Abstract
Reutealis trisperma belonging to the family Euphorbiaceae is currently used for biodiesel production, and rapid development in plant-based biofuel production has led to its increasing demand. However, massive utilization of bio-industrial plants has led to conservation issues. Moreover, genetic information on R trisperma is still limited, which is crucial for developmental, physiological, and molecular studies. Studying gene expression is essential to explain plant physiological processes. Nonetheless, this technique requires sensitive and precise measurement of messenger RNA (mRNA). In addition, the presence of internal control genes is important to avoid bias. Therefore, collecting and preserving genetic data for R trisperma is indispensable. In this study, we aimed to evaluate the application of plastid loci, rbcL, and matK, to the DNA barcode of R trisperma for use in conservation programs. In addition, we isolated and cloned the RtActin (RtACT) gene fragment for use in gene expression studies. Sequence information was analyzed in silico by comparison with other Euphorbiaceae plants. For actin fragment isolation, reverse-transcription polymerase chain reaction was used. Molecular cloning of RtActin was performed using the pTA2 plasmid before sequencing. We successfully isolated and cloned 592 and 840 bp of RtrbcL and RtmatK fragment genes, respectively. The RtrbcL barcoding marker, rather than the RtmatK plastidial marker, provided discriminative molecular phylogenetic data for R Trisperma. We also isolated 986 bp of RtACT gene fragments. Our phylogenetic analysis demonstrated that R trisperma is closely related to the Vernicia fordii Actin gene (97% identity). Our results suggest that RtrbcL could be further developed and used as a barcoding marker for R trisperma. Moreover, the RtACT gene could be further investigated for use in gene expression studies of plant.
Introduction
Perennial Reutealis trisperma, also known as Kemiri sunan or Philippine Tung, is a member of the family Euphorbiaceae. It is native to the Philippines; however, became prominent in Southeast Asia eventually. This woody and non-edible plant has been introduced in several Asian and South American regions, including Indonesia, Malaysia, China, India, Cuba, and Dominica. 1 Although it is reported to grow naturally in humid areas, R trisperma is also known to be drought-tolerant. Therefore, it also grows well in critical or unfavorable areas. 2 Together with other tropical plants, 3 R trisperma is known as a medicinal plant. Tigliane diterpenoids isolated from R trisperma have exhibited anti-HIV properties. 4 In addition, the potential of R trisperma as a bioresource for biodiesel production have been demonstrated.5,6 The kernels are extracted for their oil and produce 50 to 52 wt%. 5 Furthermore, the conversion of R trisperma oil to biodiesel through esterification and transesterification has been reported to meet the ASTM standard D6751. 5
Massive utilization of fossil fuels for transportation, industry, and many anthropogenic activities contributes to environmental issues, including air pollution, human health damage, and global climate change.7,8 Therefore, the development of alternative renewable energy sources is promising. This could reduce global carbon emissions and might be sustainable as renewable bioresources. 9 Some non-edible plant-based bioenergy have been studied, including Jatropha curcas, 10 Ceiba pentandra, 11 Calophyllum inophyllum, 12 Azadirachta indica, 13 Hevea brassiliensis, 14 Melia azedarach, 15 Nicotiana tabacum, 16 Pongamia pinnata, 17 Sapindus mukorossi, 18 Simmondsia chinensis, 19 Madhuca indica, 20 Datura stramonium, 21 Carberra odollam, 22 and Silybum marianum. 23 Recently, the potential of various tropical and non-edible plants, such as R trisperma, for biodiesel production has been reported. 5 However, exploitation of the Philippine Tung plant as energy bio-sources risks its habitat loss. As a decline in the R trisperma population worldwide has been reported, it is listed as a vulnerable plant. 24 Therefore, conservation efforts are necessary. Species identification is an important step in conserving vulnerable plants and strengthening species conservation regulations. 25 A promising method to identify plant species is through DNA barcoding, which requires genetic information of the plant. As genomic information on R trisperma is still limited, 26 developing barcoding markers for this species is necessary. Some molecular markers have been used for barcoding plant DNA, including rbcL, matK, trnH-pbsA, ITS, and ITS2. However, the Consortium for the Barcode of Life (CBOL) suggested matK and rbcl as the preferred plant barcoding region. 27 A suitable locus for plant DNA barcoding depends on the taxonomic group of the plants. 28 In addition, the discriminative power of the markers varies across plant species. 29 Therefore, in this study, we aimed to evaluate the use of both plastid rbcL and matK loci as plant DNA barcoding markers for R trisperma.
Despite its conservation issues, R trisperma, as a new source of biodiesel, has received much attention in agronomy and bioenergy-related gene expression studies. Quantification of gene expression might provide information on the activity of biodiesel-related genes or other biological activities and mechanisms, for which quantitative PCR (qPCR) is considered a reliable method. However, certain parameters should be carefully considered, including the concentration of RNA samples, PCR components, and complementary DNA (cDNA) synthesis and efficiency of the reference genes. 30 Reference genes are used to reduce quantification errors by normalizing the targeted gene expression with constitutive expression of the internal control.
Suitable reference genes for gene expression studies in biofuel plants have been screened. Zhang et al 31 suggested the use of actin and TUB8 as reliable reference genes for gene expression studies in Jatropha curcas. Karuppaiya et al 32 also validated some reference genes, including Actin, Tubulin beta-5 chain (TUB), Elongation factor 1-alpha (GAPDH1), and Ubiquitin-conjugating enzyme E2 32 (UBI). Nonetheless, to date, no study has reported a search for a reference gene in R trisperma. Therefore, this study also aimed to isolate an Actin fragment gene as a candidate reference gene. Although the effectiveness of the Actin housekeeping genes as reference genes should be further elucidated, this study might serve as a preliminary study for further gene expression profiling in R trisperma.
Housekeeping genes are frequently used as reference controls. This is mainly owing to their abundance in eukaryotic cells, and their continuous expression at every stage of development and in all tissues. 33 Some of these encode actin protein, tubulin, GAPDH, and 18S ribosomal RNA (rRNA). However, specific and optimal reference genes often depend on the species, tissue type, and treatments.34,35
Actin protein, a globular structure with a mass of approximately 42 kDa, is found in abundance and highly conserved in almost all higher organisms. 36 These proteins play an important role in forming tissues and providing mechanical support for cells, thereby determining cell shape, cell movement, and cell division.37,38 Actin is also important in plants for cell morphogenesis, cell elongation, as a cell wall component, and for the growth of root hairs, trichome cells, pollen tubes, and apical meristems. 39 Actin genes have been reported to be expressed stably in various tissues at all developmental stages. 40 Some gene expression studies in potatoes, 41 soybeans, 42 rice, 43 and in leaves, flowers, stems, and roots of Carthamus tinctorius 44 used the actin gene as the reference standard in qPCR analysis. Nevertheless, the R trisperma Actin has not been isolated and is not yet available in the database. This study was conducted to perform DNA barcoding of R trisperma using the rbcL and matK genes, as well as to isolate the R trisperma Actin-coding gene fragment. This study might serve as a preliminary study for selecting reliable reference standards for further transcriptomic studies.
Materials and Methods
Plant materials
R trisperma used in this study was provided by the Purwodadi Botanical Garden of the National Research and Innovation Agency, Indonesia. Fresh young R trisperma leaves were collected in situ from the botanical garden and stored at –20℃ for genomic DNA (gDNA) isolation and total RNA extraction. All fresh samples were ground in liquid nitrogen before being subjected to gDNA and total RNA extraction.
Data collection of Actin and primer design
Actin gene sequence data were retrieved from the National Center for Biotechnology Information database (https://www.ncbi.nlm.nih.gov/genbank/). The selected members of the Euphorbiaceae family included Ricinus communis (AY360221.1), Jatropha curcas (HM044307.1), Vernicia fordii (JQ680035.1), and Hevea brasiliensis (JF775488.1). The sequence data of the four species were downloaded in the FASTA format, and sequence alignment was performed using MEGA11. The DNA alignment results showing conserved regions were used as the basis for designing the primers (Suppl. 1). Both forward and reverse primers contained 20 to 24 bp selected from the conserved region. The selected primers were then analyzed for secondary structure, primer dimer, melting temperature, and GC content using primer analysis and Tm Calculator tool (https://sg.idtdna.com/calc/analyzer).
Nucleic acid extraction and first-strand cDNA synthesis
gDNA extraction was performed to obtain nuclear, chloroplast, and mitochondrial DNA for R trisperma DNA barcoding analysis. gDNA extraction was carried out using ZR™Plant/Seed DNA MicroPrep (Zymo Research) as per the manufacturer’s protocol. The samples used for gDNA extraction were obtained from 150 mg of R trisperma leaves, which were ground using liquid nitrogen prior to extraction. 45 Meanwhile, we used Plant Total RNA Mini Kit (Genaid) to isolate the total RNA from R trisperma seeds, as described by Jadid et al. 46 The plant endosperm was prepared and ground in liquid nitrogen. We measured the concentration and quality of the DNA and total RNA extracts using NanoDrop™. The A260/A280 ratio was used to evaluate the quality of the DNA and RNA extracts. Approximately 0.2 µg of total RNA was used as a template for cDNA synthesis using ReverTra Ace™ qPCR RT (Toyobo) according to the kit’s instructions.
DNA amplification and sequencing
DNA barcoding was carried out by amplification of the extracted gDNA using universal primers and TOYOBO KOD FX Neo (TOYOBO). The primers used included RbcL forward (5′ ➔ 3′): ATGTCACCCACCAACAGAGACTAAAGC, RbcL reverse (5′ ➔ 3′): GTAAAATCAAGTCCACCRCG, matK forward (5′ ➔ 3′): ACCCAGTCCATCTGGAAATCTTGGTTC, and matK reverse (5′ ➔ 3′): CGTACAGTACTTTTGTGTTTACGAG. The amplification parameters consisted of an initial denaturation at 95℃ for 3 min, followed by 35 cycles of denaturation at 98℃ for 10 s, primer annealing at 53℃ for 30 s, and extension at 68℃ for 45 s. A final extension was performed at 68℃ for 5 min to complete the PCR process.
Reverse transcription-polymerase chain reaction (RT-PCR) amplification of the Actin gene was carried out according to the manufacturer’s instructions using GoTaq® Green Master Mix (Promega). Actin fragment isolation was performed by using PCR. The samples were denatured initially at 95℃ for 2 min, followed by 35 cycles of amplification, which comprised denaturation at 95℃ for 30 s, annealing at 55℃ for 30 s, and extension at 72℃ for 90 s, with final extension at 72℃ for 5 min. The primers used for amplification were 10ACTks (5′
Molecular gene cloning of Actin fragment
The purified Actin fragment was cloned into a pTA2 vector (Toyobo, TAK-101). Subsequently, the pTA-Actin plasmid was integrated into Escherichia coli DH5α competent cells (Zymo Research, T3007). 47 The transformed bacteria were then cultured in Luria Bertani (LB) medium supplemented with ampicillin. Blue-white bacterial screening was performed to select bacterial colonies harboring Actin fragments. Eight bacterial colonies were randomly selected for colony PCR analysis using T3 and T7 promoter primers (KOD FX Neo [Toyobo]). 48 The expected bacterial colonies were cultured in an LB liquid medium, followed by plasmid extraction.
DNA sequencing
DNA sequencing was performed using an ABI PRISM® BigDye™ Terminator Cycle Sequencing Kit (Applied Biosystems). Sequencing of the rbcL and matK genes fragments was performed using the universal primers rbcL and matK. Plasmid sequencing was carried out at using T7 promoter primer (5′-TAATACGACTCACTATAGGG-3′) and T3 promoter primer (5′-AATTAACCCTCACTAAAGGG-3′).
Phylogenetic analysis
The nucleotide bases from the sequencing were then evaluated using nucleotide BLAST (https://blast.ncbi.nlm.nih.gov/) to determine gene similarity within the tested plant species. The nucleotide sequences obtained from BLAST analysis were downloaded and subsequently aligned with the Actin (ACT) R trisperma gene using the ClustalW Alignment algorithm (BioEdit v7.0.5). The phylogenetic tree construction was compiled using the MEGA11 application with neighbor-joining (NJ) and a 1000× bootstrapping value.
Results and Discussion
DNA barcoding using R trisperma rbcL and matK genes
The rbcL and matK gene fragments were detected as 650 and 850 bp, respectively, (see Figure 1). Subsequently, we purified the DNA via gel extraction according to the expected DNA size before performing DNA sequencing. The purification process washes DNA from remnants of salt, protein, and polyphenolic compounds. 49

Electrophoregram of PCR products of rbcL and matK Reutealis trisperma gene fragments.
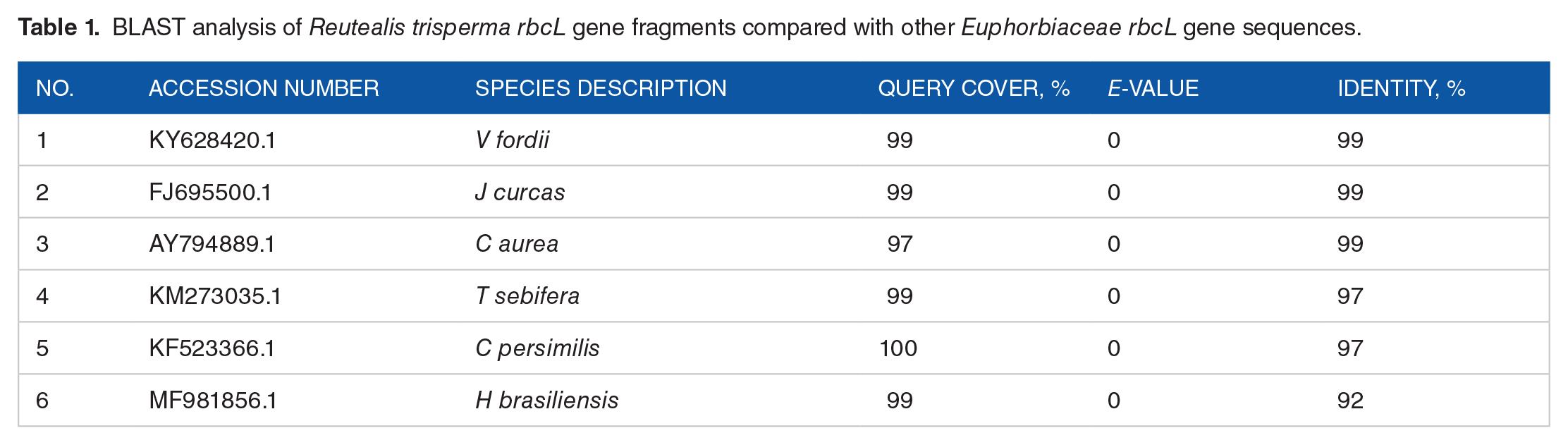
The sequencing results were analyzed for gene sequence homology using BLAST (see Table 1). BLAST analysis showed an E-value of 0 for some Euphorbiaceae members, indicating high similarity of rbcL gene among this plant family. The resulting query coverage value was 97% to 100%, with a similarity (Identity) value of 97% to 99%, compared with other Euphorbiaceae species. The high similarity value indicated high homology of the R trisperma rbcL sequence with rbcL from other members of the Euphorbiaceae family. The highest and lowest similarity values were with the V. fordii rbcL gene (accession number KY628420.1, 99%) and H. brasiliensis rbcL gene (accession number MF_981856.1, 92%), respectively.
BLAST analysis of Reutealis trisperma rbcL gene fragments compared with other Euphorbiaceae rbcL gene sequences.
The matK gene sequence was analyzed using BLAST. It showed results identical to several species from the Rutaceae and Zingiberaceae families (see Table 2), indicated by an E-value of 0. The resulting query coverage value ranged from 90% to 100%, with a resulting similarity value of 97% to 99%. These results indicate that the matK nucleotide sequence of R trisperma is highly homologous to those of several species from the Rutaceae and Zingiberaceae families. The highest and lowest similarity values were shown to the Citrus maxima matK gene (accession number KR073223.1, 99%) and Naringa cranulata matK gene (accession number AB762385.1, 97%), respectively.
BLAST analysis of Reutealis trisperma matK gene fragments compared with other Euphorbiaceae matK gene sequences.

The rbcL and matK phylogenetic trees for DNA barcoding were constructed using MEGA11, which describes the evolutionary relationship between sample organisms. 50 Using the NJ algorithm, we compared the R trisperma sequence with other organisms from the BLAST results. The NJ method best estimates the branch length that most closely reflects the distance between sequences. 51
Two main clusters were identified in the phylogenetic tree analysis of the rbcL gene (see Figure 2). The first cluster comprised species from the Euphorbiaceae family, namely, RtrbcL, VfrbcL, CarbcL, TsrbcL, CprbcL, HbrbcL, and JcrbcL. The second cluster comprised DzrbcL and CmrbcL. Our data indicate that RtrbcL belongs to a monophyletic group with other Euphorbiaceae families. This result is supported by the unique characteristics of the Euphorbiaceae family, which contains white to red latex and has a box-shaped fruit.1,52
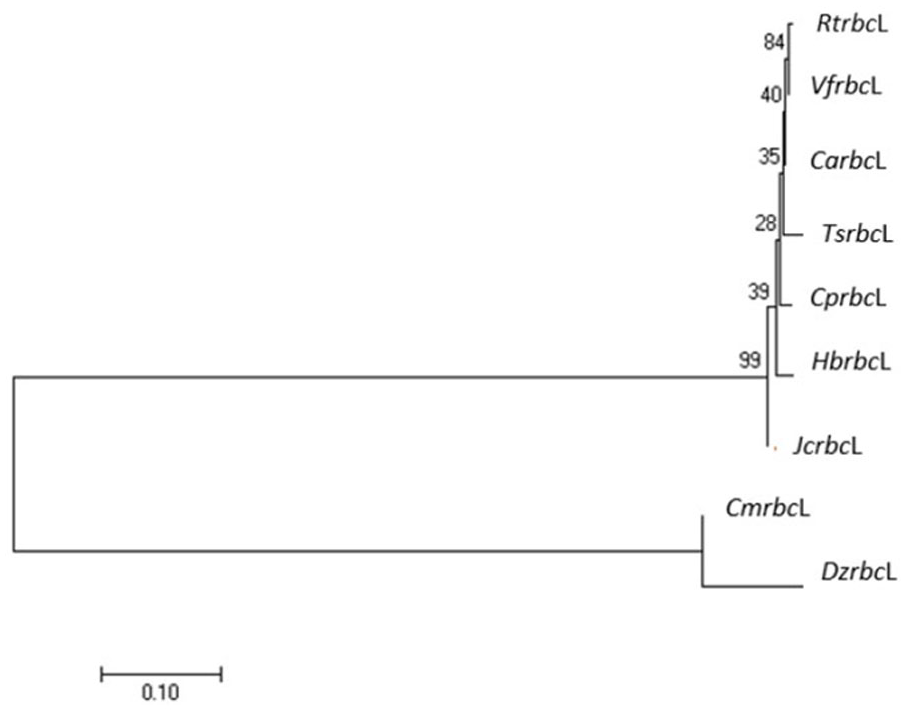
Phylogenetic tree of RtrbcL with several species of the family Euphorbiaceae.
A comparison of the R trisperma matK gene with those of other Euphorbiaceae species was also displayed in the constructed phylogenetic tree based on BLAST analysis (see Table 2). To construct the tree, additional species from the Euphorbiaceae family were added, namely, V fordii (accession number NC_034803.1), Jatropha curcas (accession number NC_012224.1), H brasiliensis (accession number HQ606140.1), and R communis (accession number AB233767.1). The matK phylogenetic tree analysis (see Figure 3) revealed two main clusters. The first cluster comprised RtmatK, CmmatK, ZomatK, CumatK, CdmatK, CjmatK, NcmatK, SbmatK, and AsmatK. In contrast, the second cluster comprised matK from species groups of the other Euphorbiaceae family— VfmatK, JcmatK, HbmatK, and RcmatK. RtmatK is in a cluster with several species from the Rutaceae and Zingiberaceae families.

Phylogenetic tree of RtmatK with several Euphorbiaceae species.
Some DNA barcoding studies have used multi-locus DNA barcoding markers, such as intergenic regions (trnH-pbsA), nuclear ribosomal ITS, or other chloroplast genomes (rpoC1 and rpoB). However, CBOL has described matK and rbcl as suitable plant barcoding markers. 27 The above results revealed that the rbcL gene provides a more informative description of the molecular phylogenetics of R trisperma. The rbcL gene sequence has a better discrimination ability than the matK gene sequence, favoring molecular identification. Similar results were also described in the plant DNA barcoding study performed by Maloukh et al, 53 where matK was also unable to discriminate against plant species native to the United Arab Emirates. Furthermore, Barthet and Hilu 54 stated that the matK gene underwent nucleotide substitution three times more often than the rbcL gene.
Isolation and molecular cloning of Actin gene from R trisperma
We successfully isolated the Actin fragment from R trisperma using RT-PCR. The isolated fragment was 986 bp in size (see Figure 4). The PCR product was gel extracted and purified. The purpose of purification was to remove excess primers, dNTPs, enzymes, and short DNA amplicons resulting from incomplete polymerization, dyes, and salt residues in the PCR reaction. 55 The purification results were then used as DNA inserts in the ligation process. Ligation of the Actin gene was performed by combining the DNA fragments with a pTA2 cloning vector, resulting in a recombinant plasmid of 3967 bp. Further steps comprised transforming the pTA2 harboring the Actin fragment into DH5α E. coli competent cells, followed by inoculation of the transforming bacteria on a solid LB medium supplemented with ampicillin.

Amplification of Reutealis trisperma Actin fragment.
Positive transformants were selected using the blue-white screening method on LB agar supplemented with ampicillin, isopropyl β-D-1-thiogalactopyranoside (IPTG), and 5-bromo-4-chloro-3-indolyl-beta-D-galacto-pyranoside (X-gal). E. coli harboring the recombinant plasmid was indicated by white colonies. 56 These white colonies were further confirmed using a colony PCR. Eight white colonies were selected for colony PCR using the T3 promoter and T7 promoter primers (see Figure 5). Colony 6 had a matching target size (1173 bp). We conducted further plasmid extraction of colony 6 bacteria, followed by DNA sequencing. Our sequencing data confirmed the 986 bp amplicon size of Actin fragment. This size is sufficient to be used as a basis for quantitative real-time PCR (qRT-PCR) primers. As previously reported, the amplicon length for qRT-PCR is generally recommended to be below 150 to 200 bp. 57 Some housekeeping genes have been used as reference genes or internal standards for gene expression analysis. 58 Among those genes, previous studies have demonstrated that Actin (ACT) might serve as the most suitable candidate. A study on the effect of abiotic stress in Daucus carota L. showed that Actin demonstrated good and stable performance for qPCR analysis in heat, cold, and methyl jasmonate treatment. 59 Moreover, Jadid et al 60 used Actin as the internal standard during gene expression profiling in different vegetative organs of Jatropha curcas. Similar results were reported by Feng et al, 61 who evaluated housekeeping genes in response to abiotic stress in celery (Apium graveolens L.). Taken together, RtActin may be used in R trisperma species. Further experiments are required to evaluate and validate the performance of the Actin gene.

Validation of pTA-ACT plasmid using PCR colony.
Phylogenetic analysis of R trisperma Actin among Euphorbiaceae species
Gene sequence homology analysis was carried out using the previously obtained sequence, which was then processed using BLAST. 26 The R trisperma Actin-coding gene sequence has a query coverage value of 99% nucleotides, with a similarity value of 92% to 97% to different Euphorbiaceae species tested in this study (see Table 3). This similarity value indicates gene sequence homology. 62 The R trsiperma Actin-coding gene fragment showed the highest and lowest similarity values to the V fordii Actin-coding gene (accession number JQ680035.1, 97%) and Euphorbia lathyris Actin gene (accession number KC182753.1, 92%), respectively. Species numbers 1 to 7 belong to the Euphorbiaceae family. Species number 8 belongs to the family Fagaceae, while species 9 to 11 belong to the Malvaceae family. According to information from the actin-coding genes, these results indicate that Fagaceae and Malvaceae are closest to Euphorbiaceae.
BLAST analysis of Reutealis trisperma ACT gene fragments compared with other Euphorbiaceae ACT gene sequences.
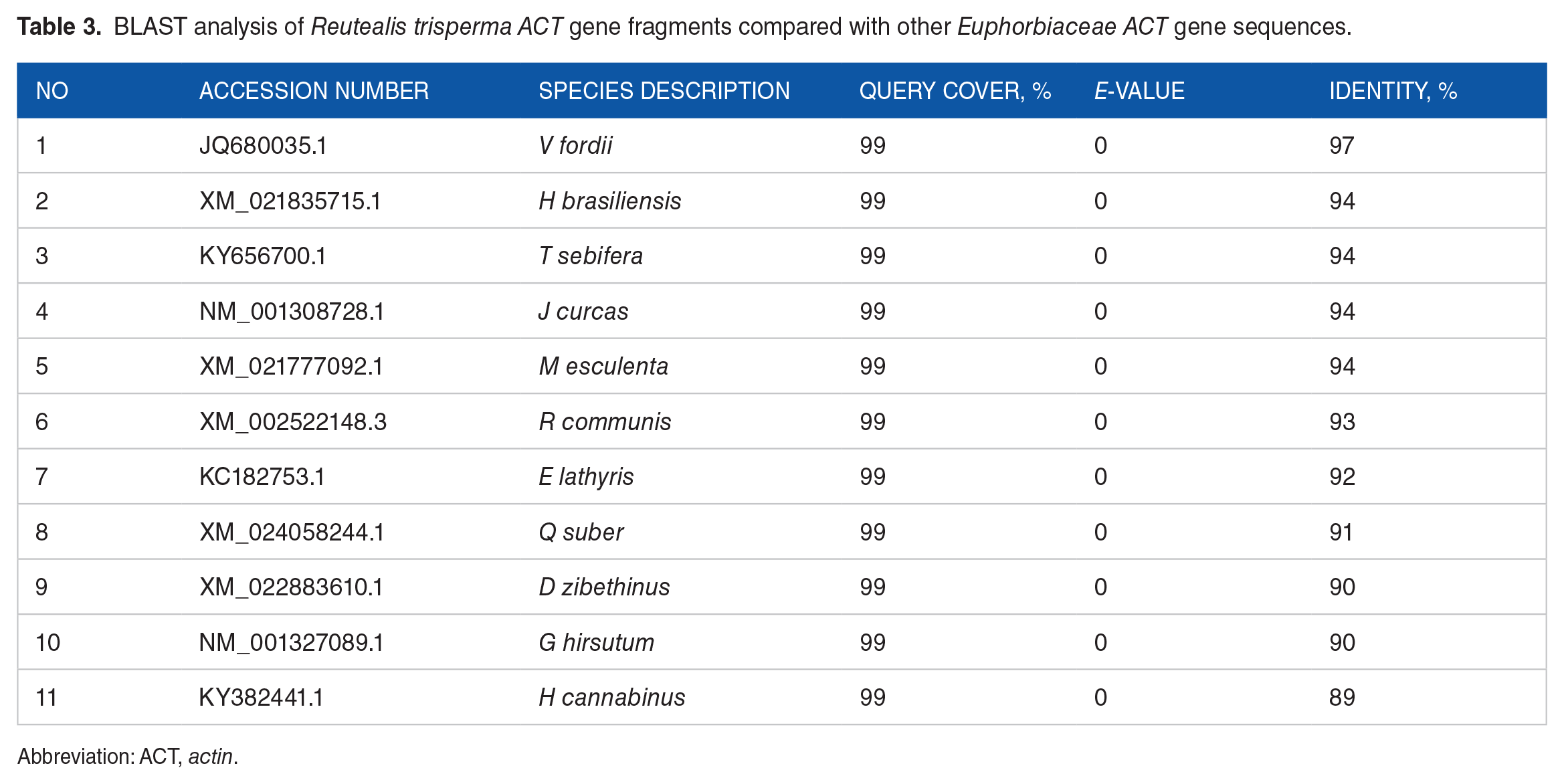
Abbreviation: ACT, actin.
The R trisperma Actin-coding gene was then compared with other Actin-coding genes of Euphorbiaceae species using phylogenetic analysis. Four species outside the Euphorbiaceae family, including Quercus suber (XM_ 024058244.1; Fagaceae), Durio zibethinus (XM_022883610.1; Malvaceae), Gossypium hirsutum (NM_001327089.1; Malvaceae), and Hibiscus cannabinus (KY382441.1; Malvaceae) were selected as outgroup. Outgroups were included in the analysis to obtain more credible information from related gene sequences. Outgroups were needed to provide polarization of characters or traits, namely, apomorphic and plesiomorphic. 63 Based on the phylogenetic tree (see Figure 6), the 11 species were separated into two main clusters. The first cluster comprised actin-coding genes from Euphorbiaceae species, including RtACT, VfACT, JcACT, TsACT, ElACT, HbACT, MeACT, and RcACT. The second cluster comprised QsACT, DzACT, GhACT, and HcACT. This result indicates that RtACT is in a monophyletic group with other Euphorbiaceae families. RtACT displayed similarity to VfACT, supported by a bootstrap value of 99%. According to Hall, 64 a clade can be trusted with bootstrap values above 90% and cannot be trusted if bootstrap values are below 25%. These results indicate that our phylogenetic tree construction is reliable, and the Actin gene can provide an informative description of the molecular phylogeny of R trisperma.

RtACT phylogenetic tree among other Euphorbiaceae species.
Conclusions
This study used two plastid markers (rbcL and matK) for the DNA barcode of R. trisperma, a promising biosource for biodiesel production. We isolated 650 and 850 bp, corresponding to rbcL and matK fragments, respectively. However, we observed that the rbcL marker is more discriminatory for the molecular identification of R trisperma plants than the matK marker. Our phylogenetic tree of the rbcL plastid marker showed that R trisperma was clustered in the same group as other Euphorbiaceae plants and displayed a high similarity to V fordii. Nevertheless, other multi-locus DNA barcoding markers, such as intergenic regions (trnH-pbsA), nuclear ribosomal ITS, or other chloroplast genomes (rpoC1 and rpoB), could be used to compare the discrimination power of the markers. Furthermore, we successfully isolated 986 bp of the Actin-coding gene, which might help in the development of gene expression analyses of R trisperma. The reference gene should be stably expressed in all plant organs because it maintains the essential cellular functions. Therefore, the actin fragment sequence obtained for differential gene expression studies of R trisperma organs should be conducted. This could provide additional information on RtACT stability as a potential reference gene. BLAST analysis showed that the gene was homologous to other Actin-coding genes of the Euphorbiaceae family and showed the highest degree of kinship with the Actin-coding genes of V fordii. Taken together, our DNA barcoding results could be used as a reference for R trisperma conservation programs. Meanwhile, validation of Actin stability and effectiveness as a reference gene for R trisperma transcriptomic analysis still needs to be further validated.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322231182768 – Supplemental material for Plastid DNA Barcoding and RtActin cDNA Fragment Isolation of Reutealis Trisperma: A Promising Bioresource for Biodiesel Production
Supplemental material, sj-docx-1-bbi-10.1177_11779322231182768 for Plastid DNA Barcoding and RtActin cDNA Fragment Isolation of Reutealis Trisperma: A Promising Bioresource for Biodiesel Production by Nurul Jadid, Nur Laili Alfina Rosidah, Muhammad Rifqi Nur Ramadani, Indah Prasetyowati, Noor Nailis Sa’adah, Aulia Febrianti Widodo and Dwi Oktafitria in Bioinformatics and Biology Insights
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors are indebted to the Directorate of Research and Community Services (DRPM) of the Institut Teknologi Sepuluh Nopember, Surabaya, Indonesia for providing financial support under the PAKERTI scheme (1658/PKS/ITS/2022).
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
NJ, NNS, and DO conceived and designed the study. NLAR, IP, MRNR, and AFW performed bioinformatics analysis. NJ and NLAR performed molecular biology experiments. NJ, NNS, and DO supervised the experiments and acquired the funding. NJ, NLAR, and MRNR drafted the article. NJ, NNS, and DO reviewed and edited the article. All authors have read and approved the final article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
