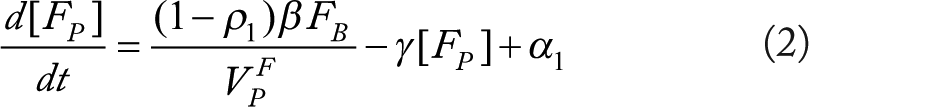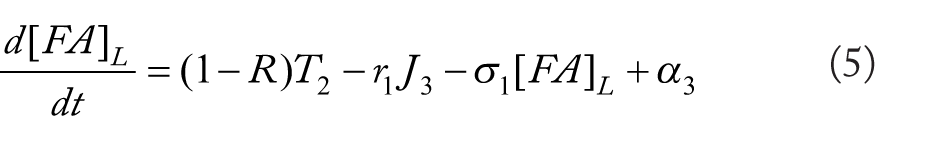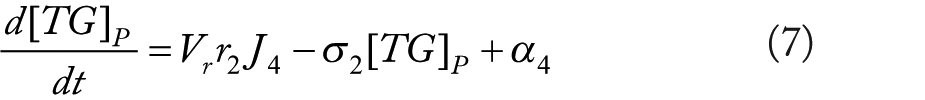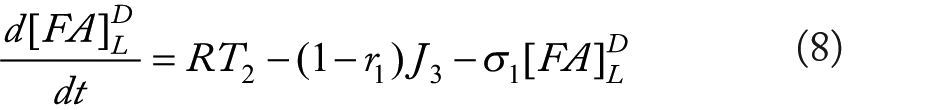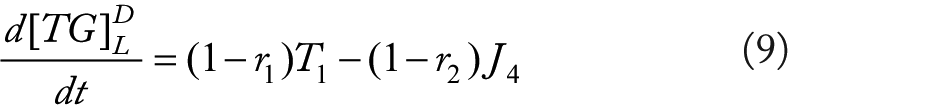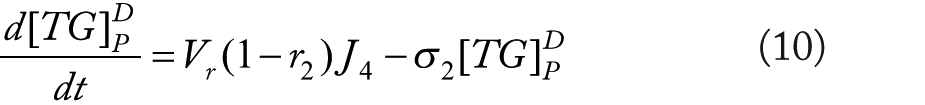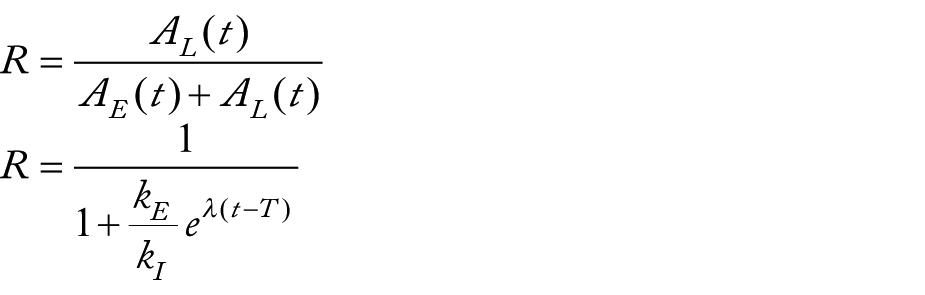Abstract
Hepatic de-novo lipogenesis is a metabolic process implemented in the pathogenesis of type 2 diabetes. Clinically, the rate of this process can be ascertained by use of labeled acetate and stimulation by fructose administration. A systems pharmacology model of this process is desirable because it facilitates the description, analysis, and prediction of this experiment. Due to the multiple enzymes involved in de-novo lipogenesis, and the limited data, it is desirable to use single functional expressions to encapsulate the flux between multiple enzymes. To accomplish this we developed a novel simplification technique which uses the available information about the properties of the individual enzymes to bound the parameters of a single governing ‘transfer function’. This method should be applicable to any model with linear chains of enzymes that are well stimulated. We validated this approach with computational simulations and analytical justification in a limiting case. Using this technique we generated a simple model of hepatic de-novo lipogenesis in these experimental conditions that matched prior data. This model can be used to assess pharmacological intervention at specific points on this pathway. We have demonstrated this with prospective simulation of acetyl-CoA carboxylase inhibition. This simplification technique suggests how the constituent properties of an enzymatic chain of reactions gives rise to the sensitivity (to substrate) of the pathway as a whole.
Introduction
De-novo lipogenesis (DNL) is a complicated hormonally regulated pathway that is active in humans in liver and adipose tissue, 1 and to a lesser extent in skeletal muscle. 2 This pathway is one mechanism for converting carbohydrates into fat, which is eventually exported, or stored, as triglycerides (TGs). Hepatic DNL has been implicated in the pathogenesis of diabetes and hepatic steatosis.3,4 In type 2 diabetics, where homeostatic mechanisms fail to control the level of glucose and gluconeogeneic precursors, 5 hepatic DNL may act as a way of diverting these substrates away from gluconeogenesis and hepatic glucose output. 6 However, the increased lipid load is an independent risk factor for cardiovascular disease. 7
A mechanistic, quantitative, understanding of the relationship between substrate load and lipid accumulation via DNL is fundamental to fully appreciating the role of DNL in disease. Fructose is a lipogenic dietary substrate that is up-taken into hepatocytes rapidly on the first pass through the liver. 8 Hence, one technique to stimulate and assay DNL is repeated oral dosing of fructose in the presence of exogenous C14 labeled acetate. Then DNL is measured as the fraction of plasma TGs that have incorporated C14 acetate, using the MIDA method. Hudgins et al. 9 reported that this technique increased de-novo lipids in the plasma by approximately 15%. The purpose of this paper is to develop and use a mathematical ‘systems pharmacology’ model of this protocol to gain a quantitative understanding of the process. This will allow us, in the future, to predict the effect of putative pharmacological agents in this context.
The development of systems pharmacology models is a time-consuming process. This is because they attempt to faithfully represent key physiological mechanisms mathematically. Hence, they are often large and complex, requiring data from multiple species and experiments. A crucial challenge within the development process is accommodating a balance between mechanistic detail and available experimental knowledge to inform that detail. Therefore, we believe that tools which resolve this tension by systematic simplification of models are particularly valuable. Often the reduction to simplified representations occurs in the development and design phase of the process. In this case it is usually difficult to retain the correspondence to physiology and experimental data.
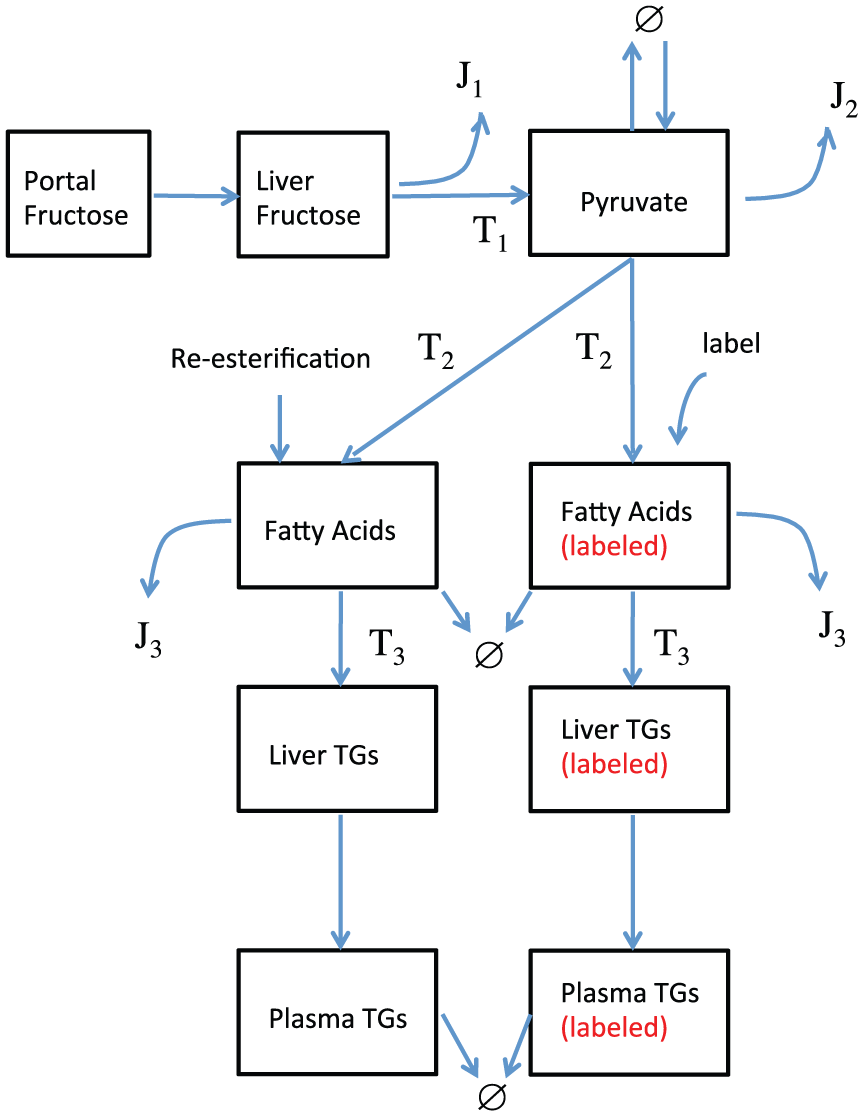
In the protocol applied by Hudgins et al., 9 it is plausible to assume that metabolism of fructose to plasma TGs can be well approximated by a linear network (Figure 1). The most direct route from fructose to plasma TGs involves 16 intermediaries: see the supplementary file. However, even in a model where each reaction is described by Hill kinetics this will give rise to at least a 32 parameter model, whereas if we seek to simply retain the boxed species (Figure 1), and construct transfer functions between them, the size of the model will be reduced dramatically. As such, it is desirable to design a simplified representative model of this process incorporating key mechanistic features and known experimental data regarding enzyme properties of the individual steps, while not modeling each enzyme explicitly.
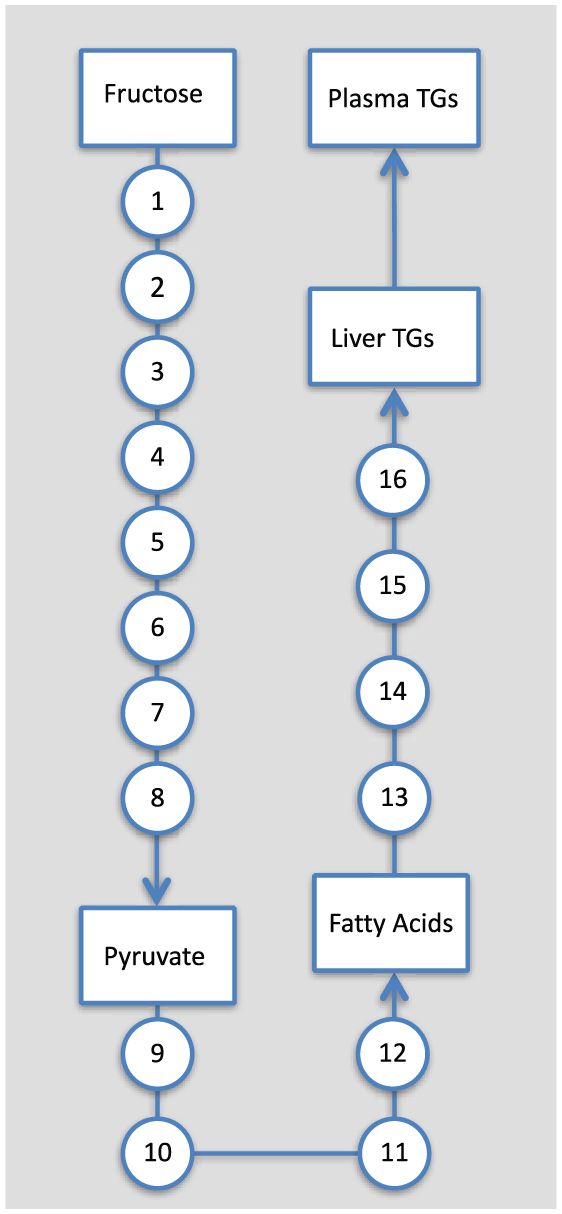
The shortest route from fructose to plasma triglycerides (TGs) includes 16 steps (see Supplementary Table 2). In the experimental conditions of Hudgins et al., 9 where hepatic fructose exposure is high for a period of approximately 8 hours, we assume that the overwhelming majority of fructose-derived TGs has been synthesized by this pathway.
Model reduction will often be an esoteric case-dependent exercise. However, there are some general approaches that have been developed. For example, one way to simplify a model is to separate time-scales, so that interactions faster than the time-scale of interest can be assumed to be in a quasi steady-state.10–12 Another approach is to reduce parameters based on Hankel singular values, and/or singular value decomposition. 13 This method ranks the parameters based on their effect on the input–output response of the model. The approach we are proposing here is a novel contribution because it uses available experimental information to establish approximate values for lumped parameters that otherwise do not have a direct biochemical and experimental meaning.
An obvious way to simplify a model is to compact linear-chain motifs (synonymously, path graph or linear tree) of a network into the input and output of the motif. If we believe the intermediaries of this graph are not rate-limiting and we do not have time-course data for those nodes, then all we are really interested in, or can constrain, is the ‘transfer function’ from the first to the last node (Figure 2).
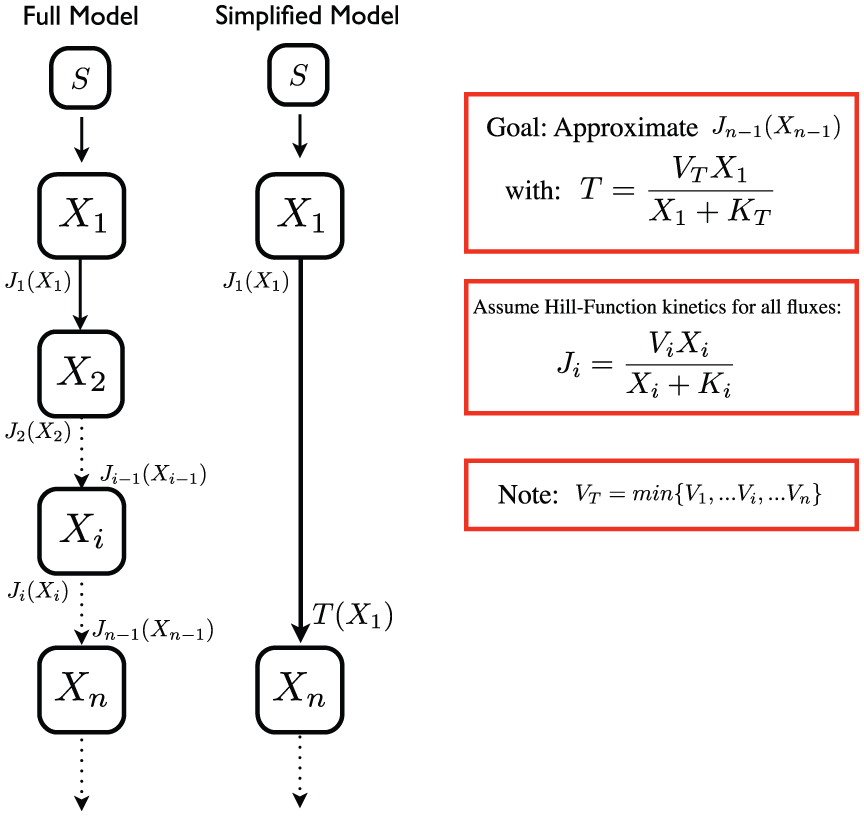
Approximating linear chains with transfer functions. Often it is desirable to reduce the ‘full model’ motif, to a ‘simplified model’ motif. In this case, the input species
For enzymatic reaction networks a biologically relevant transfer function is a Hill function, 14 which arises naturally in the case of Michaelis–Menten kinetics. This rate law is typically a reasonable assumption for metabolic networks, but for signal transduction networks, where interactions are more complex (for example, when proteins can be both enzymes and substrates) there are superior approximations. 15 Here we assume that Hill functions (with Hill coefficient one) can capture the experimental dynamics and still be predictive, even if the assumptions required for Michaelis–Menten kinetics are not precisely met. Hill functions, similar to metabolic reactions, are saturable, and can give rise to rich dynamics, 16 which we presume includes the requisite model behavior. Here we consider Hill functions with coefficient one, but our analysis shows that similar relationships to the one we develop will hold true at higher values.
Our approach to simplifying linear-chain motifs uses prior knowledge of the system under consideration. For metabolic pathways it is often the case that the maximum rate of a reaction (
A linear chain of n reactions modeled with Michaelis–Menten kinetics will have 2n parameters. By introducing a transfer function from the first to last node we can simplify this motif to being determined by either two or four parameters, depending on if we assume conservation of mass (so that the flux leaving the first node is matched by flux into the last). In this case, mass is not conserved in the model because it is ‘lost’ to the intermediary substrates that we are not considering. However, due to the linear structure of the pathway, we gain a constraint that the maximum rate,
While this simplification is often necessary, it is still highly desirable to use all prior knowledge to constrain the transfer function. As mentioned, it is unusual for detailed information regarding the
To motivate our approach for simplification we first introduce a novel model of hepatic DNL which includes the aforementioned ‘transfer functions’. Following this we describe our novel approach to parameterizing these functions, which is justified by both computational and analytical rationale.
Modeling hepatic DNL in response to high fructose levels
Model equations
The model detailed in Figure 3 was described mathematically as follows. It was assumed that all reactions could be captured by Hill functions (i.e. can be approximated by Michaelis–Menten-type kinetics). This approximation will not always be valid, and certainly the Michaelis–Menten interpretation of their meaning is not necessarily valid unless specific assumptions are met. We assume that in conditions where the pathway is operating unidirectionally (i.e. highly stimulated at the initial node) that this functional form sufficiently captures the flux from node to node.
Notation
We use F, P,
Rationale for equations (1)–(3)
These equations predict plasma and liver fructose following oral administration of fructose at discrete times given by the set
Rationale for equations (4)–(10)
The flux terms in these expression are defined as outlined in Figure 2, more specifically:
When applied,
Hudgins et al.
9
measured DNL by using C14 acetate, which identifies the de-novo lipids using the MIDA method. In the model we can consider that
where
Parameter estimates
Parameter ranges, units, and final model fits are reported in the supplementary material. Here we detail our strategy for generating estimates for the parameter ranges, and fitting the model.
For equations (1)–(3) parameters
Feasible ranges for the zeroth-order source terms in the model (
Apart from the Michaelis constants in the transfer functions (
If a range was not estimable from experimental data, a reasonably wide-range around the reported value was passed to the optimization routine: this range spanned one, two, or three orders of magnitude, the latter being assigned to the
Computational approach to estimate parameters of the transfer functions
In this section we describe a novel computational approach to estimate a value of
Below we show that with limited assumptions about the nature of the pathway and experimental conditions, a rule can be constructed to estimate the expected value,
It is imperative to point out that the rule developed in this section only provides an estimate, or bounds (depending on prior knowledge), for
Our strategy for approaching this problem was to simulate the full model in Figure 2, and attempt to fit the flux into the final node as function of the concentration of the first node (Figure 2, inset). In this manner we could attempt to establish a relationship between the set
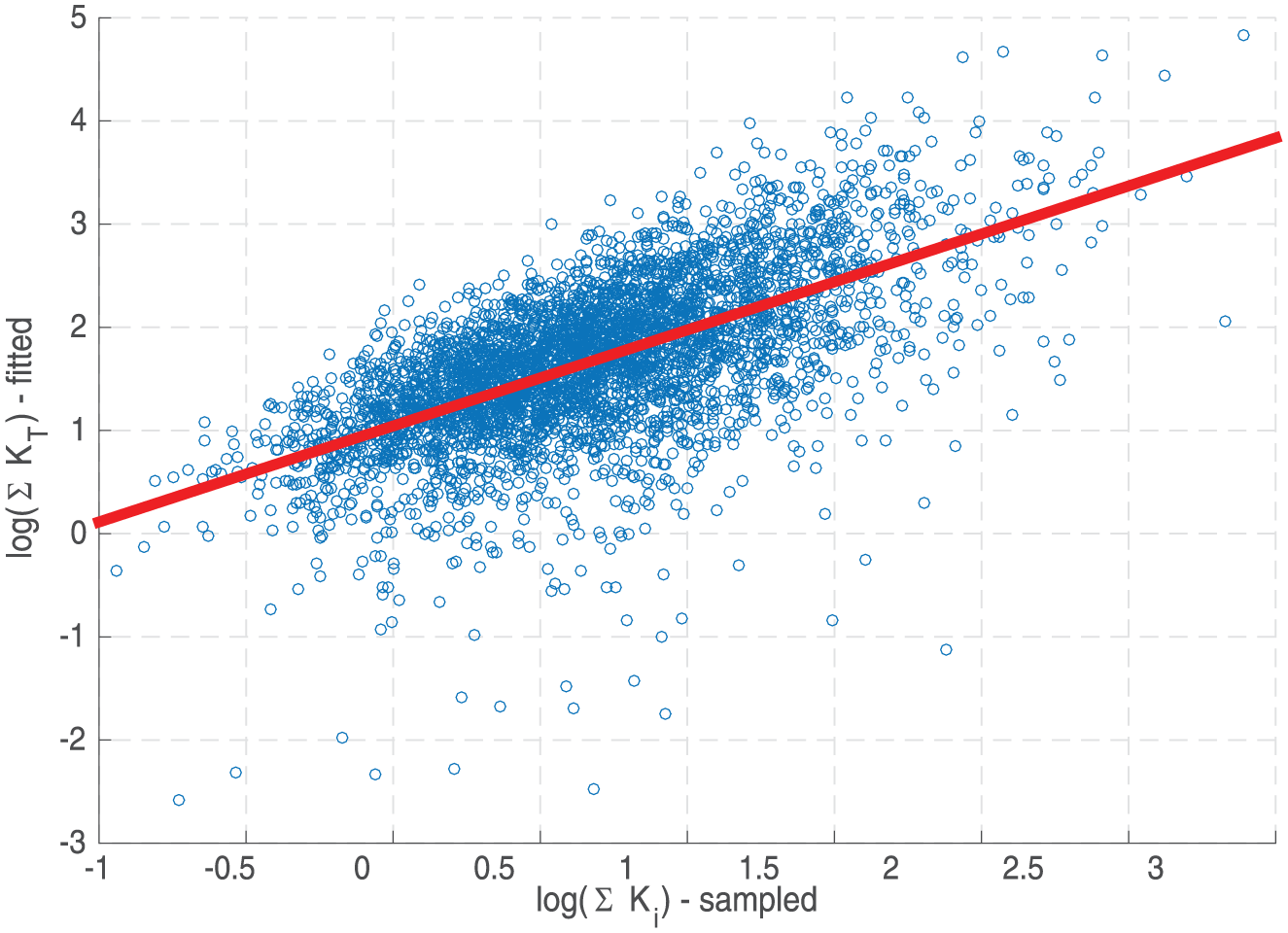
Predicting transfer function kinetic parameters from individual flux parameters. For a high stimulus level and given chain length (
To investigate how robust this relationship is, we repeated this procedure for different lengths of chains (n = 3 to 24) and stimulus strengths (over five orders of magnitude). We noticed that the correlation between
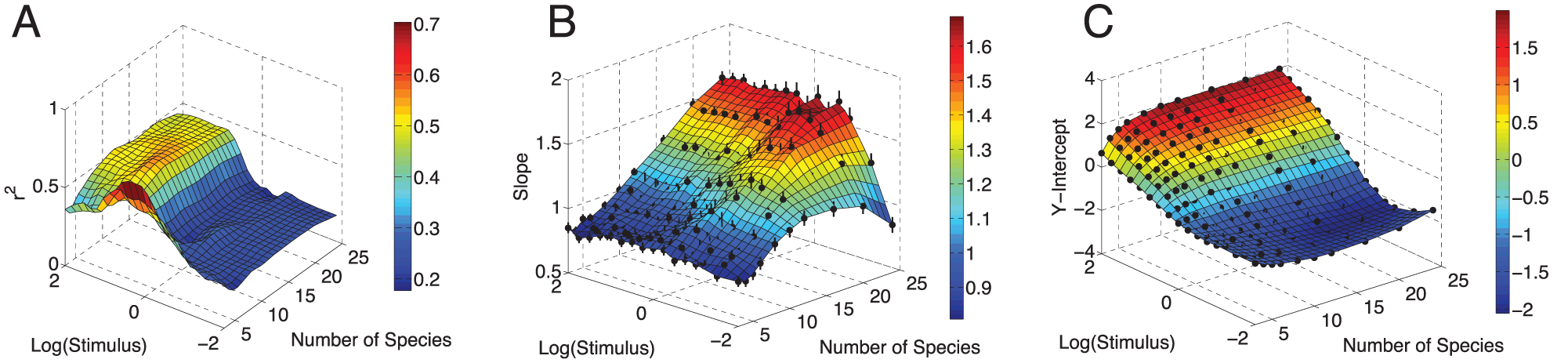
The effect of chain length and stimulus level on the relationship between
In the appendix we provide an analytical analysis of this problem, which provides a direct interpretation of why this relationship holds in a limiting case.
Simplification parameters for the hepatic DNL model
An unknown in applying this technique to the DNL model is the stimulus strength relative to the
Application to and simulation of a model of hepatic DNL
To demonstrate this technique we applied it to the model of hepatic DNL presented in Figure 5. Note that this model represents a significant simplification of the 16 enzyme path (Figure 1). The individual reactions and estimates for their
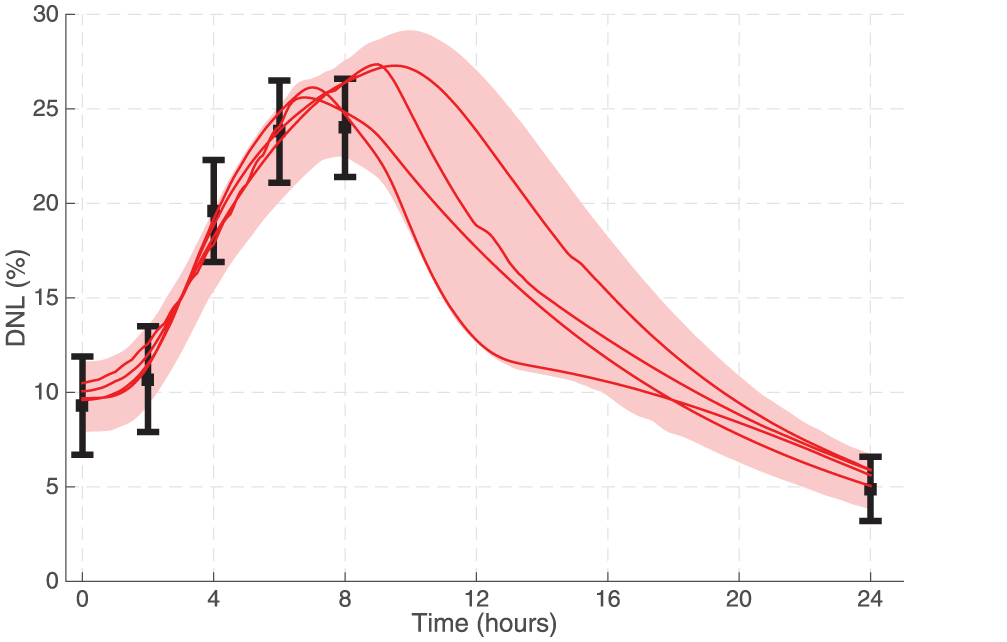
Summary of model performance compared with clinical data from Hudgins et al.9 (black points). Multiple fits (≈1800) were found that could describe the data. The upper-quartile of fits were found to cluster into four distinct groups (cluster centroid, solid red lines). The shaded region summarizes the upper and lower bounds of all fits with outliers removed (defined as the top and bottom percentiles).
Because novel inhibitors of acetyl-CoA carboxylase (ACC) have been developed,27,28 we wanted to investigate the effect of inhibiting ACC in this model. This was simulated by re-running the model with a reduction of parameter
Figure 7 shows the effect of ACC inhibition in the model. Inhibition of ACC is predicted to have a potent effect on fractional DNL as assayed by this clinical methodology. The model also predicts that in a clinical trial the variability in response will be higher for doses that induce 50% inhibition than placebo or higher doses (as shown by wider prediction intervals for the 50% case). This prediction illustrates the utility in generating multiple parameter sets to explore variability in response, and also has implications for how such a study should be conducted. However, this effect could be masked by other sources of variability.
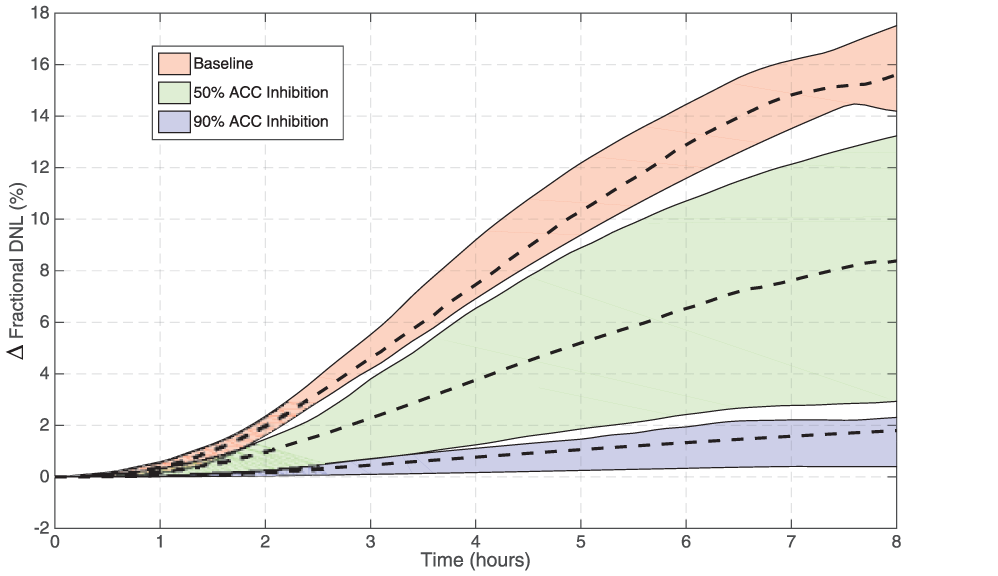
Evaluation of the model response to 50% and 90% inhibition of acetyl-CoA carboxylase (ACC) in comparison with the baseline response (shaded area is the 10–90% prediction interval). The model predicts that the fractional DNL response to fructose stimulation can be effectively modulated by inhibition of ACC. Variability in predicted response at 50% inhibition is significantly larger than the baseline and 90% case.
Since the expression of ACC can be tightly controlled, 30 it is interesting to interpret these results as the effect of modulating ACC expression to 50% or 10% of baseline (mathematically, these are the same simulations as Figure 7). In this interpretation the results supports the hypothesis that regulation of ACC by SREBP1c 30 is a crucial component of the lipogenic regulatory program controlled by SREBPC1 and regulation of ACC expression directly controls the contribution of DNL to the liver and plasma TG pools.
Discussion
Here we have developed a novel systems pharmacology model of hepatic DNL. In particular, this model is designed to support, and analyze, clinical experiments where hepatic DNL is measured. As such, this model could be a powerful tool to assess novel pharmacological agents that may affect DNL. For example it could be used to infer pharmacologically meaningful parameters such as IC50, or
As a focused simulation of a specific experimental protocol we have high confidence in the model when applied in this context, but in other contexts the model would require further validation and refinement before application. However, we believe that within this context the model is a good representation of the underlying biology because of the novel simplification approach we developed. This approach links underlying biochemical constants with a parameter of a lumped representation that hitherto had no biologically interpretable meaning.
The model we have presented here is an acute view of fructose-induced DNL and tracks a major metabolic route of fructose in these conditions. However, fructose metabolism also has a major signaling role which can alter downstream metabolic pathways. These signaling effects include activation of SREBP1c which has a profound effect on the entire lipogenic pathway. 30 Our assumption is that the acute nature of the study allows us to ignore these effects during the course of the simulated trial. However, future work could include analysis and modification of the model to describe patients from a different population, such as those with non-alcoholic fatty liver disease. This population consumes large amounts of dietary fructose and have a higher fasting DNL level. Such an analysis like could facilitate insight to the precise biochemical parameters that are inducing increased DNL in these patients, and how they might respond in this clinical protocol.
Systems pharmacology models are a useful tool to understand biological systems, disease pathophysiology, and the potential effects of pharmacological intervention. As more biological detail is added to a model it becomes less constrained and the balance between the number of parameters and available data becomes less favorable. Therefore it becomes necessary to simplify the model. However, there is a balance between redressing this while retaining enough mechanistic detail to satisfactorily address the biological questions and hypotheses of interest. In this case it is important that all available data is used to establish parameter values.
The model reduction technique applied here simplifies a model while utilizing available data about individual enzyme kinetics. This technique supplements existing approaches of constraining, or reducing, a model based on dynamic data. By bounding the search for lumped parameter values into a biologically and mathematically justified space, physiologically reasonable parameter sets can be identified. Our analysis and simulations show that the Michaelis constant of the transfer function is related to the sum of the individual Michaelis constants and the stimulus level relative to the distribution of
We have focused on this method in order to develop rules for fitting parameters. However, there is a biological interpretation of our result which generates an interesting hypothesis for future testing. We can consider the transfer function as a summary of the kinetics of the pathway. As such,
Footnotes
Appendix
Acknowledgements
We would like to acknowledge Gianluca Nucci (Pfizer Inc.) for his support and motivation of this work. We would also like to thank Theodore Rieger (Pfizer Inc.), and William C. Thompson (formerly Pfizer Inc., currently SAS Institute Inc.) for valuable input and feedback on this work.
Author Contributions
RJA performed the analysis, wrote the manuscript, and approved the final manuscript. CJM reviewed and approved the final manuscript.
Peer review:
Five peer reviewers contributed to the peer review report. Reviewers’ reports totaled 673 words, excluding any confidential comments to the academic editor.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: RJA, and CJM were employees of Pfizer Inc. during the completion and/or analyses of these studies. Pfizer Inc. has provided all funding for these studies and was responsible for the study design, data analysis, decision to publish, and preparation of the manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.