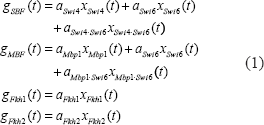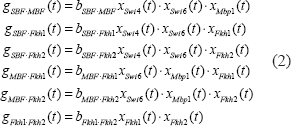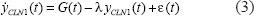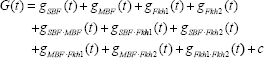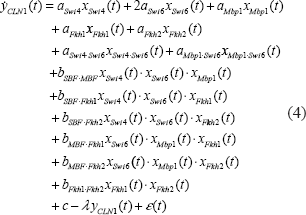Abstract
The trans-regulatory circuit is considered as the regulatory interactions between upstream regulatory genes and transcription factor binding site motifs or cis elements. And the cis-regulatory circuit is viewed as a dynamic interactive circuit among binding site motifs with their effective action on the expression scheme of target gene. In brief, gene transcription depends on the trans/cis regulatory circuits. In this study, nonlinear trans/cis regulatory circuits for gene transcription in yeast are constructed using microarray data, translation time delay, and information of transcription factors (TFs) binding sites. We provide a useful nonlinear dynamic modeling and develop a parameter estimating method for the construction of trans/cis regulatory circuits, which is powerful for understanding gene transcription. We apply our method to construct trans/cis regulatory circuits of yeast cell cycle-related genes and successfully quantify their regulatory abilities and find possible cis-element interactions. Not only could the data of yeast be applied by our method, but those of other species also could. The proposed method can provide a quantitative basis for system analysis of gene circuits, which is potential for gene regulatory circuit design with a desired gene expression.
Introduction
Due to advances in DNA microarray technology and genome sequencing, it has become possible to measure gene expression levels on a genomic scale. Microarray technology provides insight into the transcriptional state of a cell. In recent studies, the expression profiles and motif binding sites of genes in yeast have been revealed, and regulatory networks have been proposed to explain their regulatory functions (Spellman et al. 1998; Iyer et al. 2001; Simon et al. 2001; Hartemink et al. 2002; Lee et al. 2002; Harbison et al. 2004). However, in order to gain more insight into the infrastructure of the regulatory scheme inside the gene transcription, trans/cis regulatory circuit constructed by the view of the systems biology is an important topic.
Genes are always regulated by a number of upstream regulatory genes through their products, for instance, transcription factors bind to specific sites (i.e. cis elements) in the DNA promoter region. The interactions between upstream regulatory genes and cis elements are described by a trans-regulatory circuit (see Fig. 1). The DNA binding motifs, served as anchors to transcription factors, play the role of staying platforms for the assembly of multi-proteins at the output. This mechanism specifies the cis-regulatory circuit, which is viewed as a dynamic interactive circuit among binding site motifs with their effective action on the expression scheme of target gene (Fig. 1). In brief, gene transcription depends on the trans/cis regulatory circuits. Therefore, the comprehension of gene transcriptions needs to recognize the corresponding trans/cis regulatory circuits. Unfortunately, although the cis-regulatory circuit has been widely discussed in Drosophila (Berman et al. 2002) and sea urchin (Yuh and Davidson, 1996; Yuh et al. 1996; Yuh et al. 1998), not much is known about trans/cis-regulatory circuits because of their complicated interactive schemes, which are not easily detected directly by conventional experiments. That is why we want to propose a method to construct trans/cis regulatory circuits to understand gene transcription process thoroughly.
The dynamic trans/cis regulatory model of the gene transcription of target gene CLN1. xi. j (t) denotes trans regulatory function of the complex of the transcription factors i and j and gi. j (t) denotes the interaction between the cis elements i and j.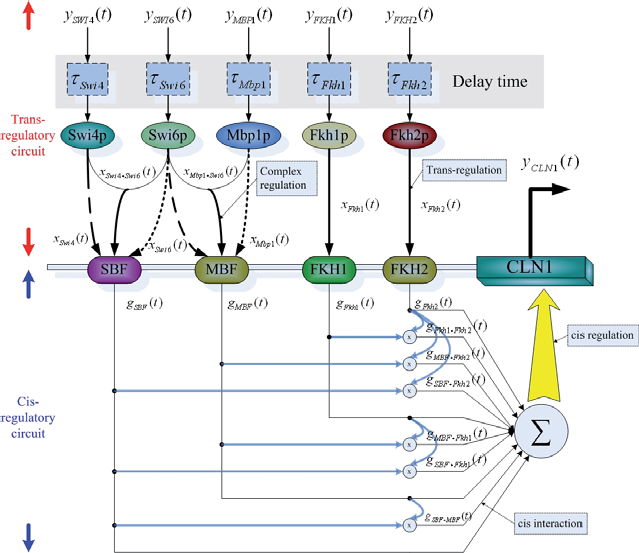
Given a processed data set, one expects to be ready to tackle the biological data interpretation problem. It is popular to use clustering, classification, and projection methods to analyze the data set. However, most analysis methods only use microarray data, not both dynamic expression time profiles of microarray data and the information of motif binding sites (Gardner et al. 2003; Chen et al. 2004; Chen et al. 2005). Furthermore, the interactions among proteins and delay from transcription to translation are not considered in their models of gene transcription. Recently, systems biology and computational biology methods have been widely considered to describe the biological functions from the dynamic system perspective (Yeung et al. 2002; Davidson et al. 2003; Tegner et al. 2003; Chen et al. 2004; Sontag et al. 2004; Chang et al. 2006). Engineering theory has also been used to know more about biological complexity (Carlson and Doyle, 1999; Yi et al. 2000; Csete and Doyle, 2002; Hasty et al. 2002; Gardner et al. 2003; Tegner et al. 2003). Here, we provide a nonlinear dynamic model to gain more insight into trans/cis regulatory circuits for gene transcription using the knowledge of microarray data, the information of motif binding sites, translation delays and protein complexes from the systems biology perspective.
In this study, a nonlinear dynamic model is proposed to describe the kinetics of trans/cis regulatory circuits for gene transcriptions of yeast genes. In the trans circuit part (see Fig. 1), the transcription regulatory functions on cis elements are described by the regulatory activities of transcription factors and reaction of complexes among transcription factors. The binding from upstream regulatory genes to the transcription factor binding sites is described by a biological sigmoid function to model the binding (activating the binding function) and no binding (inactivating the binding function) signals beyond or below some threshold of mRNA concentrations of regulatory genes. The translation time delay from the mRNA of a regulatory gene to transcription factor is also considered in our model. The reactions (cooperations) of complexes of transcription factors are represented by nonlinear interactions (Chang et al. 2006). In this study, nonlinear interactions between two proteins and among three proteins are also considered to mimic the effect of protein complexes on a gene transcription regulation process. In the cis-regulatory circuit part, the interactions of cis elements are also modeled by a nonlinear dynamic equation. The regulatory functions of double and triple nonlinear interactions among the cis elements are considered in this dynamic model. Furthermore, the decay rate is also considered to describe how the output mRNA is degraded in the target gene.
The nonlinear dynamic model is useful to describe how the upstream regulatory genes control a target gene to produce the output expression of mRNA through its trans/cis regulatory circuit. Using the information on transcription factors (Simon et al. 2001) and the experimental profiles on upstream regulatory genes and target gene (Spellman et al. 1998), the model of trans/cis regulatory circuit of target gene can be transformed to an algebraic regression equation for parameter estimation of the dynamic transcription model. With this study, the stochastic noises of microarray data are also considered to describe the uncertainties of the data. This is capable of improving the accuracy of parameter estimations in the dynamic model. The kinetic coefficients of the stochastic dynamic model, the decay rate of target gene's mRNA and the variance of noises are estimated via the maximum likelihood estimation method and the downhill simplex search method. After estimating these parameters via expression profiles of the target gene and its regulatory genes, they will be substituted into the nonlinear dynamic model to confirm the validity of trans/cis regulatory circuits. For illustration, the trans/cis regulatory circuits of the gene CLN1 are constructed in detail by the proposed method from expression profiles in the microarray data (Spellman et al. 1998) and knowledge of binding site motifs of yeast cell cycle (Simon et al. 2001). After constructing the model, we are able to use the mRNA expression profiles of regulatory genes to predict the expression profiles of the target gene and analyze the characteristics of the nonlinear trans/cis regulatory circuits to gain more insight into gene transcription, which also provides a potential method for gene regulatory circuit design with a desired gene expression.
Modeling and Identification
First of all, we construct a dynamic trans/cis regulatory circuit with two parts. One simulates the binding effects of regulatory proteins and their complexes, and the other considers interactions of cis elements in the target gene. After that, we apply the downhill simplex search method and the maximum likelihood method to estimate the dynamic parameters of the trans/cis regulatory circuit. Finally, we show the experimental data that have been used and how to preprocess it before system identification of regulatory circuits. The proposed method will provide a way for system analysis of gene regulatory circuits and a method for gene circuit design with a desired gene expression.
Dynamic model of trans/cis regulatory circuits
First, we consider a trans/cis regulatory network as a system block with several regulatory genes as inputs and a target gene as output. In our model, we use the mRNA expression data of upstream regulatory genes corresponding to transcription factors as the system input and the mRNA expression data of the target gene as the system output. For the illustration of nonlinear dynamic trans/cis regulatory circuit, an example of the target gene CLN1 is shown in Figure 1. From the binding site motif data (Simon et al. 2001), we know that the gene CLN1 has five transcription factors (Swi4, Swi6, Mbp1, Fkh1, and Fkh2). Some of the investigations (Koch et al. 1993; Uetz et al. 2000; Ito et al. 2001) show that some proteins could form a protein complex via protein-protein interaction, and then the protein complex binds to the motif of the target gene. The binding of transcription factors on binding site motifs is described by the sigmoid function to model the binding (ON) and nonbinding (OFF) through some threshold (Goldbeter and Koshland, 1981; Mestl et al. 1995; Gardner et al. 2003; Klipp et al. 2005). In the trans-regulatory circuit, we also consider this protein complex effect. The translated proteins Swi4 and Swi6 will form a protein complex SBF as a transcription factor to bind to the motif in the target gene, and proteins Mbp1 and Swi6 form another complex MBF (Iyer et al. 2001). Therefore, in Figure 1, we use linear functions and nonlinear functions to express the trans-regulatory functions of the transcription factors and their complexes, respectively. The trans-regulatory functions gSBF (t), gMBF (t), gFkh1 (t), and gFkh2 (t) of the respective cis elements SBF, MBF, Fkh1, and Fkh2 are the regulatory result of the expression profiles of transcription factors and their complexes interactions. In the trans-regulatory circuit, these trans-regulatory functions on cis elements, which are generated from upstream transcription factors and their interactive complexes, are shown as follows
After considering the trans effects of regulatory proteins and their protein complexes on cis elements, we also take into account the cis effects of regulatory interactions of cis elements on the transcription of coding region of the target gene. We suppose that the interaction function of two cis elements could be represented by the product of trans-regulatory functions from their transcription factors. From Figure 1, the interactions of cis elements are represented as follows:
where gi. j(t) denotes the interaction between cis elements i and j. The coefficient bi. j denotes the corresponding interactive ability.
The trans-regulatory functions in Equation (1) and their cis interactions in Equation (2) will lead to the transcription of gene CLN1 with mRNA expression profiles yCLN1(t) in Figure 1. The transcriptional behavior of cis regulatory circuit will be described by the following stochastic dynamic equation.
where the nonlinear transcriptional regulatory function G(t) is denoted as
The biological meaning of Equation (3) is that the change of mRNA level in the gene CLN1 is the synthesis due to the transcriptional regulatory function G(t) and the degradation –λyCLN1(t). The mRNA expression of the gene CLN1 will increase if the synthesis regulatory function G(t) is greater than the degradation λCLN1(t). Otherwise, it will decrease. Substituting Equations (1) and (2) into Equation (3), we get the dynamic equation of trans/cis regulatory circuit in the gene transcription processing of the gene CLN1 as follows
In the above dynamic equation of gene transcription, xi(t) denotes the expression profiles of the binding transcription factors. The nonlinear term
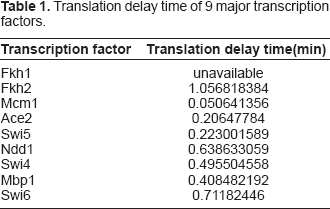
Translation delay time of 9 major transcription factors.
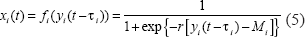
where r denotes the transition rate of the sigmoid function, Mi denotes the mean of mRNA expression level of the regulatory gene i, and τ i denotes the time delay of the translation time from mRNA to transcription factor (protein) for the regulatory gene i. In Figure 1, yi(t) and xi(t) represent the mRNA expression profile and the binding regulation function of the corresponding transcription factor on its motif binding site, respectively. The biochemical meaning of Equation (5) is that the regulation of transcription factor xi(t) on the binding site motifs is between ON (binding) and OFF (no binding) signal with some transition region dependent on beyond or below some threshold of mRNA expression level of the regulatory gene after a time delay τ i , which is available in Arava et al. (Arava et al. 2003). The sigmoid function can also be considered as a method to normalize the expression profiles of regulatory genes between 0 and 1 to model the binding and no binding, which has been successfully employed to describe the binding of regulatory gene (Chen et al. 2004; Klipp et al. 2005; Chang et al. 2006).
Hence, based on the mRNA expression time profiles and the translation delay time, the dynamic equation of trans/cis regulatory circuit for the gene transcription of CLN1 is described by
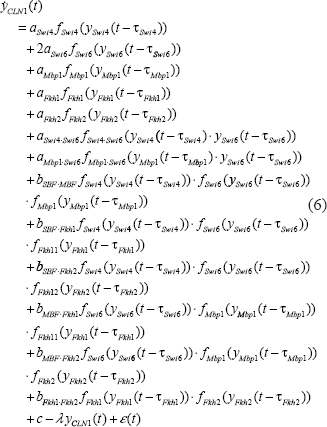
Based on the above analysis, in general, a block diagram to construct trans/cis regulatory circuit of gene transcription is shown in Figure 2, and the nonlinear dynamic model of trans/cis regulatory circuit of gene transcription of any gene of interest in yeast can be described as follows
Block diagram to construct a dynamic trans/cis regulatory circuit for gene transcription.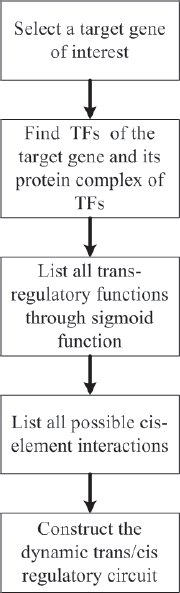
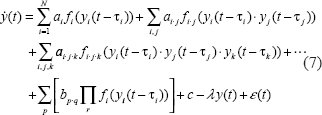
Unlike the previous dynamic gene regulatory models only with linear regulatory terms (Gardner et al. 2003; Chen et al. 2004; Chen et al. 2005), in our model, nonlinear interactions among two proteins and three proteins are also considered in our gene expression dynamic model. Furthermore, the delay from transcription to translation is also considered in our dynamic model. According to the information on protein complexes and motif binding sites in the promoter region of the gene, y(t) denotes the mRNA expression profiles of a gene of interest and yi(t), i ∊ {1 2 … N}, denote the mRNA expression profiles of upstream regulatory genes. With sampling expression in the next section, Equation (7) can be rewritten as.
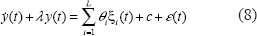
The next step is to identify the parameters θi, λ, c, and the covariance σ2 of noise from mRNA microarray data. The parameter estimations of θi, λ, c, and σ2 are achieved by the combination of downhill simplex search algorithm and maximum likelihood parameter method using the Methods in the sequel. After the parameter estimation is finished, we could construct a trans/cis regulatory circuit for any gene of interest via mRNA microarray data and the motif binding site information of transcription factors and their complexes.
Experimental data
We use the yeast microarray hybridization data of Spellman et al. (Spellman et al. 1998) as our mRNA expression profiles. They have many experimental methods to reset the yeast cell cycle to measure mRNA expression profiles for the whole genome comprehensively. Here, we use the experimental cell cycle data of the “α factor”. The information of motif binding sites in a target gene is from Simon et al. (Simon et al. 2001), Lee et al. (Lee et al. 2002) and Harbison et al. (Harbison et al. 2004), and then the protein complex data are obtained from Simon et al. (Simon et al. 2001) and the MIPS database. In the study of Simon et al. (Simon et al. 2001), there are 9 major transcription factors of cell cycle genes in yeast, so we use these 9 major transcription factors as the regulatory input of our dynamic model, which could also avoid overfitting in parameter estimation due to only 18 time points in expression profiles. Using the above information, we could construct dynamic models as Equation (7) for cell cycle genes and then identify the parameters from mRNA microarray data by the proposed parameter estimation algorithm methods.
We also include the translation delay time to our dynamic model. The data of translation delay time is available from Arava et al. (Arava et al. 2003).
Results
Reconstruction of microarray raw data by the gene regulatory circuits
We use the “α factor” microarray data of Spellman et al. (Spellman et al. 1998) and information on 9 major transcription factors from Simon et al. (Simon et al. 2001) to construct our nonlinear dynamic trans/cis model. In the study of Simon et al. (Simon et al. 2001), there are 769 cell cycle genes, but only 189 cell cycle genes have at least one motif binding site of 9 major transcription factors (P-value < 0.0015). Therefore, we could construct dynamic trans/cis regulatory circuits for these 189 cell cycle genes.
After estimating the parameters of dynamic trans/cis regulatory circuit for each gene, we compare the actual expression profiles with the constructed expression profiles of 189 genes by the cluster analysis and visualization tool (details can be found in Methods). The comparison can be seen in Figure 3 and we the correlation coefficient of these two data is 0.7276. However, we found that 80 cell cycle genes have only one motif binding site of 9 major TFs. These genes may have other motif binding sites which are not of the 9 major transcription factors, or they have post-transcription to dominate their expression profiles. For example, FUN26, SHL7, STE2, AGA2, YBL111C, YBL112C, and YBL113C have many other motif binding sites, which are not in the 9 major transcription factors (Lee et al. 2002; Harbison et al. 2004). Thus these 9 transcription factors may not play dominating roles in these genes, and the performances of predicted expression profiles of these genes by dynamic trans/cis regulatory circuits may not be very satisfactory. After removing 80 genes with one motif binding site from 189 genes, the dynamic trans/cis regulatory circuits of 109 cell cycle genes with at least two motif binding sites of these 9 transcription factors are considered to be more accurate. The actual profiles and the predicted profiles by the dynamic trans/cis regulatory circuits are shown in Figure 4, and their correlation coefficient is 0.8502. We also use the chi-square method to test the hypothesis of trans/cis circuit by comparing our predicted data and real microarray data. We found that 176 of 189 target genes cannot be rejected by the proposed trans/cis circuit by 95% significance, and other 13 target genes cannot be tested by the chi-square method because the number of their parameters is more than the number of their expression profiles data.
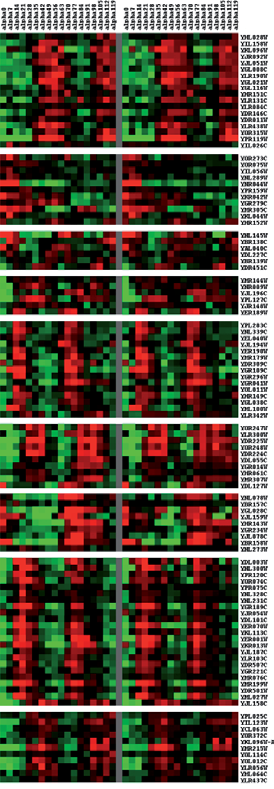
Comparison between the experimental mRNA expression profiles and those predicted by the proposed model. The experimental mRNA expression profiles of 189 cell cycle genes are at left side, and the profiles which are generated by the predicted dynamic regulatory circuits are at right side. Genes represented by red tonalities are over expressed and those represented by green ones are down regulated. The correlation coefficient of both profiles is 0.7276. Comparison between the experimental mRNA expression profiles and those predicted by the proposed model. The experimental mRNA expression profiles of 109 cell cycle genes are at the left side, and the profiles predicted by the dynamic regulatory circuits are at the right side. And the correlation coefficient of both profiles is 0.8502.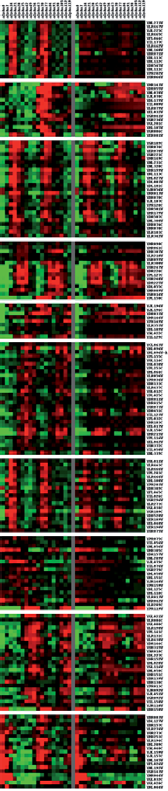
Test of shuffled microarray data
In order to test the accuracy of our model, we shuffled Spellman et al.'s “a factor” microarray data (Spellman et al. 1998) of expression profiles by random. After shuffling the microarray data, we applied our method to construct another new trans/cis regulatory circuit and then used the new regulatory circuit to generate profiles. For the 189 target genes with at least one motif of 9 major TFs, the correlation coefficient between shuffled profiles and their corresponding profiles predicted by new regulatory circuits via shuffled data is 0.1143 (shown in supplementary Figure S1), but the correlation coefficient between actual profiles and their corresponding predicted profiles is 0.7276. Furthermore, for the 109 target genes with at least two more motifs of 9 major TFs, the correlation coefficient between shuffled profiles and their corresponding profiles predicted by shuffled data is 0.5939 (shown in supplementary Figure S2), but the correlation coefficient between actual profiles and their corresponding predicted profiles is 0.8502. Obviously, only data generated by real biological systems could be identified well by the proposed model. From the view of system identification, a proper system model will lead to a good system identification if overfitting could be avoided. In our case, the proposed model is applicable to the trans/cis regulatory circuit because the correlation coefficient of real microarray data is much larger than that of shuffled microarray data. So we can say that our dynamic model is proper for the trans/cis regulatory circuit of gene expression program of yeast.
Regulatory activities of TFs in the target gene
Parameters of dynamic trans/cis regulatory models of CLN1, UTR2, and MNN1 based on Equation (4).
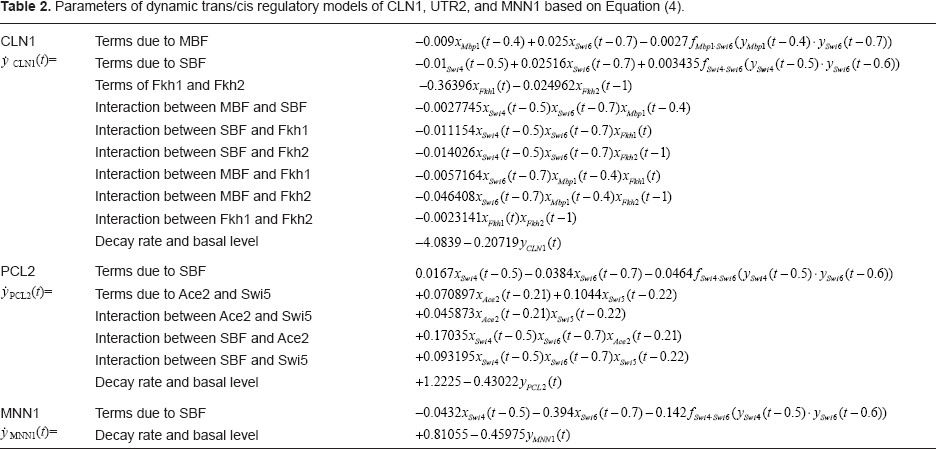

Our model also provides another important estimation of interactions among the cis elements. All possible interactions among the cis elements are considered, and then their kinetic parameters are estimated by expression profiles of microarray data. These estimated kinetic parameters could not only tell us the possibilities of these interactions of cis elements, but also the significance of these interactions. For example, we consider 5 cis-element interactions in target gene CLN1. From the kinetic parameters estimated, there are three parameters larger than the others. We may conclude that these three terms, MBF · Fkh2, SBF · Fkh1, and SBF · Fkh2, could be the three possible interactions among real cis elements.
With the same transcription factor binding to different target genes, different kinetic parameters of this transcription factor determine different regulatory effects on the target genes. A larger parameter of the same transcription factor binding to different target genes means that it plays a more important role and is more sensitive than those binding to another target gene. From our estimated kinetic parameters, we can compare these effects on different target genes with the same transcription factor. The G1 cyclin PCL2, which associates with Pho85p cyclin-dependent kinase (Cdk) to contribute to entry into the mitotic cell cycle, is regulated by SBF, Ace2, and Swi5. (Simon et al. 2001). On the kinetic parameters of PCL2 (see Table 2), the effect of transcription factor SBF on PCL2 is dominated by Swi6 and Swi4 · Swi6. For another target gene MNN1, required for addition of alpha1,3-mannose linkages to N-linked and O-linked oligosaccharides, the effect of transcription factor SBF on MNN1 is also dominated by Swi6 and Swi4 · Swi6, which can be seen in SGD database. However, the kinetic parameters of SBF on PCL2 (Simon et al. 2001) are all smaller than those of SBF on MNN1. We can consider that SBF has more regulatory effect on MNN1 than PCL2 and the sensitivity of SBF on MNN1 is more than that on PCL2. After simulation by changing the expression profiles of Swi4 or Swi6, of which SBF is composed, we find that the result is the same as we discussed above.
Possibilities of cis-element interactions
The possible cis element interactions which appear within the two-sided 90% confidence interval (there are 30 cis-element interactions within the two-sided 90% confidence interval).
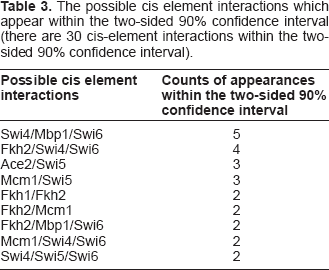

For the complex interactions of Swi4/Mbp1/Swi6, both SBF, the complex of Swi4 and Swi6, and MBF, the complex of Mbp1 and Swi6, control the transcription of G1/S cyclin genes, and many genes, Cdc6, Swi4, Clb6, Swe1, and Cln1, are both bound by SBF and MBF at the same time. Therefore, it is possible that the activators (Swi4/Mbp1/Swi6) have a cis-element interaction, and the possibility of their relationship has been reported by Kato et al. (Kato et al. 2004). The other two possible cis-element interaction terms (Fkh2/Swi4/Swi6 and Ace2/Swi5) calculated by our method may also exist. A few genomic analyses have indicated the involvement of SBF and Fkh1/Fkh2 in S phase (Lee et al. 2002), and SBF and Fkh2 most probably regulate budding, cell-wall synthesis, and spindle-related genes in S phase (Kato et al. 2004). Ace2 and Swi5 are a pair of TFs of yeast that regulate the expression of many cell cycle-specific genes, including Sic1, an inhibitor of Cdc28 protein kinase, Rme1, a regulator of meiosis, and Ash1, a regulator of meiosis (Doolin et al. 2001). In recent studies, Ace2 and Swi5 cooperate to induce the expressions of a subset of genes, but the antagonistic interaction of Ace2 and Swi5 was found (Doolin et al. 2001). With 82% identical DNA binding domains, Ace2 and Swi5 bind to the same DNA sequence (McBride et al. 1999), and it is possible that proteins compete for access on these promoters, but only one activates transcription (Doolin et al. 2001). Therefore, one partner of Swi5 and Ace2 sometimes can have a stronger contribution towards regulation, and the antagonistic interaction of Ace2 and Swi5 found is not surprising.
Discussion
Our method uses mRNA expression profiles, protein complexes, translation time delay, and the information of binding site motif to construct a dynamic trans/cis regulatory circuit to gain more insight into the gene expression of yeast. It is more precise to use these kinds of data to construct a nonlinear stochastic regulatory model for the gene transcription. Furthermore, a stochastic regulatory model can easily describe the properties of change, interaction, and uncertainty in mRNA expression profiles. Recently, Vu and Vohradsky also used nonlinear dynamic model to infer the transcriptional regulators of target genes (Vu and Vohradsky, 2007). They successfully identified possible transcriptional factors of yeast cell cycle, which shows the power of nonlinear dynamic models.
Constructing the trans/cis regulatory circuits of the cell cycle genes of yeast is useful to quantify the influence of transcription factors on their target genes, and then the possibilities of cis element interactions could be found from the statistical perspective. The confirmation of these possible cis element interactions would be a direction of further research. In addition, the proposed method can provide a quantitative basis for system analysis of gene circuit and give a scheme for gene circuit design with a desired gene expression in the future. Not only could the data of Saccharomyces cerevisiae be applied by our method, but those of other species also could. Any species with their mRNA expression profiles and the information of binding site motif can be applied using this method to construct their trans/cis regulatory circuits.
However, in this study, we found that the mRNA expression profiles of several genes generated by the predicted dynamic trans/cis regulatory circuit have a little difference from their original mRNA expression profiles. It may be because 9 transcriptional factors are not enough in these cases in which the genes may not be controlled dominantly by 9 TFs. We may use ChIP-chip data (Harbison et al. 2004) to replace the 9 major TFs from Simon et al. (Simon et al. 2001) to construct the transcriptional regulatory circuit. 204 DNA-binding transcriptional factors are found in yeast (Harbison et al. 2004), which may be useful for this elaboration of the transcriptional regulatory circuit. Furthermore, only 18 time points of mRNA expression profiles are used to estimate the parameters of dynamic model in each gene, which are not enough to get precise parameter estimation because of the large number of kinetic parameters to be estimated, which will easily lead to overfitting. If we include all possible 204 TFs into our model, the number of parameters that we want to estimate may be so large that overfitting would happen. Consequently, we only choose the most important 9 TFs in our model to avoid the overfitting in parameter estimation. If there are more time point profiles of mRNA expression and information of binding site motifs, the accuracy of the dynamic trans/cis regulatory circuit could be improved by the proposed method.
In conclusion, unlike the convention linear dynamic models, this study provides a nonlinear stochastic dynamic trans/cis regulatory circuit for any species via mRNA expression profiles, the information of binding site motif, protein complexes, and translation time delay to gain more insight into transcriptional regulatory infrastructures of gene expression program of yeast. The results are confirmed by a statistical hypothesis test.
Methods
Parameter estimation of dynamic trans/cis regulatory circuit
The dynamic trans/cis regulatory circuit of a gene of interest can be described by the dynamic Equation (7) according to the mRNA expression profiles of upstream regulatory genes and their nonlinear interactions. The parameters of the dynamic trans/cis circuit are then specified to meet the practical input/output microarray data of the regulatory genes and the target gene.
In order to avoid the interruption of high frequency noise with the use of the derivative information y(t), the solution of the dynamic Equation (8) is expressed by the following integration equation.
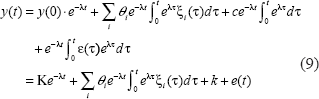
Noise of y(t) is written with
In Equation (9), ξi(t) is a combinative regulatory function of the regulatory genes. We use a third order polynomial cubic spline method to approximate ξ
i
(t) through microarray data with 18 time points, and then use the partial integration method to calculate
For ξ i (t) function, we use the cubic spline method to approach it. Cubic spline is a method of using a third order polynomial to approach every four data points (Faires and Burden, 1998). So we can rewrite the function ξ i (t) as follows.
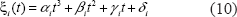
From Equations (9) and (10), we can use partial integration to refine Equation (9) as follows.
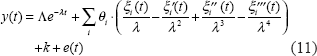
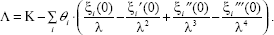
After applying the cubic spline method to solve and rewrite the dynamic Equation (9), we use a search algorithm, downhill simplex search method, for parameter estimations (Jang et al. 1997). However, this search method is not very efficient. Therefore, we combine the downhill simplex search method and maximum likelihood method to estimate the parameters. We use the downhill simplex search method to estimate the nonlinear parameter λ at first and then use the maximum likelihood method to estimate linear parameters K, θ i , and k.
After estimating λ by downhill simplex research method, we use the method of maximum likelihood to estimate the other linear parameters. Equation (11) can be written as the following regression form.
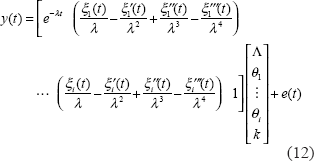
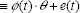
If we have a lot of microarray data points to process by the method above to get values of
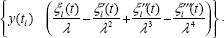
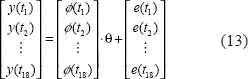
For simplicity, we can further define the notations Y, Φ, and E to represent Equation (13) as follows.

In Equation (13), we assume noises e(ti) at different time points as independent random variables of normal distribution with zero mean and unknown variance σ2, i.e. the variance of E is Σ = E{EET} = σ2I, where I is a unit matrix. In this study, a maximum likelihood parameter estimation method will be used to estimate θ and σ2 from microarray data of regulatory genes and the target gene (Johansson, 1993). If E is assumed to be normally distributed with N elements, its probability density function is of the following from
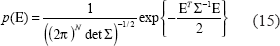
Under Equation (14) we have the likelihood function
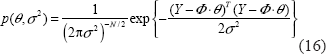
Equation (16) can be considered as a function of parameters θ and σ2. We want to specify θ and σ2 to maximize the likelihood function in (16). It is practical to take the logarithm of their likelihood function, and then we have the following log-likelihood function as follows.
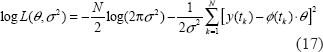
Here we expect the log-likelihood function to have the maximum at
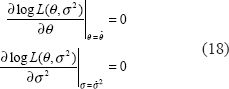
The estimated parameters 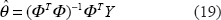
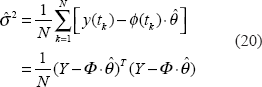
After estimating The combined method of the downhill simplex search and the maximum likelihood estimation for λ and θ iteratively.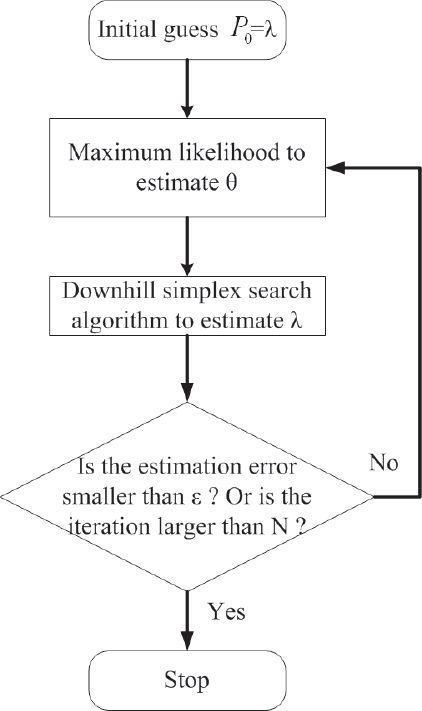
Comparison between actual/constructed expression profiles
We use the cluster analysis and visualization tool (Cluster and Treeview) written by Michael Eisen (http://rana.lbl.gov/EisenSoftware.htm) to compare the actual expression profiles with the constructed expression profiles. When clustering the expression profiles, we use hierarchical clustering methods (Eisen et al. 1998).