Abstract
In metazoans, microRNAs (miRNAs) are essential regulators of gene expression, affecting critical cellular processes from differentiation and proliferation, to homeostasis. During miRNA biogenesis, the miRNA strand that loads onto the RNA-induced Silencing Complex (RISC) can vary, leading to changes in gene targeting and modulation of biological pathways. To investigate the impact of these “arm switching” events on gene regulation, we analyzed a diverse range of tissues and developmental stages in zebrafish by comparing 5p and 3p arms accumulation dynamics between embryonic developmental stages, adult tissues, and sexes. We also compared variable arm usage patterns observed in zebrafish to other vertebrates including arm switching data from fish, birds, and mammals. Our comprehensive analysis revealed that variable arm usage events predominantly take place during embryonic development. It is also noteworthy that isomiR occurrence correlates to changes in arm selection evidencing an important role of microRNA distinct isoforms in reinforcing and modifying gene regulation by promoting dynamics switches on miRNA 5p and 3p arms accumulation. Our results shed new light on the emergence and coordination of gene expression regulation and pave the way for future investigations in this field.
Introduction
Metazoan gene expression is tightly regulated by a myriad of molecular regulators that determine cell fate. Among them, microRNAs (miRNAs) have emerged as key players in regulating a variety of biological processes, including embryo development (such as gastrulation, hematopoiesis, and tissue formation,1-3 as well as homeostasis, including responses to environmental influences 4 and diseases.5,6 MiRNAs are a large class of non-coding RNAs that regulate gene expression post-transcriptionally, primarily by binding to the 3′ untranslated region (3′UTR) of target mRNAs,7,8 although functional repression through interaction with the coding sequence (CDS) and 5′UTR 9 regions of the mRNA has also been reported. Thanks to their particular binding affinity to targets, miRNAs can promote an extensive spectrum of interactions, enabling a single miRNA to regulate multiple genes, while a single gene can be regulated by several miRNAs.10,11
During the biogenesis of miRNAs, 1 strand of the miRNA duplex is preferentially loaded onto the RNA-induced Silencing Complex (RISC), thereby targeting mRNA sites with reverse complementarity, while the other strand undergoes predominant degradation.12,13 This preferentially loaded strand, known as the canonical or dominant strand, is generally consistent across tissues and developmental stages and for this reason it was considered the functional miRNA molecule. However, examination of microRNA transcriptomic data from various studies investigating distinct biological traits and states9-11 revealed that a minor fraction of mature miRNAs can still originate from the opposite arm. These sequences were previously labeled as passenger or miR* sequences. Under certain circumstances, arm switching events – the process in which the RISC loads the passenger strand instead of the canonical strand – may occur, 14 leading to fluctuations in the distribution of canonical mature miRNAs and passenger transcripts. Interestingly, for a large fraction of miRNA precursor hairpins, both arms originate substantial amounts of functional transcripts, currently designated as mature 5p and 3p miRNAs.
This process can significantly impact the miRNA’s targeted genes and their biological functions, as the miRNA-target pairing is crucial for gene regulation. 15 Although the mechanisms driving arm switching are not fully understood, the thermodynamic properties and 5′ nucleotide identity of the miRNA duplex have been shown to play a key role. 16
Arm switching events have been reported within tissues, organs, or developmental stages of several living vertebrates, including Homo sapiens, 17 Mus musculus, 18 Oryctolagus cuniculus, 19 Gallus gallus, 20 and Oreochromis niloticus. 21 Switches in arm usage were also depicted between invertebrate species, such as Drosophila melanogaster and Tribolium castaneum. 14 These events have also been linked to diseases and imbalances in cell homeostasis, such as various types of cancer.22-25
MiRNAs can also exhibit a diverse range of isoforms, known as “isomiRs,” that arise from a variety of mechanisms including single nucleotide substitutions, indels, 3′ end non-templated additions, and 5′ or 3′ cleavage shifts. 26 These variants result from alternative Dicer and Drosha cleavages, leading to size variations of canonical miRNAs and potentially changing their thermodynamic properties and 5′ nucleotide identity. 26 IsomiRs are not randomly distributed and be functionally active as partners of canonical miRNAs in several human tissues.27,28 Importantly, 5′ and 3′ end isomiRs can regulate distinct targets and are expressed in a tissue- and developmental-specific manner, indicating a coordinated regulation of miRNA biogenesis and function. 29
In this study, we conducted a comprehensive genome-wide analysis of microRNA arm switching in zebrafish to understand the impact of strand selection on gene regulation. Using bioinformatics, we identified distinct patterns that associate the incidence of arm switching with different developmental stages in vertebrates. Furthermore, our findings suggest an association between arm switching events and isomiR profiles, shedding light on the potential impact of isomiRs on the regulation of gene expression
Material and Methods
RNAseq data
All experiments were approved by the Ethics Committee on Animal Use (CEUA protocol no. 1008 - 2017). To identify the arm switching profiles, small RNA-seq data were obtained from several adult tissues and developmental stages of zebrafish. Adult tissue samples were collected from SRP041544 study, 30 which performed an RNA sequencing on an Illumina HiSeq 2000 machine using TruSeq v3 cBot and SBS kits. The data provided (n = 3) from the ovary (with 8.8; 14.3 and 25.9 million reads per replicate), testis (17.4; 20.2 and 14.2 million reads per replicate), eye (9.4; 5.7 and 2.9 million reads per replicate), heart (38.1; 34.8 million reads per replicate), male brain (5.1; 6.5; 10.5 million reads per replicate), female brain (2.3; 3.3; 12.6 million reads per replicate), male gut (16.2; 8.9; 3.1 million reads per replicate), female gut (13.9; 8.3; 7.5 million reads per replicate), male liver (18.4; 9.4; 4.6 million reads per replicate) and female liver (5.1; 8.2; 1.6 million reads per replicate). Embryonic samples were obtained from SRP028895, 31 which was performed at the Genome Technology Core of Vanderbilt University using the Illumina sequencing platform, and contained small RNA-seq data (n = 2) from different developmental stages: 256 cells (2.5 hours post-fertilization—hpf) with 3 196 331 reads, sphere (4-hpf) with 13 680 619 reads, shield (6-hpf) with 4 928 631 reads, and 24-hpf with 8 158 340 reads.
Sample treatment, miRNA identification, and 5p/3p arms characterization
Firstly, we run a quality control analysis to ensure data quality using the FastQC tool (v0.11.5 32 ). After, adaptor sequences were trimmed using the cutadapt tool (v1.14 33 ), and the reads were converted from the fastq to the fasta format using FASTX-Toolkit (v2.8.1). Reads were then counted, collapsed to unique reads, filtered by length (16-23 nts) and complexity (⩾3 distinct nucleotides), and reads matching other non-coding RNAs such as rRNAs and tRNAs were removed, using the filter module of UEA small RNA Workbench (v3.2, release 19 34 ).
Treated sequences were mapped to the zebrafish genome vGRCZ10 using Mapper script and miRNAs were identified with miRDeep (v2.0.0.7 35 ). To perform miRNA identification, pre- and mature miRNAs from zebrafish, and mature miRNAs from other species were obtained from miRBase (v22 36 ) and MirGeneDB (v2.0 and v2.137-39). These sequences were used as a reference, allowing a maximum of 1 mismatch outside the seed region.
Once miRNAs were identified, 5p and 3p arms from each miRNA were characterized. In this approach, pre-miRNAs were divided into 2 subsections with 30 nucleotides overlap. Each identified read was then mapped to one of these 2 sections using the Shortstack tool (v1.1.2 40 ), and accordingly classified as the 5p or 3p arm. Reads mapped only to the overlap sections were filtered and removed from the analysis.
Data normalization and arm switching events identification
The expression of the identified 5p and 3p reads from all samples were normalized with the TMM method (trimmed mean of M values 41 ) using the edgeR package (R Bioconductor). After normalization, 5p and 3p expression ratios of each miRNA were obtained and arm switching events were identified.
For the arm switching identification, we only considered events in which we observed a twofold difference between 5p and 3p arms in at least 2 tissues, changing the predominant arm between them, and when we had at least 10 mapped reads on the raw read counts on both tissues. 21
Target prediction and functional enrichment analysis
The target prediction analysis was performed only on miRNAs with identified arm switching events. For this analysis, we used the union method to combine the results obtained from three different tools: miRanda ( 10 www.microrna.org), RNA22 (v2, 42 https://cm.jefferson.edu/rna22/), and TargetsScan ( 43 www.targetscan.org, ensuring the best performance regarding specificity and sensibility of the analysis. 44 The results were further filtered by the presence of the predicted targets in the tissue (or embryonic developmental stage) of interest. To do this, RNAseq data from brain and 24hpf24-hpf samples were obtained and the FPKM (Fragments Per Kilobase Million) of target mRNAs were calculated. We considered only predicted targets of FPKM ⩾ 5,45,46 and mRNAs with lower values were discarded.
Functional enrichment analysis of the predicted targets was performed using the g: Profiler tool 47 using the multi-query function and the significance threshold method g:SCS threshold (0.05) searching for Biological Process (BP), Cellular Context (CC), and Molecular Function (MF) (Gene Ontology 48 ), and also Biological Pathways (KEGG 49 ).
IsomiRs identification
Bam files obtained from Shortstack alignment were loaded in Tablet software 50 for graphical visualization of the reads and identification of canonical reads and isomiR patterns. Pre-miRNA subsections generated during miRNA 5p and 3p miRNA characterization were used as reference.
Results
Overview of RNAseq data
From the analysis of RNAseq data from 2 previously published reports examining miRNA expression from both adult and developmental stages from zebrafish,30,31 we obtained about 508 million raw reads that had passed quality control analyses after removing sequencing adaptors. From the 498 million remaining reads, about 163 million passed through all filtering steps and were aligned against the zebrafish genome for miRNA identification.
Read distribution analysis revealed that ~70% had a size range of 21 to 23 nucleotides (nts, Table S1). The prevalent size for most samples was 22 nts, except for the male and female brains, in which the prevalent size was 23 nts. Gonadal samples also had an additional peak at 25 nt, typical of piwi-RNAs (piRNAs). PiRNAs are abundant in gonadal tissues, acting to silence transposons and retrotransposons, 51 thereby contributing to the protection, maintenance, integrity, and stability of the germ line genome. 52
miRNA identification and detection of arm switching events
From the 2 expression data sets, we identified 1288 mature 5p- and 3p-miRNAs that originated from 644 precursor sequences. A total of 578 were derived from known pre-miRNA sequences, and 66 originated from putative novel pre-miRNAs (Tables S2 and S3).
Through a careful examination of the miRNA expression profiles, we identified arm switching events in 14 pre-miRNAs—13 from known pre-miRNAs (ie, dre-mir-27b-1, dre-mir-27b-2, dre-mir-92a-1, dre-mir-92a-2, dre-mir-135b, dre-mir-137-1, dre-mir-137-2, dre-mir-153a-1, dre-mir-153a-2, dre-mir-153b, dre-mir-222a, dre-mir-2188, dre-mir-31) and a single arm switching in the novel pre-miRNA dre-mir-n001 (Figure 1). The 5p/3p ratio values for each identified arm switching event can be found in Table S4. The arm switching events identified could be further observed in 4 major comparisons detailed below.

MA plot of zebrafish arm switching events. The Y-axis represents the 5p/3p arms ratio. Positive values represent 5p arm predominance while negative values represent 3p arm predominance. Red lines represent the twofold threshold for a bona fide arm switching. The X-axis represents the expression intensity of the miRNAs. Triangles represent developmental stages and circles, adult tissue.
The first comparison involved events between developmental stages and adult tissues, contemplating all events identified (Figure 1). Remarkably, half of the differential arm prevalence observed occurred between 24-hpf embryos and the adult brain.
The second comparison comprised miRNA arm switching events between distinct developmental stages, where changes in arm prevalence were detected for dre-mir-27b-2, dre-mir-135b, dre-mir-92a-1, dre-mir-153a-1, dre-mir-153a-2, dre-mir-222a, and dre-mir-2188 (Figure 2). Notably, except for dre-miR-153a-1/2, which exhibited no expression at 2.5-hpf (a normal feature considering the dynamic recruitment of miRNAs by cells), the miRNA 5p/3p ratio at the same period corresponded to that observed in adult samples (Figure 1), with major changes in arm usage occurring between 2.5-hpf and the immediately subsequent 4-hpf sphere stage. After this very early developmental period, no consistent arm usage patterns were observed for the distinct miRNAs.

5p/3p ratio of arm switching events during the 4 developmental stages investigated. Red dashed lines represent the twofold threshold for a bona fide arm switching.
The third comparison involved sex-biased variant arm usage, identified in dre-mir-31, which exhibited an inverted 5p/3p ratio between the testis and ovary (Figure 1). Interestingly, this was the unique miRNA arm switching event between 2 adult tissues.
Finally, the fourth comparison involved arm switching events associated with 2 miRNA paralogs (Figure 3) with differential arm usage observed between dre-miR-92a-1 and dre-miR-92a-2 at 24-hpf, and between dre-mir-153a-1-3p and dre-mir-153b-5p in the brain sample (Figure 3).
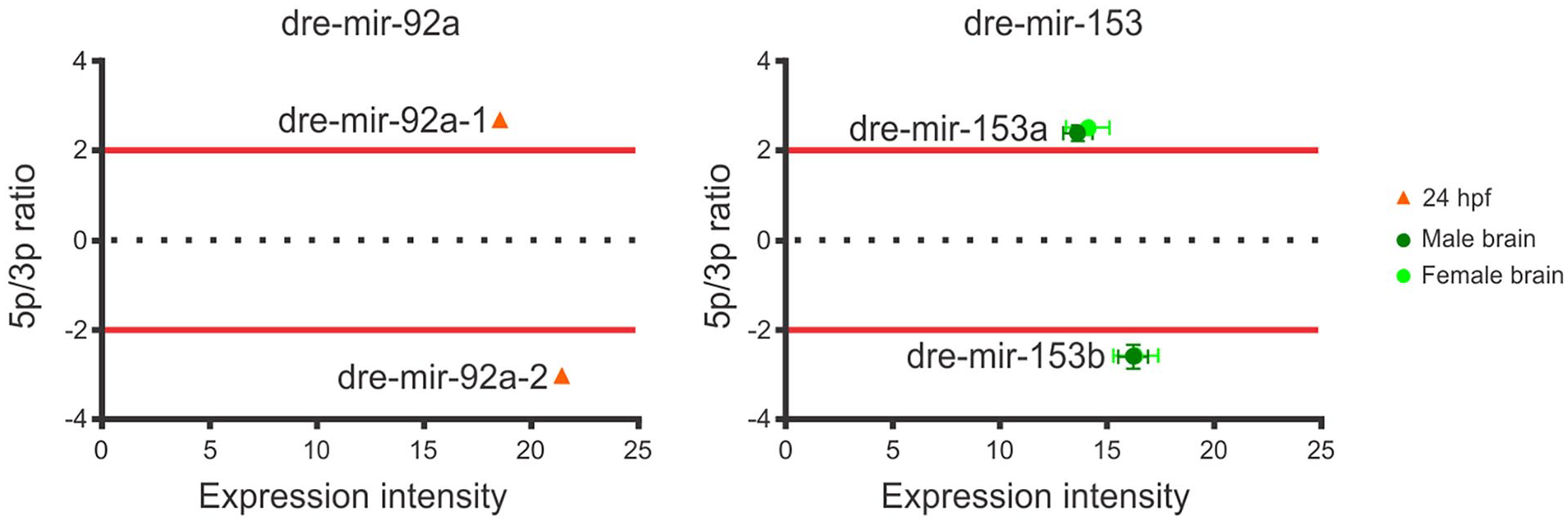
Arm-switching events between paralog copies of miRNAs dre-mir-92a and dre-mir-153. Red lines represent the twofold threshold for a bona fide arm switching.
Target prediction and functional enrichment data
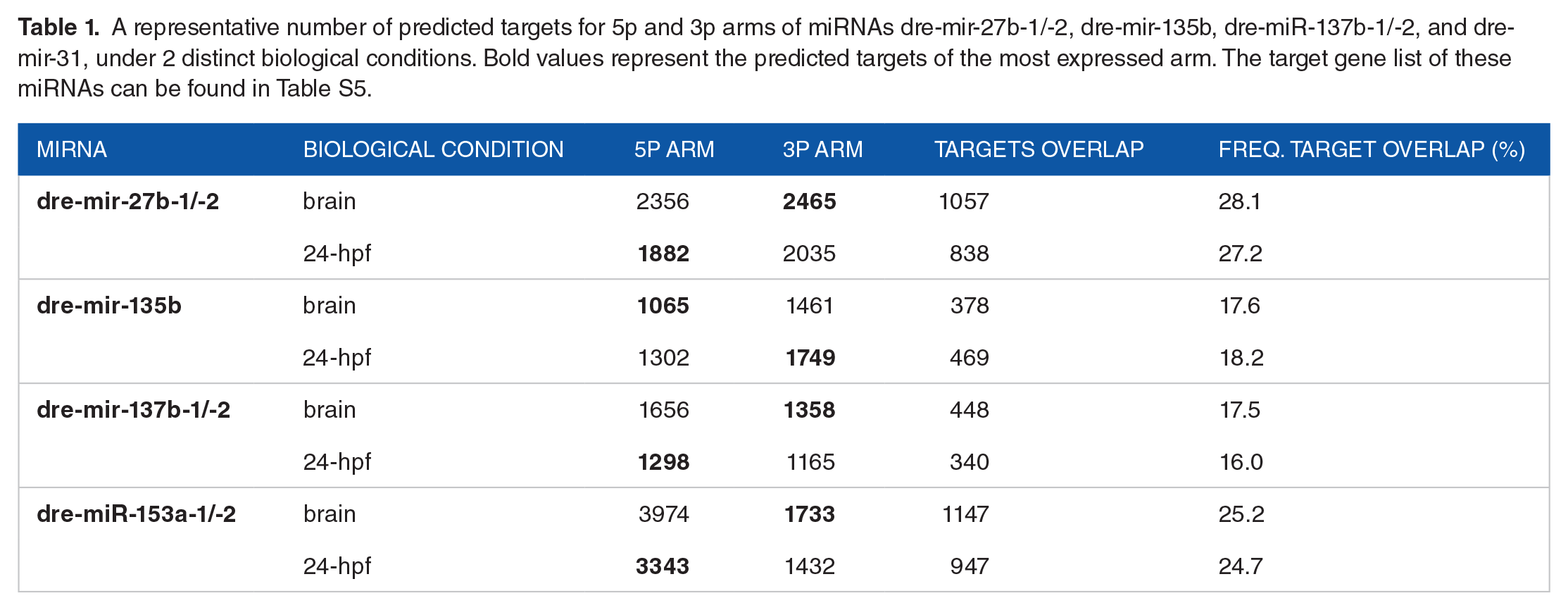
Because 50% of zebrafish arm switching events were identified between 24-hpf embryos and adult brains, we exclusively performed target prediction, functional enrichment, and isomiR analysis in these tissues. Selecting 2 tissue types for comparison enabled us to properly identify any prospective correlations between enriched targets and isomiR profiles with differential arm usage with higher accuracy. Target prediction analysis returned a total of 16 593 and 13 681 potential targets of dre-mir-27b-1/-2, dre-mir-135b, dre-mir-137-1/-2, and dre-mir-153a-1/-2 expressed in the brain and 24-hpf tissues respectively. Consistent with previous studies,14,15 our analysis revealed a relatively low degree of target overlap between the 5p and 3p arms of miRNAs, which exhibited between 71,9% and 84,0% of non-overlapping targets. Additionally, we did not find any correlation between the prevalence of a specific arm for a given tissue and the number of its predicted targets (Table 1).
A representative number of predicted targets for 5p and 3p arms of miRNAs dre-mir-27b-1/-2, dre-mir-135b, dre-miR-137b-1/-2, and dre-mir-31, under 2 distinct biological conditions. Bold values represent the predicted targets of the most expressed arm. The target gene list of these miRNAs can be found in Table S5.
Our findings also revealed an interesting aspect of miRNA regulation, not only do the 5p and 3p arms regulate distinct sets of targets, but they also differ in the functional categories of the targets they regulate (Figure 4 and Table S6). Notably, we observed the most significant discrepancies in Molecular Functions and Biological Pathways, while for Biological Processes and Cellular Components, although there are some processes controlled exclusively by 1 arm, there are still common terms observed, implying that the choice of miRNA arm utilization can have a substantial impact on the downstream molecular functions and pathways that are targeted for regulation.

Graphic visualization of dre-miR-27b functional enrichment on 24-hpf embryo. Red dots = molecular function. Yellow dots = biological process. Green dots = cellular components. Pink dots = biological pathways. The point position on the Y-axis indicates P-value. The point position on the X-axis represents specific terms. Dot diameter represents the number of genes associated with the term. Light circles of each color represent terms found without significant P-value. Detailed results from functional enrichment analysis can be found in Table S6.
For dre-miR-27b expression at 24-hpf, only 1 Molecular Function (transcription coregulator activity; GO: 0003712) and 1 KEGG pathway term (Notch signaling pathway; KEGG: 04330) was enriched for both the 5p and 3p arms. However, 11 of the 34 enriched cellular context terms and 22 of the 119 biological process terms could be observed on both arms. Notably, only the dre-miR-27b-5p arm, which is the most expressed at 24-hpf, displayed exclusively enriched terms for Molecular Function, Cellular Context, and Biological Process, while 2 KEGG terms (ie, Insulin signaling pathway; KEGG: 04910, and Terpenoid backbone biosynthesis; KEGG: 00900) were uniquely enriched on the 3p arm (Figure 4 and Table S6).
When examining the brain tissue, where the dre-miR-27b-3p is the prevalent arm, we observed 4 out of the 21 Molecular Functions and 1 out of the 4 KEGG terms (Endocytosis; KEGG: 04144) enriched on both arms. For cellular context and biological process, 17 out of 46 and 37 out of 115 enriched terms, respectively, were found for both arms. Notably, unlike the 24-hpf embryo sample, there were a greater number of terms enriched exclusively to either 5p or 3p in the brain tissue (Table S6). Additional results for functional enrichment analysis of other miRNAs expressed in the 24-hpf embryo and brain tissues can be found in Table S6.
Correlation between arm switching events and isomiR expression
IsomiRs are miRNA variants that differ from the predominant miRNA due to modifications including single nucleotide substitutions, insertions, deletions, 3′ end non-templated additions, and/or 5′ and/or 3′ cleavage shifts during biogenesis. 21 These variations in the miRNA sequence can affect the stability of the miRNA-miRNA* duplex and alter the initial nucleotide identity of each arm, which can impact arm selection. Therefore, we analyzed the isomiR patterns present in the identified arm switching events to evaluate their impact on arm selection. Our analysis identified several modifications in the expressed transcripts’ sequences, both in template and non-template reads, characterizing typical isomiRs (Figure 5). The majority of isomiRs probably originated from distinct Dicer and Drosha cuts during miRNA biogenesis, whereas a small fraction of them (~0.72%) corresponded to non-template isomiRs, such as single nucleotide modifications.

IsomiR patterns of dre-miR-153a-2 expressed in the brain. Several patterns of isomiRs were identified on both arms, such as reads with insertions and deletions on both 5′ and 3′ ends, 3′non-templated insertions, and single-nucleotide mutations.
It is noteworthy that we found identical isomiRs in both embryos at 24-hpf and adult brain tissues, whereas tissue-specific isomiRs were rare. However, such shared identical isomiRs exhibited significant differences in their relative expression levels between the 2 tissues, strong enough to change the representative isoform of at least 1 of the mature arms for 5 out of 7 miRNAs analyzed (Table 2). These findings suggest that arm switching is accompanied by a shift in the expression profile of mature miRNAs, including changes in the representative isoform. For dre-miR-135b, the 5p arm is less expressed than the 3p arm at 24-hpf, and its representative isoform is a 23-nucleotide-long. However, in the brain, an arm switching event takes place, leading to an increased expression of the 5p arm relative to its 3p counterpart. This change is accompanied by a shift in the representative isoform of dre-miR-135b-5p, resulting in a shorter variant with a missing adenine at the 3′ region (Table 2).
Representative isoform from 5p and 3p miRNA armsa,b at 24-hpf embryos and adult brain.

Nucleotides shadowed in orange color were exclusively found on the representative read of a single biological sample category.
Reads in bold refer to the highest expressed miRNA arm.
Our data also revealed tissue-specific isomiR expression patterns. Specifically, the representative isoforms identified at 24-hpf were generally longer than those in the adult brain. We also observed a higher frequency of non-template isomiRs in the brain (~1.29%) than at 24-hpf (0.29%). Interestingly, we observed a unique pattern in dre-miR-135b, where the predominant read of dre-miR-135b-3p in the 24-hpf sample had an additional adenine in its 5′ portion compared to its counterpart in the brain sample. IsomiRs with variations in the 5′ region are rare and can lead to a phenomenon known as “seed shifting,” which alters the sequence of the seed region responsible for target recognition. 53
Discussion
Arm switching is a conserved mechanism, but events are rare and mainly species-specific
MicroRNA arm switching has been identified in numerous species, including mice,18,54 chickens, 20 nematodes, 55 insects, 14 and fish. 21 However, each study has revealed a different set of miRNAs exhibiting changes in arm prevalence, and only a limited number of events have been identified so far. A possible caveat for direct comparison of data from these studies is that most of them have employed distinct next-generation technologies for assessing miRNA reads abundance (eg, 454 pyrosequencing, ABI SOLiD, or Illumina “Solexa” sequencing). Similarly, biological variability across samples can confound the detection of miRNA arm switching events in small RNA-seq data. Nonetheless, even experiments using the same approach (ie, Illumina technology) and sample source (ie, skeletal muscle) have also obtained distinct findings. For instance, a large-scale analysis of miRNA expression in Nile tilapia identified arm switching in only 9 miRNAs out of the 368 miRNA loci analyzed. 21 Similarly, in the current study, arm switching was observed in only 14 miRNA loci out of 674 on zebrafish. Interestingly, none of the miRNAs having changes in arm usage were common to zebrafish and Nile tilapia, despite both studies being experimentally compatible in strategy and sequencing technology.
A high-throughput study of mouse miRNAs also reported 21 arm switching events (from 506 miRNAs identified) occurring across several developmental stages (ES, e7.5, e9.5, e12.5, and newborn), testis, ovary, and brain of adult animals. 54 Again, none of the identified arm switching events were shared in Nile tilapia or zebrafish. In addition to these large-scale analyses, other experiments have also described arm switching in specific contexts. For example, miR-100 and miR-125 have the 5p arm as the most expressed in mammals, but the 3p arm has dominant expression in some tissues. 56 Four miRNAs (miR-135a-2, miR-30e, miR-219, and miR-30c) were found to undergo arm switching during chicken development, 20 while miR-30e was observed to switch arms between stomach and spleen in mice. 18 Arm switching events were also identified in nematodes (Caenorhabditis elegans, C. briggsae, C. remanei, and Pristionchus pacificus 55 ) and insects (Drosophila melanogaster and Tribolium castaneum. 14 Interestingly, reports have shown that while the overall predominant arm for vertebrates is the 5p, invertebrates usually have the 3p as the prevalent arm, 54 suggesting that a major change in arm prevalence may have occurred at some point in metazoan evolution.
It is notorious that despite an increasing number of studies searching for and identifying arm switching events, the number of events identified comprises only a very small fraction of the total miRNA repertoire. These data reveal that although arm switching is a conserved mechanism among metazoans, they have rare and occasional occurrences on a species-based level. Additionally, results show that the cases observed for 1 species usually cannot be identified in others (Figure 6), demonstrating that arm switching is mainly species-specific and may be related to particularities of the cells that the miRNAs are expressed in, which widely vary between 1 species to another. An interesting exception is the miR-30 family, where arm switching events of mir-30c were reported for chicken and Nile tilapia, while miR-30e-1 events were identified in chicken, Nile tilapia, and mice (Figure 6).

Distribution of arm switching events among species. Green, red, blue and yellow segments represent arm switching events discovered on the mouse, zebrafish, Nile tilapia, and chicken, respectively. Black squares represent arm switching events identified only on 1 species. Light gray square (miR-30c) represents an arm switching event identified on 2 species (chicken and Nile tilapia). A dark gray square (miR-30e-1) represents an arm switching event identified on 3 species (mouse, chicken, and Nile tilapia).
Arm switching is mainly related to organism development rather than tissue specification
Our analysis revealed arm switching events for 14 miRNAs across 4 different biological comparisons. Firstly, we identified arm switching between embryos and adults, indicating a potential role in the transition from early development to adulthood. Secondly, we found arm switching events occurring between distinct early developmental stages, suggesting a possible involvement in developmental progression. Thirdly, we observed arm switching between sexes, specifically between ovary and testis, highlighting a potential role in sexual differentiation. Finally, we identified arm switching events between miRNA paralogs, which may be indicative of functional divergence during evolution. Notably, our findings are consistent with previous reports of arm switching in Nile tilapia 21 and mouse. 56 Although the specific miRNAs involved were different, these studies also identified arm switching events associated with developmental processes. Altogether, these data highlight the importance of arm switching in regulating gene expression during development and emphasize the possibilities for further research on this topic.
In addition to identifying arm switching between embryos and adults, our analysis also detected 7 events when comparing early developmental stages (Figure 2). These developmental stages originally sequenced by 31 and revisited by us, represent critical periods during embryonic development of vertebrates and exhibit notable variations in gene expression. At the 256-cell stage, most RNAs are of maternal origin and are eventually replaced by endogenously transcribed RNAs during the subsequent sphere stage. During the shield stage, the formation of the 3 germ layers occurs, and by 24-hpf, most organs are already formed. 57
Some of the investigated miRNAs (ie, miR-92, miR-2188, miR-153, miR-137 and a possible novel miRNA here named frer-miR-n001) showed a 5p/3p ratio at 2.5-hpf of the miRNAs similar with the ratio observed in adult tissues. This data may be explained by the fact that, until the 256-cell stage, all mature miRNAs present in the embryo are of maternal origin, 31 and therefore, the 5p/3p ratio at this stage is expected to mirror that observed in the ovary. An interesting exception is dre-mir-31, with the 5p/3p ratio at 2.5-hpf being opposite to the 1 found in the ovary (Figure 1). We hypothesize that distinct regulatory elements present between the ovary and 2.5-hpf could be promoting this phenomenon (eg, differential decay rates promoted by target-directed miRNA degradation—TDMD 25 ). However, further research is needed to elucidate the mechanisms underlying this exception. At the 4-hpf (sphere) stage, the embryonic machinery begins to produce endogenous miRNAs, while maternal RNAs start degradation.31,58 Not surprisingly, we observed a shift in the prevalence of 5p/3p arm ratios between the 2.5-hpf and 4-hpf periods (Figure 3), indicating a possible correlation of these arm switching events with the onset of endogenous embryonic transcription.
For dre-miR-27b paralogs, there was a very similar expression profile during embryonic stages, with a higher prevalence of the 5p arm compared to the 3p arm at 2.5-hpf. Then arm expression gradually lowered and became near equivalent at 4-hpf, ceasing expression at 6-hpf and reappearing at 24-hpf with 5p arm again dominant. For dre-miR-2188 and dre-miR-135b, after the arm switching event between 2.5 and 4-hpf, the 5p/3p ratio stabilized with the 3p arm being prevalent in subsequent periods. Regarding dre-miR-222a, the 5p arm was dominant at 2.5-hpf, but an arm switching event made the 3p arm the main functional transcript at 4-hpf. From 6-hpf onward, both arms were equivalently expressed. The complete arm switching event for dre-miR-92a-1 also occurred over a longer period. At 2.5-hpf, there was a higher prevalence of the 3p arm. At 4-hpf, both arms were expressed relatively equally, and finally, at 6-hpf, the 5p arm became prevalent.
An interesting observation was the detection of arm switching in dre-mir-31 between zebrafish ovary and testis (Figure 1). Previous studies have shown that miR-31 plays a crucial role in embryonic gonadal differentiation in chickens. 59 During the onset of sexual dimorphism in chickens, mir-31 is predominantly expressed in males, whereas later, after differentiation, there is a balance in the overall expression of this miRNA between the sexes. 59 Intriguingly, high levels of expression of dre-miR-31 were detected only during the early stages of development (2.5-hpf and 4-hpf) and in the adult testis and ovary tissues. At stages 2.5-hpf, 4-hpf, and testis, there was a prevalence of the 3p arm, while in the ovary, there was a higher expression of the 5p arm (Figure 1). Although zebrafish sexual differentiation is a polygenic event, sexual reversal analyses and DNA site mapping experiments suggest the presence of a ZZ/ZW system, similar to that found in birds. 59 Computational analyses of target prediction indicate that, in birds, mir-31 regulates components of the TGF-β signaling pathway, which plays a fundamental role in gonadal development.60,61 Therefore, it is plausible that dre-miR-31 may act similarly during the development of zebrafish reproductive structures.
The last biological comparison identified in our analysis examined arm switching events between miRNA paralogs. We observed a variation in the arm selection between dre-mir-92a-1 and dre-mir-92a-2 at 24-hpf, where the most prevalent arm was, respectively, 5p and 3p for each paralog. Additionally, for dre-mir-153, there was a significant switch in the prevalence of the 5p and 3p arms between dre-mir-153a and dre-mir-153b in the brain. In dre-mir-153a-1, the 3p arm was predominant, whereas in dre-mir-153b, the 5p arm was the primary transcript (Figure 3). The regulatory role of miRNAs is highly conserved across different organisms, but any imbalance can disrupt homeostasis. The emergence of parallel copies of a miRNA can alleviate the selective pressure to maintain the expression of a single copy, which could facilitate arm switching events. 14 proposed 2 types of arm 14 switching events following gene duplication. The first type occurs when both arms are expressed, and the appearance of a parallel copy can lead to sub-functionalization, where each copy specializes in 1 arm, resulting in increased expression of opposite arms in each copy. The second type occurs when 1 arm is predominantly expressed. In this case, gene duplication followed by an arm switching event can lead to the neofunctionalization of the new copy. In both conditions, the selective pressures may result in different miRNA genes, with low sequence identity, despite the common ancestry. 62 In fact, gene duplication events resulting in subfunctionalization and neofunctionalization are major sources for the emergence of novel miRNAs. 63
Target repertoire and function changes according to the prevalent arm strand
The 5p and 3p arms of miRNAs exhibit differences in their sequences, particularly in the seed region, which is the primary site for miRNA binding to its mRNA target. 64 Thus, we hypothesized that the 2 arms could interact with their target genes in 2 ways: (i) each arm would regulate a distinct set of genes, or (ii) the arms would interact with the same genes, but at different binding sites. Our analysis revealed a moderate overlap between the specific targets predicted for each arm (Table 1), suggesting that the 5p and 3p arms of the same miRNA are predominantly associated with the regulation of distinct biological functions. These findings support previous reports that also observed limited overlap in the target gene lists for their analyzed miRNAs.14,15
In Drosophila (Drosophila melanogaster), the miR-100/10 family predicted targets were fewer for the canonical than for passenger arm, 14 leading to the authors hypothesis that there might be selective pressure against the presence of matching sites for the most expressed arm. However, our data shows that this is not true in all situations. The dre-miR-27b-3p (24-hpf), dre-miR-135b-3p (brain), dre-miR-137b-5p (brain), and dre-miR-153a-5p (brain) arms have more predicted targets than their counterparts, although they are the most expressed in these tissues. Conversely, the dre-miR-27b-5p (24-hpf), dre-miR-135b-5p (brain), dre-miR-137b-3p (brain), and dre-miR-153a-3p (brain) arms are canonical and have the least number of targets (Table 1). The discrepancies observed in the outcomes of the 2 studies may be attributed to the difference in target prediction methodologies. 14 considered the global prediction of these miRNAs, without taking into account their tissue-specific expression, while our study compared the expression of miRNA arms in different tissues and revealed that the proportion of 5p and 3p arms can fluctuate depending on the tissue type (Table 1). For instance, in the brain, the canonical arm of some miRNAs—such as dre-miR-135b, dre-miR-137b, and dre-miR-153a—have fewer targets than the passenger arm, whereas in 24-hpf embryos, the canonical arm of the same miRNAs has a greater number of targets than the passenger arm. These results suggest that the miRNA target repertoire is influenced by the cellular context, and therefore, such selective pressure against canonical arm interaction sites may not apply in all situations.
Our analysis revealed that the biological roles of each arm’s targets also differ from each other. While some Biological Process terms show similarities, significant divergences were observed in Molecular Functions and Biological Pathways enriched (Figure 4). These findings demonstrate that the 5p and 3p arms have the potential to exert a prevalent regulatory separation, with distinct arms regulating distinct Molecular Functions and Biological Pathways even when they are inserted in the same Biological Process. It is worth noting that the identification of highly enriched terms in both arms, such as “developmental process” (GO: 0032502) and “anatomical structure development” (GO: 0048856), observed in the miRNA enrichment data of dre-miR-27b at 24-hpf (Figure 4), may stem from a potential recurring methodological bias in the functional enrichment technique. The Gene Ontology consortium assigns multiple terms to each gene, with each term having 1 or more associated parental terms. As a result, the enrichment of different specific terms can lead to the enrichment of shared parental terms in both arms. Another noteworthy observation is that, despite the divergences in Molecular Functions and Biological Processes of the targets, there was a higher degree of similarity in the Cellular Components where the targets of both arms act (Figure 4). This finding is by the biogenesis process of miRNAs, where both arms originate from the same pre-miRNA and reinforces the hypothesis that arm switching events could be a result of alterations that took place during miRNA biogenesis.
IsomiR occurrence may play an important role in arm switching formation
The observation that changes in the 5p/3p ratio across different tissues are often associated with alterations in the representative isoform of at least 1 miRNA arm implies that isomiRs could play a role in the formation of arm switching. Modifying the signature of the representative isoform may potentially affect both the relative stability of the ends of the duplex and the identity of the initial nucleotide of each arm, which are known to be critical features in arm selection by RISC, as noted by Kim et al. 16 In fact, several studies have established that changes occurring at distinct points of miRNA biogenesis may interfere with duplex stability and, ultimately, promote variations in arm usage. Alterations as early as DROSHA alternative processing on pri-miRNAs are sufficient to produce mature miRNAs with variable end instabilities, 65 while modifications on the basal and apical junctions of pri-miRNAs can promote such alternative DROSHA cleavage sites, 66 thereby influencing arm selection.
In humans, it was demonstrated that an arm switching event in miR-324 was regulated by alternative processing during its biogenesis. 16 Uridylation of the pre-miRNA by the terminal uridylyl transferases TUT4 and TUT7 causes a shift in the position of DICER on the pre-miRNA, resulting in changes in the cleavage sites. 16 This alternative processing produces non-template isomiRs, resulting in the selection of the 3p arm instead of its 5p counterpart in the miRNA-miRNA* duplexes. In fish, 67 found size variants of 9 miRNAs that had mature arms differentially expressed along several developmental stages of the Atlantic halibut. These authors suggested that those isomiRs could provide diversification of targets in miRNA transcriptional regulation during development but did not discuss arm switching per se. On the other hand, isomiRs discovered in the Atlantic salmon showed a great overlap of targets with the canonical forms of the miRNAs, indicating that miRNAs isoforms may not have such an impact on target diversity, except when the allelic variation comprised the seed region of the miRNA gene. 68 In fact, seed shifts observed in such isomiRs are a potential factor interfering in variable read count accumulation between 5p and 3p mature arms in fish that deserves to be deeply investigated.
Together, the data reported by these authors and our findings regarding the correlation between isoform prevalence and arm usage variations bring new insights into the impact of isomiRs in promoting arm switching. However, some limitations of our analysis refer to the current availability of a restricted number of datasets. Larger and more diverse sample sizes are needed to further confirm our results. Also, functional studies are desirable to better investigate such interaction and for deeper validation of specific miRNA isoforms association to arm switching events under variable contexts. For instance, qPCR or gene knockdown/overexpression experiments of mature 5p and 3p miRNAs after manipulation on pri- and pre-miRNA sequences could explore how modifications in the sequence or stability of isomiRs influence the selection of 1 arm over the other. Moreover, our study was focused on specific developmental stages or tissue types, limiting our ability to generalize findings across other stages or tissues. Given the upcoming availability of novel datasets, a broader analysis across multiple stages and tissues would provide a more comprehensive understanding of the relationship between isomiRs and arm switching. Understanding the role of isomiRs in arm selection could lead to the development of new strategies for manipulating miRNA activity and provide novel insights into the regulation of gene expression in various biological contexts.
Conclusions
In summary, our study sheds light on the rare yet significant occurrence of miRNA arm switching across metazoans. Despite its infrequency, arm switching plays a crucial role in developmental processes, spanning embryonic stages to adulthood, and even between miRNA paralogs. We highlight the species-specific nature of arm switching, suggesting its close association with unique cellular contexts. Moreover, our analysis reveals distinct regulatory roles for 5p and 3p arms, with limited overlap in their target gene sets, indicating their differential biological functions. Additionally, we propose a potential link between isomiRs and arm switching, suggesting that variations in isoform prevalence may influence arm selection during miRNA biogenesis. Our findings underscore the complexity of miRNA regulation and its implications for gene expression dynamics. Further research into the mechanisms underlying arm switching promises to deepen our understanding of gene regulatory networks across species and tissue.
Supplemental Material
sj-xlsx-1-evb-10.1177_11769343241263230 – Supplemental material for MicroRNA Transcriptomes Reveal Prevalence of Rare and Species-Specific Arm Switching Events During Zebrafish Ontogenesis
Supplemental material, sj-xlsx-1-evb-10.1177_11769343241263230 for MicroRNA Transcriptomes Reveal Prevalence of Rare and Species-Specific Arm Switching Events During Zebrafish Ontogenesis by Arthur Casulli de Oliveira, Luiz Augusto Bovolenta, Lucas Figueiredo, Amanda De Oliveira Ribeiro, Beatriz Jacinto Alves Pereira, Talita Roberto Aleixo de Almeida, Vinicius Farias Campos, James G Patton and Danillo Pinhal in Evolutionary Bioinformatics
Footnotes
Acknowledgements
We are grateful to the technical staff of the Genetics sector and the members of the LGEM lab for their invaluable support throughout this research study.
Author Contributions
Conceptualization, A.O., and D.P.; methodology, A.O., L.B., and L.F.; data curation, A.O., L.B., L.F., A.R., and B.P.; writing – original draft preparation, A.O., V.C., J.P., and D.P.; writing – review and editing, A.R., B.P., J.P., T.A., and D.P.; funding acquisition, V.C. and D.P. All authors read and approved the final manuscript.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from Sao Paulo Research Foundation (FAPESP; #2017/17510-2, #2018/05484-0, and #2019/15494-5), Rio Grande do Sul Research Foundation (FAPERGS; #19/2551-0002283-2) National Council for Scientific and Technological Development (CNPq; #167444/2017-4, #433746/2018-1, #136152/2018-0, and #315139/2020-0), and by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES; Finance Code 001).
Ethics Approval
Experimental protocols were approved by the local Ethics Committee on Animal Use (CEUA protocol no. 1008 - 2017) at the Institute of Biosciences of Botucatu.
Data Availability Statement
Additional data that support the findings of this study are available from the corresponding author, D.P., upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

