriplex lentiformis. The pipeline integrates a fine-tuned Mask Region-based Convolutional Neural Network (Mask R-CNN) segmentation model trained on 176 plant images and achieves high performance despite the small training data set (Dice coefficient = 0.781). We quantitatively compare the fine-tuned Mask R-CNN model to Meta AI’s Segment Anything Model (SAM) and evaluate natural language prompts using Grounded SAM and the Leaf-Only SAM post-processing pipeline for refining segmentation outputs. Our findings highlight that transfer learning on a specialized data set can still outperform a large foundation model in domain-specific tasks. In addition, we integrate QR codes for automated sample identification and benchmark multiple QR code decoding libraries, evaluating their robustness under real-world imaging conditions like distortion and lighting variation. To ensure accessibility, we deploy the pipeline as a user-friendly Streamlit web application, allowing researchers to analyze images without deep learning expertise. By focusing on practical deployment in addition to model performance, this study provides an open-source, scalable framework for plant science applications and addresses real-world challenges in automation and usability by the end-researcher.
Keywords
Introduction
Plant phenotyping involves studying complex traits such as growth dynamics, shoot and leaf structure, root architecture, visible stress symptoms, and yield. It requires measurement of specific quantitative parameters that define these traits. 1 Plant phenotyping plays a critical role in plant science and agriculture, especially in breeding crops that are both productive and resilient. As climate change intensifies, plant phenotyping is increasingly important in developing crops capable of withstanding extreme conditions, such as heat and drought.2,3 Furthermore, it provides insights into how plants adapt to unfavorable environmental conditions, including poor soil quality, nutrient deficiencies, or pollutants. By understanding these mechanisms, phenotyping aids in developing crops with enhanced resistance to environmental stressors. In addition, it supports the selection of plant species suitable for phytoremediation efforts at restoring contaminated environments.4,5
Image-based plant phenotyping has gained traction as a non-destructive method for measuring and studying plant traits, requiring the development of precise techniques for extracting meaningful data from images. 6 One key challenge is leaf segmentation, which involves the extraction of leaf boundaries from images to determine leaf area, count, and further analyze leaf properties. The Leaf Segmentation Challenge (LSC), first held in 2014, aimed to advance the state of the art in leaf segmentation analysis and facilitate the development of new algorithms. To support this challenge, they released the Computer Vision Problems in Plant Phenotyping (CVPPP) data set of rosette plant images, 7 which was expanded in 2017 to include ground truth segmentation masks and led to significant advancements in deep learning–based segmentation techniques. 8 Today, it remains a widely used data set in leaf segmentation research. 6 ,9 -11 Notably, the CVPPP data set consists of rosette-structured plants, which have a more regular and symmetrical leaf arrangement. Similarly, leaf segmentation studies focusing on top-down images of live plants also tend to feature rosette species, including the KOMATSUNA, Growliflower, and Sugar Beets data sets.11,12 In contrast, fewer tools have been developed or evaluated for non-rosette species, leaving a gap in segmentation approaches for plants with more variable growth patterns.
Among deep learning-based approaches, convolutional neural network (CNN)-based methods, particularly Mask Region-based Convolutional Neural Network (Mask R-CNN), have been most widely adopted for instance segmentation in plant phenotyping.11 -15 The Mask R-CNN is a region-based CNN that unifies object detection and instance segmentation in a 2-stage architecture. In the first stage, a Region Proposal Network (RPN) identifies candidate object regions. In the second stage, these regions are classified, their bounding boxes refined, and a pixel-level segmentation mask is predicted for each instance. This integration of detection and segmentation makes Mask R-CNN well-suited for tasks that require precise delineation of individual objects, such as leaf segmentation. Its widespread adoption has been supported by the availability of pretrained models and modular, user-friendly implementations in open-source libraries like Meta AI’s Detectron2 and Hugging Face’s timm (PyTorch Image Models).16,17 Beyond CNN-based methods, alternative leaf segmentation approaches, such as LeafMask 10 and self-supervised methods, 18 have also been explored.
Unlike CNN-based models that rely on transfer learning—fine-tuning a pretrained network for a specific task—Meta AI’s Segment Anything Model (SAM) is a foundation model designed for general-purpose, promptable segmentation across diverse tasks and domains. 19 The SAM is trained on a massive data set of 1.1 billion masks across 11 million images and can generate masks for a wide range of objects without additional training or manual annotations. Its architecture consists of 3 main parts. First, it uses a Vision Transformer (ViT) to encode the image into a rich embedding, capturing spatial and contextual information across the entire image. Second, it has a prompt encoder that turns user prompts, such as points or bounding boxes, into embeddings that guide what region to segment. Third, a lightweight decoder takes both the image and prompt embeddings and generates a segmentation mask.
Because SAM does not label segmented objects, identifying objects of interest requires either supplying prompts manually, using another detection algorithm to guide segmentation, or applying a post-processing pipeline to classify and label the resulting masks. As a result, many methods combine SAM with other techniques. For example, the Leaf-Only SAM pipeline developed by Zhang et al 13 for tobacco leaf segmentation uses several post-processing steps to isolate leaf masks from SAM’s segmented outputs using properties like color and shape. Grounded SAM combines SAM with Grounding DINO (DETR (DEtection TRansformer) with Improved deNoising anchOr boxes) to support natural language prompts for segmentation, offering an alternative to point or bounding box prompts for guiding SAM’s outputs.20,21
While SAM’s flexibility enables it to work across diverse data sets, CNN-based models typically achieve higher accuracy when applied to data that closely resembles their training set, making them more effective for specialized tasks and data sets. In addition, transfer learning–based approaches like CNNs are often faster at inference than large foundational models like SAM.
While deep learning models like Mask R-CNN and SAM provide powerful segmentation capabilities, their adoption is limited by technical barriers, as they require programming and machine learning expertise. To bridge this gap, user-friendly tools are essential for broader accessibility and widespread adoption. Tools like LeafMachine2 help address this need by providing a Streamlit-based web interface for segmentation and leaf analysis, leveraging a pretrained model trained on a large data set of herbarium images 14 However, its performance on non-herbarium images remains uncertain, as live plant images present different challenges than flat leaves, particularly in experimental setups like ours, which captures Atriplex lentiformis (A lentiformis) during germination and early growth, a stage characterized by high morphological variation. This variability necessitates custom-trained models that can adapt to structural changes across developmental stages while still providing a user-friendly interface for researchers and practical examples for training custom models.
To address these challenges, we developed an open-source, end-to-end deep learning pipeline for leaf segmentation using a data set of A lentiformis, a native species used in phytoremediation projects in the southwestern United States.22 -24 In this study, we focus on 3 major challenges in real-world plant phenotyping workflows: (1) achieving accurate segmentation of non-rosette plant species, (2) evaluating transfer learning vs foundation model performance on a small, specialized data set, and (3) lowering the barrier to adoption through a user-friendly interface. As part of the pipeline, we incorporated QR code–based sample identification to facilitate automated image tracking and organization. Our objectives are to quantitatively compare Mask R-CNN and Segment Anything–based methods, test the robustness of QR code–based image tracking under real-world conditions, and deliver a reproducible, containerized tool accessible to both deep learning experts and domain researchers.
Methods
Pipeline overview
Our system consists of 4 main components:
A fine-tuned Mask R-CNN model implemented using Detectron2, trained on our data set to handle the morphological variability of A lentiformis. The model’s performance is benchmarked against 2 Segment Anything-based methods, offering insights into the effectiveness of transfer learning compared to a large foundational model.
QR code–based data management, streamlining image tracking and organization throughout the segmentation process.
A Docker-based training and evaluation environment, using Jupyter notebooks, allowing researchers with deep learning expertise to fine-tune models or adapt the pipeline to new data sets in a containerized setup that works across different operating systems, reducing dependency conflicts.
A Streamlit-based web application, providing a user-friendly interface for applying the trained model without requiring programming or deep learning expertise.
Data source
This image processing pipeline was developed for a phytoremediation study of plant establishment in metal-contaminated mine tailings. The study focused on the early development of A lentiformis, to assess its tolerance to high metal concentrations. Seeds were planted in clear plastic rhizoboxes (12 × 12 × 1.5 cm) filled with various combinations of soil, compost, and mine tailings, creating a gradient of metal toxicity. Starting 2 weeks after germination, plant shoots were imaged weekly for 6 weeks to monitor growth. The JPEG images were taken with a Canon EOS M6 Mark II camera (Canon, Tokio, Japan) with a MACRO 28 mm lens using an aperture of f/5.6, an exposure time of 1/125 seconds, and an ISO of 200. The camera was mounted on a tripod and positioned above the plant, which was placed on a custom platform angled at 45 degrees. A 2 × 2 cm red paper reference square was included in each image for scale reference and RGB value calibration. To ensure uniform lighting conditions, 2 softboxes were used. In addition, leaf counts were recorded weekly.
The images were routinely analyzed and processed using the following procedures. Image files were renamed manually. Leaf area (cm2) was calculated using the open-source software EasyLeafArea, 25 which utilizes RGB values and the red reference square. When darker or less visible leaves were not detected by the software, manual adjustments were made using Adobe Photoshop. This involved enhancing the green color of the leaves with a selection tool and removing or masking any interfering objects present in the image.
The data set used in this study demonstrates extensive diversity in leaf morphology, pigmentation, and structural arrangement. A representative sampling of this diversity is illustrated in Figure 1. Small sprouting leaves, in the early stages of growth, exhibit distinct morphology and coloration (Figure 1A). More mature leaves, with a typical structural form but strikingly vivid red pigmentation, showcase natural color variations (Figure 1B). Additional variations, such as curled leaves, are depicted in Figure 1C. Furthermore, the complexity of leaf arrangements, including overlapping structures, is highlighted in Figure 1D.

A sampling of images from the data demonstrates the wide variety of leaf types. (A) Small sprouting leaves. (B) Typical structure but very red leaves. (C) Curled leaves. (D) Regular structure with many overlapping leaves.
Data preparation
The processing pipeline automates leaf segmentation and data management, including image renaming and organization. To improve efficiency and accuracy, QR codes were used for sample identification, ensuring each image contains a machine-readable identifier linked to the specimen. Each plant’s unique sample ID was encoded in a QR code, affixed to the rhizobox, and captured in every image.
This setup enables automatic association of images with their sample ID and timestamp, streamlining data organization and reducing human error. By eliminating manual file tracking, QR codes enhance scalability, making the approach well-suited for large-scale experiments. While background objects may introduce complexity, machine-readable identifiers remain essential for accurate and reproducible data processing.
Labeling criteria
An initial data set of 176 plant images (6960 × 4640) was manually annotated using Label Studio, an open-source data labeling software, hosted via a Docker image to facilitate multiuser collaboration. The pipeline required detecting 3 object classes: leaves, QR codes encoding the sample ID, and 2 × 2 cm red reference squares used for metric area calculations. These were labeled as leaf, qr, and red-square, respectively. Some annotated images are shown in Figure 2. QR codes and red reference squares were annotated with tight bounding boxes. Both whole and partial leaf masks were labeled, and small sprouting center leaves were labeled as single leaves. Cotyledons, the first leaves that emerge during seed germination, were classified as leaves. Overall, the data set consisted of 1055 annotations: 797 leaves, 119 red squares, and 139 QR codes. Leaf growth stages were not distinguished in this study.

Examples of annotated images showing 3 object classes: leaf, qr, and red-square. The red-square class corresponds to 2 × 2 cm red reference squares used for metric area calculations. The qr class represents the QR codes encoding sample IDs. The leaf class includes both whole and partial leaves, with small sprouting center leaves considered as a single leaf.
Model training and evaluation
Model training was conducted with an NVIDIA A-100 GPU within a gpu-jupyter Docker container, which enables GPU calculations in Jupyter notebooks in a reproducible environment and includes popular machine learning packages. 26 We utilized Detectron2, a library for object detection and segmentation, and FiftyOne, which facilitates data set management and evaluation and provides interactive visualization of predictions compared to ground truth masks.16,27
We evaluated Mask R-CNN, 28 a widely used baseline, for instance, segmentation, along with 2 methods utilizing the annotation-free SAM. 19 Full model parameters and Jupyter notebooks for all methods are available in the GitHub repository. 29
Mask Region-based Convolutional Neural Network
We fine-tuned a Mask R-CNN model with a ResNet-50-FPN backbone using Detectron2. ResNet-50 is a widely used convolutional network that provides a good balance between model depth and computational efficiency, making it suitable for moderate-scale segmentation tasks. Training was initialized from weights pretrained on the Common Objects in Context (COCO) data set—a large-scale benchmark containing over 200 000 labeled images and 80 object categories. 30 Pretraining on COCO helps the model start with general visual features learned from diverse natural scenes, improving both convergence speed and final performance, particularly when fine-tuning on smaller or domain-specific data sets.
Training was conducted for 5000 iterations with a batch size of 4 and a learning rate of 0.00025. A batch size of 4 was used to balance training stability with GPU memory constraints, as the input images were high-resolution (6960 × 4640). We applied 5-fold cross-validation to maximize use of the limited data and reduce the risk of overfitting. Data augmentation included resizing, horizontal flipping, and brightness jitter. The learning rate remained constant throughout training.
Model performance was evaluated using several metrics:
Dice coefficient: We computed the Dice coefficient, a similarity metric used in computer vision to quantify how well the predicted leaf regions align with the ground truth annotations to assess the model’s ability to correctly segment the leaves while minimizing false positives and false negatives.
mAP (Mean Average Precision): mAP is a standard metric for evaluating object detection models, measuring both detection accuracy and localization precision. It calculates the average precision (AP) at different Intersection over Union (IoU) thresholds and then averages across all object categories. In this study, mAP reflects the model’s ability to correctly detect individual leaves while minimizing false positives and false negatives.
Pearson’s Correlation Coefficient (R): Pearson’s R was calculated to evaluate the linear relationship between the predicted and manually measured leaf area. This measure helps determine how closely the model’s predictions match the leaf area values obtained manually.
Kendall Tau (τ): Kendall tau was used to evaluate the correlation between the predicted and manually counted leaves. This metric is appropriate given that manual leaf counts were taken directly from the plant, while the predictions from top-down images may be impacted by overlap and occlusion, making the rank order of the counts more meaningful than exact values.
Segment Anything
Although Segment Anything does not require annotations like Mask R-CNN does, we utilized the existing annotations to crop each image to the plant’s bounding box before applying the model. This cropping minimized the background, reduced image size and computational load, and decreased the likelihood of false detections from background objects. Segment Anything, when used without a prompt, generates numerous masks, often supersets or subsets of each other, necessitating post-processing to isolate the masks of interest. Cropping the image proved to be the most effective method for eliminating many irrelevant masks.
Leaf-Only Segment Anything Model
We tested 2 approaches using the SAM. The first was a modified version of Zhang et al’s 13 Leaf-Only SAM, which applies SAM without prompts and removes non-leaf and overlapping masks in a post-processing step. Our implementation has 3 modifications:
Modified Overlapping Mask Removal: In the original notebook, we found that the full plant mask was not always removed by the Leaf-Only SAM pipeline in the test data set. In addition, in our data set, some masks containing other masks were not being filtered out, leading to duplicate mask regions. To address this, we relaxed the constraints on overlapping mask removal by checking for subset relationships and removing the larger mask.
Refined HSV Thresholds: We adjusted the HSV color thresholds to better match the leaf color in the A lentiformis data set.
Visualization: We added a visualization step to the notebook, allowing users to display all detected masks or inspect individual masks. This made it easier to evaluate results and identify duplicate or overlapping masks.
Grounded Segment Anything Model
The second approach was Grounded SAM, a pipeline that utilizes Grounding DINO for text prompt object detection, followed by Segment Anything to generate masks from the Grounding DINO results. 20 ,31 -33 We initially tested Grounding DINO’s ability to generalize using the simple text prompt “leaf,” but qualitative evaluation revealed frequent failures in detecting certain leaf types, particularly those with non-standard shapes or colors, textures, or occlusions. To improve recall, we conducted prompt tuning, introducing descriptive keywords targeting specific failure cases and varying the number of keywords to assess their effect on detection performance. However, since Grounding DINO’s performance remained suboptimal despite refinements, we did not pursue exhaustive prompt tuning. Instead, we focused on practical adjustments based on qualitative evaluation to determine the feasibility of text-based leaf detection. A detailed analysis of prompt variations and their impact is provided in the Results section.
After prompt tuning, we applied a post-processing pipeline to detect and remove overlapping masks—similar to the approach used in Leaf-Only SAM. This helped address cases where Grounding DINO mistakenly identified the entire plant as a single leaf. This refinement ensured that only individual leaves were passed to Segment Anything for mask generation.
User-friendly web application
To make this pipeline accessible to all scientists, regardless of their machine learning expertise, we designed a user-friendly web application using Streamlit, a “low-code” platform that expedites the development of web applications in Python. 34 This application serves a dual purpose; it allows the user to run the machine learning pipeline and provides an interface to visualize the predictions. The application performs leaf segmentation using the fine-tuned Mask R-CNN model and renames images by sample ID and date using the decoded QR code and EXIF data. This could be expanded to organize files into folders or manage metadata for the data set.
Our pipeline is built on CyVerse, an open science cyberinfrastructure that supports reproducible and shareable data analysis. 35 The images are stored in the CyVerse Data Store, and the Streamlit app is deployed as a CyVerse VICE (Visual Interactive Computing Environment) app, which interacts directly with the Data Store. This design allows users to upload their data, launch the app, and run the leaf segmentation pipeline without installing packages or setting up their own computing environment. External researchers can create a free CyVerse account and use the published app or create their own pipeline. Free accounts include a limited amount of data storage and compute, with additional resources available through subscription tiers. While CyVerse offers a robust platform for hosting the pipeline, the workflow can also be deployed independently if preferred. Because the app is fully Dockerized, deployment is platform-agnostic.
The end goal of the automated workflow is illustrated in Figure 3. Once a model is trained, researchers can upload their images to CyVerse, launch a web application to select images for processing, and visualize the results. A brief user guide is provided in the following section for those interested in applying the pipeline to their own data.
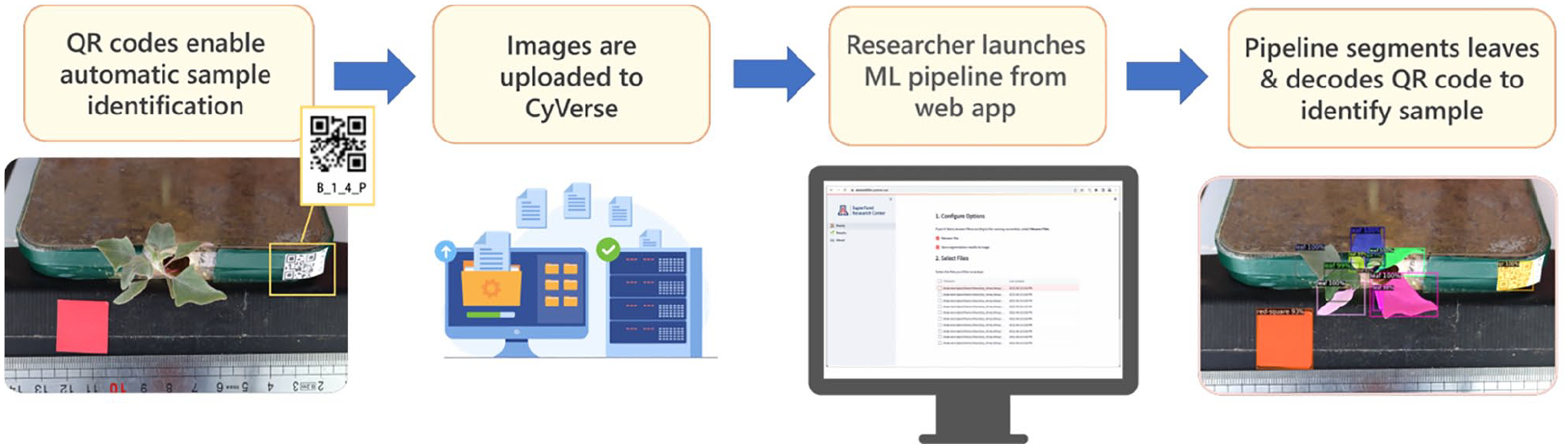
The automated workflow. (1) Unique QR codes are generated for each sample and affixed to the sample, present in each image. (2) Images are uploaded to the CyVerse Data Store. (3) Using a simple web app, the researcher can select images to process through the Machine Learning (ML) pipeline. (4) Leaves are segmented with the ML model and the QR codes are decoded and used to perform data management tasks, like renaming the files and attaching metadata.
Getting started
To begin using the pipeline with your own image data, you will first need polygon mask annotations of leaves and other objects of interest. These can be created using LabelStudio or similar annotation tools and should be exported in COCO JSON format, which is required by the training pipeline.
Our data—including images, annotations, manual leaf measurements, and Mask R-CNN model weights—are available on the CyVerse Data Store, accessible via the Discovery Environment. 36 Individual files are available for download from the Discovery Environment, but to download the entire data set, please see the CyVerse Documentation for the Data Store. 37
To train and evaluate Mask R-CNN, Leaf-Only SAM, and Grounded SAM, see the ua-srp-dmc/leaf-segmentation GitHub repository 29 for container setup instructions and Jupyter notebooks. An NVIDIA GPU is required for model training.
To run the Streamlit web application for leaf segmentation and visualization, there are 2 options:
Run Locally: Use Docker and access it via localhost. Detailed setup instructions are available in the ua-srp-dmac/leaf-detect Github repository. 38
Use the published VICE app on the CyVerse Discovery Environment. 39 This requires a free CyVerse account. Test data are included for demonstration purposes, and instructions for running the app on CyVerse are available in the GitHub repository.
Inference with Detectron2 can run on a CPU for small-scale data sets, but a GPU is recommended for improved efficiency or processing large data sets. If using a CPU, 32 GB of RAM and 16 cores are recommended for high-resolution images. Inference can run at lower specifications for testing purposes or smaller images but with reduced performance.
Results
Mask Region-based Convolutional Neural Network model performance
The Mask R-CNN model was evaluated by comparing its predictions against both the annotated ground truth data and the measurements collected manually using Easy Leaf Area following manual enhancement of the plant images’ greenness. The evaluation metrics included precision, recall, and the Dice coefficient, as well as mAP, which together provide a comprehensive assessment of the model’s predictive performance.
For leaf area, the model achieved an AP of 0.881 and an average recall of 0.705 compared to the ground truth annotations. The Dice coefficient was 0.781 averaged across the 5 folds, reflecting the model’s competence in most cases. The mAP, however, was much lower at 0.430. The low mAP suggests that while the model performs well in detecting and segmenting larger leaves, it struggles with smaller leaves or fails to segment overlapping leaves accurately. Evaluation metrics are shown in Table 1. Figure 4 summarizes the segmentation results and highlights cases where the model encountered difficulties. Figure 5 shows the training loss curve for one of the 5 folds; the others followed similar patterns. The training loss decreased steadily and plateaued toward the end of training, indicating stable convergence.
Evaluation metrics for Mask R-CNN, Leaf-Only SAM, and grounded SAM compared to ground truth annotations (here, the results are reported for Grounding DINO prompt “leaf”).


Mask R-CNN segmentation results. The first row demonstrates accurate segmentation results from Mask R-CNN. The second row provides examples of less successful Mask R-CNN results where the model: (a) failed to detect smaller, thinner leaves and incorrectly labeled a piece of the plastic rim as a leaf and (b) predicted the folded backs of the leaves as separate leaves. In other cases, the backs of folded leaves were not detected at all. (c) Detected overlapping leaves as a single leaf, and the masks did not cleanly segment blurred leaves.

Training loss curve during fine-tuning of the Mask R-CNN model on the Atriplex lentiformis data set. The total loss decreased rapidly during early iterations and plateaued by ~4000 iterations, indicating stable convergence using COCO-pretrained weights and a batch size of 2 over 5000 iterations.
Comparison with manual methods
In Figure 6, the leaf count and area results predicted by the Mask R-CNN model are compared to those collected using the manual method described in section “Data source.” Measurements correlated strongly between the 2 methods, with R = 0.993 for leaf area (Figure 6A) and R = 0.889 for leaf count (Figure 6B).

Mask R-CNN results compared to manual procedure for leaf area (A) and leaf count (B). Leaf area and leaf count were highly correlated (R = 0.993 for leaf area and R = 0.889 for leaf count). However, the model showed reduced consistency in leaf count due to several factors: small leaves at the center of larger plants not being detected, blurred leaves preventing individual segmentation, partially occluded leaves being missed, and curled or folded leaves being mistakenly identified as multiple leaves.
Leaf-Only Segment Anything Model
For leaf area segmentation, Leaf-Only SAM achieved a Dice coefficient of 0.615 and mAP of 0.435, with a precision of 0.692 and a recall of 0.553. A summary of results from all methods is presented in Table 1. The most frequent source of error was in the color-based post-processing step, where leaves were often filtered out due to challenges in defining accurate “green” color thresholds. This difficulty was exacerbated by leaf characteristics such as the presence of white salt bladders, which affected the average color of many leaves. Figure 7 shows the example segmentation results from Leaf-Only SAM compared to Grounded SAM, illustrating common failure cases and qualitative differences between the 2 methods.

Leaf-Only SAM vs Grounded SAM results. From left to right, (A) Leaf-Only SAM identified all leaves, while Grounded SAM combined the masks of 2 overlapping leaves. (B) Grounded SAM successfully detected individual leaves, while Leaf-Only SAM missed several leaves, which got filtered out in post-processing on color. It was challenging to find values that resulted in all leaves being detected while not picking up background objects. (C) Leaf-Only SAM approach does not filter out all background objects, while Grounded SAM does not pick up the center leaves.
Grounded Segment Anything Model
To evaluate how well Grounding DINO could generalize across different leaf morphologies, we initially used the simple text prompt “leaf,” which yielded the highest precision across all methods (0.956). However, its recall was notably low (0.398), missing many actual leaf instances and resulting in incomplete segmentation. This is reflected in a Dice coefficient of 0.562 and mAP of 0.300, indicating that despite high precision, overall detection coverage was limited. Qualitative analysis revealed that Grounding DINO struggled with detecting certain leaf types, particularly:
Cotyledons, curled leaves, sprouting leaves, and angled leaves, as well as occluded or blurred leaves.
Red leaves, including when the curled underside was visible.
Inconsistent detection of visually similar leaves, where some were detected while others were missed. This issue was especially pronounced in highly textured leaves with salt bladders, where detection appeared arbitrary.
To address this, we implemented a prompt tuning step, introducing descriptive keywords targeting failure cases, such as “long narrow leaf” for cotyledons, “leaf with white speckles” to capture surface texture variations, and “red leaf” to capture color variation. We also varied prompt length, testing both simple and complex prompts with more keyword phrases. Performance metrics for each prompt variation are summarized in Table 2.
Prompt tuning results from Grounding DINO.
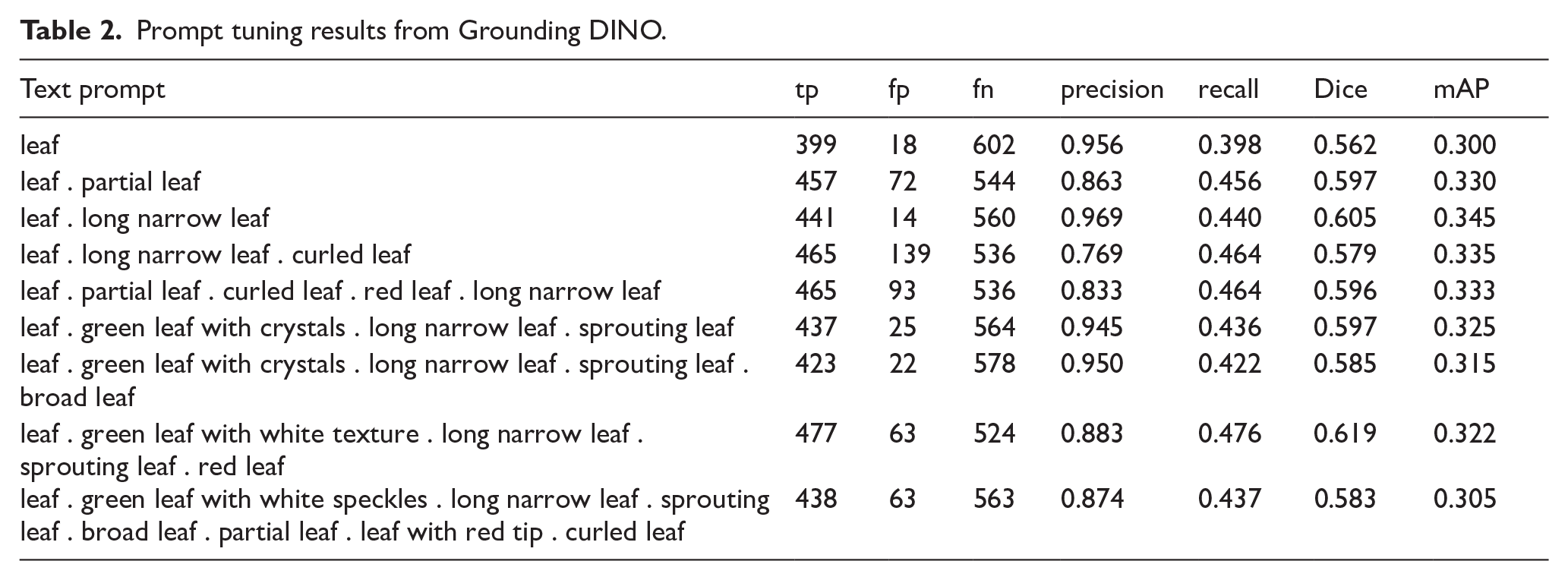
Prompt tuning led to some improvements. The addition of phrase “long narrow leaf” improved segmentation performance across the board, increasing precision to 0.969, recall to 0.440, mAP to 0.345, and Dice to 0.605. The highest Dice coefficient (0.619) was achieved using the prompt included 5 keyword phrases: “leaf . green leaf with white texture . long narrow leaf . sprouting leaf . red leaf,” suggesting that adding targeted descriptors improved segmentation overlap with ground truth. However, this came at the cost of lower precision (0.883), indicating a trade-off between detection completeness and false positives. Increasing prompt complexity beyond a certain point had diminishing returns. The longest prompt tested with 7 keyword phrases, “leaf . green leaf with white speckles . long narrow leaf . sprouting leaf . broad leaf . partial leaf . leaf with red tip . curled leaf,” lowered precision to 0.874 and mAP to 0.305, indicating that excessive detail introduced noise without substantial gains in recall.
While prompt tuning helped improve Grounding DINO’s recall, its overall performance remained inferior to fine-tuned Mask R-CNN and Leaf-Only SAM, suggesting that text-based object detection alone may not be sufficient for high-quality leaf segmentation.
QR code detection
We evaluated 4 open-source QR decoding libraries: OpenCV, 40 pyzbar, 41 pyboof, 42 and QReader, 43 and 1 commercial solution, Dynamsoft Barcode Reader (DBR). 44 All libraries are python-based. These libraries were tested on the 172 images in the data set containing QR codes. Images were cropped to the bounding boxes of the detected QR code with a border of 500 pixels before being processed by each of the libraries, since not all libraries have robust detection capabilities.
The success rate of each library on the 172 images is shown in Figure 8. The commercial solution, DBR, successfully decoded the most QR codes, achieving an 83.72% success rate, followed by QReader, which achieved a 73.54% success rate. Overall, 86.05% of the 172 QR codes were able to be decoded by at least one of the libraries, with the success rate decreasing to 77.33% when considering only the open-source libraries. This suggests that in a small number of cases, the lower-performing libraries were able to decode QR codes that the higher-performing libraries failed to.

Comparison of success rates of Python-based QR decoding libraries. Commercial Dynamsoft Barcode Reader (DBR) successfully decoded 84% of the QR codes, closely followed by open-source QReader at 74%. 86% of the QR codes were able to be decoded by at least one of the libraries.
Streamlit web application
Figure 9 shows the Streamlit web application for using the Mask R-CNN model. Users can upload their data to the CyVerse Data Store and launch the app in the CyVerse Discovery Environment, where they can select files to process and visualize the results. By exposing the machine learning pipeline through a web application, users without machine learning or programming expertise can use the model.
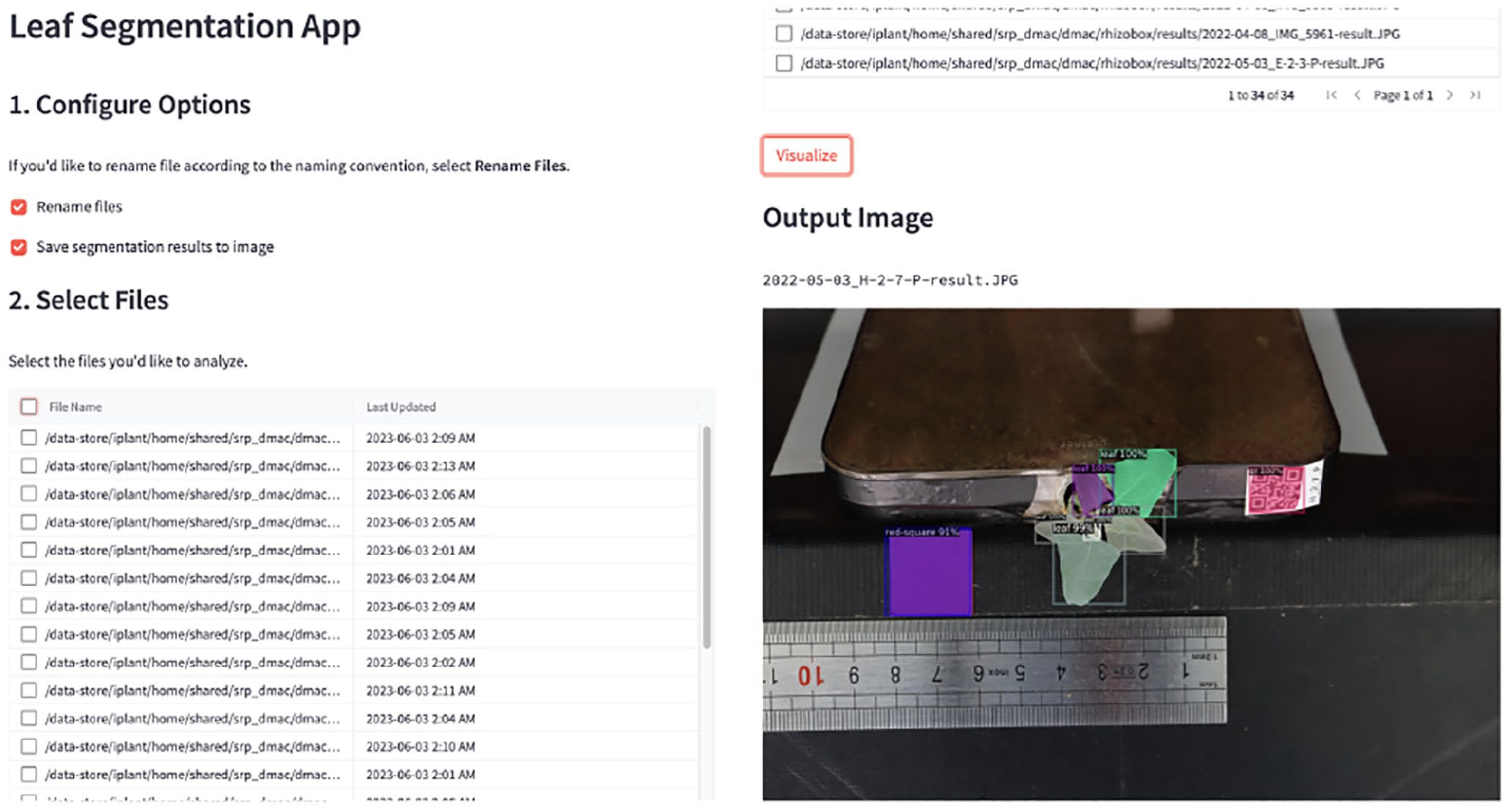
Streamlit web application for running the Mask R-CNN model. The app allows users to select images to process and visualize the segmentation results. Streamlit allows for quick conversion of machine learning workflows into web applications, reducing technical barriers to machine learning.
Discussion
In this study, we developed an accessible deep learning pipeline for leaf segmentation, enabling researchers—including those without machine learning expertise—to automate segmentation for a non-rosette species, A lentiformis. By leveraging QR code–based sample tracking, a user-friendly Streamlit interface, and containerized computing environments with Docker, our approach enhances reproducibility and accessibility in plant phenotyping. Much of the existing leaf segmentation literature focuses on rosette plants with relatively uniform leaf structures. In contrast, A lentiformis presents high morphological diversity, particularly during germination and early growth stages, introducing challenges such as leaf overlap, variation in size, shape, and color, and environmental factors like curling, folding, and soil interference. Our study evaluates Mask R-CNN and Segment Anything–based approaches on this complex plant structure, offering insights into segmentation challenges beyond well-studied rosette species.
Model performance and data set limitations
Despite the limited data set size (176 annotated images) and the substantial variation in leaf characteristics, our fine-tuned Mask R-CNN model achieved an average Dice coefficient of 0.781, demonstrating strong segmentation performance for A lentiformis. However, it only achieved a .043 mAP, highlighting challenges in precise instance detection and detection of small leaves. The Leaf-Only SAM relied heavily on “greenness” to distinguish leaf from non-leaf regions, but defining color thresholds that captured leaves without including background objects proved difficult for A lentiformis, due to the wide variation in color and the presence of white salt bladders on the leaves that affected the average color. Grounded SAM performed well on different types of leaves depending on the prompt. For example, using the prompt “leaf” identified larger leaves well but failed to identify most small leaves and blurred or angled leaves pointing toward the camera. Introducing more detail such as “long narrow leaf” or “sprouting leaf” improved identification of smaller leaves, but also caused leaves previously detected with “leaf” to be missed in some cases.
The small data set size highlights a broader issue in plant phenotyping: the time-intensive nature of manual annotation. Since segmentation accuracy depends heavily on high-quality training data, expanding the data set would likely enhance model performance, particularly in handling variations in leaf size, structure, and shape, including curling and folding; texture and developmental stage differences; and environmental factors such as soil conditions and color variations. Notably, our results suggest that even with a relatively small data set, high-quality annotations combined with transfer learning can produce strong segmentation outcomes. Compared to annotation-free methods like SAM and Leaf-Only SAM, which showed lower Dice and recall scores, our fine-tuned Mask R-CNN model achieved significantly better performance. This highlights that when annotations are accurate and tailored to the task, transfer learning remains a state-of-the-art approach for domain-specific segmentation.
Beyond data set limitations, image quality impacted segmentation accuracy. A primary challenge was blurring caused using a macro lens, which, while essential for capturing intricate leaf details, reduced depth of field. As plants grew taller, maintaining focus on the entire plant became increasingly difficult, leading to unclear leaf boundaries, overlapping leaves, and occlusions. In addition, background artifacts such as the rhizobox, tape, and other experimental elements occasionally introduced false detections, leading to segmentation errors.
One key lesson from this study was the importance of having a highly detailed annotation protocol. While we followed a predefined protocol, it did not sufficiently account for challenges such as blurring and zoom-level inconsistencies. This led to some variability in annotations, which in turn affected model performance. For future work, annotation guidelines should explicitly define when to include or exclude blurred leaves, establish rules for labeling overlapping structures, and ensure consistent representation of small or emerging leaves. Standardizing zoom levels and developing criteria for handling ambiguous cases would further reduce inconsistencies, ultimately leading to higher-quality training data and improved segmentation outcomes.
Integrating automated labeling approaches could also help address annotation challenges by increasing efficiency and consistency. Semi-supervised methods, where models generate initial segmentations refined by human annotators, could significantly reduce annotation time while maintaining accuracy. Future work should focus on increasing data set diversity by incorporating more images across different growth stages and environmental conditions, experimenting with lighting conditions and imaging distance, and exploring machine learning–assisted annotation to improve data set size and scalability for broader applicability.
Reproducibility and computational accessibility
Our pipeline was developed with reproducibility in mind, ensuring that researchers can experiment with common segmentation models in a containerized notebook-based environment. By using a pre-configured, version-controlled computing environment, we eliminate the common challenges associated with dependency management and software compatibility. This approach lowers the barrier to entry for plant scientists who may want to experiment with deep learning techniques but lack extensive programming experience, allowing them to focus on model evaluation and data set adaptation rather than technical setup. Our Jupyter notebooks include training and evaluation of a Mask R-CNN model trained on A lentiformis, and comparisons of 2 SAM-based approaches: Leaf-Only SAM and Grounded SAM, exploring annotation-free segmentation methods. In addition, the notebooks include evaluation and integration with FiftyOne, an interactive tool for visualizing results.
To extend accessibility beyond a code-based environment, we Dockerized a Streamlit web application, providing a graphical interface for segmentation analysis. This allows users without programming expertise to upload images, run segmentation models, and visualize results interactively. Dockerizing the Streamlit app ensures a consistent deployment across different systems, regardless of the host OS, while eliminating manual environment setup.
To optimize computational efficiency, we deployed our Streamlit app on CyVerse, which supports:
On-demand resource allocation, allowing researchers to launch computing resources only when needed, rather than requiring continuous provisioning. This ensures that the Streamlit application utilizes computing resources efficiently only during active analysis, reducing unnecessary computational costs.
Collaborative access, enabling research teams to share data, models, and applications seamlessly within a unified platform, further enhances reproducibility and accessibility.45,46
Overall, our findings demonstrate the practical feasibility of adapting deep learning methods to field-relevant plant species while highlighting common limitations related to data quality, annotation consistency, and deployment. By fully containerizing the pipeline—from training and evaluation to the web-based interface—this work supports reproducible and accessible phenotyping workflows.
Future work
To further improve the study, several avenues could be explored. Expanding the annotated data set to include more examples across a wider range of developmental stages and diverse soil and imaging conditions would help the model generalize more effectively. Experimenting with imaging parameters, such as lighting and distance from the subject, may help mitigate challenges related to blur, shadowing, and occlusion. Exploring semi-automated labeling techniques could accelerate data set expansion while reducing the burden of manual annotation. Further refinement of prompt engineering strategies for foundation models like SAM, especially for small or partially obscured leaves, may improve segmentation accuracy. On the deployment side, further streamlining the training process and making the web interface more flexible—eg, by supporting multiple models and allowing users to adjust parameters—could make the application more broadly usable beyond the current research setting.
Conclusions
In conclusion, this research introduces an end-to-end machine learning system designed to automate leaf segmentation in image-based plant phenotyping. Through the implementation of a Mask R-CNN model and the integration of QR codes for sample tracking, coupled with a user-friendly web interface, this system breaks down technical barriers, enabling researchers without deep machine learning expertise to independently process their data. While the model, tested on A lentiformis images, yields a high Dice coefficient, it does not yet replace the manual segmentation method due to data collection and annotation inconsistencies. Nevertheless, by sharing our methodology and tools, we aim to facilitate the adoption and adaptation of automated pipelines, which free up the scientist for critical thinking and the pursuit of new discoveries.
Footnotes
Acknowledgements
We thank Priyanka Kushwaha and Tomasz Wlodarczyk for their assistance in carrying out the plant science experiments and collecting plant images. ChatGPT was used in the refinement of text. No data were generated or modified using AI.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Author contributions
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by National Institute of Environmental Health Sciences (NIEHS) Superfund Research Program Grant P42 ES004940.
Competing interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
The data sets generated and/or analyzed during the current study are available in the CyVerse Data Store. 36
