Abstract
MicroRNAs are small non-coding RNA molecules that are produced in a cell endogenously. They are made up of 18 to 26 nucleotides in strength. Due to their evolutionary conserved nature, most of the miRNAs provide a logical basis for the prediction of novel miRNAs and their clusters in plants such as sunflowers related to the Asteraceae family. In addition, they participate in different biological processes of plants, including cell signaling and metabolism, development, growth, and tolerance to (biotic and abiotic) stresses. In this study profiling, conservation and characterization of novel miRNA possessing conserved nature in various plants and their targets annotation in sunflower (Asteraceae) were obtained by using various computational tools and software. As a result, we looked at 152 microRNAs in Arabidopsis thaliana that had already been predicted. Drought tolerance stress is mediated by these 152 non-coding RNAs. Following that, we used local alignment to predict novel microRNAs that were specific to Helianthus annuus. We used BLAST to do a local alignment, and we chose sequences with an identity of 80% to 100%. MIR156a, MIR164a, MIR165a, MIR170, MIR172a, MIR172b, MIR319a, MIR393a, MIR394a, MIR399a, MIR156h, and MIR414 are the new anticipated miRNAs. We used MFold to predict the secondary structure of new microRNAs. We used conservation analysis and phylogenetic analysis against a variety of organisms, including Gossypium hirsutum, H. annuus, A. thaliana, Triticum aestivum, Saccharum officinarum, Zea mays, Brassica napus, Solanum tuberosum, Solanum lycopersicum, and Oryza sativa, to determine the evolutionary history of these novel non-coding RNAs. Clustal W was used to analyze the evolutionary history of discovered miRNAs.
Keywords
Introduction
Sunflower (Helianthus annuus) belongs to the Asteraceae family. By cloning method, 700 types of miRNA were identified in plants; in 2012, miRNA was identified in Arabidopsis thaliana. All the processes of miRNA targets are based on coding and non-coding sequence. 1 Previous studies show that RNA polymerase play important role in the transcription of the miRNA gene. 2
In recent years, for the identification of miRNA scientists have used high throughput sequencing and computational analysis techniques. 3 Almost all scientists have concluded that microRNAs involved in the regulatory function of flowering and non-flowing plants are conserved. 4 Plants are damaged by 2 types of environmental stresses categorized as biotic and abiotic stress. Damage to living organisms by living organisms like parasites, bacteria, viruses, and fungi is known as abiotic stress as well as damage to a living organism by the source of nonliving factors called abiotic stress. 5 To describe abiotic stress, we should study the function of different organisms that survived in different environments. Stress always affects the plant’s tissues. plants need enough water for their sufficient growth. Up and down movement of water expand plant cells, which causes plant growth. Enough amount of water expands the plant’s cells and transfers the minerals from the soil to the tip of the leaves but stress causes an imbalance in the plant’s routine processes.
According to research every year, we lost 50% of our food production due to abiotic stress. Abiotic stress affects plants’ fruits, crops, metabolism, respiration processes of plants, and at the end plant’s seeds. Seeds are used for next generation; hence, unhealthy seeds affect further production. 6 H. annuus typically refer to annual species that tend to spread rapidly and can become aggressive. 7 The plant family improvement depends on the genetically resistant varieties, seed productivity, modern cultivation, and biotic-abiotic stress tolerance. Plant improvement can be assessed by studying its genetic makeup and sowing in a different location. 8 For the human, it acts as an important source of nutrients. Nutritionally, it is the main source of vital nutrients inkling carbohydrates, proteins, and dietary fibers, and provides almost 20% of the dietary energy supply. According to the miRNA database, H. annuus contains a total of 6 precursors and 7 mature microRNA that are compared with A. thaliana for the identification of novel microRNA. 9 A. thaliana contained 205 precursors and 384 mature non-coding RNA. All data regarding miRNA are present in miRBase.
Methodology
Many tools are used as comparative genomics approaches to achieve novel and interesting information about miRNAs in plants and animals. In the initial step, identify sequences and reference sequences, and download them from the microRNA Registry Database. This miRBase Pre-miRNA database was available at https://www.mirbase.org/ freely. Pre-miRNA potential candidates were predicted by subjecting the downloaded mature and precursor miRNAs sequence through the Basic Local Alignment Tool. For this purpose, nucleotide BLAST available freely at Genbank of the National Center for Biotechnology Information (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE_TYPE=BlastSearch) was used. The miRNA* sequences, both mature and precursor, were subjected to BLAST against H. annuus expressed sequence tags (ESTs) sequentially using the BLASTn program following a maximum of up to 4 mismatches with miRNAs*.
EST single-tone selection
BLASTn program was used and the setting of the parameters was adjusted as expect values, 1000; low complexity, the sequence filter, database, others; organism, H. annuus; program selection, somewhat similar sequences; and all other parameters, by default. To identify the coding part of miRNA, we used BLASTx. BLASTx highlights coding regions. 10
Prediction of miRNAs secondary structure
MFOLD, a secondary structure prediction tool was used to produce a stem-loop structure for the initially identified potential H. annuus. All the initial candidate sequences that failed to develop stable secondary structures were discarded. MFOLD software updated as UNAfold http://www.unafold.org/ and then clicked on MFOLD and then selected application (RNA fold form Version 2.3). 11
UNAfold → MFOLD → Application → RNA fold form version
The setting of MFOLD parameters was adjusted as RNA sequence, linear; folding temperature, 37°C, ionic concentration, 1 mol/L of National Center for Biotechnology Information (NCBI) having no divalent ions; percent sub-optimality number, 5; maximum interior loop size 30.
Conservation and phylogenetic analysis
Clustal W was selected for phylogenetic analysis. Clustal W is used for multiple sequence alignment or the alignment of more than one sequence available at https://www.genome.jp/tools-bin/clustalw.
Results
New potential miRNAs in sunflower
In all research, we predicted 152 miRNAs that performed regulatory process against drought-tolerant stress in A. thaliana. miRNAs that respond to drought stress are 156a, 156b, 156c, 156d, 156e, 156f, 158a, 159a, 164a, 164b, 165a, 165b, 166a, 166b, 166c, 166d, 166e, 166f, 166g, 168a, 168b, 169a, 170, 171a, 172a, 172b, 173, 156b, 319a, 319b, 169b, 169c, 169d, 169e, 169f, 169g, 169h, 169i, 169j, 169k, 169l, 169m, 169n, 171b, 171c, 172c, 172d, 339a, 339b, 394a, 394b, 397a, 397b, 398a, 398b, 398c, 399a, 399b, 399c, 399d, 399e, 399f, 400, 401, 402, 403, 404, 408, 159g, 156h, 158b, 159c, 319c, 164c, 172e, 417, 418, 414, 415, 416, 419, 420, 426, 427a, 427b, 427c, 827, 830, 833a, 835, 836, 837, 839, 841a, 842b, 844, 845a, 846, 848, 850, 851, 847, 855, 854a, 854b, 854c, 856, 857, 858, 859, 860, 861, 865, 845b, 870, 1886, 1888a, 2933a, 2933b, 2936, 3434, 774b, 4221, 854e, 5021, 5024, 5025, 5026, 5029, 5641, 5642a, 858b, 833b, 156i, 156j, 5652, 5653, 5654, 5655, 5656, 5658, 5188, 1888b, 5666, 5996, 8121, 8165, 8170, 8178, 8180, and 8181. All these miRNA sequences are reported in A. thaliana.
Prediction of novel miRNA
In A. thaliana, after identifying the 152 microRNA sequences, we performed local alignment of these sequences by BLASTn with the parameters of somewhat similarities and H. annuus in organisms and selected those BLAST sequences that were 80% to 100% similar. So in local alignment, we got 19 sequences of A. thaliana against H. annuus. The accession numbers of these 19 sequences are XR_002550943, XR_002552183, XR_002552875, XR_002554296, XR_002556375, XR_002562643, XR_002570335, XR_002574508, XR_2575503, XR_002579567, XR_002582121, XR_002587840, XR_002549854, XR_002592285, XR_004862999, XR_004863001, XR_004865012, XR_004869948, and XR_004890935 (see in Table 1).
Novel microRNA in Helianthus annuus that is predicted with the reference of Arabidopsis thaliana sequences.

These 19 sequences code for different miRNAs that were novel. In our research, we predicted 12 novel miRNAs in sunflowers against A. thaliana. These novel predicted miRNAs are MIR156a, MIR164a, MIR165a, MIR170, MIR172a, MIR172b, MIR319a, MIR393a, MIR394a, MIR399a, MIR156h, and MIR414 (see in Table 2).
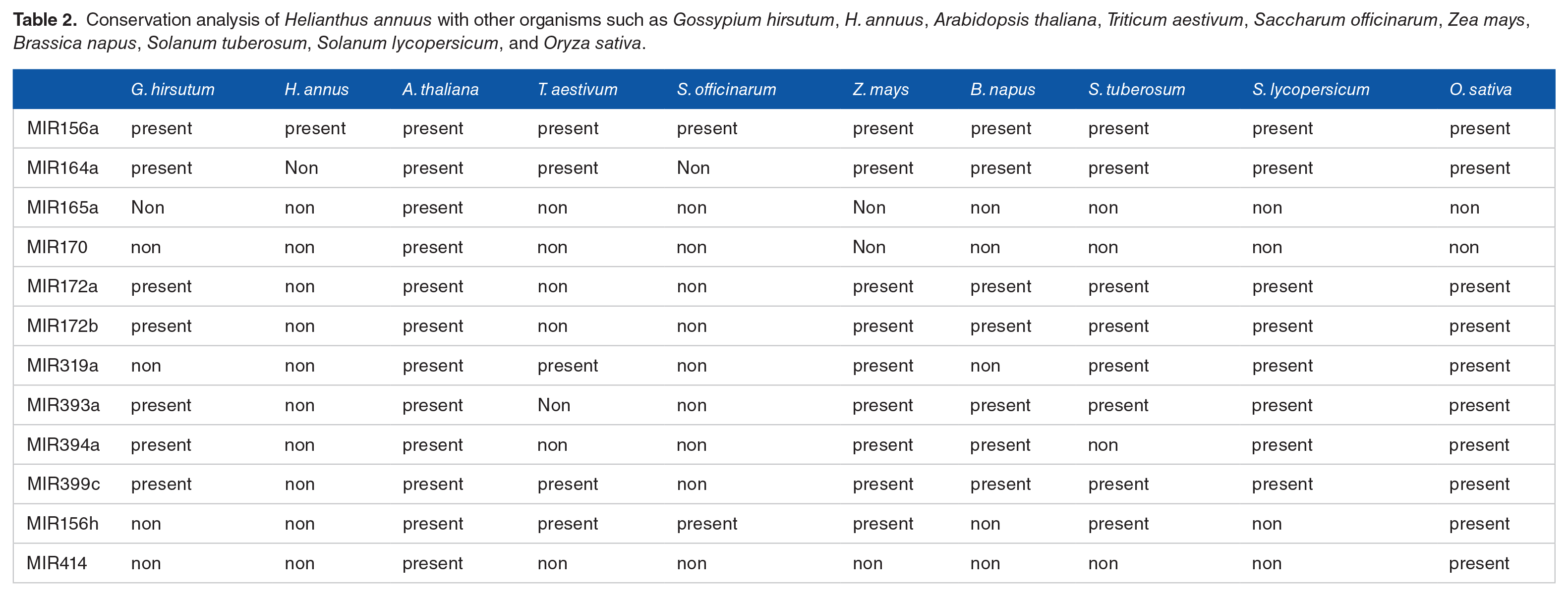
Conservation analysis of Helianthus annuus with other organisms such as Gossypium hirsutum, H. annuus, Arabidopsis thaliana, Triticum aestivum, Saccharum officinarum, Zea mays, Brassica napus, Solanum tuberosum, Solanum lycopersicum, and Oryza sativa.
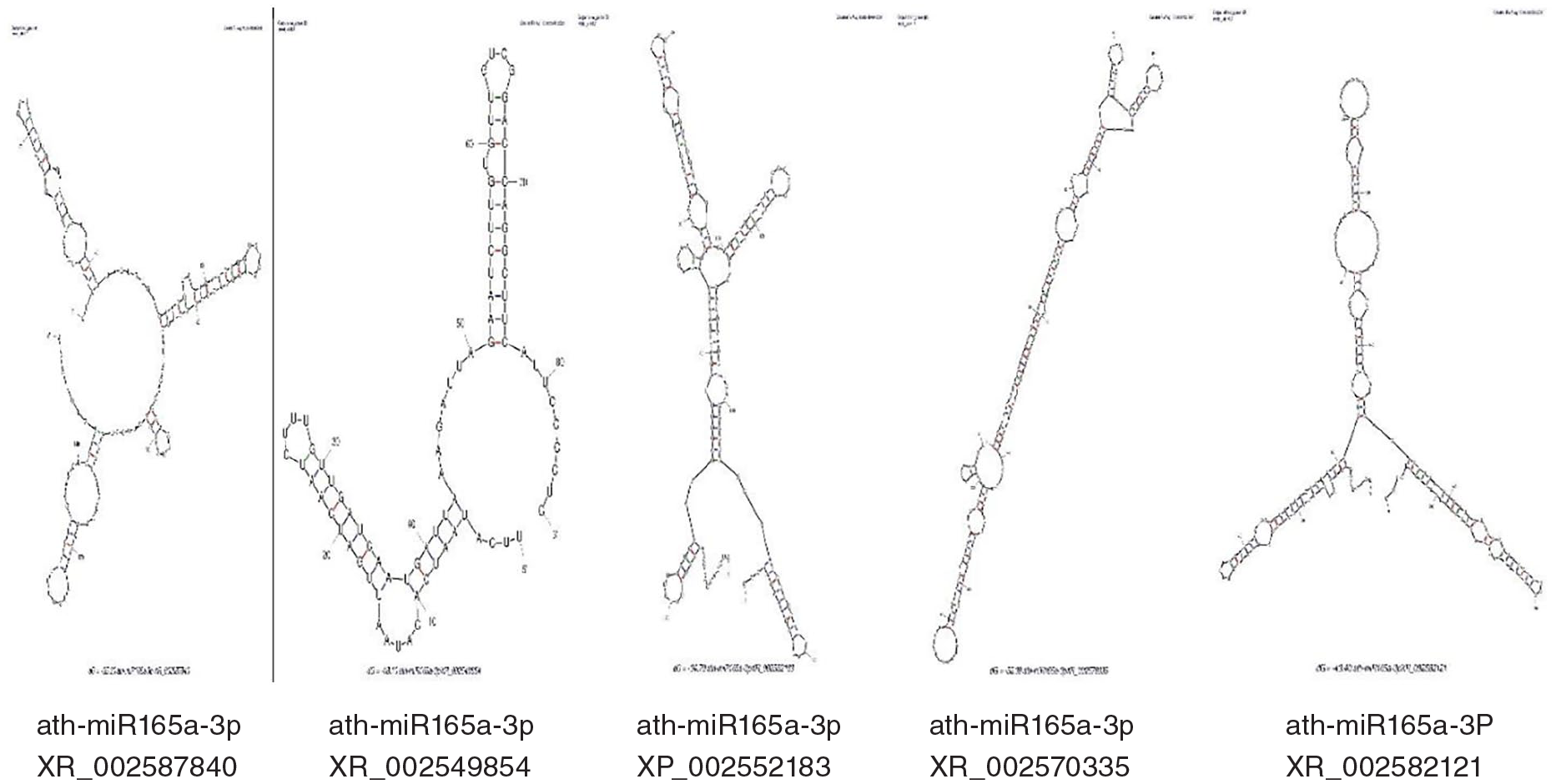
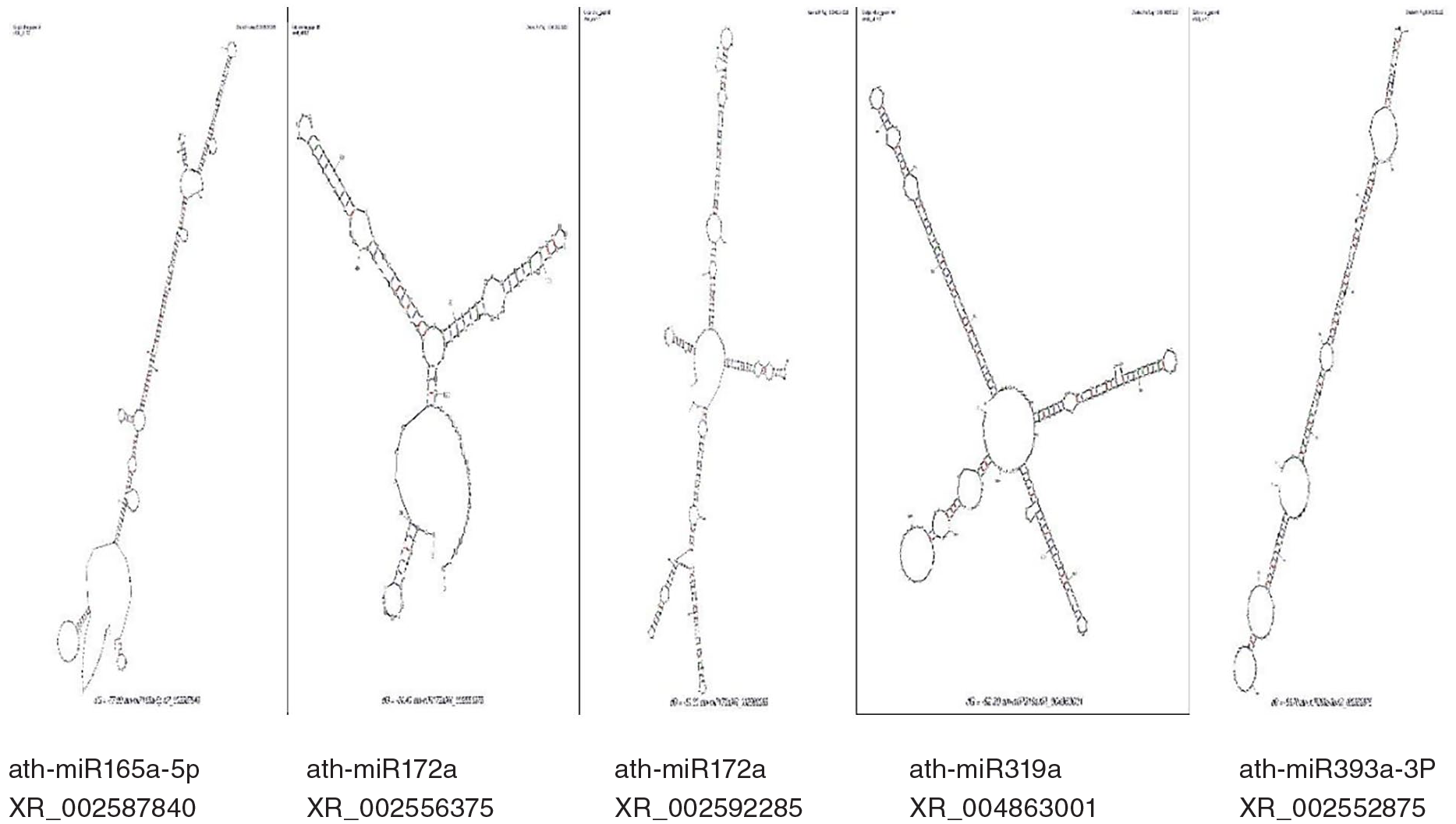
Novel sunflower miRNAs structures predicted from A. thaliana
In our research, we predicted 12 novel microRNAs in the sunflower that respond against drought stress (MIR156a, MIR164a, MIR165a, MIR170, MIR172a, MIR172b, MIR319a, MIR393a, MIR394a, MIR399a, MIR156a, and MIR414). Here, we explain the miRNAs structures on the base of both 5 and 3 prime by MFOLD.




Ancestral conservation of H. annuus
In addition, novel sunflower precursor miRNAs were selected for conservation studies. For this process, all novel miRNAs selected to evaluate their conservation in other plants. For conservation studies, we selected 10 different types of plants like Gossypium hirsutum, H. annuus, A. thaliana, Triticum aestivum, Saccharum officinarum, Zea mays, Brassica napus, Solanum tuberosum, Solanum lycopersicum, and Oryza sativa. Using the same process, many researchers have determined precursor conservation and phylogenetic analysis in different plants.
Phylogenetic analysis of non-coding microRNAs
The study of the evolutionary history of a species or a group of organisms or a particular characteristic of an organism. Here we have done phylogenetic analysis by clustal W.
Phylogenetic analysis of MIR156a
The phylogenetic analysis of mir156a is described in Figures 1 and 2. According to the MIR156a phylogenetic analysis, H. annuus with the Accession number “XR_002550943” and “XR_4869948” shows close relation with T. aestivum, B. napus, A. thaliana, G. hirsutum, O. sativa, Z. mays, and S. officinarum.

MIR156a.

MIR 156.
Phylogenetic analysis of MIR164a
The phylogenetic analysis of mir164a is described in Figure 3. According to the phylogenetic analysis of MIR164a, H. annuus with the Accession number “XR_002562643” shows close relation with T. aestivum, B. napus, A. thaliana, O. sativa, Z. mays, S. lycopersicum, and G. hirsutum and shows a distance relationship with S. tuberosum.

MIR164a.
Phylogenetic analysis of MIR172b
The phylogenetic analysis of mir172b is described in Figure 4. According to the phylogenetic analysis of MIR172b, H. annuus with the Accession number “XR_004862999” shows close relation with A. thaliana, Z. mays, S. lycopersicum, S. tuberosum and shows a distance relationship with G. hirsutum.

MIR172b.
Phylogenetic analysis of MIR172a
Phylogenetic analysis of mir172a is described in Figures 5 to 7. According to the phylogenetic analysis of MIR172a, H. annuus with the Accession number “XR_002592285” and “XR_002556375” shows close relation with A. thaliana, Z. mays, S. lycopersicum, O. sativa, B. napus and with “XR_004865012” shows close relationship with G. hirsutum.

MIR 172a (XR_002592285).

MIR172a (002556375).

MIR172a (004865012).
Phylogenetic analysis of MIR319
The phylogenetic analysis of mir319 is described in Figure 8. According to the phylogenetic analysis of MIR319, H. annuus with the Accession number “XR_004863001” shows close relation with A. thaliana, S. lycopersicum, and T. aestivum.

MIR319(XR_004863001).
Phylogenetic analysis of MIR393a
The phylogenetic analysis of mir393 is described in Figure 9. According to the phylogenetic analysis of MIR93a, H. annuus with the Accession number “XR_002552875” shows close relation with A. thaliana, G. hirsutum, O. sativa, S. tuberosum, and Z. mays.

MIR393a(XR_002552875).
Phylogenetic analysis of MIR394
The phylogenetic analysis of mir393 is described in Figure 10. According to the phylogenetic analysis of MIR94, H. annuus with the Accession number “XR_002554296” shows a distance relation with A. thaliana, G. hirsutum, O. sativa, B. napus, S. lycopersicum, and Z. mays.
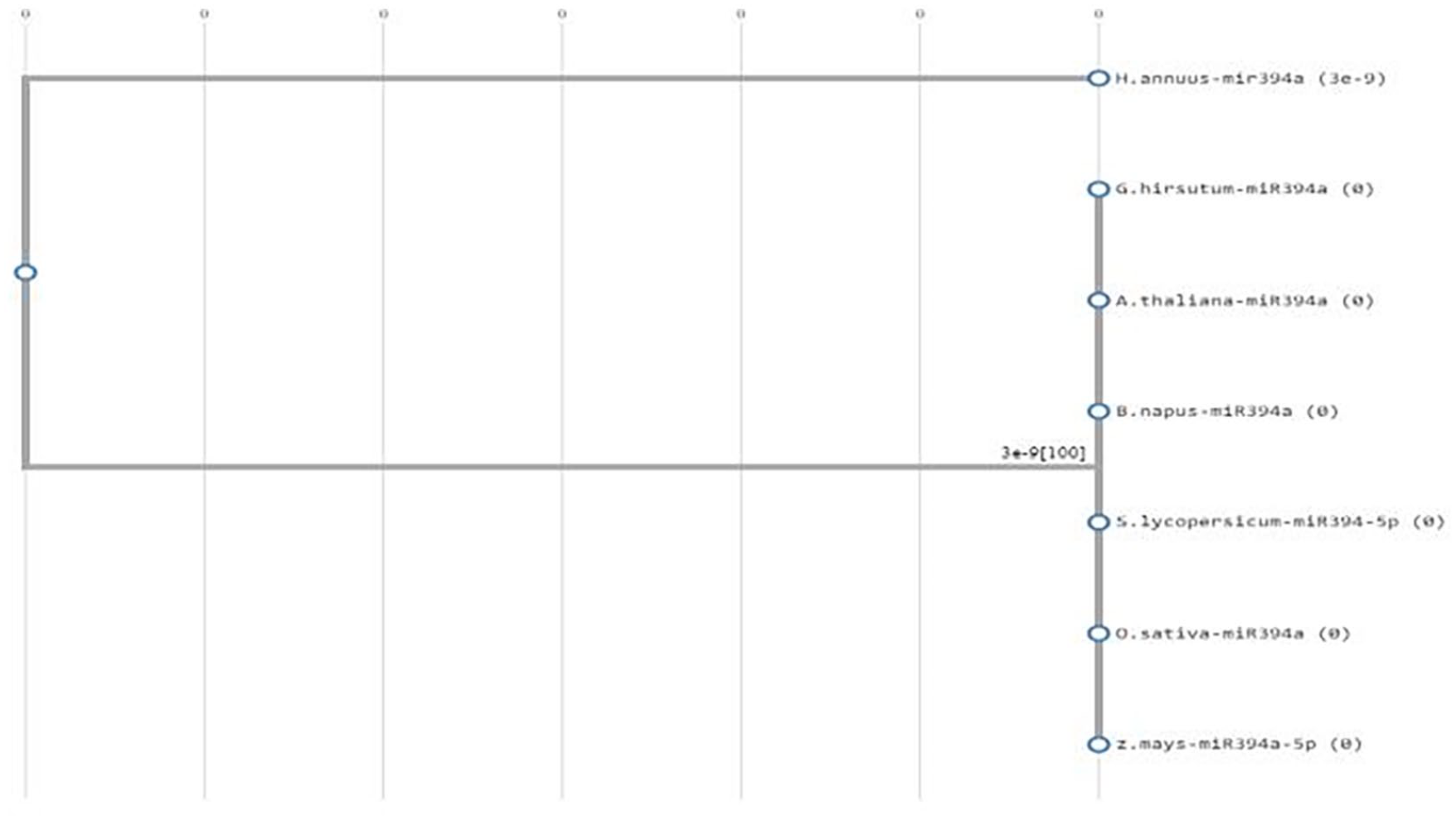

XR_002554296.
Phylogenetic analysis of MIR399
The phylogenetic analysis of mir399 is described in Figure 11. According to the phylogenetic analysis of MIR94, H. annuus with the Accession number “XR_002574508” shows a distance relation with A. thaliana, G. hirsutum, O. sativa, B. napus, S. lycopersicum, and Z. mays.

MIR399(XR_002574508).
Discussion
Sunflower is the fourth biggest oil-seed crop in the world. The seeds of sunflowers are used in food as well as their dried stalk is used as fuel. It has previously been used as an ornamental plant and was also used in ancient ceremonies. 12 Moreover, different parts of sunflowers are used in body painting, decorations, and making dyes for the textile industry. Its oil is used in the manufacturing of margarine and salad dressings, and cooking. With roasted seeds, a coffee type could be made. In industry, it is used in cosmetics and paints. Due to its lack of anti-nutritional factors and high nutritional values, it is a potential source of protein for human consumption. Due to its metabolic, physiological, and morphological adaptation strategies, the sunflower is one of the most important oil-seed crops and is resistant to various abiotic stresses. This crop is of special interest for its adaptation to limited water availability, high temperatures, high salinity, and heavy-metal concentrations in soil. The dried stems which are used for fuel contain potassium and phosphorous which can be composed and returned to the soil as fertilizer. 13
MiRNAs arise from primary longer RNA transcripts that include a self-complementary fold-back, from which the mature miRNAs are excised. They are short RNA molecules containing 19-24 nucleotides in size.14,15 They are familiar as regulators of gene expression by binding to open reading frames (ORF) or untranslated regions (UTR) of specific mRNAs, targeting them for directing or cleavage translation inhibition at the mRNA level. It has been demonstrated that around 60% of protein-coding genes are targets of miRNAs and are modulated by these small RNAs.
miRNAs are derived from hairpin pre-miRNA from which both miRNA and the imperfectly complementary miRNA* strands are released. Their sequences are not conserved between plants and animals, and even not have been seen in fungi. Many miRNAs within the kingdom have an ancient origin, some being completely conserved among sunflower, Arabidopsis, rice, and even liverworts, mosses, and hornworts. 16
By regulating gene expression, miRNA plays a vital role to regulate the developmental processes of organisms. 17 The negative regulation of miRNAs in gene expression in both plants and animals has been demonstrated. 18 MicroRNAs have been revealed to modulate diverse developmental processes, including polarity, identity, and organ separation, and to regulate their function and biogenesis. 18 In our study, we used the miRBase database to find miRNAs from A. thaliana that were tolerant against drought stress that was 152 in strength and then performed local alignment of these miRNAs against sunflower and found 12 novel miRNAs (MIR156a, MIR164a, mir165a, mir170, mir172a, mir172b, mir319a, mir393a, mir394a, mir399c, mir156h, and mir414). The secondary structure of these 12 novel miRNAs, including forward and reverse strands, was forecasted by MFOLD software by using default parameters. Later, conservation and phylogenetic analysis were done by selecting 10 different organism type such as G. hirsutum, H. annuus, A. thaliana, T. aestivum, S. officinarum, Z. mays, B. napus, S. tuberosum, S. lycopersicum, and O. sativa. In our study, all novel miRNAs are present in only A. thaliana.
In our study, mir-156a is present in all species; mir-164 is present in all except H. annus and S. officin; 19 mir-165a and mir-170 are present in only A. thaliana; mir-172a and mir172-b are present in all except H. annus, T. aestiyum, and S. officin; 20 mir-319a is present in all except G. hirsutum, H. annus, S. officin, and Z. mays; mir-393a is present in all except H. annus, T. aestiyum, and S. officin; mir-394a is present in all except H. annus, T. aestiyum, S. officin, and B. napus; mir-399c is present in all except H. annus and T. aestiyum; mir-156h is present in all except G. hirsutum, H. annus, B. napus, and S. lycopersicum; and mir-414 is absent in all except A. thaliana, and O. sativa.
Conclusions
Twelve novel miRNAs (MIR156a, MIR164a, mir165a, mir170, mir172a, mir172b, mir319a, mir393a, mir394a, mir399c, mir156h, and mir414) were identified against drought stress in A. thaliana. We targeted these miRNAs to cope with the drought tolerance in sunflowers. In our study, all novel miRNAs are present in only A. thaliana. Moreover, different parts of sunflowers are used in body painting, decorations, and making dyes for the textile industry. Its oil is used in the manufacturing of margarine and salad dressings, and cooking. With roasted seeds, a coffee type could be made. In industry, it is used in cosmetics and paints. The improvement method also increases the production of sunflowers and benefits economically.
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research study is self-funded.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
All authors have contributed to this study.
Availability of Data and Materials
The data associated with a paper are available on demand through email contact of co-author
Ethical Approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Consent for Publication
This study is based on research.
