Abstract
Background:
A recent COVID-19 pandemic has resulted in a large death toll rate globally and even no cure or vaccine has been successfully employed to combat this disease. Patients have been reported with multi-organ dysfunction along with acute respiratory distress syndrome which implies a critical situation for patients and made them difficult to breathe and survive. Moreover, pathology of COVID-19 is also related to cytokine storm which indicates the elevated levels of interleukin (IL)-1, IL-6, IL-12, and IL-18 along with tumor necrosis factor (TNF)-α. Among them, the proinflammatory cytokine IL-6 has been reported to be induced via binding of severe acute respiratory syndrome coronavirus 2 (SARS)-CoV-2 to the host receptors.
Methodology:
Interleukin-6 blockade has been proposed to constitute novel therapeutics against COVID-19. Thus, in this study, 15 phytocompounds with known antiviral activity have been subjected to test for their inhibitory effect on IL-6. Based on the affinity prediction, top 3 compounds (isoorientin, lupeol, and andrographolide) with best scores were selected for 50 ns molecular dynamics simulation and MMGB/PBSA binding free energy analysis.
Results:
Three phytocompounds including isoorientin, lupeol, and andrographolide have shown strong interactions with the targeted protein IL-6 with least binding energies (−7.1 to −7.7 kcal/mol). Drug-likeness and ADMET profiles of prioritized phytocompounds are also very prominsing and can be further tested to be potential IL-6 blockers and thus benficial for COVID-19 treatment. The moelcular dynamics simulation couple with MMGB/PBSA binding free energy estimation validated conformational stability of the ligands and stronger intermolecular binding. The mean RMSD of the complexes is as: IL6-isoorientin complex (3.97 Å ± 0.77), IL6-lupeol (3.97 Å ± 0.76), and IL6-andrographolide complex (3.96 Å ± 0.77). In addition, the stability observation was affirmed by compounds mean RMSD: isoorientin (0.72 Å ± 0.32), lupeol (mean 0.38 Å ± 0.08), and andrographolide (1.09 Å ± 0.49). A similar strong agreement on systems stability was unraveled by MMGB/PBSA that found net binding net ~ −20 kcal/mol for the complexes dominated by van der Waal interaction energy.
Conclusion:
It has been predicted that proposing potential IL-6 inhibitors with less side effects can help critical COVID-19 patients because it may control the cytokine storm, a major responsible factor of its pathogenesis. In this study, 3 potential phytocompounds have been proposed to have inhibitory effect on IL-6 that can be tested as potential therapeutic options against SARS-CoV-2.
Introduction
Recent outbreak of coronavirus disease 2019 (COVID-19), caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) an enveloped RNA virus, primarily infects human respiratory system but may also attack urogenital system, nervous system, digestive system, and circulatory system. 1 In the Nidovirales order, SARS-CoV-2 belongs to Coronaviridae family. Virus was named as corona virus because of the presence of crown like spikes on its outer surface (corona in Latin means crown). 2 In the past 2 decades, thousands of deaths are caused by infectious and lethal coronaviruses including severe acute respiratory syndrome (SARS) coronavirus and Middle East respiratory syndrome (MERS) coronavirus. 3 The recent outbreak of COVID-19 was discovered in late December 2019 in Wuhan, China, where pneumonia of a mysterious cause was initially diagnosed in the patients. 4 To contest this virus, there is an urgent need to explore the SARS-CoV-2 pathogenicity mechanism and how it manipulates the immune system. 5 It has been reported that SARS-CoV-2 infection results in an exaggerated immune response in some patients regulated by excessive release of circulating cytokines termed as cytokine release syndrome (CRS). 6 Cytokine release syndrome is found to be one of the influential reason of major deterioration of COVID-19 patients leading to multiorgan failure. 7 This has also been termed as “cytokine storm” and is mostly indicated by increased levels of interleukin (IL)-6, IL-1, IL-2, IL-10, tumor necrosis factor (TNF)-α and interferon (IFN)-γ. 8 Interleukin (IL)-6 having significant proinflammatory properties plays crucial role in acute respiratory distress syndrome (ARDS), systemic inflammation, pneumonia related to respiratory failure, and many other adverse effects. 9 Interleukin-6 levels are diligently linked to the severity of COVID-19 infection, and its high level has been observed in patients with respiratory dysfunction. 10 Interleukin-6 level is associated with mortality risk while its mean values were observed to be more than 3 times higher in patients with complicated COVID-19 compared with those with noncomplicated disease. 11 Therefore, IL-6 blocking agents have been effectively used for the treatment of patients with hyper inflammatory states and also approaches intended at impeding this cytokine have been rapidly endeavored. 12
Many plant-derived natural compounds namely phytochemicals might provide a preliminary opinion for the use of plant extracts in IL-6 inhibition. Phytochemicals can provide an affluence of chemical diversity, with anti-inflammatory actions, and thus may have effectiveness as therapeutic agents against COVID-19 infection. Plant could provide different and cost-effective source of drugs that can standardize IL-6 levels. 13 Presently, there is a converted interest in the hunt of new phytochemicals with anti-inflammatory activity to reduce the menace of COVID-19. Numerous studies accentuate the importance of phytochemicals as an anti-inflammatory agent thus inhibiting the level of IL-6 in various coronavirus diseases. The use of phytochemicals as anti-IL-6 may thus be an obliging strategy for reducing the side effects of COVID-19. 14
This study has been designed to elucidate the potential of potent antiviral phytocompounds as IL-6 inhibitors. During infection, there is a rapid activation of T-lymphocytes into pathogenic T-helper cell and produce granulocyte-macrophage colony-stimulating factor (GM-CSF). This environment of cytokines may induce inflammatory monocytes with an increased expression of IL-6 as well as stimulate inflammation. Then these inflammatory monocytes and pathogenic T-helper cells in large number may enter in pulmonary circulation and damage lung functions which causes difficulty in breathing and may lead to mortality. 15 All observations suggest that severe lungs pathology may associated with excessive noneffective immune response. Thus to block or inhibit inflammatory/cytokine storm IL-6 may play very effective role and can be a promising treatment for the patients of COVID-19 infection.16,17 In this study, phytochemicals isoorientin, lupeol, and andrographolide are found to be potential phytocompounds to inhibit or regulate the IL-6 and targeted 3-dimensional (3D) structures of protein was retrieved from Protein Data Bank (PDB). The plant-based phytocompounds were survey from literature and then selected as antiviral compounds The aim of the in silico study is to identify the efficacy of (isoorientin, lupeol, and andrographolide) production that can be tested as potential candidates against COVID-19 infection.007A
Materials and Methods
Sequence retrieval
Sequence and structure of IL-6 were retrieved from PDB under PDB iD (1alu). Furthermore, the retrieved model of targeted protein was subjected to UCSF Chimera 1.12 to detach and remove water molecules from the protein, thus preparing it for docking. In addition, energy minimization was performed by UCSF Chimera using steepest descent approach with conjugate gradient 1000 runs having Amber force-field. 18
Phytocompound selection
After extensive literature, 15 phytocompounds (8-epicatechin, afzelechin, andrographolide, azadirachtin, catechin, isoorientin, isovitexin, eucalyptol, kaempferol, lupeol, marmin, melanoxetin, nimbin, quercetin, and wedelolactone) were selected based on their antiviral activity. 19 The 2D structures of selected phytocompounds were drawn by ChemDraw and retrieved as PDB format. 20 Energy minimization was performed against all selected phytocompounds using UCSF Chimera 1.12. Furthermore, the top 3 phytocompounds were selected on the basis of binding affinities (Kcal/mol). The Lipinski’s rule of 5 was determined by using mCule server. 21 ADMET (Absorption, Distribution, Metabolism, Excretion, and Toxicity) properties were calculated by admetSAR server. 22
Interaction studies
Molecular docking studies were executed to identify the protein-ligand interactions and binding conformational behavior within active pocket of targeted protein IL6. Before docking experiments, active sites of IL6 were identified by literature 23 and online tool using Galaxy binding site server. 24 Grid was set around the active site residues of selected protein. AutoDock Vina was used to perform docking experiment. 25 The 100 models were generated to observe the best binding conformations. The generated docked complexes were visualized and analyzed by UCSF Chimera 1.12 and Discovery studio, 26 respectively.
Molecular dynamics simulations
To obtain detailed dynamics and mechanistic analysis, all docked complexes were subjected to molecular dynamics simulation using AMBER 14 simulation package. 27 Initial libraries were generated using Antechamber program for all 3 compounds. To integrate the docked complexes in TIP3P water box of size 12 Å, along with ff14SB force field (which is required to describe the molecular interactions within the system), leap program was used. To neutralize the systems, counter ions were added to the hydrated complexes. Gradually, energy of the systems was minimized, resulting in relaxation of all hydrogens for about 500 cycles followed by 1000 cycles of water box minimization with a restraint of 200 kcal/mol Å2. The systems were then again minimized for 1000 cycles with a restraint of 5 kcal/mol Å2 on carbon alpha atoms. After that, nonheavy atoms were minimized for up to 300 cycles with a restraint of 100 kcal/mol Å2. Once, minimization accomplished, systems were then heated to 300 K for 20 picoseconds (ps) with a time step of 2 femtoseconds (fs) and 5 kcal/mol Å2 restraint on carbon alpha atoms. Langevin dynamics have been used to maintain the overall temperature by keeping the gamma ln value of 1.0. 28 To constrain bonds with hydrogen atoms, SHAKE was employed, whereas heating the system was accomplished by Canonical ensemble (NVT). 29 Complexes were then again equilibrated for 100 ps with 2 fs time steps along with SHAKE for H-bonds, NPT to maintain pressure (utilizing isotropic positions) and Langevin dynamics for temperature scaling. 30 For another 50 ps, same parameters were used to reduce the carbon restraint by 1 kcal/mol Å2 after every 10 ps. Afterward, systems equilibrate itself for 1 ns using the same conditions. NVT ensemble coupled with Berendsen temperature coupling algorithm was used for production run. 31 SHAKE was again used to apply on hydrogen bonds with a threshold value of 8.0 Å for nonbonded interactions. For 50 ns, production run was carried out with 2 fs time intervals. The initial velocity in each of the simulation run was kept random. The simulation trajectories were analyzed using CCPTRAJ program 32 of AMBER package and were visualized in visual molecular dynamics (VMD) version 1.9.3, 33 and depictions were captured. 34 Root mean square deviations (RMSD) and root mean square fluctuations (RMSF) analysis of complexes were also performed to evaluate systems stability.
Binding free energy calculation
Binding affinities of the optimal docking conformations obtained as a result of molecular docking were calculated by using MMGB/PBSA method. 35 The AMBER MMPBSA.py model was selected to perform the calculations. 35 Four protein-inhibitor complexes and their binding modes were used as representatives to calculate binding free energy employing Prime v3.5 using following formulas:
where ΔE MM, ΔG sol, and −TΔS represents the gas-phase MM energy, the solvation free energy, and conformational entropy, respectively. Internal ΔE having all bond, angle, and dihedral energies along with electrostatic internal energies (ΔEele) and the van der Waal’s (ΔEvdw) were considered in ΔEMM. ΔG was the sum of electrostatic solvation energies, whereas GB or PB models were used to calculate the polar contribution. The nonpolar energy was assessed by evaluating solvent accessible surface area (SASA). As entropy calculation is computational expensive, therefore, it was not included in the study.
Results
Structural and functional evaluation of phytocompounds
The basic biochemical, ADMET properties and the chemo-informatics analysis were assessed to check their therapeutic behavior. Supplementary Table 1 shows that isoorientin, lupeol, and andrographolide have molecular masses of 434.34, 426.71, and 350.44 g/mol, respectively, comparable to standard value (<5000 g/mol). Moreover, the Lipinski’s rule of 5 (RO5) analyses showed that phytocompounds (wedelolactone, catechin, and quercetin) possess better HBA and HBD values which are significantly justified their drug like behavior (Supplementary Table 1).
Interactive residues of prioritized phytocompounds with IL-6.

Interaction analysis
Molecular docking is best computational approach to explore the active sites of protein and conformational position of ligands within active pocket of targeted protein. The generated docked complexes from molecular docking studies were examined on the basis of binding affinities (Kcal/mol), molecular interactions as well as bonding interactions. The lowest binding energy value depicts the best conformational position of ligand within the active region of targeted protein IL-6. The docking results showed 3 phytochemicals, including isoorientin, lupeol, and andrographolide show best interactions with the active site of IL-6. The best possible orientations of 3 prioritized phytocompounds have been shown in Figure 1. The interactive residues of each compound with IL-6 are listed in Table 1. All the 3 prioritized phytocompounds including isoorientin, lupeol, and andrographolide were found to bind at the same active region of protein IL-6. Key residues in protein-ligand docked complex which can play key role in the inhibition of viral activity have also been demonstrated. In addition, the hydrogen and hydrophobic analysis of molecular interactions were further analyzed on the basis of binding interactions and behaviors of receptor-ligand complex. In isoorientin-IL6 docked complex, couple of hydrophobic bonds (Leu-147, Lys-144 with distance bond length 3.68 Å and 4.20 Å respectively) and single hydrogen bond were observed at Leu-140 with 1.93 Å distance (Figure 1A). Similarly, Leupeol-IL6 docking complex, couples of hydrophobic interactions were identified at Leu-147, Leu-151 with distance bond length 5.39 Å and 5.20 Å, respectively (Figure 1B). Likewise, in docked complex andrographolide-IL6 results showed that a single hydrogen bond (Ser-91 with bond distance length 2.18 Å) was observed with 2 hydrophobic interactions (Leu-147, Leu-151 with bond length 5.10 and 5.20 Å, respectively) (Figure 1C). The compounds interact with different sites of IL6 which are involved in direct or indirect contacts with the IL-6 receptor (α-chain) and gp130 (β-chain) which might act as potential blockers of functional hexameric IL-6 receptor complex. 36

Docked complexes and interacting residues of IL-6 with (A) isoorientin, (B) lupeol, and (C) andrographolide. Red dotted lines represent hydrogen bonds and blue lines show hydrophobic bonds.
Three phytocompounds including isoorientin, lupeol, and andrographolide exhibited good interactions with IL-6 protein and were further analyzed to check their binding behavior against target protein. Their 2D structures have been represented in Figure 2. Isoorientin showed better binding affinity against IL-6 as compared to all other phytocompounds with the binding energy of −7.7 Kcal/mol.
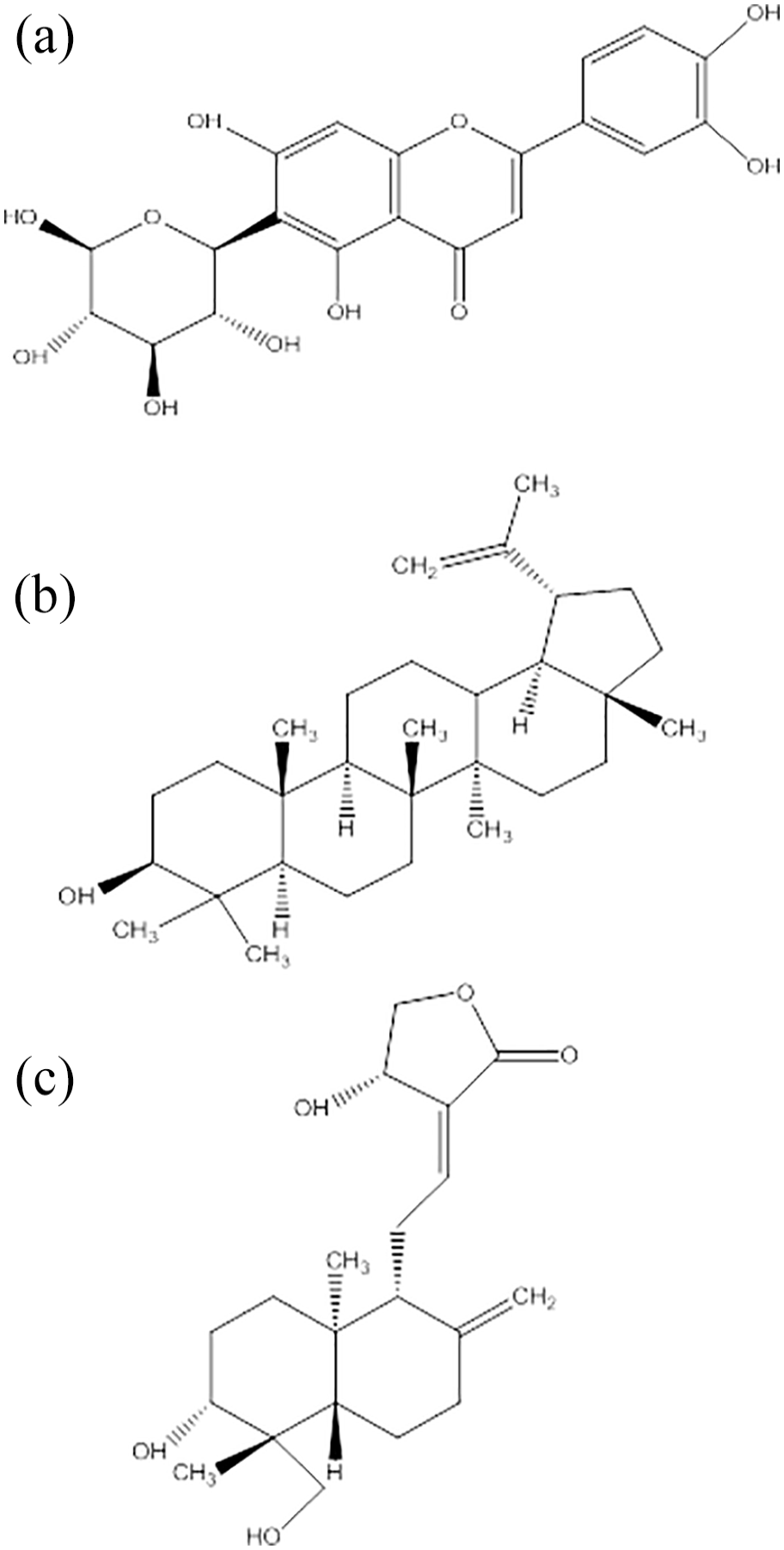
2D chemical structures of prioritized phytocompounds (A) isoorientin, (B) lupeol, and (C) andrographolide.
Molecular dynamics simulation
The conformational stability, structural integrity, and dynamics of all 3 complexes were elucidated by running MD simulation for the period of 50-ns at 300 K. First, all atom carbon alpha atoms RMSD compared to the first snapshot which was treated as a reference was estimated that revealed the mean RMSD values as follow: IL6-isoorientin complex (3.97 Å ± 0.77), IL6-lupeol (3.97 Å ± 0.76), and IL6-andrographolide complex (3.96 Å ± 0.77). A very stable nature of dynamics of the complexes was noticed, which demonstrated strong strength of intermolecular interactions (Figure 3A). In addition, compounds binding mode stability was further affirmed by assaying ligand RMSD (Figure 3B). Isoorientin (maximum, 1.6 Å, and mean 0.72 Å ± 0.32) and lupeol (maximum, 0.80 Å and mean 0.38 Å ± 0.08) binding conformation was found as highly stable, whereas andrographolide (maximum, 1.96 Å, mean, 1.09 Å ± 0.49) was reported to show conformational flexibility. Visual inspection demonstrated that compound achieved a more stable binding mode than the original predicted by molecular docking. Once, a new stable pose achieved the compound was noticed to have stable conformation till rest of the simulation time (Figure 4). Next, radius of gyration (Rg) was calculated for all 3 complexes to look for IL6 structural compactness: higher Rg indicates less tight packing of secondary structure elements and vice versa (Figure 3C). The mean Rg values for complexes are as follows: IL6-isoorientin (26.64 Å ± 0.713499), IL6-lupeol (27.26 Å ± 0.86), and IL6-andrographolide (27.05 Å ± 0.83). These values reflect highly packed nature of the systems and thus very stable behavior of the receptor in the presence of compounds.

Simulation trajectories analysis: (A) receptor RMSD versus time, (B) inhibitors RMSD verses time, and (C) Rg versus time.

Andrographolide binding conformation at the start (tan) and end (sky blue) of simulation time. The receptor can be depicted in dark olive green.
Estimating MMGB/PBSA binding free energy of complexes
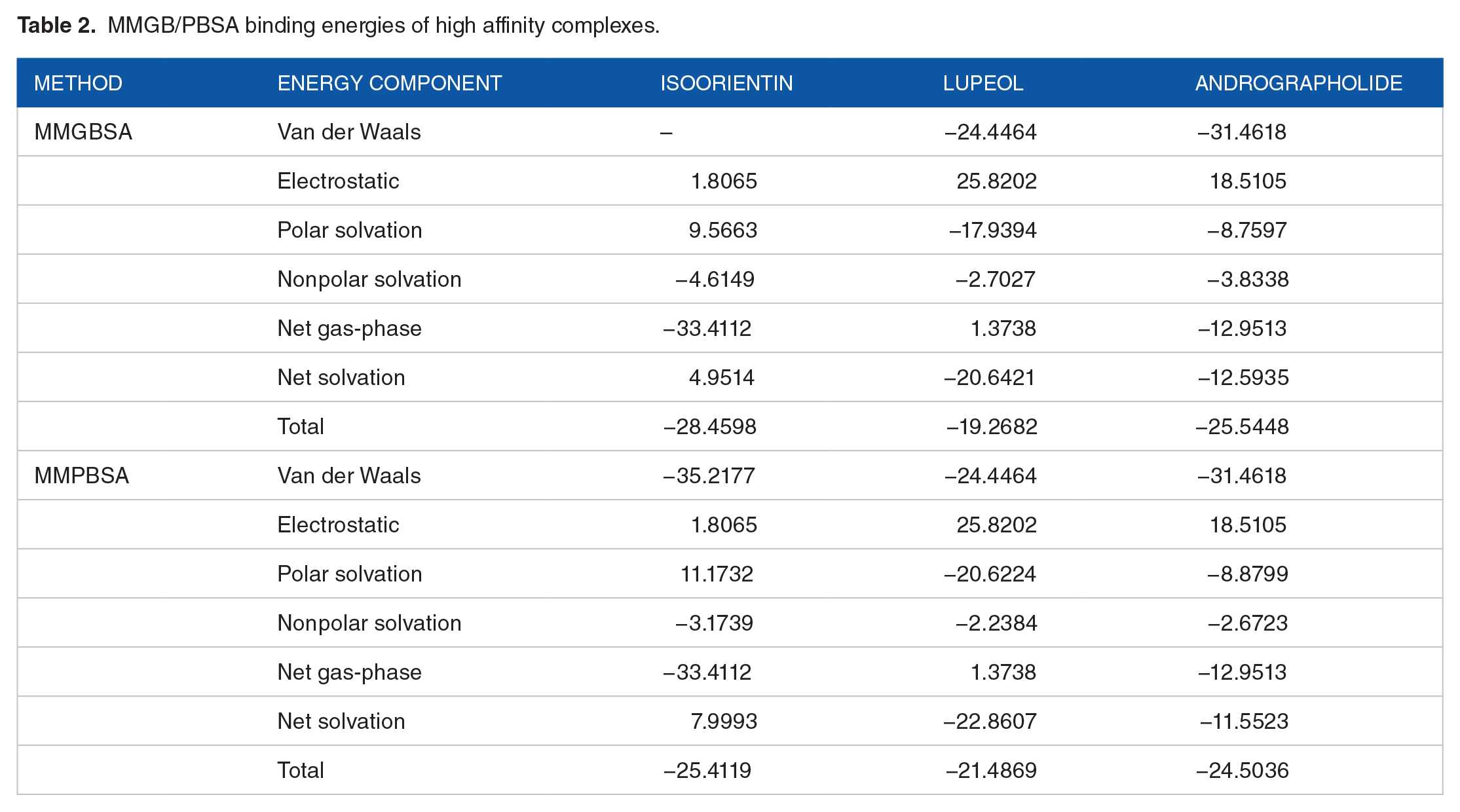
Quantitative assessment of the compounds binding to the IL6 was estimated using the MMGB/PBSA methods (Table 2). Van der Waals’s energy of the molecular mechanic’s force field of all 3 complexes in both approaches reports to be favorable and contributes significantly to the protein’s docked conformation stability. The electrostatic energy is insignificant in compounds, and IL6 interaction, therefore less important in the binding strength. The polar energy of isoorientin in both MMGBSA and MMPBSA participates less in binding while contributing considerably to lupeol and andrographolide binding. Compound isoorientin was rescored to have favorable lower binding energy in the gas-phase (−33.4112 kcal/mol in both MMGBSA and MMPBSA) in contrast to nonfavorable delta solvation energy (4.9514 kcal/mol in MMGBSA and 7.9993 in MMPBSA). The net energy of IL6-isoorientin complex is −28.4598 kcal/mol in MMGBSA and −25.4119 kcal/mol in MMPBSA. IL6-Lupeol complex, on the other side, illustrated favorable solvation energy and not gas-phase energy, and IL6-andrographolide complex demonstrated that both gas-phase energy and solvation energy are essential in compound binding to the receptor. The IL6-lupeol complex’s net binding energy is −19.2682 kcal/mol in MMGBSA and −21.4869 in MMPBSA, and andrographolide complex is −25.5448 kcal/mol in MMGBSA and −24.5036 in MMPBSA.
MMGB/PBSA binding energies of high affinity complexes.
Discussion
Through the membrane receptor cells, tissues carry out their specific functions after stimulation. Frequently such receptors are not only used to transmit signals regarding cell cycle activation and progression but may also mediate the signal of cell apoptosis before it gets a chance of differentiation and proliferation. A classic example can be of lymphocytes. Interaction of receptor and antigen results in initiation of signaling cascade which leads to cell differentiation and proliferation when costimulatory signal provided simultaneously. If apoptosis is not inhibited/stopped, it will result in cell death. Mature T-cells attain costimulatory device, if there would be an inhibition of co-stimulation ligation of antigen to receptor results in mature T-cell death. Apparently, effective activation signals require effective inhibition of cell death. 37 In viral clearance T-cells play very vital role with cytotoxic T-cell that may secrete different molecule and IFN-γ to remove virus from the host. 38 Meanwhile to clear the pathogen, helper T-cells promote the ability of B-cells and T-cells. 39 Although virus-mediated continuous stimulation may induce exhaustion of T-cells and results in reduced capability of cytokines production as well as function. 40 In COVID-19 patients decrease in both total T-cell, cytotoxic T-cells, and helper T-cells has been observed. 41
In response to infection, cytokines production increases and this phenomenon of excessive inflammatory reaction known as cytokine storm may contribute to develop ARDS and multiple organ dysfunction syndrome as well as respiratory viral infection setting.34,42 SARS-CoV-2 has been observed to significantly increase the level of IL-2, IL-7, IL-10, TNF-α, G-CSF, IP-10, MCP-1, and MIP-1A. 43 Consistent with this, Diao et al 41 also found the increase concentration of different cytokines such as TNF-α, IL-6, and IL-10 in COVID-19 patients. All these observations suggest an inverse correlation between the number of T-cells and the level of IL-6, IL-10, and TNF-α, also suggest that these cytokines stimulate T-cell reduction in COVID-19 patients. A potent pyrogenic cytokine IL-6 produced in response to infection or tissue damage has an essential role to modulate host immune response as well as it has a critical role in the progression of virus. 44 During an infection, IL-6 might be considered as important cytokine along with IL-1 and TNF-α. 45 Recent studies describe that severity of disease and its outcomes in COVID-19 infected patients are associated with characteristics of immune response. Various components of inflammatory cascade and IL-6 induce host defense against infection. Although increased production of IL-6 may result in acute severe systemic inflammatory response named as cytokine storm. Moreover, an important proinflammatory cytokine like TNF-α may interact with its specific receptor and results in T-cell apoptosis, as increased expression of receptor for TNF-α has been seen in aged T-cells. Various studies demonstrate low number of T-cells in patients over 60 years which indicates that in these patients TNF-α can be directly responsible for reduction of T-cells in these patients. 27 Similarly, another study found elevated level of different cytokines such as IL-2, IL-6, IL-7, IL-10, interferon-γ-inducible protein, granulocyte-colony stimulating factor, macrophage inflammatory protein 1 alpha, and TNF-α. 30 These observations suggest that cytokine response magnitude and characteristics are associated with the pathogenesis of disease. Thus, it has been shown that dysregulated and persistent release of IL-6 is associated with pathogenicity of infection. Blockage of IL-6 has been shown to be effective against this infection.31,46 As it is hypothesized that IL-6 may play very vital role in serious adverse outcomes in patients infected with SARS-CoV-2 pneumonia, as well as the inhibition of IL-6 could be an appropriate therapeutic target for these patients. 47
Computational approaches can provide rapid and reliable solutions drug discovery and have been contributing significantly in the field since long.48,49 Various studies suggest that use of bioinformatics analysis and techniques can decrease the risk factor and cost related to novel drug discovery. 50 They have number of benefits as compared to conventional approaches such as time saving, efficiency, less cost, and also identify the side effects of drug utilizing different bioinformatics tools. 51 In current years, due to the advance advantages of herbal medicines, interests of pharmaceutical industry in natural products are increased. 52 Meditational plants are best source for dietary supplements, while its active compounds of plants play important role in treatment, preventing from number of diseases, and they are more effective as compared to pharmaceuticals drugs. 53 Previous studies showed that plant-derived phytocompounds have been effective activity against viral diseases. 54
In this study, interaction analysis of 15 phytocompounds (8-epicatechin, afzelechin, andrographolide, azadirachtin, catechin, isoorientin, isovitexin, eucalyptol, kaempferol, lupeol, marmin, melanoxetin, nimbin, quercetin, and wedelolactone) has been performed against targeted protein (IL-6). The basic idea was to discover a potent and efficient IL-6 inhibitor with antiviral activity from the prioritized phytocompounds with least side effects. Three phytocompounds showed strong antiviral activity on the bases of strong interaction, drug-likeness properties (Table 3), and binding affinity with IL-6 (Supplementary Table 1). The interaction analysis and binding affinities proved that all top 3 phytocompounds are effective against targeted protein IL-6.
ADMET properties of top 3 selected phytocompounds.

Prioritized phytocompounds against targeted protein IL-6 may inhibit the activity of viral mechanism thus reducing the pathogenicity of SARS-CoV-2. Molecular docking studies of active compounds and drug-likeness properties proved that these compounds are strong candidate for antiviral drug against corona virus. Even among the prioritized phytocompounds, lupeol and andrographolide have more favorable results to be used as therapeutic compounds than isoorientin, but still all these are computational predictions and need experimental validations.
Conclusions
The current in silico study is performed to recognize the therapeutic effect of plant-based compounds against corona virus. Three phytocompounds including isoorientin, lupeol, and andrographolide have been prioritized, which can display effective antiviral characteristics against SARS-CoV-2.
Supplemental Material
sj-pdf-1-bbi-10.1177_11779322211021430 – Supplemental material for Inhibitory Potential of Phytochemicals on Interleukin-6-Mediated T-Cell Reduction in COVID-19 Patients: A Computational Approach
Supplemental material, sj-pdf-1-bbi-10.1177_11779322211021430 for Inhibitory Potential of Phytochemicals on Interleukin-6-Mediated T-Cell Reduction in COVID-19 Patients: A Computational Approach by Arif Malik, Anam Naz, Sajjad Ahmad, Mansoor Hafeez, Faryal Mehwish Awan, Tassadaq Hussain Jafar, Ayesha Zahid, Aqsa Ikram, Bisma Rauff and Mubashir Hassan in Bioinformatics and Biology Insights
Footnotes
Acknowledgements
The authors acknowledge the IMBB department of The University of Lahore, Lahore campus for their administrative support.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Authors’ Note
Sajjad Ahmad is now affiliated to the Department of Health and Biological Sciences, Abasyn University, Peshawar, Pakistan.
Author Contributions
AM and AN designed the study. AN along with MH, SA, MH, FMA, THJ and AZ carried out the computational analysis and designed the first draft. AI, MH and BR helped in improving and revising the manuscript. All authors read and agree to all the data provided in the manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

