Abstract
The detection of polymorphisms in genes that control livestock reproduction could be highly beneficial for identifying and enhancing economic traits. One of these genes is pentraxin 3 (PTX3), which affects the reproduction of sheep. Therefore, this study investigated whether the variability of the PTX3 gene was related to the litter size of Awassi and Hamdani ewes. A total of 200 ewes (130 Awassi and 70 Hamdani) were used for genomic DNA extraction. Polymerase chain reaction was used to amplify the sequence fragments of exons 1, 2, 3, and 4 from the PTX3 gene (Oar_v4.0; Chr 1, NC_056054.1), resulting in products of 254, 312, 302, and 253, respectively. Two genotypes, GG and GT, were identified for 302 bp amplicon. A novel mutation was discovered through sequence analysis in the GT genotype at position g.22645332G>T. The statistical analysis revealed a significant association between single nucleotide polymorphism (SNP g.22645332G>T; Oar_v4.0; Chr 1, NC_056054.1) and litter size. The presence of the SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1) genotype in ewes resulted in a significant difference compared to ewes with GG genotypes. The discrepancy became apparent in several aspects, including litter sizes, twinning rates, lambing rates, litter weight at birth, and days to lambing. There were fewer lambs born to ewes with the GG genotype than to ewes with the GT genotype. The variant SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1) has positive effects on the litter size of Awassi and Hamdani sheep. The SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1 has been associated with an increase in litter size and higher prolificacy in ewes.
Introduction
Sheep reproductive efficiency is directly linked to their fecundity.1,2 Fecundity is determined by ovarian folliculogenesis, which regulates the proliferation and differentiation of granulosa, theca, and germ cells. 3 This process is influenced by the interaction of local hormones and granulosa growth factors, such as pentraxin 3 (PTX3). 4 PTX3 is a glycoprotein produced by granulosa cells surrounding the oocytes, also known as cumulus cells, following the stimulation of preovulatory follicles with luteinizing hormone or human chorionic gonadotropin. 5 This protein, a member of the pentraxin superfamily, plays a critical role in stabilizing and completing the formation of cumulus-oophorus complexes (COCs), which are essential for ovulation. 6 According to Chang et al, 7 PTX3 is expressed during the preovulatory period in cumulus and mural granulosa cells, indicating that the formation of the COC and the extracellular matrix are closely related processes. In addition to its role in determining female fertility, 8 PTX3 is important in transporting the cumulus oophorus-oocyte complex to the oviducts and in determining successful fertilization. 9 A successful pregnancy also relies on the expression and production of PTX3. Research by Zhang et al 10 has shown that cows in early pregnancy experience an increase in PTX3 levels, demonstrating its importance in maintaining corpus luteum function in domestic ruminants during this critical period.
This protein is a product of the PTX3 gene, which consists of three exons located on chromosome 1 in sheep (NCBI Reference Sequence NC_056075.1) and on chromosome 1 in caprine. 11 Multiple research studies have linked PTX3 polymorphism to phenotypic traits in mammals. Onteru et al 12 investigated PTX3 gene polymorphism in pigs to determine their association with meat traits. According to their findings, high levels of fat deposition are associated with the results. According to Ilie et al, 11 the single nucleotide polymorphism (SNP; g.108076746C>T) in PTX3 goat breeds has been associated with the remarkable ability to resist mastitis and parasites in the gastrointestinal system. Essa et al 13 have discovered three significant SNPs (C189T, C364G, and C488A) within the PTX3 gene, which are closely associated with resistance to infection in Holstein and Montbéliarde dairy cows. In terms of reproduction, deleting the PTX3 gene in mice leads to female infertility, which results from abnormal ovulation of cumulus cells and oocytes. 14 According to May et al, 15 variations in the human PTX3 gene (specifically the rs6788044 SNP) have an impact on female fertility and production. The results of these previous studies suggest that the PTX3 gene product plays a critical role in female reproduction. However, no studies have been conducted yet on the association between PTX3 polymorphism and reproductive traits in small ruminants. Reproductive traits in ewes are regulated by genes with both major and minor effects. Major prolificacy genes, such as GDF9, BMP15, BMPR1B, and B4GALNT2, have been reported in numerous indigenous sheep breeds.16,17 In addition, researchers have advocated either the identification of candidate genes controlling prolificacy in indigenous sheep18,19 or the introgression of major prolificacy genes into indigenous low-prolific sheep breeds. 20 As a result, the PTX3 gene is an excellent candidate for genetic and breeding-assisted selection. To the best of our knowledge, there have only been limited studies on the association between PTX3 polymorphism and livestock productive traits. However, there is a lack of research exploring the impact of these genetic variations on Awassi and Hamdani litter size. The prevalence of Awassi and Hamdani sheep is high in most Middle Eastern countries.21,22 The morphological characteristics of each breed distinguish them from one another. Hamdani and Awassi exhibit distinct nucleic acid sequence variations in response to external environmental challenges.1,23 Even though Awassi sheep are known for their adaptability, 24 the Hamdani breed is an exceptional animal that thrives in its natural environment, providing a bountiful abundance of meat, wool, and milk. However, they are known as reproductive seasonality and have lower reproductive rates than Karakuls and Assafs.21,25,26 Based on these findings, the study fills this knowledge gap by providing genotypic information and reporting new associations. The use of genetic markers to assess economic traits could also be possible in the future. It is essential to further explore this subject and uncover the potential influence of PTX3 polymorphism on sheep litter size. Therefore, this study investigated whether genetic variations in the PTX3 gene affect sheep litter size.
Materials and Methods
Sheep population
The research conducted between July 2022 and September 2023 was approved by the animal ethics committee (Agri, No. 01, 7, 22). This approval was based on international guidelines for animal care and use. This study involved 130 Awassi ewes (57 single births and 73 twin births) and 70 Hamdani ewes (42 single births and 28 twin births) that were sexually mature and healthy. The ewes weighed between 40 and 60 kg and were aged between 3 and 4 years. They were selected at random from Babylon and Karbala stations. Both animal breeds differed genetically and geographically. The animals were fed concentrated feed and had access to fresh water at all times. All aspects of lambing were measured at the breeding stations, including the twinning rate, the lambing rate, and the survival rate.
DNA extraction and PCR
Genomic DNA was extracted from 5 ml of sheep blood collected from the jugular vein using rapid salting-out methods. 27 The Primer 3 online program 28 was employed to design PCR primers based on the sequence of the ovine PTX3 (Oar_v4.0; Chr 1, NC_056054.1). To identify the optimal conditions for PCR amplification, we used a Bioneer premix and the Eppendorf thermal gradient apparatus from Germany. The PCR was performed, and thermal gradients were applied to determine the optimal amplification conditions, as detailed in Table 1. The amplification conditions were as follows: 5 minutes of initial denaturation at 94 °C; 30 seconds of denaturation at 94 °C; 45 seconds of annealing; 30 seconds of extension at 72 °C, for a total of 30 cycles; and 5 minutes of final extension at 72 °C followed by storage at 4 °C. 29 Electrophoresis of PCR products on 2% agarose gels was performed and images were captured using a Chemidoc Gel Imager (Bio-Rad, Hercules, California). 30
The oligonucleotide primer sets are designed for the amplification of the ovine PITX3 gene (Oar_v4.0; Chr 1, NC_056054.1).
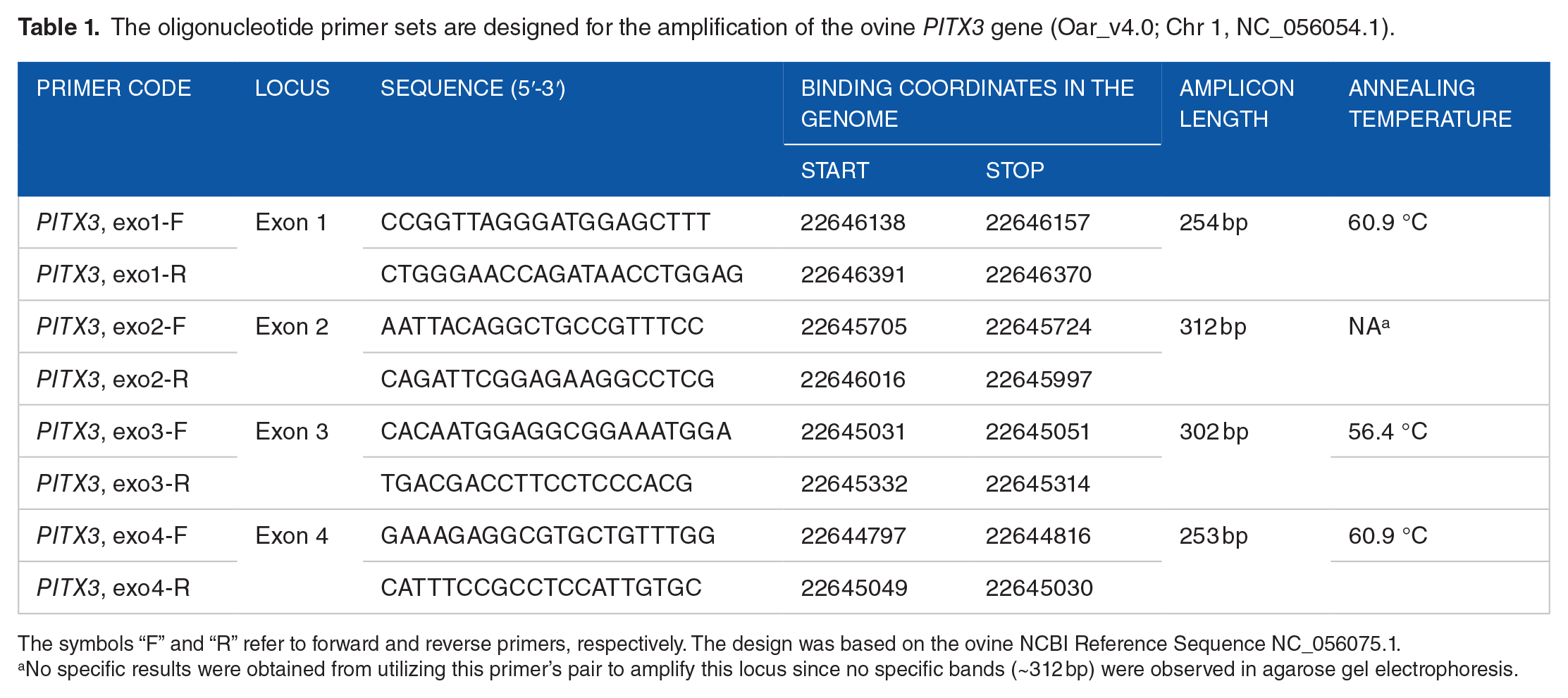
The symbols “F” and “R” refer to forward and reverse primers, respectively. The design was based on the ovine NCBI Reference Sequence NC_056075.1.
No specific results were obtained from utilizing this primer’s pair to amplify this locus since no specific bands (~312 bp) were observed in agarose gel electrophoresis.
Genotyping analysis
The genotype of each PCR product was determined according to Mohammed et al. 31 About 10 μL of PCR product were added to 10 μL of single-strand conformation polymorphism (SSCP) loading buffer dye (Bio-Rad, Hercules, California). To effectively preserve the conformation of single-stranded DNA, each sample was carefully denatured for 7 minutes and then promptly transferred to ice for 10 minutes. 32 Non-denaturing 12% polyacrylamide gels were run at room temperature for 4 h/200 mA/100 V. 33 A DNA sequence variation was identified by electrophoresing SSCP gels, followed by fixing and staining, as described by Byun et al. 34 After detecting the polyacrylamide gel bands using SSCP, each observed an electrophoretic SSCP pattern was sequenced using Sanger sequencing (ovine reference genome v4.0; NC_056054.1). The sequencing results were analyzed using SnapGene Viewer 4.0.4 (http://www.snapgene.com) and BioEdit 7.1 (DNASTAR, Madison, USA) to identify potential SNPs. The Ensemble Genome Browser version 96 was used to assess the novelty of the observed variants. The online prediction software for RNA folding Web Server (http://rna.tbi.univie.ac.at/cgi-bin/rnawebsuite/rnafold.cgi) was used to examine the impact of genotypes on PTX3 gene mRNA secondary structure.
Data analysis
The genotype and allele frequencies of the participants were determined using PopGen32 version 1.31. 35 Our next step was to calculate the Hardy-Weinberg equilibrium (HWE) and then determine the polymorphism information content (PIC) as described by Botstein et al. 36 IBM SPSS version 23.0 (New York) was used to conduct an association analysis of PTX3 genotypes as follows:

where Yijkl = phenotypic value (litter size), μ = mean, Gi = fixed effect of ith genotypes (i = GG, GT), Bj = fixed effect of jth breed (j = 1, 2), Pk = fixed effect of kth parity (k = 1, 2, 3), and eijkl = random residual error. A significant difference was determined by the Tukey-Kramer test at the 0.05 and 0.01 level. An analysis of three reproductive traits was conducted using the chi-square test: lambing rate, survival rate, and litter size. Lambing rate (%) = (Number of ewes lambing / Number of ewes mated) × 100. Litter size = Total number of lambs birth / Number of ewes lambing. Twinning rate (%) = (Number of ewes lambing twins / Number of ewes lambing) × 100. The factors of interaction, lambing season, station, and age were evaluated, and those that were not significant were dismissed.
Results
Genotyping, sequencing of PTX3 genes, and genetic diversity
Four DNA fragments, 254, 312, 302, and 253 bp in length, along with their flanking regions, were amplified from all coding regions of the PTX3 gene (Figure 1A). No distinct PCR bands were observed on agarose gels for the PCR amplicons of 312 bp designed to amplify exon 2. However, PCR products covering the remaining three exons showed greater specificity. PCR-SSCP analysis of exon 3 revealed two distinct genotypic patterns (Figure 1B). Sequencing analysis of the 302 bp amplicons confirmed that the SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1) occurred only in one of the SSCP variants. The identified SSCP variants showed two genotypes: GG and GT, as confirmed by the presence of homozygous G/G patterns and heterozygous G/T patterns resulting from this nucleic acid substitution (Figure 1C). The online prediction software RNA fold Web Server was utilized to predict the secondary structure of the mRNA of the PTX3 gene. The genotypes with the SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1) mutation showed modified secondary structures of PTX3 (Figure 1D).

A schematic diagram of the PTX3 gene-based PCR-SSCP-sequencing strategy in Awassi and Hamdani ewes. (A) PCR-specific primer pairs were designed for the amplification of 254, 312, 302, and 253 bp in exon 1, exon 2, exon 3, and exon 4, respectively. (B) PCR-SSCP genotyping revealed two genotypes in exon 3: homozygous and heterozygous. (C) DNA sequencing electropherograms of the detected genotypes revealed the presence of one SNP, g.22645332G>T, in heterozygous GT genotype within exon 3. (D) Predicting the secondary structure of mRNA in the PTX3 gene for different genotypes. mRNA indicates messenger RNA; PCR, polymerase chain reaction; SNP, single nucleotide polymorphism; SSCP, single-strand conformation polymorphism; MEF, minimum free energy.
Concerning genetic diversity, the frequencies of the PTX3 gene genotypes and alleles of Awassi and Hamdani sheep, as well as their Hardy-Weinberg results, are presented in Table 2. Both the Awassi and Hamdani populations showed a significant deviation from HWE, as evidenced by a chi-square value with a significance level of P ⩽ .05. Observed heterozygosity was higher than the expected heterozygosity in the studied breeds, indicating genetic polymorphism. Moreover, the PIC values obtained were less than 0.5, indicating a moderate level of genetic variability within the ovine PTX3 gene. The classification was determined based on the PIC value, below 0.25 in the low range, between 0.25 and 0.5 in the medium range, and values above 0.5 in the high range.
Genetic diversity of the PITX3 gene (Oar_v4.0; Chr 1, NC_056054.1) in Awassi and Hamdani ewes detected by PCR-SSCP.
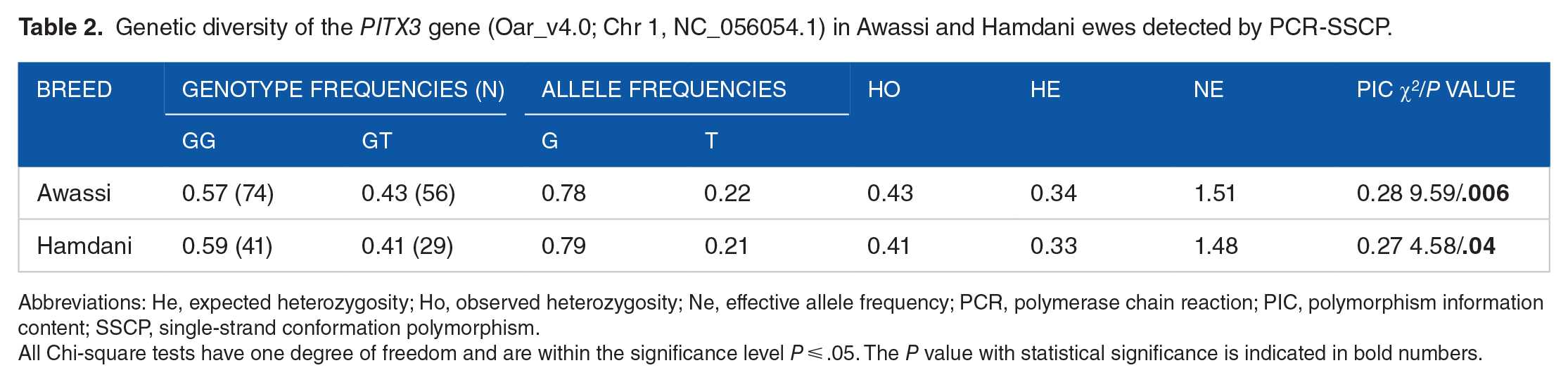

Abbreviations: He, expected heterozygosity; Ho, observed heterozygosity; Ne, effective allele frequency; PCR, polymerase chain reaction; PIC, polymorphism information content; SSCP, single-strand conformation polymorphism.
All Chi-square tests have one degree of freedom and are within the significance level P ⩽ .05. The P value with statistical significance is indicated in bold numbers.
Association analysis
The association analysis of the SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1) revealed no significant difference (P ⩾ .01) in survival rates between individuals with the GG genotype and those with the GT genotype in both breeds. At the same genetic locus, the GT genotype showed a significant association (P ⩽ .01) with higher litter sizes, twinning rates, lambing rates, litter weight at birth, and fewer lambing days compared to those with the GG genotypes. The average number of lambs per ewe with GT genotypes was 1.75 in Awassi ewes and 1.62 in Hamdani ewes, compared to lambs in GG genotypes. As a result, the SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1) was positively associated with these traits (see Tables 3 and 4).
The association between PITX3 (Oar_v4.0; Chr 1, NC_056054.1) genetic polymorphism and reproductive performance in Awassi and Hamdani ewes.

The P value with statistical significance is indicated in bold numbers.
Least squares means ± SE of the litter traits and PITX3 (Oar_v4.0; Chr 1, NC_056054.1) genetic polymorphism in Awassi and Hamdani ewes.
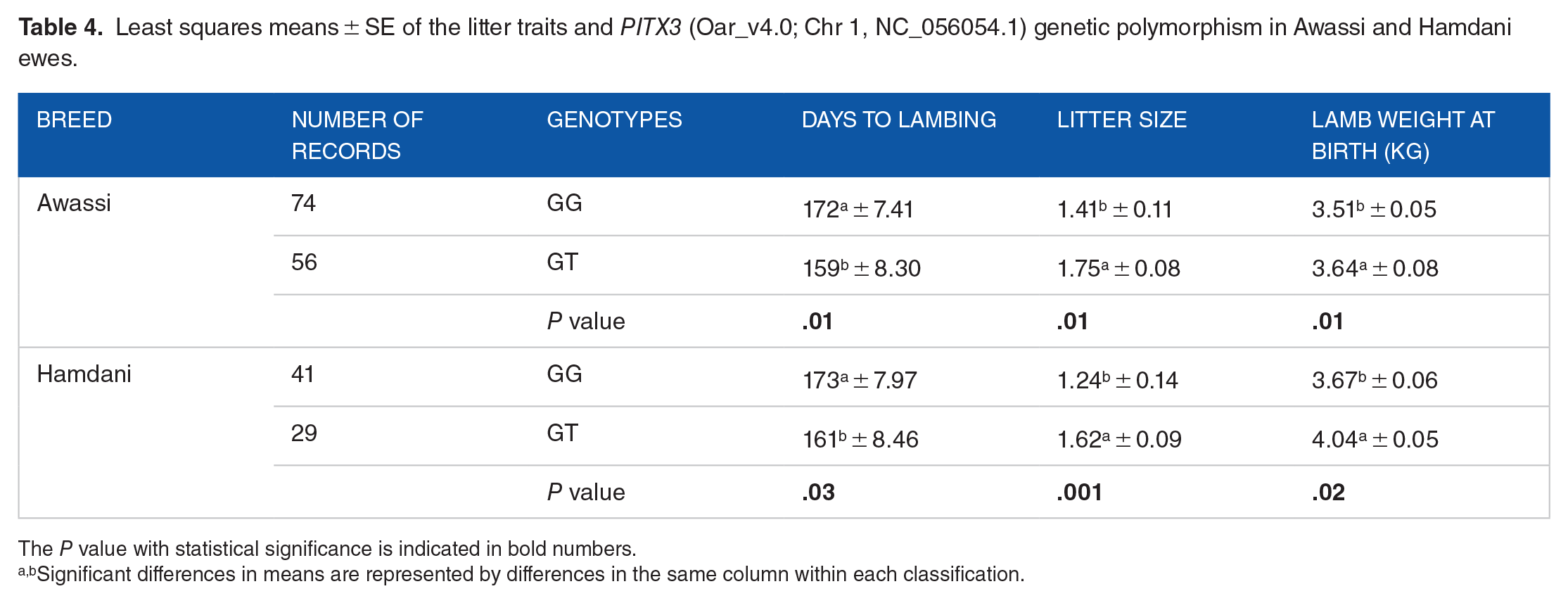

The P value with statistical significance is indicated in bold numbers.
Significant differences in means are represented by differences in the same column within each classification.
Discussion
In this study, we found that the SNP g.22645332G>T; Oar_v4.0; Chr 1, NC_056054.1 contained two genotypes (Figure 1), and the PIC values showed a moderate level of polymorphism, suggesting genetic variation (Table 2). The PTX3 gene has been the subject of numerous studies indicating genetic variability in livestock. In Danish Holstein cows, the SNP rs109126926 on BTA1, specifically an intron variant, is located within the protein-coding gene PTX3. 37 In addition, Ilie et al 11 identified polymorphic SNP (g.108076746C>T) located in the intron of PTX3 in goat breeds. Furthermore, in Holstein and Montbéliarde dairy cows, three SNPs (C189T, C364G, and C488A) have been identified within the exonic region. 13 However, there is no previous literature on PTX3 variation in Awassi and Hamdani sheep.
In the context of SNP bioinformatics, a silent mutation at position 122 Arg in exon 3 of the PTX3 gene had a significant effect on litter size (Figure 1). Although this mutation is silent at the protein level, it has a strong linkage disequilibrium with a nearby causal mutation. 16 The silent mutation studied in this case suggests that although silent mutations can result in proteins with identical amino acid sequences, they may exhibit varying structural and functional characteristics. 38 Synonymous mutations do not alter the encoded message; however, RNA splicing could potentially affect protein production, and modification of regulatory sequences could alter gene expression. 39 Furthermore, this scenario may directly impact the translation of mRNA into PTX3 protein, subsequently affecting the transmission of the hedgehog signal. In a study by Ahmed et al, 40 it has been revealed that a silent mutation can have a significant impact on the regulation of post-translational gene expression, especially when interacting with RNA-binding proteins. This interaction can significantly influence multiple crucial aspects of genome functionality. These factors have a significant impact on processes such as protein folding, the interaction between RNA and functional sites, alternative splicing, and the ability of proteins to recognize RNA-binding sites. 41
Statistically, the GG genotype exhibited lower litter sizes, twinning rates, lambing rates, litter weight at birth, and longer days to lambing compared to the GT genotype. Mutation g.22645332G>T; Oar_v4.0; Chr 1, NC_056054.1 of the PTX3 gene was found to have a positive effect on litter size (Tables 3 and 4). PTX3 is a molecular marker that has been associated with fertility in farm animals. 42 The role of PTX3 in female fertility is crucial, as it transports cumulus oophorus-oocyte complexes into the oviducts and influences the success of fertilization. 43 The occurrence of dizygotic twinning among Gambians is likely attributed to this mechanism. 9 Ovaries and oocytes express a wide variety of transforming growth factor-β (TGF-β) members, each making a distinct contribution to mammalian reproduction. 44 Basavaraja et al 45 observed that various members of the TGF-β superfamily regulate PTX3 in granulosa cells through distinct signaling pathways. The signaling pathway involving SMAD1/5 and SMAD4 regulates the production of PTX3 through ALK2/3-dependent signaling. 10 Controlled ovarian stimulation revealed that PTX3 expression is high in granulosa cells after ovulation, and could serve as a marker of oocyte maturation. 46 The PTX3 gene is also activated by the fecundity gene, which is essential for the expansion of cumulus cells during oocyte maturation and before ovulation, 47 and it plays a critical role in cumulus matrix production. 5 Aside from its critical role in female reproductive processes, the findings of this study suggest that PTX3 is a potential gene associated with ewe prolificacy. This is supported by the association analysis of different genotypes with litter size in Awassi and Hamdani sheep. The low prevalence of the silent SNP could also account for the low prolificacy of the Awassi and Hamdani breeds. As a result, PTX3 could impact the litter size of sheep breeds. A significant improvement in sheep fertility rates could result from this initiative in the future.
Conclusion
A novel SNP, known as SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1), was discovered in the PTX3 gene, as confirmed through the Ensembl Genome Browser. This variation was primarily observed in individuals with heterozygous GT genotypes. Ewes with the GT genotype showed significant correlations with increased litter sizes, enhanced twinning rates, elevated lambing rates, and reduced days to lambing compared to ewes with the GG genotype. The results highlight that ewes with the SNP g.22645332G>T (Oar_v4.0; Chr 1, NC_056054.1) exhibit increased litter size and higher productivity compared to those without it. However, further studies with a larger sample size and more extensive records are necessary. In addition, genome-wide association studies are expected to be useful for identifying genetic markers associated with prolificacy in Awassi and Hamdani sheep.
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Tahreer M Al-Thuwaini: Conceptualization, writing-original draft, supervision, methodology. Faris S Imran: Methodology.
