Abstract
Reproductive traits are affected by many factors, including ovarian function, hormones, and genetics. Genetic polymorphisms of candidate genes are associated with reproductive traits. Several candidate genes are associated with economic traits, including the follistatin (FST) gene. Thus, this study aimed to evaluate whether the genetic variations in the FST gene are associated with the reproductive traits in Awassi ewes. The genomic DNA was extracted from 109 twin ewes and 123 single-progeny ewes. Therefore, 4 sequence fragments from the FST gene were amplified using polymerase chain reaction (PCR) (exon 2/240, exon 3/268, exon 4/254, and exon 5/266 bp, respectively). For a 254 bp amplicon, 3 genotypes were identified: CC, CG, and GG. Sequencing revealed a novel mutation in CG genotypes c.100C > G. The statistical analysis of c.100C > G showed an association with reproductive characteristics. Ewes carrying the c.100C > G had significantly (P ⩽ .01) lower litter sizes, twinning rates, lambing rates, and more days to lambing compared with CG and CC genotypes. Logistic regression analysis confirmed that the c.100C > G single-nucleotide polymorphism (SNP) is responsible for decreasing litter size. According to these results, the variant c.100C > G negatively affects the traits of interest and is associated with lower reproductive traits in Awassi sheep. As a result of this study, ewes carrying the c.100C > G SNP have lower litter size and are less prolific.
Introduction
Reproductive traits play a significant role in sheep productivity.1,2 Genetics and the environment influence reproductive traits. Sheep reproductive traits are regulated predominantly by genetic factors.3-5 Among the candidate genes associated with reproduction in livestock, follistatin (FST) is located on chromosome 7 of caprine, chromosome 20 of cattle, and chromosome 16 of sheep, which is composed of 6 exons.6,7 This gene encodes FST protein, is a monomeric glycoprotein (31-39 kD), that binds receptors on cysteine-rich cells and microenvironments. 8 It is expressed widely in the ovary, skin, and skeletal muscle. 6 The primary function of FST is to bind and neutralize activin through its binding properties.9,10 Since FST binds to activin, it inhibits follicle-stimulating hormone (FSH) secretion by blocking its receptor (Act-RII) from combining with it.8,11 Through suppressing FSH secretion from the anterior pituitary, FST controls granulosa cell differentiation, follicular growth, atresia, and luteinization processes. 12 In livestock species, FST variations affect sheep productivity. 13 According to the findings of Ma et al, 6 2 single-nucleotide polymorphisms (SNPs) (Chr 16. 25 633 662 A > G; Chr 16. 25 633 569 T > C) are associated with traits associated with wool quality in Chinese Merino sheep. In White Leghorn and Aseel chickens, haplogroups KF921968.1, KF921969.1, MK455102, and MK45510 are associated significantly with growth traits (P < .05). 10 Recently, a study by Zhu et al 7 revealed that SNP_chr7-65652612, located at the 3′ untranslated regions (3′ UTR ) of FST-like-4, significantly relating to litter size in Dazu black goats of the first and second parity. However, the relationship between FST variants and reproductive traits in Awassi sheep has not been investigated.
The Awassi breed is dominant mainly in Middle Eastern countries. 14 Despite their ability to cope with harsh conditions in unfavorable situations, 15 Awassi sheep have been reported to be less prolific than Karakul and Assaf sheep in the Middle East. 16 Most breeders in the Middle East are concerned about Awassi sheep’s low reproductive performance, which drives efforts to improve their reproductive quality. The FST gene may be useful in improving breeding efficiency in Awassi sheep. Therefore, this study examined possible associations between FST polymorphisms and reproductive traits in Awassi sheep by examining polymorphisms within its coding regions.
Materials and Methods
Animal
The study was conducted between July 2021 and April 2022 at Al-Qasim Green University according to international guidelines (Agri, No. 015,7, 20). The study included 232 ewes of 3 to 4 years of age that were sexually mature and multiparous. The ewes were classified as 123 singletons and 109 twins according to their reproductive history, weighing 40 to 60 kg. Therefore, 2 stations—Babylon and Karbala—were randomly selected for the ewes. In the studied flocks, 10 to 12 rams were randomly allocated to mate with about 20 to 25 ewes per ram, with male identification recorded. Veterinarians confirmed the rams’ health and demonstrated their fertile status, as they previously produced offspring. A record of lambing dates and litter sizes was kept from 160 to 200 days after the ram was introduced. All animals were fed individually and kept in similar nutrition conditions. The animals were fed daily a concentrated diet composed of 59% barley, 40% bran, and 1% salt in proportion to their body weight of 2.5%. However, 3 kg alfalfa and 1 kg of straw were also fed to each animal. All animals had access to fresh water at all times. Several reproductive traits were recorded at the breeding stations, including twinning rate, lambing rate, days to lambing, survival rate, and litter size.
DNA and polymerase chain reaction
Genetic analysis was performed by collecting blood from the sheep’s jugular vein. Rapid salting-out method was used for the extraction of genomic DNA. 17 NCBI Primer-BLAST 18 was used to amplify the FST genetic sequence for all 232 animals. Polymerase chain reaction (PCR) experiments were performed with Bioneer PreMix (50 μM deoxynucleotide triphosphates [dNTPs], 10, 30, and 1.5 mM Tris-HCl, KCl, and MgCl2, respectively, and 1 U Top DNA polymerase). The optimal PCR amplification conditions were determined using a thermal gradient device (Eppendorf, Germany) (Table 1). The PCR reaction was performed at 94°C for 4 minutes, then 30 cycles (30 seconds each) of 94°C, annealing, and 72°C were conducted. 19 A Chemidoc Gel Imager (Bio-Rad, USA) was used to image agarose gel images of PCR products electrophoresed on agarose gels (2%). 20
The oligonucleotide primer sets designed for the amplification of the ovine FST gene. The symbols “F” and “R” refer to forward and reverse primers, respectively. The design was based on the ovine NCBI Reference Sequence NC_056069.1.
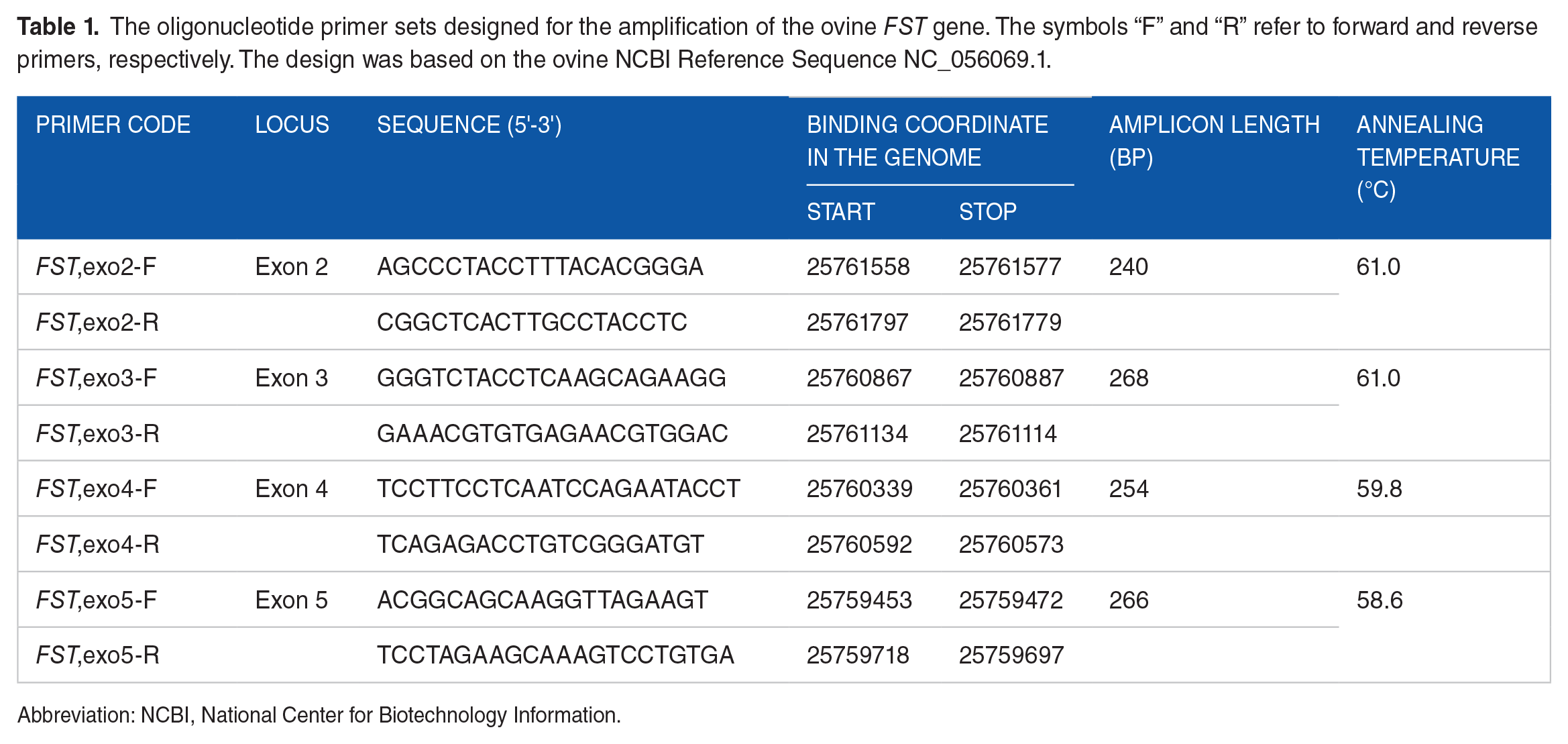
Abbreviation: NCBI, National Center for Biotechnology Information.
Single-strand conformation polymorphism
A genotype-based analysis was performed using the methodology described in Al-Thuwaini et al. 21 Denaturing loading buffers were added equally to each PCR amplicons (95% formamide, 0.05% xylene cyanol, and 20 mM EDTA, pH 8). A PCR amplicon was denatured for 7 minutes, then placed on wet ice and stored for 10 minutes. Samples were loaded in a 0.5 TBE buffer in polyacrylamide gels with a neutral denaturant. Following this step, the gels were electrophoresed at room temperature for 4 hours with 200 mA and 100 V. According to Byun et al 22 protocol, gels were stained using the rapid staining protocol.
DNA sequencing
Sequence laboratories (Macrogen, Geumcheon-gu, Korea) performed downstream reactions on all 232 animals once single-strand conformation polymorphism (SSCP) bands were detected on polyacrylamide gels. A sequence of the FST gene was obtained from the NCBI (https://www.ncbi.nlm.nih.gov). DNA polymorphisms within genotypes were visualized and edited with SnapGene Viewer 4.0.4 and BioEdit 7.1 (DNASTAR, Madison). To determine the novelty of observable variants, Ensembl genome browser 96 was used (https://asia.ensembl.org/index.html).
Statistical analysis
A genotype and allele frequency analysis was performed using PopGen32, version 1.31. 23 The Hardy–Weinberg equilibrium (HWE) was calculated, and the polymorphism information content (PIC) was determined according to Botstein et al. 24 Genotype associations were analyzed using IBM SPSS 23.0 (NY, USA) as follows

where Yijk are phenotypic traits, μ is mean, Gi is the fixed effect of ith genotypes (i = CC, CG, GG), Pj is the fixed effect of jth parity (j = 1, 2, 3), and eijk is random residual error. Statistical significance was determined at .05 with the Tukey–Kramer test. A chi-square test was conducted on 2 reproductive traits: lambing rate and birth type. The FST polymorphisms and litter size were analyzed using logistic regression. Model results in the preliminary analysis did not show any effect on interactions, lambing season, and age, so these were excluded from the calculation.
Results
Genotyping of FST gene, sequencing analysis, and genetic diversity
Overall, 4 FST gene coding regions corresponding to 240, 268, 254, and 266 bp were screened with PCR designs (Figure 1A). Electrophoresis showed monomorphic migration of all PCR-SSCP patterns containing amplicons covering exons 2, 3, and 5. A 254-bp amplicon designed for exon 4 showed 3 different PCR-SSCP patterns (Figure 1B). Sequencing reactions showed that only one of the SSCP variants contained the c.100C > G SNP, confirming the exon 4 heterogeneity. As a result of this nucleic acid substitution, CC, CG, and GG were assigned to the detected SSCP variants with homozygous C/C and G/G patterns, and the heterozygous C/G patterns found in the PCR-amplified products (Figure 1C). According to SNP bioinformatics analysis, the silent SNP p.205Tyr= had negative influences on the FST protein (Figure 1D).
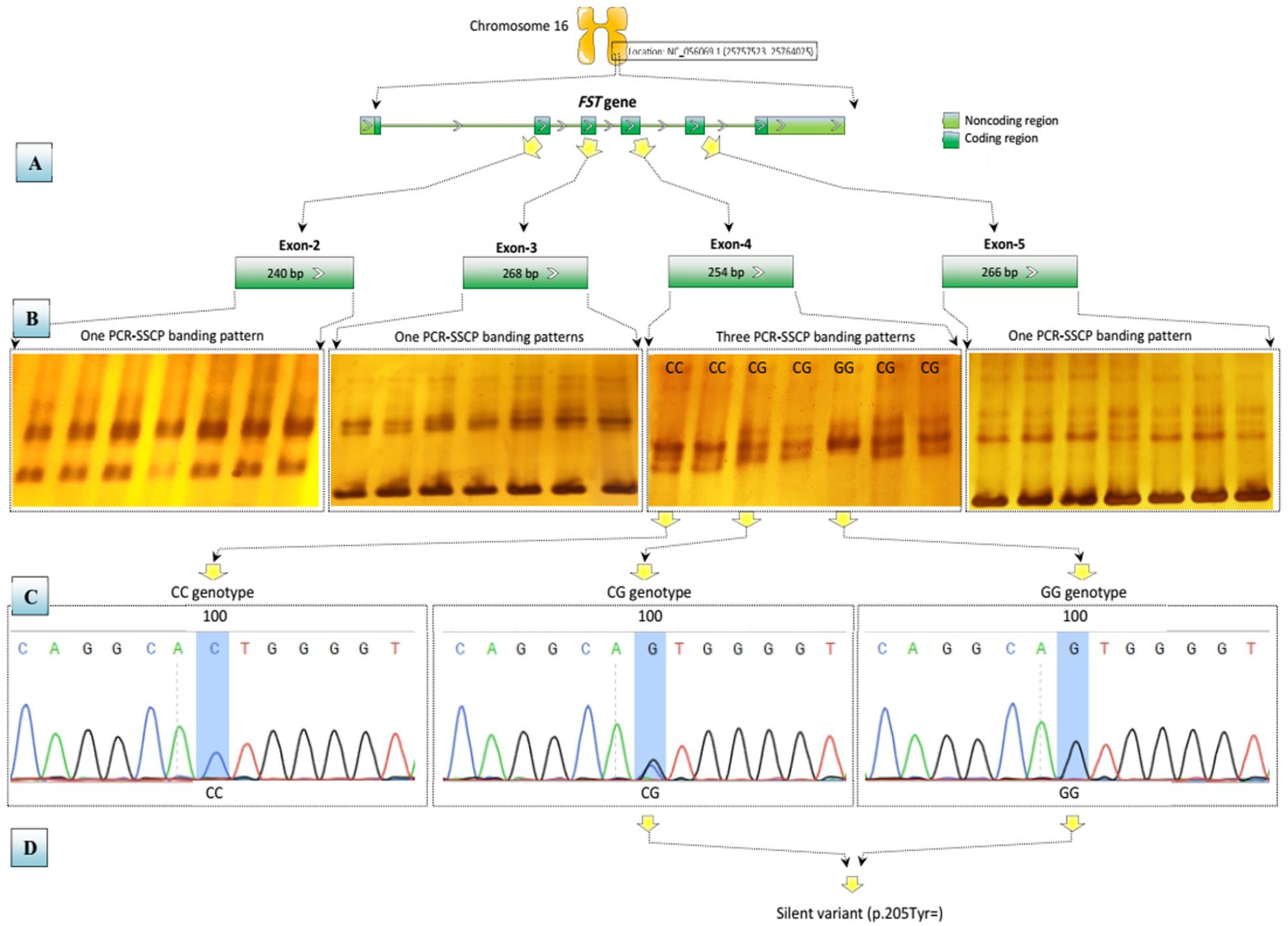
A schematic diagram for the FST gene-based PCR-SSCP-sequencing strategy within Awassi ewes. (A) PCR design of 4 PCR specific primers pairs for the amplification of 240, 268, 254, and 266 bp in exons 2, 3, 4 and 5, respectively. (B) PCR-SSCP genotyping, in which only exon 4 showed 3 genotypes, homozygous and heterozygous. (C) DNA sequencing electropherograms of the detected genotypes, in which 1 SNP, c.100C > G, was detected in the exon 4 in the heterozygous CG genotype. (D) SNP bioinformatics analysis of the observed silent p.205Tyr= SNP in the FST gene.
Genetic diversity analysis of FST revealed that CC genotype dominated the study population with a frequency of 0.48 (n = 111). The frequency of GG genotype was 0.35 (n = 82), followed by 0.17 (n = 39) for CG genotype. Based on PIC results (0.25 < PIC < 0.50), this study showed moderate polymorphism levels at the c.100C > G locus. Compared with HWE, a chi-square test indicated distinct differences (P ⩽ .05) in the FST polymorphisms (Table 2).
Genetic diversity of the FST gene in Awassi ewes detected by PCR-SSCP.

All chi-square tests have 2 degrees of freedom and within the significance level P ⩽ .05.
Abbreviations: χ2, chi-square; He, expected heterozygosity; Ho, observed heterozygosity; n, number of individuals; Ne, effective allele frequency; PCR, polymerase chain reaction; PIC, polymorphism information content; SSCP, single-strand conformation polymorphism.
Association analysis of FST gene with reproductive traits
In the association analysis between p.205Tyr= SNP locus and reproductive traits, CC genotype ewes, and CG and GG genotype ewes did not differ significantly (P ⩾ .05) concerning survival rate. Meanwhile, at the same p.205Tyr= SNP locus, individuals with CC genotype had statistically higher (P ⩽ .01) litter size, twinning ratio, lambing rate, and fewer days to lambing than individuals with CG and GG genotypes (Table 3). The logistic regression analysis provided further insight into the relationship between p.205Tyr= SNP and litter size. Ewes with the CC genotype had 1.65 lambs per animal compared with CG and GG genotypes. In this case, p.205Tyr= SNP adversely affected these traits.
The association between FST genetic polymorphism at locus c.100C > G and reproductive performance in Awassi ewes. (A) Reproductive performance for each genotype is shown. (B) Logistic regression analysis of FST genotypes and litter size in Awassi ewes is shown.

The P-value with statistical significance is indicated in bold numbers. Twinning rate (is the propensity to have more twin litters per 100 ewes at the same average lambing percentage). Number of days to lambing (number of days between the joining of rams until the subsequent lambing).
Survival rates of the lambs (the numbers of dead lambs were determined until weaning). Litter size (the number of lambs born per ewe lambing).
Abbreviations: CI, confidence interval; LSM ± SE, least square means ± standard error; β, regression coefficients.
Significant differences in means represent by differences in the same column within each classification.
Discussion
Polymorphism of FST gene and SNP bioinformatics
Molecular genotyping and genetic diversity analyses revealed polymorphisms in exon 4 of the FST gene in Awassi sheep. In livestock, FST gene variations have been investigated in several studies. According to Ma et al, 6 sequencing analysis of Chinese Merino sheep (Junken type) identified 7 SNPs in the FST gene. Dushyanth et al 10 reported polymorphisms in the FST gene coding region (exons 2 and 5) of native American (Aseel) and white leghorn chickens. The SNaPshot software is used by Zhu et al 7 to detect 26 SNPs and 1 deletion in UTRs from 186 Dazu black goats. However, there is a lack of literature on FST polymorphism and Awassi sheep. Accordingly, this study will provide genotypic details and new associations applicable to selecting sheep for marker-assisted selection, enabling future programs to measure functional traits more effectively.
Concerning SNP bioinformatics, the results revealed that silent mutation p.205Tyr= SNP negatively affected reproductive traits. This situation could be explained by the fact that silent mutations have previously been shown to be involved in biological processes. 25 Although silent mutations do not change amino acids, they can interfere with RNA splicing, affecting protein production negatively. Furthermore, it can alter regulatory sequences and lower gene expression. 26 According to Manning and Cooper, 27 silent variants can influence RNA synthesis and translation initiation. Changes in RNA secondary structures can affect many functions of the genome, including protein folding, accessibility of RNA to functional sites, and alternative splicing. 28 However, silent SNPs have been rarely reported to affect phenotypic traits in the literature.
Association analysis of the FST genotypes with reproductive traits of Awassi ewes
There was a highly significant (P ⩽ .01) association between the SNP and reproductive traits in those ewes with the p.205Tyr= SNP had higher litter size, twinning ratio, lambing rate, and fewer days to lambing than ewes with CG and GG genotypes. Mammalian FST controls steroidogenesis, oocyte development, and follicular cell division and differentiation. 29 A previous study by Lee et al 30 suggested that FST derived from the oocyte is crucial for determining the competence of oocytes in cattle by speeding up their development into blastocysts. Based on O’Connell et al, 31 activins and FST facilitate luteinization and conceptus implantation by interfering with multiple reproductive processes. Activin-FST influences the production of gonadotropin receptors and steroid hormones, which are responsible for the differentiation and growth of antral follicles. 32 FST, activin, and inhibin interact with autocrine/paracrine to regulate anterior pituitary FSH secretion. 33 Activin internalization and degradation are inhibited by FST, decreasing their bioavailability and preventing FSH secretion. 34 High levels of FSH and luteinizing hormone (LH) affect ovarian follicle development and maturation, resulting in increased litter size.35,36 In Dazu black goats, FST polymorphisms are associated with litter sizes. 7 Despite this, there has been little study of the relationship between FST variants and reproductive traits in livestock, and little is known about FST polymorphisms and their impact on sheep reproduction. As a result of these results, FST appears to be a promising candidate gene for sheep marker selection.
Conclusions
There was an association between genotype variations of the FST gene and reproduction in Awassi sheep. Animals of the CC genotype had higher twinning rates, higher lambing rates, and shorter days to lambing than animals of the CG and GG genotypes. These traits are adversely affected by the silent variant p.205Tyr= SNP. Identifying genetic polymorphisms in the FST gene can be recommended to improve sheep reproductive performance using a marker-assisted selection program.
Footnotes
Acknowledgements
The authors express great appreciation for the ewes provided by sheep stations in Babylon and Karbala.
Author Contributions
All authors contributed equally.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
