Abstract
Background:
Inflammatory bowel diseases (IBDs) are complex, multifactorial disorders that comprise Crohn’s disease (CD) and ulcerative colitis (UC). Recent discoveries have brought much attention to the genetic predisposition of patients with IBD. Here we evaluate the interaction between IBD genetic risk factors susceptibility and CD occurrence in an IBD pediatric patient population, performing a clinical exome survey.
Methods:
From February 2018 to April 2019, we collected blood samples from 7 pediatric patients with IBD concerns from several collaborating health centers and/or hospitals. Blood samples were processed by extracting and sequencing DNA for a clinical exome survey. Shophia-DDM-v3-4 platform allowed sequenced reads alignment on hg19 genome as well as genetic variant calling. Both IBD risk and pathogenic genetic variants covered by at least 20 reads were selected for subjacent analysis.
Results:
Normality and Bartlett tests of both risk and pathogenic genetic variants suggested random and heterogeneous distribution of these variants in this group of IBD pediatric patients.
Conclusion:
Our study revealed specific genetic variants linked to CD susceptibility, autoimmune and/or innate immunodeficiency as well as to metabolic defects, as favoring factors of IBD, suggesting the valuable role of next generation sequencing (NGS) approaches in IBD molecular diagnostic procedures.
Keywords
Introduction
The inflammatory bowel disease (IBD) is a group of gastrointestinal disorders including Crohn’s disease (CD) and ulcerative colitis (UC), which are distinguished by the distribution of chronic inflammatory changes. The inflammation in UC is continuous and restricted to the mucosal layer of the colon, whereas CD is characterized by segmental transmural lesions that can affect any part of the gastrointestinal tract. The gastrointestinal tract is always solicited in regulating the gut microbial flora, controlling immune responses to food chemical substances antigens as well as maintaining homeostasis. It is well known that homeostasis disturbances contribute to risk of occurrence of IBD. Of note, the innate immune response is indispensable for maintaining homeostasis, and abnormal innate immune activity is deeply involved in the pathogenesis of IBD. Research in this field has made substantial advancements in recent years.1
-3 Genome-wide association studies (GWAS) have identified over 200 loci associated with IBD,4,5 providing many potential therapeutic targets. The large majority of these loci are common to CD and UC, implicating abundant pathways, notably the pro-inflammatory interleukin (IL)-23 pathway.
6
CD predominant loci include nucleotide-binding oligomerization domain-containing protein 2 (
Materials and Methods
IBD patient population
Seven pediatric patients with diagnosis of IBD were included in the present clinical exome diagnostic analysis. Patients diagnosed with different types of inflammatory bowel disease were enrolled from February 2018 to August 2018; samples were collected in the reference center and shipped to our laboratory for a clinical exome sequencing analysis. The overall research included 4 research institutes (IRCCS) and 8 Italian universities. Here, we reported and summarized general and clinical features regarding processed IBD pediatric patients (Table 1).
Summary of generic and clinical data features of processed inflammatory bowel disease pediatric patients.

Abbreviation: IBD, inflammatory bowel diseases.
DNA library preparation and exome sequencing
Whole blood samples from 7 unrelated patients were collected in EDTA. Genomic DNA (gDNA) was extracted by QIAamp DNA Mini Kit Qiagen following manufacturer’s instructions. Genomic DNA was loaded into the spectrophotometer UV/Vis Infinite M200 TECAN to verify purity of the samples. Then, we used 200 ng of DNA for library preparation with SOPHiA GENETICS Clinical Exome Solution. Briefly, gDNA is fragmented and ligated to Dual Index Adapters, and then fragments with the correct size (300-700 bp) are selected using the AMPure XP beads. We assessed libraries quality by using Quant-iT PicoGreen dsDNA Reagent (Life Technologies) consisting of a calibration curve by quantifying serial dilution of standard DNA. For this purpose, we mixed 1 μL of library sample with 99 μL of TE Buffer and 100 μL of PicoGreen working solution (1:200 in TE Buffer) and quantified. Four individual libraries (300 g each) from 4 patients were pooled and lyophilized in the Eppendorf Concentrator 5301 at 45°C, then the pellet is hybridized to the xGen Lockdown Probes for 4 hours; the hybridization product is then bound to streptavidin beads prior to polymerase chain reaction (PCR) amplification with KAPA HiFi HotStart ReadyMix (Roche). The PCR product is cleaned up through binding with AMPure XP magnetic beads and finally resuspended in IDTE buffer to be stored at −20°C. Before sequencing, the sample was quantified with Qubit 4 Fluorometer; the pool molarity was determined, and sample was diluted to reach the concentration of 10 pM. Pair end sequencing was performed on MiSeq (Illumina) platform.
Reads quality control and aligned sequences statistic
In this section, we summarized read quality control item for each processed inflammatory bowel disease (IBD) pediatric patients. Exome sequencing were executed by Miseq machine by using Illumina technology, performing DNA pair end sequencing. Read quality control as well as base calling statistic were automatically generated by sequencer machine. We summarized reads and base calling statistical survey (Table 2).
Descriptive statistic and quality control analysis of aligned read sequences.

Clinical genomic data processing by Sophia platform
SOPHiA DDM is the platform of choice for clinicians performing routine diagnostic testing. This platform performs sequence data analysis by including alignment up to calling annotated facilitated variants. So, after login to SOPHiA DDM platform, we processed in loading raw DNA sequence FastQ file. Loaded raw DNA sequence are automatically and immediately processed and aligned on GRCh37/hg19 reference genome, using patented advanced algorithms and machine learning approaches assuring the highest accuracy of subjacent genomic analysis. After this step, processed data are visualized on SOPHiA DDM platform. SOPHiA AI an algorithm of SOPHiA DDM platform allows processed data interpretation by pre-classifying variants in different pathogenic classes from highly pathogenic to benign. We directly interpret the variants and generate a variant report all that within one tool. Next, we exported variant VCF files in an external environment (R software) for subjacent variant filtering as well as statistical analysis. Risk factor and/or pathogenic clinical variants for IBD were filtered for target regions covered at least 20 reads, as well as by checking SNP variant on online mendelian inheritance in man (OMIM) inerrant status (heterozygous and/or homozygous status) concordance. In addition, we checked and compared for each selected SNP variants, associated biological functions by using several genomic databases (ie, Gene Card and Ensembl).
Statistical analysis
Data (selected IBD risk factor as well as pathogenic genetic variants) were statistically processed in R programming environment. We checked for IBD SNP risk factor variants heterogeneity as well as normality distribution by applying respectively Bartlett and Shapiro normality tests. Indeed, risk factors under ClinVar parameter as reported in genomic variation as it relates to human health with sequencing depth at least to 20 reads were processed for above-mentioned variance homogeneity as well as normality tests. We analyzed IBD pediatric patient’s interaction by achieving Z-score and
Results
Assessment of IBD risk factor SNP variants variance homogeneity in processed IBD pediatric patients
Because of IBD multifactorial features, we assessed variance homology with selected IBD-related SNP variants by performing a Bartlett test. In total, 157 IBD genetic variants, under ClinVar parameter as reported in genomic variation as it relates to human health, covered by at least to 20 read sequences were selected for present variance homogeneity test. Of note, Bartlett test assessing homogeneity of risk factor variance in analyzed IBD pediatric patient population revealed statistically significant Bartlett’s K-squared = 100.71 at a freedom degree (
Bartlett variance homogeneity test out came by comparing IBD patient population each to other by processing genetic risk factor variants.
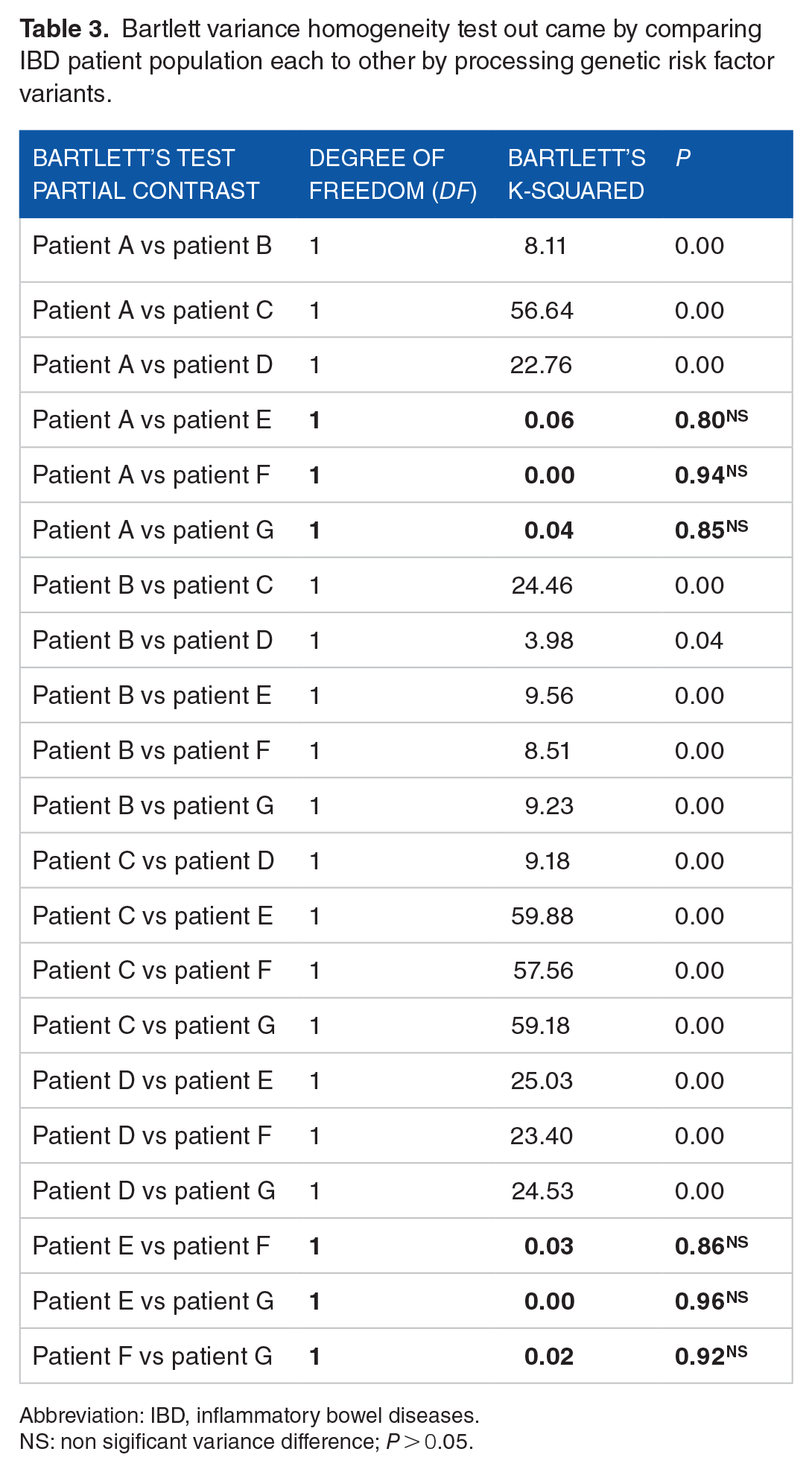
Abbreviation: IBD, inflammatory bowel diseases.
NS: non sigificant variance difference;
Inflammatory bowel disease pediatric patients clustering analysis by processing IBD risk factor SNPs variants
We investigated 91 non-redundant risk factor variants in 7 children with IBD. Indeed, analysis of selected genetic variants by Z-score clustering suggested a high heterogeneity in the analyzed IBD pediatric patient population (Figure 1A). Z-score analysis of genetic variants showed that IBD pediatric patients’ heterogeneity behavior clustered in two groups, suggesting a relative resemblance of IBD pediatric patients A, E, F, and G as compared with patients B, C, and D (Figure 1). Furthermore,

IBD pediatric patient population clustering analysis by processing recurrent risk factor genetic variants (SNPs) by Z-score (A) and
List of the top risk factors and/or pathogenic genetic variants recurrently recorded in analyzed inflammatory bowel disease (IBD) pediatric patients cover by at least 20 reads.

Abbreviations: IBD, inflammatory bowel diseases; SNP, single nucleotide polymorphism.
Homozygote variant.
Evaluation of risk factor variant features distribution in the IBD pediatric patient population
Venn diagram analyses of genetic variants in IBD patients revealed high heterogeneity and random distribution of selected risk factor variants in IBD pediatric patient population (Figure 2). Indeed, normality test (Shapiro normality test) by exhibiting Shapiro coefficient; W = 0.54 at

(A) Venn diagram assessing risk factor variants (SNP) data distribution in processed inflammatory bowel disease (IBD) pediatric patients. (B) Density plot measuring risk factor SNP variants normality in processed IBD pediatric population.
Genetic pathogenic variant in evaluating IBD pediatric patient population diagnosis
Next, we assessed the occurrence of genetic pathogenic variants in children with IBD. The study showed in patient A the synonymous rs10065172 heterozygous (c.313C > T/ p.(Leu105=) variant in IRGM gene and in patient C the missense homozygous variant rs1805010 (c.223A > G / p.(Ile75Val)) in IL4R gene, which are associated with inflammatory bowel disease and immunodeficiency syndrome, respectively (Table 2). Interestingly, in patient D we identified the above-mentioned pathogenic variants (rs10065172 and rs1805010), together with the MBL2-related rs5030737 heterozygous variant (c.154C > T/p.(Arg52Cys)), which is associated with partial mannose-binding protein deficiency. Of note, both patients A and F carried the pathogenic variant rs33995883 (p.(Asn2081Asp)) of LRRK2 gene, which confer increased risk to develop CD. In addition, we identified both the heterozygous variant of IL4R gene (rs1805010 variant) and the mannose binding protein deficiency variant (rs5030737) in patients F and G. In patients E and F, we identified an intronic NOD2 variant (rs5743289: c.2717 + 158C > T), although with low coverage average, which was linked to both Yao syndrome and CD. Besides gene variants associated with immune response, we have also identified in children with IBD pathogenic variants associated with metabolic disorders. In patients D and F, we identified several metabolism disorder disease pathogenic genetic variant such as, rs696217 (c.211C > A/p.(Leu71Met)) in GHRL gene, rs3733402 (c.428G > A/p.(Ser143Asn)) in KLKB1 gene and rs351855 in FGFR4 gene. All of them are involved in metabolic syndrome, obesity age at onset, body fat distribution, cancer progression and tumor cell motility regulation. In addition, genetic variants in PQBP1 (rs606231195: INDEL mutation resulting in frameshift) and in SLC6A20 gene (rs17279437) causing metabolic dysfunction were identified in patient F (Supplementary Tables 1 and 2).
Discussion
Our study suggests that risk factor variants associated with metabolic disorders, autoimmune or CD susceptibility can be linked to distinct clusters of IBD patients. Numerous studies have shown that multiple factors, including genetic susceptibility, microbiota, and immune system functions can influence IBD pathogenesis.19
-22 Our study revealed rs2066844 (NOD2) and rs2241880 (ATG16L1) IBD susceptibility risk factor variants in A and F patients. In addition, our study revealed a CTLA4 variant in patient A with IBD. These results are in agreement with previous articles which reported the link between genetic variants in ATG16L, NOD2, CARD15, IBD5, CTLA4, TNFSF15, IL2R genes, and CD.23
-27 Indeed, CTLA4 gene is a member of the immunoglobulin superfamily which is expressed on the surface of helper T cells and is essential in the control process of intestinal inflammation by CD25+/CD4+ regulatory T cells.
28
We have also observed that the genetic variant of ATG16L1 gene (rs2241880) was heterozygous in 3 patients while homozygous in an additional one, which is associated with highly increased risk for CD. Some of the genetic variants identified in patients with IBD are related to common biological pathways. In particular, the IBD-associated variants in NOD2, ATG16L1, and IRGM genes affect cellular autophagy processes and bacterial clearance in innate immune cells, and may influence bacterial composition of the gut in patients with IBD. Analysis of the functional effects of the exonic synonymous rs10065172 variant in
In conclusion, we report a correlation of IBD CD occurrence and autoimmune with metabolic disorder susceptibility in a small pediatric IBD patient population by performing a clinical exome analysis. Our observations suggest that both (1) autoimmune and/or innate autoimmune autophagy vulnerability and (2) metabolic disorders can influence the risk to develop IBD in children. Despite the limited number of patients analyzed, our study promotes the use of next generation sequencing as a reasonable and reliable tool to identify genetic risk factors which can help IBD diagnosis in children.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322211055285 – Supplemental material for Genetic Variants Assessing Crohn’s Disease Pattern in Pediatric Inflammatory Bowel Disease Patients by a Clinical Exome Survey
Supplemental material, sj-docx-1-bbi-10.1177_11779322211055285 for Genetic Variants Assessing Crohn’s Disease Pattern in Pediatric Inflammatory Bowel Disease Patients by a Clinical Exome Survey by Dago Dougba Noel, Pinelli Marinella, Giacomelli Mauro, Serena Ilaria Tripodi, Alessia Pin, Arrigo Serena, Bramuzzo Matteo, Fuoti Maurizio Giuseppe, Alvisi Patrizia, Calza Stefano, Alberto Tommasini and Badolato Raffaele in Bioinformatics and Biology Insights
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants NET-2013-02355002 and RF-2016- 02362384 from Ministero della Salute (Italia) to RB. NDD and SC were supported by research grants from the Italian Ministry of University (PRIN projects n. 20178S4EK9).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Prof. BR set up the experiment. Dr DDN wrote the paper. Prof. BR revised the manuscript. Dr MP prepared the DNA library for the exome sequencing. Dr DDN performed bioinformatics and biostatistical analysis. Dr GM give a contribution interpreting genetic and genomic results. Dr SIT provided an assistant in understanting patients clinical features. All co-authors read and approved final version of the paper.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
