Abstract
Eukaryotic non-coding regulatory features contribute significantly to cellular plasticity which on aberration leads to cellular malignancy. Enhancers are cis-regulatory elements that contribute to the development of resistance to endocrine therapy in estrogen receptor (ER)-positive breast cancer leading to poor clinical outcome. ER is vital for therapeutic targets in ER-positive breast cancer. Here, we review and report the different regulatory features present on ER with the objective to delineate potential mechanisms which may contribute to development of resistance. The UCSC Genome Browser, data mining, and bioinformatics tools were used to review enhancers, transcription factors (TFs), histone marks, long non-coding RNAs (lncRNAs), and variants residing in the non-coding region of the ER gene. We report 7 enhancers, 3 of which were rich in TF-binding sites and histone marks in a cell line-specific manner. Furthermore, some enhancers contain estrogen resistance variants and sites for lncRNA. Our review speculates putative models suggesting potential aberrations in gene regulation and expression if these regulatory landscapes and assemblies are altered. This review gives an interesting perspective in designing integrated in vitro studies including non-coding elements to study development of endocrine resistance in ER-positive breast cancer.
Keywords
Introduction
Promoters, enhancers, and insulators are some regulatory features located on eukaryotic non-coding genomic regions that contribute to the spatiotemporal development of cells. 1 One such important feature is the enhancer element which when bound by specific proteins expedite gene expression regardless of their distance, orientation, and position from the associated target gene.2,3 Interaction of enhancer with regulatory proteins form a complex gene regulatory network that determines cell identity during development.4,5 Chromatin immunoprecipitation, microarrays, and high-throughput sequencing have helped to elucidate chromatin architecture and associated enhancer function. 6
Chromatin signature of enhancers helps to understand basic characterization of enhancer organization and cell-type specific activities. 7 Methylation marks on histones near enhancers correlate to its function in different cell types.8,9 Genes that are actively transcribed show histone H3 monomethylated at Lys 1 (H3K4me1) and acetylated at Lys 27 (H3K27ac),10-12 but developmental genes that are poised for future activation contain repressive trimethylated at Lys 27 (H3K27me3) near enhancers. 10 DNA methylation is important in determining enhancer landscape and eviction of transcription factors (TFs) contributing to cell-type specific enhancer activity 13 and gene expression. 14 The TF identity, binding site, and number reflect the complexity of the enhancer regulatory network. 15 Active enhancers also transcribe a class of non-coding RNA, enhancer RNA (eRNA) which assist in enhancer-promoter looping, chromatin modification, and regulation of gene expression. These eRNAs exhibit tissue and lineage specificity and are markers of cell state and function. 2 In some cases, an aggregation of multiple enhancers, termed as super enhancer, can contribute to gene expression patterns leading to a robust phenotype.16,17 Super enhancer 18 is located near genes that have high H3K4Me1, H3K27Ac marks, and p300 protein occupancy. 19 They are known to produce higher levels of eRNA which is responsible for cell identity, cell fate, transcription in cancer cells, and tumor heterogeneity. 2 The presence of disease associated non-coding variants on enhancers is estimated to explain a large proportion of heritability in comparison to coding variants. 20 Mutations in enhancer lead to loss of potential motif for TF-DNA interaction and aberrations in epigenetic modifications leading to enhancer-associated gene dysregulation.17,19
Estrogen receptor (ER) signaling is key driver of ER+ breast carcinogenesis and inhibition of ER signaling is the main focus of ER+ breast cancer therapy. 21 Intricate mutations in the ER pathway lead to development of endocrine resistance. 21 Those changes may be associated with ERα modifications either at the genetic, regulatory, or protein level. Evidence shows interaction of genomic features at pre- or post-transcription to contribute to the development of resistance. Specially enhancer reprogramming driven by assembly of TFs contribute to endocrine resistance.22-24 Studies have shown estrogen mediated epigenetic remodeling of enhancers at ERα binding in endocrine resistance breast cancer. 25 Also, reprogramming of enhancer by TF assemblies supports endocrine-resistant breast cancer 26 and breast cancer-specific epigenomic landscape contribute to dysregulated oncogene expression in breast tumorigenesis. 27 These studies report role of regulatory features contributing to development of endocrine resistance. However, there is limited information on the dynamics of DNA regulatory elements on estrogen receptor 1 (ESR1), estrogen receptor estrogen response element (ER-ERE) signaling, and how variation in enhancer landscape connects to chromatin reconfiguration leading to altered gene expression. Organization of the gene regulation is multi-tiered which contribute as target sites to address ER modulation. 28 This study uses ENCODE data base and data mining to review genomic regulatory features present on ER1 gene to illustrate the non-coding regulatory features, enhancers, variants, TFs, and methylation signatures on ER which can contribute to its modulation. In addition, it provides insights to cohesively integrate this information to understand development of endocrine resistance.
Methods
Regulatory elements present on Estrogen receptor 1 and its features
The University of California Santa Cruz (UCSC) Human Genome Browsers of builds GRCh37/hg19 at chr6:151,921,777-152,102,547 and GRCh38/hg38 at chr6:151,599,479-151,779,568 was evaluated. The Encyclopedia of DNA Elements (ENCODE) Project29,30 tracks were used to identify the gene regulatory elements on the ESR1 gene. The UCSC GeneHancer 31 track features were overlaid on the above coordinates to identify DNA regulatory elements corresponding to ESR1 gene. Further DNA regulatory tracks such as histone marks (H3K4Me1, H3K4Me3, H3K9Me3, H3K27Ac), chromatin state segmentation, and transcription factor-binding sites (TFBSs) were mapped on the GeneHancer identified regulatory sequence on ESR1. Tracks specific for cell lines were used to integrate and compare the chromatin states and TF binding at the regulatory region. The human genome build, GRCh38/hg38 was used to evaluate potential candidate cis-regulatory elements (cCREs) at ESR1. Flowchart illustrating the various databases used to annotate features on ESR1 in Supplementary Figure 1.
Single nucleotide polymorphisms on Estrogen receptor 1 regulatory elements
The single nucleotide polymorphisms (SNPs) located on the non-coding DNA regulatory elements on ESR1 was evaluated using literature search, Human Gene Mutation Database (HGMD), 32 and 1000 Genome Phase 1 and Phase 3 available from the UCSC hg19 genome build. SNPs present across the 7 enhancers were tabulated, evaluated, and characterized based on their association with estrogen resistance, clinical relevance, and allele frequency. Regulome Database 33 was used to investigate the chromatin accessibility of these SNPs.
Enhancer RNA signatures on Estrogen receptor 1 regulatory element
Enhancer RNA in Cancer (eRIC) database 34 was used to identify the eRNAs expressed in ⩽1 mb of ESR1 gene in The Cancer Genome Atlas (TCGA) Breast invasive carcinoma (BRCA) cell type. The eRIC database was used to confirm the eRNA target genes in BRCA cells.
Result
Estrogen receptor 1 gene contains Enhancer regulatory elements
We report 7 regulatory marks present at chr 6:151,921,777-152,102,547 genomic region. Six were annotated to be double elite enhancers and 1 was a promoter. Elite enhancers are DNA regulatory elements that are evidenced from more than 1 source, whereas double elite enhancers are association of multiple elite enhancers (Figure 1). The first enhancer (GH06J151615) was located on CCDC170 gene, 4 enhancers (GH06J151629, GH06J151631, GH06J151633, GH06J151666) were present at the intergenic region, and the last enhancer (GH06J151763) was present in the second intron of ESR1 gene. Interaction of enhancers GH06J151615, GH06J151629, GH06J151631, and GH06J151633 was in the forward direction (solid line), whereas enhancers GH06J151666, GH06J151689, and GH06J151763 showed reverse directed interaction (dotted line) shown in Figure 1. ENCODE scored strengths of the genomic regulatory elements based on the literature, number of unique TFs, and overlaps with ultra-conserved non-coding genomic elements. GH06J151633 showed the highest score of 11.03, whereas GH06J151631 showed the lowest confidence score of 4.5 (Table 1).
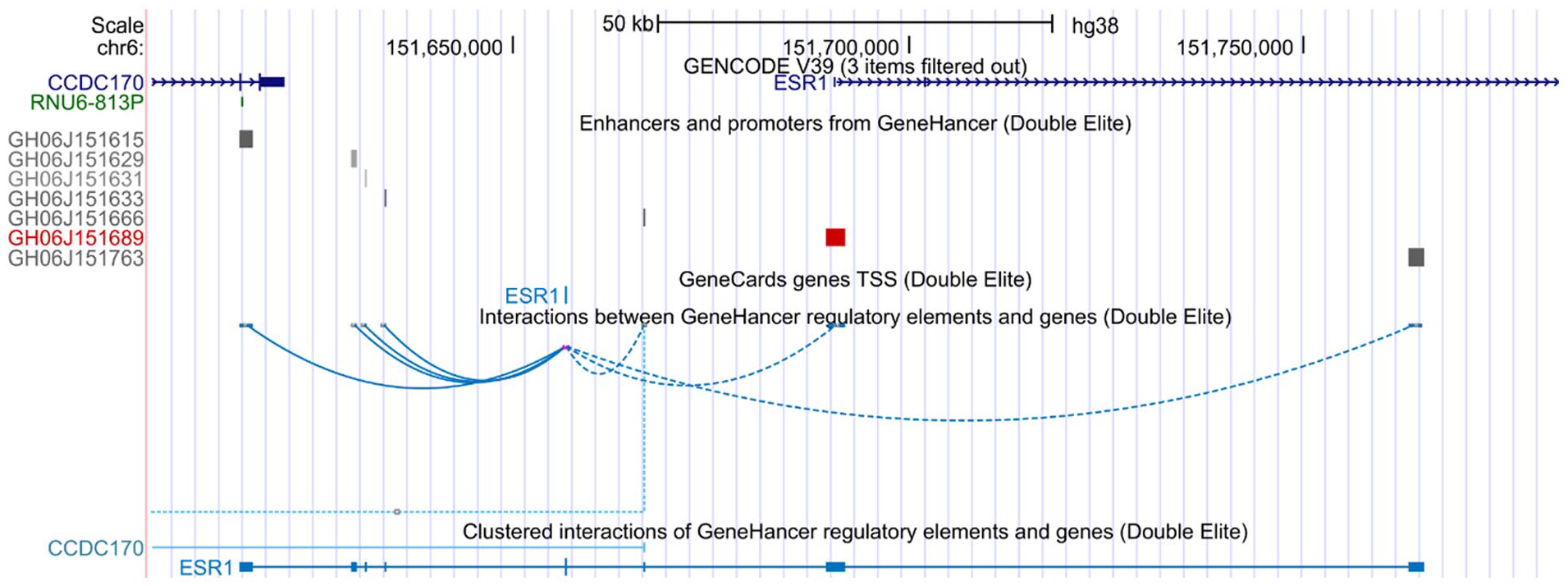
Double elite features and regulatory regions on ESR1 gene loci 6q25.1 region of human genome using ENCODE on UCSC Browser. A total of 7 double elite regulatory elements were observed using GeneHancer database available in UCSC ENCODE Browser using Human Hg38 version near the vicinity of ESR1 gene. GH06J151615 enhancer was seen in CCDC170 gene, whereas GH06J151629, GH06J151631, GH06J151633, and GH06J151666 were observed in the intergenic region. GH06J151689 was the promoter and GH06J151763 was present in the second intron of the transcript variant 4 of the ESR1 gene.
Elite enhancer coordinates, enhancer interactions, GWAS catalog, transcription factor, total number of SNPs.

Abbreviations: GWAS, Genome-wide association studies; NA, not available; SNP, single nucleotide polymorphism; TFBS, transcription factor-binding site; UCSC, University of California Santa Cruz.
Table provides a cumulative overview of the identified enhancers and its associated factors such as Transcription factors, GWAS catalog, and number of SNPs found in the enhancer. A total of 7 elite enhancers were reported using hg19 version of GeneHancer available in UCSC Genome Browser. The lowest confidence was seen for enhancer GH06J151631, and the highest confidence score was for the enhancer GH06J151689. The enhancers showed interactions with various non-coding RNAs, CCDC170 and ESR1. Also, SNPs rs6929137 and rs9397435 were found in enhancers—GH06J151615 and GH06J151629, respectively.
Regulatory elements on Estrogen receptor 1 interact with different genes
Six of the DNA regulatory elements on ESR1 interacted with both CCDC170 and ESR1 genes except GH06J151763, which interacted only with ESR1. In addition, enhancers GH06J151666, GH06J151689, and GH06J151763 interacted with Spectrin repeat containing nuclear envelope 1 (SYNE1), GH06J151689 with Zinc Finger and BTB domain containing 2 (ZBTB2) genes, and GH06J151615 with an lncRNA such as lnc-RMND1-2 (Table 1).
Histone marks on regulatory elements of Estrogen receptor 1
Enhancers associated with histone marks, H3K4Me1, H3K4Me3, H3K9Me3, and H3K27ac help initiate active transcription. 35 Enhancers GH06J151615, GH06J151633, GH06J151631, and GH06J151763 reported histone H3K4Me1 peaks in human hepatic cancer cells (HepG2). Enhancers GH06J151631 and GH06J151763 showed active H3K4Me1 and H3K4Me3 marks in human mammary epithelial cells (HMECs). Moderate H3K4Me1 and H3K9Me3 peaks were observed in GH06J151615, GH06J151629, and GH06J151763 in human embryonic stem cell (H1-hESC) cells (Figure 2).

Chromatin and histone state across 7 enhancers. Each cell line illustrates peaks for histone marks, H3K4Me1, H3K4Me3, H3K9Me3, and H3K27ac. (A) GH06J151615, (B) GH06J151629, (C) GH06J151631, (D) GH06J151633, (E) GH06J151666, (F) GH06J151689, and (G) GH06J151763. The first panel of tracks shows the horizontal bar of chromatin states retrieved from the ChromHMM database across the 3 cell lines. The second panel of vertical peaks colored in H1-hESC (green), HMEC (black), and HepG2 (pink).
Transcription factor-binding sites on Estrogen receptor 1 regulatory elements
Enhancers consist of dense clusters of TFBSs which are cell-type specific and involve in modulation of gene expression. 36 Five of the ESR1 regulatory elements (GH06J151615, GH06J151633, GH06J151666, GH06J151689, and GH06J151763) report cell-type TFBS (Figure 3). A total of 19 TFs show binding to enhancer GH06J151615 of which CEBPB and RXRA were reported in H1-hESC cells, whereas HNF4A and HNF4G were reported in hepatic cells. The GH06J151633 enhancer was contained binding site for 13 TFs. A TF SMC3, which is a DNA repair protein binds to GH06J151633 enhancer in H1-hESC cells, whereas FOXA1 and TEAD4 TFs are bound to it in HepG2 cells. The type of TF and enhancer TFBS is cell-type dependent. ZNF263, a zinc finger TF repressor, is bound to GH06J151666 enhancer in HMECs. The ESR1 regulatory element GH06J151689 is a promoter, with 39 TFs. Interestingly, the IKZF1 a chromatin remodeling factor bound to GH06J151689 specifically in HI-hESC cells, but bound to KAP1, a nucleosome remodeling protein in HMEC cells. Finally, GH06J151763, an ESR1 intronic enhancer, bounds to 23 TFs. It specifically bound to SP4 in H1-hES cells, but in HMEC cells, it bound to KAP1 and to SMC3, a DNA repair protein in HepG2 cells (Figure 3).

Transcription factor binding to the ESR1 DNA regulatory elements. Using UCSC we identified several tissue specific and common transcription factors binding to ESR1 enhancers. 19 TFs were bound onto GH06J151615 (A), 13 TFs on GH06J151633 (B), 39 TFs on the promoter (D), and 23 TFs on the intronic enhancer, GH06J151763 (E). Singleas seen binding to GH06J151666 enhancer (C). Enhancers GH06J151629 and GH06J151631 did not harbor any transcription factor.
Variants on Estrogen receptor 1 Enhancer elements
A total of 4 databases—1000 genome Phase 1 and Phase 3, 37 ClinVar, 38 HGMD, 32 and literature search with terms, “Breast cancer AND estrogen receptor AND Genetic”—were used to find and annotate the variants across the 7 enhancers on ESR1 (Supplementary Table 1). Overall, 245 variants were identified of which 12 were responsible for estrogen resistance and sum of 179 variants were located across the 7 regulatory elements of ESR1 gene. However, only rs6929137 and rs9397435 were present on GH06J151615 and GH06J151629 enhancers and were linked to estrogen resistance (Table 2). The potential function of rs6929137, rs9397435, and rs140068132 located on enhancers were analyzed using Regulome DB and SNP functionality scored based on its chromatin state in H1-hESC, HMEC, and HepG2 cell lines (Supplementary Table 2). The rs6929137 present on GH06J151615 showed strong transcription activity in H1-hESC cells, low activity in mammary epithelium cells, and enhancer activity in HepG2 cells. The SNP, r9397435 on GH06J151629 reported enhancer activity in H1-hESC and HMEC cells but showed strong transcription activity in HepG2 cells. Similiary, rs140068132 present on GH06J151633 reported low activity in H1-hESC cells, repressed state in HMEC cells and enhancer activity in HepG2 cell (Supplementary Table 2). Distinct variants across different enhancer show different regulatory potential across different cell lines.
Curative table of SNPs responsible for estrogen resistance along population and enhancer status.
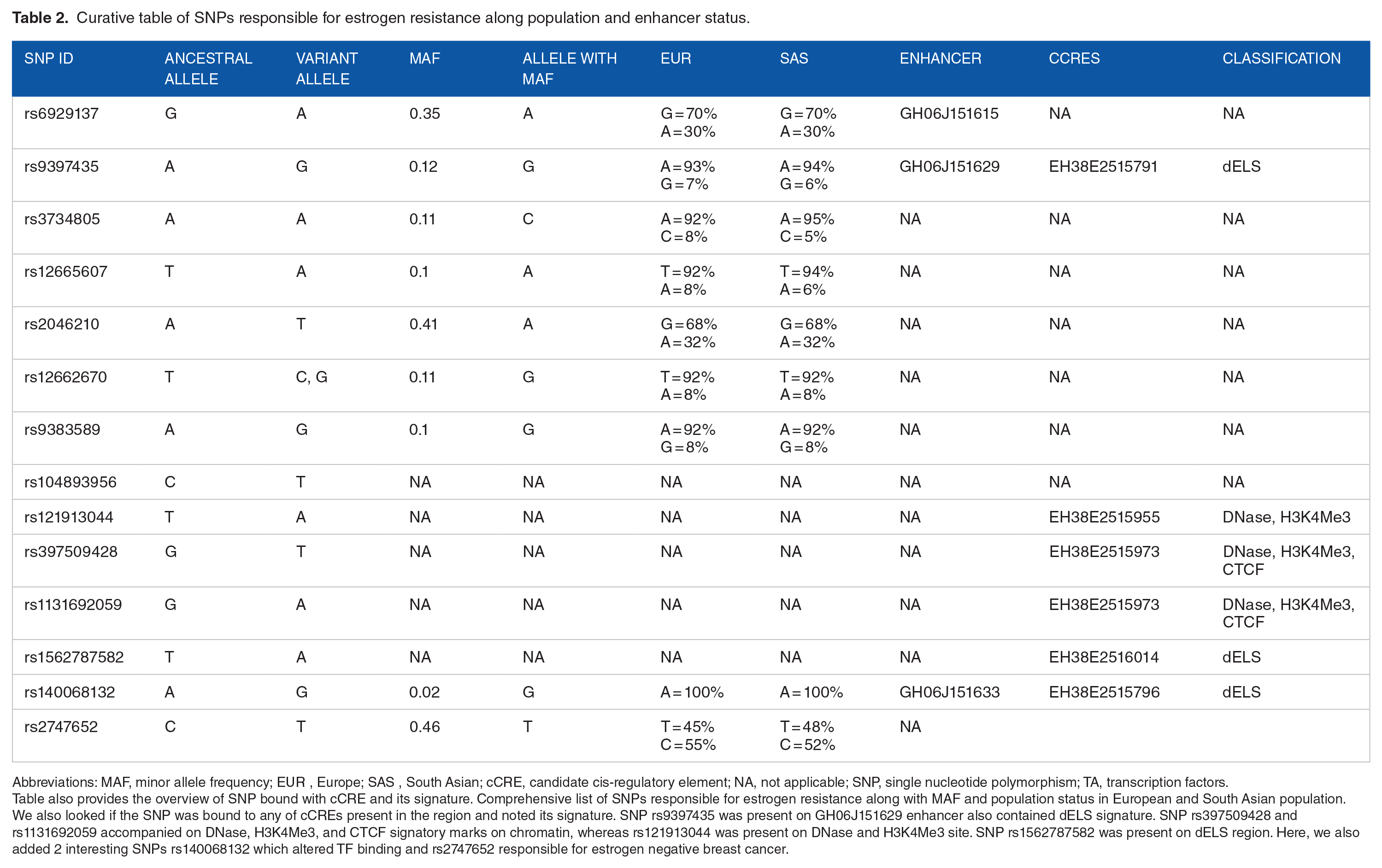
Abbreviations: MAF, minor allele frequency; EUR, Europe; SAS, South Asian; cCRE, candidate cis-regulatory element; NA, not applicable; SNP, single nucleotide polymorphism; TA, transcription factors.
Table also provides the overview of SNP bound with cCRE and its signature. Comprehensive list of SNPs responsible for estrogen resistance along with MAF and population status in European and South Asian population. We also looked if the SNP was bound to any of cCREs present in the region and noted its signature. SNP rs9397435 was present on GH06J151629 enhancer also contained dELS signature. SNP rs397509428 and rs1131692059 accompanied on DNase, H3K4Me3, and CTCF signatory marks on chromatin, whereas rs121913044 was present on DNase and H3K4Me3 site. SNP rs1562787582 was present on dELS region. Here, we also added 2 interesting SNPs rs140068132 which altered TF binding and rs2747652 responsible for estrogen negative breast cancer.
Candidate cis-regulatory elements on estrogen receptor 1 enhancer elements
The cCREs are sequences on non-coding DNA that regulate transcription of neighboring genes. We identified 156 potential cCREs on ESR1 at chr6:151616119-152130808 (hg38) (Supplementary Table 3). Majority (88/156) of cCREs were classified as distal enhancer-like sites (dELS). A total of 6 of 14 SNPs present on cCREs had clinical significance. The variants rs9397435, rs121913044, and rs397509428 were linked to estrogen resistance and rs121913043 produced mutant ER alpha protein.39,40 The rs121913044 was located on DNase and H3K4Me3 signatures, whereas rs397509428 resided on CTCF and H3K4Me3 sites (Table 2) (Supplementary Figure 2.)
Enhancer RNA on estrogen receptor 1 regulatory element
Enhancer RNAs are lncRNA transcribed from DNA sequence of regulatory regions. The eRIC database reported 7 eRNA regulating ESR1 from the TCGA BRCA cell-type expression database. The eRNA ENSR00000205343 was reported to be transcribed from the position, chr6: 151763273-151765000 containing GH06J151763 and cCREs with the mean expression of 3.14 units (Figure 4A). The ENSR00000205343, ENSR00000205344, and ENSR00000323738 were seen to be transcribed from cCREs containing dELS and CTCF bound signatory marks. All the 7 eRNA identified in BRCA cell type was known to act on ESR1 and its neighboring genes such as Required for Meiotic Nuclear Division 1 (RMND1) and ZBTB2 (Figure 4B).

cCRE on enhancer GH06J1151763. (A) Using UCSC, we identified 4 cCREs found on the enhancer GH06J151763. All the 4 cCREs had the distal enhancer-like sites and CTCF-binding signatures. (B) Seven eRNAs were identified using eRIC database. All the 7 eRNAs were found to act on ESR1, RMND1, and ZBTB2 genes. Interestingly, eRNA ENSR00000205343 was transcribed by GH06J151763 enhancer. We also looked if any cCREs were present on the eRNA transcribed region and found that except ENSR00000205366 eRNA all the other eRNAs contained cCREs in them. eRNA ENSR00000323747 showed the highest mean expression of 13.74, and ENSR00000205344 and ENSR00000323738 showed the lowest mean expression of 2.885 in BRCA cell lines retrieved from TCGA database which was available in eRIC database.
Discussion
The non-coding DNA vastly outnumber coding DNA supporting their role as designers of eukaryotic complexity. Enhancers are DNA regulatory elements which regulate transcription of one or more genes regardless of its orientation or relative distance from the target promoter. 41 They contribute significantly to cell growth, differentiation, and disease. Estrogen plays an important role in breast cancer development and endocrine therapy prevents the effect of estrogen on breast cancer cells. Endocrine therapy significantly reduced relapse rate by half in breast cancer patients; however, 1 of 3 patients still develop hormone resistance and relapse. 42 Hormone resistance in breast cancer is an acquired phenotype in which the cells stop responding to hormone treatment. The molecular mechanism driving this resistance in ER-positive breast cancer patients is poorly understood. Recent studies report global reprogramming of estrogen responsive regions in the DNA could contribute to this insensitivity. Here, we use the UCSC Genome Browser, ENCODE, and associated databases to review and report DNA regulatory elements on ESR1 genomic region to postulate the role of genomic regulatory marks contributing to hormone resistance.
The genetic regulatory elements that define endocrine response in ESR1-positive breast cancer patients has remained poorly understood. The spatial organization of the genome inside the cell nucleus is pivotal to cell function. 43 Non-coding genomic regions harbor specific nucleotide sequence favoring certain interaction and bindings in its 3-dimenisonal (3D) conformations that may facilitate certain repertoire of gene expression leading to development of resistance.
Using data mining approaches, we review and report 7 double elite regulatory elements spanning across ESR1 and CCDC170 genes at intergenic, promoter, exonic, and non-coding intronic regions. These enhancers formed clusters and enact as a super enhancer 44 (Supplementary Table 4). Furthermore, they are sites for lncRNA binding and facilitate gene expression. 45 The UCSC database tracks, ChromHMM, histone marks, TFs, SNPs, cCREs, and eRNA illustrate the genomic complexity of ESR1 regulation.
The 7 ESR1 regulatory sequences hosted histone marks and transcriptional factors which varied across cell type example some enhancers active in normal mammary epithelial cells but not in hepatoma and vice versa. Similarly, enhancers at intergenic and on introns of ESR1 contained active histone marks, whereas those on exon showed no activity.
There was no reported TF binding on 2/7 enhancers. Intergenic variants rs6929137 and rs9397435 are responsible for estrogen resistance. The rs140068132 variant found at TF binding site was present on GH06J151633 enhancer and is known to be associated with substantially lower risk of breast cancer, particularly ER-negative breast cancer.46,47 ESR1 enhancers contained cCREs at sites of transcription initiation in breast invasive carcinoma (BRCA) cell type. The variants rs121913044 and rs397509428 which were associated with estrogen resistance were located at CREs harboring DNase, histone, and CTCF marks. We speculate that the development of estrogen resistance could be due to loss/alteration of above chromatin signatures.
The human phenotype, normal or disease is a complex interaction of genes and regulatory elements in a certain genomic conformation. Most genes in the human genome are likely regulated by multiple enhancers. Transcriptional enhancers are required for the differential expression of the human ER in breast cancers. 48 Development of estrogen resistance could be contributed by features present within the ESR1 genomic landscape such as SNPs, CREs, and histone marks, which contribute to ESR1 regulation/expression. ERα is well-known TF involved in maintenance of female reproductive system, bone mass and plays an important role in protecting cardiovascular and central nervous systems. Genomic studies comparing normal, MCF-7, tamoxifen-resistant, and fulvestrant-resistant cells showed loss/altered CTCF and ERα-binding sites with development of estrogen resistance. 49
However, till date there are limited studies on DNA regulatory marks of ESR1 to understand its role in development of estrogen resistance. This study reviews the regulatory regions and its associated features speculating its potential role in ESR1 activity. Driver mutations directly impact the protein product; however, variations in the enhancers might affect the TF signature and gene expression profile. 50 We believe the ESR1 DNA regulatory signatures and mutations could contribute to the development of estrogen resistance during long-term hormone therapy.
Using our observations, we put forth 3 assumptions to better understand the development of estrogen resistance attributed by non-coding DNA regulatory features. In the first assumption, the mutation in enhancer results in loss of TF binding consequently led to reduced transcription. In the second assumption, the loss of TF binding site results in a modified mRNA production, and in the third assumption, the entire enhancer landscape is perturbed due to loss of histone modification resulting in modified/altered enhancer landscape leading to production of mutated mRNA, no mRNA, and/or loss of enhancer-promoter interaction. These assumptions mentioned could be the repercussion of long-term exposure to tamoxifen which altered cytosol metabolite meliu leading to change in chromatin conformation and enhancer landscape. At present, the study is limited only to in silico analysis and no in vitro studies have been conducted to validate these assumptions.
Conclusions
We review the potential role of non-coding DNA regulatory elements present on ESR1 in regulating estrogen resistance in breast cancer. This study can be used for development of better diagnostic test in identification of estrogen resistance. We postulate the role of intergenic enhancer and its features in contributing to estrogen resistance. In this study, we speculate that change in enhancer landscape and spatio-temporal gene expression results in the development of estrogen resistance over the course of hormone therapy.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322231167971 – Supplemental material for Bioinformatic Evaluation of Features on Cis-regulatory Elements at 6q25.1
Supplemental material, sj-docx-1-bbi-10.1177_11779322231167971 for Bioinformatic Evaluation of Features on Cis-regulatory Elements at 6q25.1 by N Sreekar and Smeeta Shrestha in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-2-bbi-10.1177_11779322231167971 – Supplemental material for Bioinformatic Evaluation of Features on Cis-regulatory Elements at 6q25.1
Supplemental material, sj-xlsx-2-bbi-10.1177_11779322231167971 for Bioinformatic Evaluation of Features on Cis-regulatory Elements at 6q25.1 by N Sreekar and Smeeta Shrestha in Bioinformatics and Biology Insights
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contribution
SN – Literature and database search; SS – designed study and write manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
