Abstract
Dengue fever is still a major threat worldwide, approximately threatening two-fifths of the world’s population in tropical and subtropical countries. Nonstructural protein 5 (NS5) methyltransferase enzyme plays a vital role in the process of messenger RNA capping of dengue by transferring methyl groups from
Introduction
Dengue is a mosquito-borne infection found in the tropical and subtropical regions. In recent years, the transmission has increased in urban and semiurban areas and has become a major health problem of the international community. Approximately 2.5 billion people are at risk of infection, of which almost 40% of them live in tropical or subtropical countries such as Africa, Southeast Asia, the Americas, and the Pacific.1–3 This disease is caused by dengue virus (DENV) vector or carrier through female Aedes mosquitoes, such as
Dengue virus is a single-stranded RNA virus which belongs to the
The NS5 protein, which is known as the largest protein encoded by the
Drug design and discovery is a process that involves many disciplines, such as medicinal chemistry, pharmacology, biochemistry, and computational biology. Previously, researchers conducted a trial-and-error method to find compounds that can act as inhibitors or potentially be developed as a lead drug, which is time-consuming and cost-intensive. Today, with the help of computers, drug design can be more efficient.19,20 In the process of developing new drug molecules, virtual screening (VS) is a powerful tool that can be significantly involved as part of the computer-aided drug design and development (CADDD) method. Virtual screening can be used to assess whether known compounds, which are retained from any source of a molecular database, are possible to become lead compounds for a specific target. 21 One method that is frequently used together with the VS process is molecular docking simulation; this simulation is often used to predict the small molecule’s binding orientation of the targeted protein in the hope of obtaining potential drug candidates based on the predicted binding affinities and activity.22–24 Moreover, the CADDD process is frequently followed by molecular dynamics simulation. This simulation is useful to predict the complex stability of the ligand and its target protein, although the whole process itself is computationally expensive and time-consuming.25,26
Several studies related to the inhibition of NS3 and NS5 proteins, either in silico or in vitro, have been conducted earlier.13,16,27–36 The modification of certain chemical compounds, supported by the advancement of technology in bioinformatics and medicinal chemistry fields, can be done to find the structure that can be used as a new, potential antiviral drug candidate. In this study, we tried to prove that the modified compound of SAH can inhibit the NS5 methyltransferase enzyme of DENV using molecular docking and molecular dynamics simulation. The computational ADME-Tox (absorption, distribution, metabolism, excretion, and toxicity) test would be conducted as well to eliminate further the remaining potential ligand.
Materials and Methods
The methodology of this experiment has been further modified based on our established research pipeline.37–39 In this research, we used several kinds of offline and online software. Molecular Operating Environment (MOE) 2008.10,40,41 ACD/Labs ChemSketch 12.01, PerkinElmer ChemBioDraw 13.0, VegaZZ 3.0.5, CAESA (Computer-Assisted Estimation of Synthetic Accessibility, v2.4; Symbiosis Inc, Lanham, MD, USA), Toxtree 2.5.0, 42 and OSIRIS Property Explorer 43 were the offline and online software that were used to conduct this research, respectively.
Preparation of the standard ligands and the modified SAH ligands
In this study, the modified SAH ligands and the standards were drawn in 2-dimensional format using ACD/Labs ChemSketch 12.01 software, which later can be saved in .mol (MDL format Molfile). All these ligands were further optimized using the VegaZZ software before they were entered into the MOE database. In this software, optimization of several ligands occurred, such as geometry optimization and 3-dimensional (3D) structure-energy minimization. All ligands were optimized using MMFF94x force field with root mean square (RMS) gradient of 0.001. The other parameters were adapted to the existing default of MOE 2008.10 software. The optimized ligands were later stored in .mdb database format.
Preparation of the NS5 methyltransferase DENV
The sequence data of NS5 methyltransferase enzyme were searched through the National Center for Biotechnology Information (NCBI) database. 44 The sequence was stored in FASTA format. The entire sequence was further input into ClustalW2 server (http://www.ebi.ac.uk/Tools/clustalw2). On this server, the multiple sequence alignment (MSA) was performed between NS5 methyltransferase enzyme sequences, which resulted in the most viable NS5 methyltransferase enzyme sequence for the 3D structure template, based on its highest score. The 3D structure of this enzyme can be later downloaded from the Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB) through the http://www.rcsb.org/pdb/home/home.do.
The downloaded PDB data were then opened with MOE 2008.10 software for further optimization. The optimization process of NS5 methyltransferase protein begins by removing the water molecules and metal atoms, followed by protonation of the protein using “Protonate 3D” default protocol. Then, the energy minimization of the protein was done by selecting AMBER99 force field in the gas phase solvation. The RMS gradient of 0.05 kcal/mol Ǻ was selected, whereas the rest of parameters used the standard defaults that already exist. Then, the minimized NS5 protein structure was saved in .moe format.
Molecular docking simulation of NS5 methyltransferase and modified SAH ligands
The process of molecular docking simulation between enzyme and ligand begins by opening the previously optimized protein structure in .moe format and the ligand database in .mdb format. After choosing the selected amino acid residues using “SiteFinder” feature, the docking process was conducted by selecting Compute → Simulations → Dock feature. The docking area was chosen based on the “SiteFinder” result, whereas the “Triangle Matcher” placement method and London dG scoring function were selected during the docking process. The docking results were further validated with postplacement refinement, using the GBVI/WSA dG method. The docking simulation was conducted twice, first with 30 times and second with 100 times of placement retention process. The other parameters were adjusted based on a default setting. The docking results were stored in .mdb format. The best enzyme-ligand complex was selected based on its free binding energy value, inhibition constants, and molecular interactions.
Computational ADME-Tox and synthetic accessibility prediction
In this study, the remaining ligand candidates were screened through their ADME-Tox properties using various software, namely, OSIRIS Property Explorer (www.organic-chemistry.org), Toxtree (toxtree.sourceforge.net), and ADME-Tox (http://ilab.acdLabs.com). These software analyzed the druglikeness properties, prediction of ADME-Tox properties, and carcinogenicity/mutagenicity prognosis of the ligands, respectively. Furthermore, the best ligands which have the best binding affinity to the NS5 methyltransferase protein would be checked for their synthetic accessibilities using CAESA software.
Molecular dynamics simulation
In this study, the molecular dynamics simulation was performed using MOE 2008.10 software. The preparation begins with the NS5 methyltransferase-ligand preparation, which involves energy minimization and geometry optimization. The parameters set in the energy minimization step were almost similar to those in the molecular docking simulation; the only difference is the Generalized Born solvation model which was set in this step. This simulation consists of 3 major steps: initialization, equilibration, and production. The simulation was conducted with Nosé-Poincaré-Anderson (NPA) algorithm and canonical ensemble (NVT). In the initialization step, the simulation was performed for 100 ps at 300 K, whereas in the equilibration step, the simulation was conducted at 300 K, gradually increasing to 310 K for 100 ps. Finally, the production step was implemented for 5000 ps (5 ns). The simulation results were later analyzed for determining the NS5 methyltransferase-ligand complex stabilities.
Results and Discussion
Preparation of the standard ligands and the modified SAH ligands
The molecular structure of SAH compound consists of 5 functional groups, including 2 amines, 2 alcohols, and a carboxylic group. These groups can be modified by other functional groups to alter its polarity or enhance its druglikeness. However, some groups, such as anhydride, epoxide, acyl halide, thiol, thiourea, aliphatic imine, and acrylamide, are potentially toxic and harmful because they are likely to undergo metabolic activation.45,46 The NS5 methyltransferase enzyme has 2 binding sites that are connected by a Y-shaped slit. The first one is the SAM-binding (methyl group donor) site, whereas the second is the RNA-cap–binding site. 47 In this study, the modification of the SAH ligand was conducted based on the polarity and shape of the SAM-binding site (Figure 1). The SAM-binding site itself consists of 14 amino acids (Ser56, Lys61, Cys82, Gly86, Trp87, Thr104, Lys105, Asp131, Val132, Phe133, Asp146, Ile147, Lys181, and Glu217) that could be selected as a drug target for NS5 methyltransferase. 48 Therefore, the modification of SAH ligand should fit with the SAM-binding site. The functional groups used in the SAH modification are alcohols, esters, aldehydes, ketones, amides, amines, and carboxylic acids. These groups were chosen because they are commonly found in the marketed drugs and used in the drug design and development studies. 49 Based on the molecular structure, the SAH modification took place at 5 different functional group sites that are mentioned earlier in this article (Figure 2).
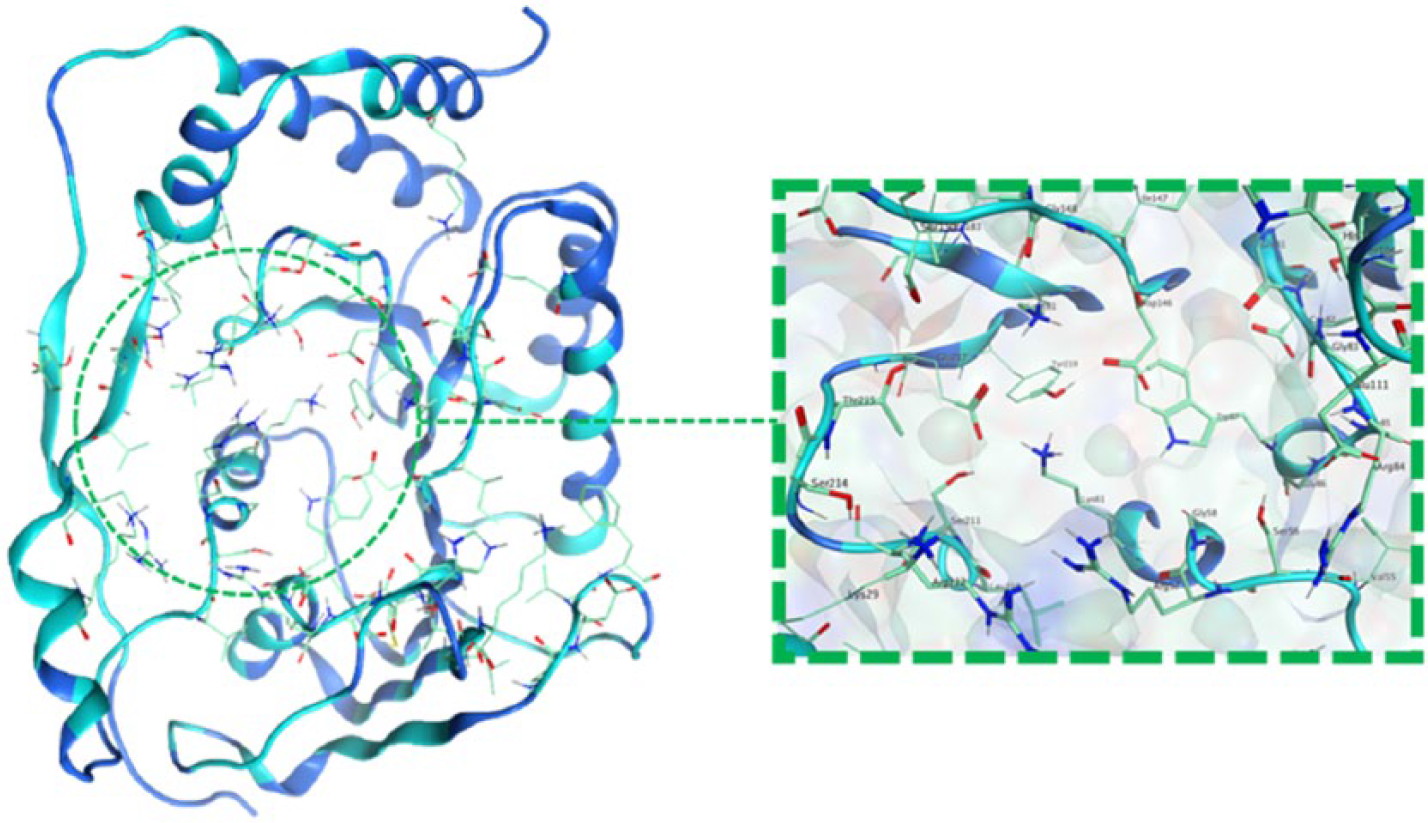
The 3-dimensional structure of NS5 methyltransferase enzyme (Protein Data Bank ID: 2P41) and the
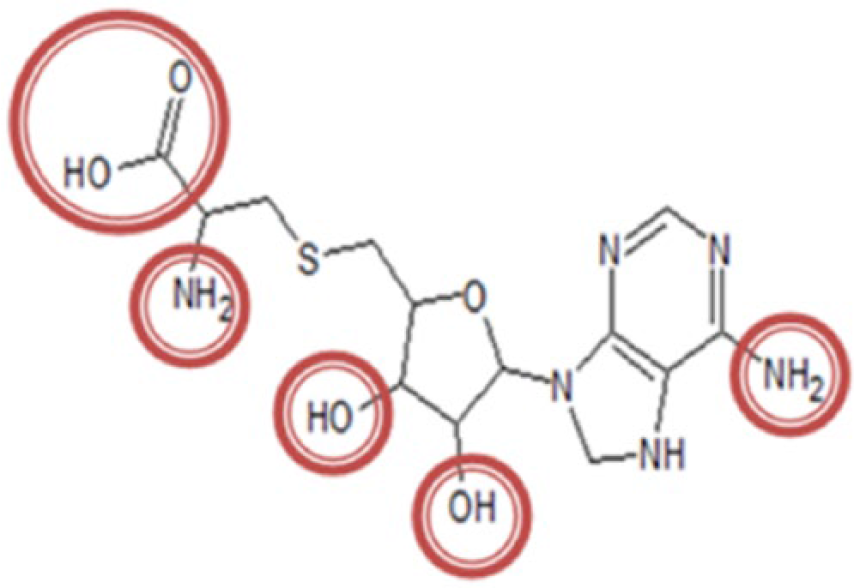
The place of functional group modification of
Before the SAH modification was done and drawn, we made a list of the modification of the respective functional group to simplify the drawing process, and so, there are no identical compounds in the resulting 3460 SAH-based ligands. Also, 3 standard ligands were drawn: SAH, SAM, and CTWYC ligand (a cyclic peptide compound); the former 2 compounds were used as a natural substrate of NS5 methyltransferase enzyme, whereas the latter compound was the best ligand from our previous studies. 50 Each ligand was drawn using ChemSketch ACD/Labs software and stored in MDL Molfiles (V2000) (.mol) format. Then, all ligands were optimized using the VegaZZ software before the docking process proceeded.
The ligand optimization was further performed using MOE 2008.10 software; this optimization includes geometry optimization and energy minimization. Furthermore, the optimization of the ligand was done using MMFF94x force field; this was done because this force field is frequently used to optimize the small-molecule structures that can be used later for molecular docking purpose. Then, the “Partial Charge” protocol was conducted as well to nullify the binding energy that occurs outside the system, as well as balancing the charge’s system so that the docking process can be run smoothly.
Afterward, energy minimization was done to eliminate the undesirable interaction in the ligand structure, such as unfit, yet unlikely van der Waals forces, and to minimize the steric effect that may occur in the system. The minimization was done at an RMS gradient of 0.001 kcal/Å. 51 In general, the aim of ligand optimization itself is to eliminate any bad contact in the system and to generate the actual geometry of structure that may occur in nature.
Preparation of the NS5 methyltransferase DENV
Searching of NS5 methyltransferase enzyme sequences
The NS5 methyltransferase enzyme sequences are searchable at NCBI Web site, with “dengue methyltransferase” as the keywords in “protein sequences” selection. In this step, we obtained thousands of sequences that match the keywords, but we further eliminated these sequences with only 12 sequences left because we wanted to use “Chain A dengue methyltransferase” as this sequence has the crystal structure in RCSB PDB database and is widely used in the dengue research field. These 12 sequences were saved in FASTA format so that it can later be used for the MSA step.
Multiple sequence alignment
The aim of the MSA step is to align all the leftover sequences to obtain the best sequence that will be used in the later process. This alignment was done using ClustalW software via European Molecular Biology Laboratory (EMBL) European Bioinformatics Institute Web site (http://www.ebi.ac.uk). Based on the result of alignment, we obtained 7 PDB sequences that have high similarity among the 12 sequences, which were 2P3L, 2P40, 2P3O, 2P41, 2P3Q, 2P1D, and 1L9K. Of these sequences, we chose 2P41 52 because this PDB has the lowest resolution (at 0.18 nm). In addition, the 2P41 PDB structure has SAH ligand as well, which can be used later for molecular docking simulation purposes.
The structure optimization of the NS5 methyltransferase enzyme
Similar to the ligand optimization process, the optimization of the NS5 methyltransferase enzyme was done using MOE 2008.10 software. In this particular step, we first removed any water molecules and sulfate ions
Molecular docking simulation of NS5 methyltransferase enzyme and SAH-based ligands
Molecular docking simulation is a computational method that is used to describe the molecular interaction between a molecule (ligand) and its receptor, either a protein or an enzyme. The purpose of this simulation is to predict the ligand activity toward its receptor and to filter any compounds that can be used to interact with the receptor to become the potential inhibitors of the protein/enzyme.22,54
In this study, the molecular docking simulation was performed using MOE 2008.10 software. The docking parameter involves placement, rescoring, retain, and refinement. We used Triangle Matcher as the placement parameter; this parameter serves to show a random movement of the ligand, based on charge and spatial fit, in the active site of the enzyme to produce the optimal bond orientation. 55 Also, this parameter is better than the “Alpha Matcher” because it can produce the pose/conformation which is more accurate and systematic. 56 Furthermore, London dG was used as the rescoring parameter. It is used to measure the biological activity of the ligand and the target protein by calculating the binding energies based on their molecular interaction from each pose/conformation that is generated in the software. 57
The determination of the complex structure was done by selecting the smallest Gibbs free binding energy (ΔGbinding) from the docking simulation result. The most negative ΔGbinding result indicates that the ligand conformation is the most stable, yet favorable complex conformation above all. From 3460 ligands that we tested in this simulation, we selected 98 ligands that have the lowest ΔGbinding from the standards. Afterward, we ran another docking process to validate the docking results that we obtained previously. This led to 3 ligands that have the lowest ΔGbinding among others. Table 1 shows the ΔGbinding and inhibition constant (pKi) of the 3 best ligands, as well as 3 standard ligands that we tested in this study (Figure 3).
ΔGbinding and pKi value from the docking results.
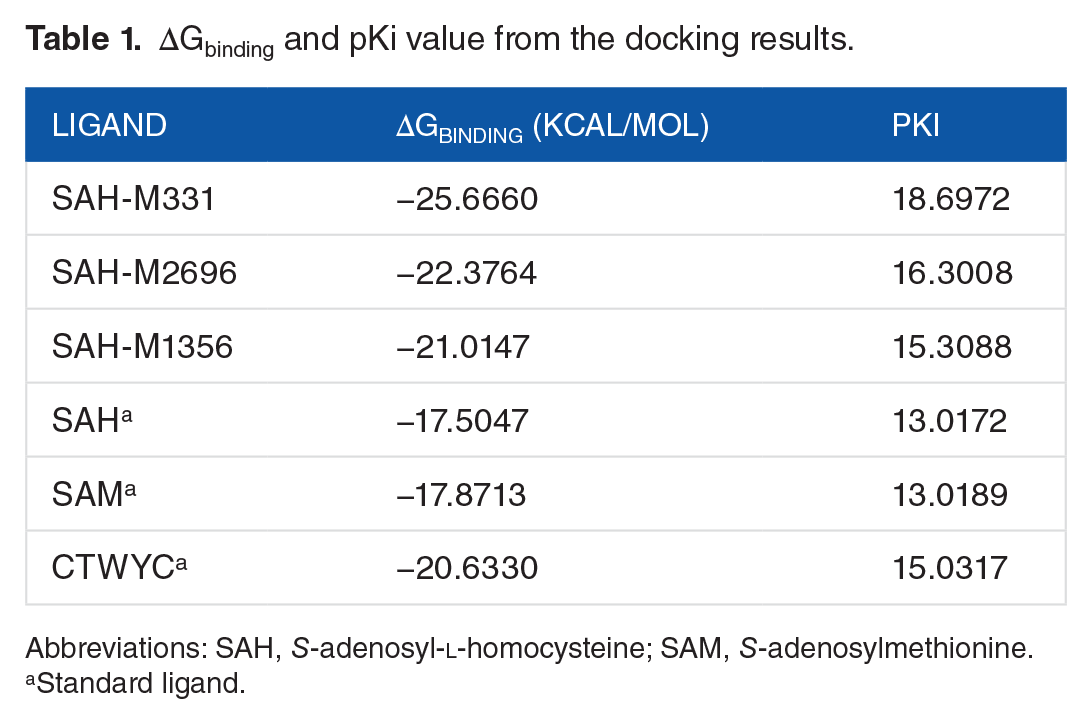
Abbreviations: SAH,
Standard ligand.
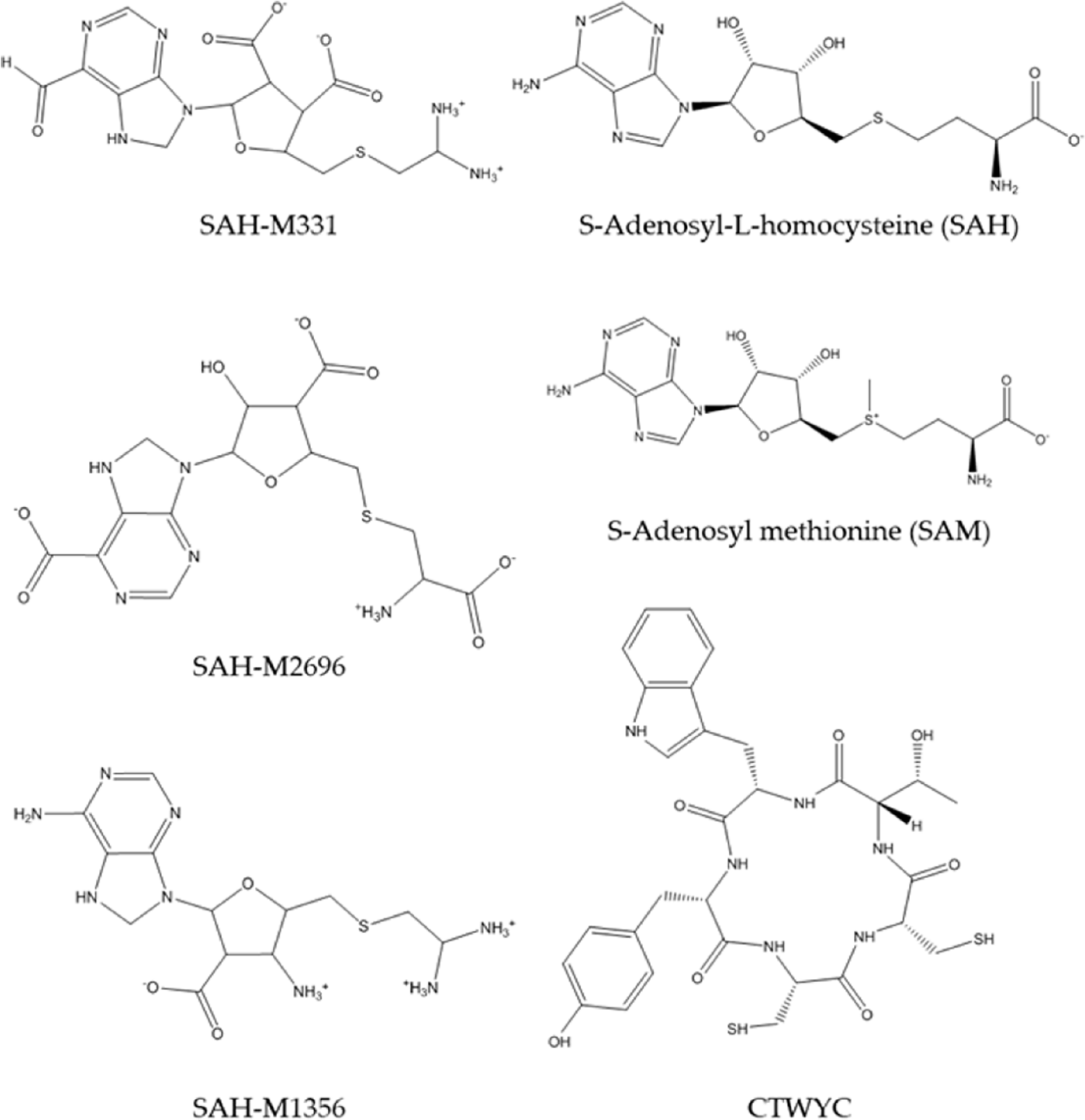
The chemical structures of the standards and the best ligands from molecular docking simulation.
Besides Gibbs free binding energy and the inhibition constants, the molecular interaction of the enzyme and the ligand can also be observed. This molecular interaction includes hydrogen bonds, which are defined as the intermolecular force that occurs between an electronegative atom and hydrogen atom that covalently binds to high electronegativity atom. 58 Based on the docking results between the modified SAH ligands with the active site of NS5 methyltransferase enzyme, the molecular interaction is as shown in Table 2.
Ligand interaction with the SAM-binding site of NS5 methyltransferase enzyme.
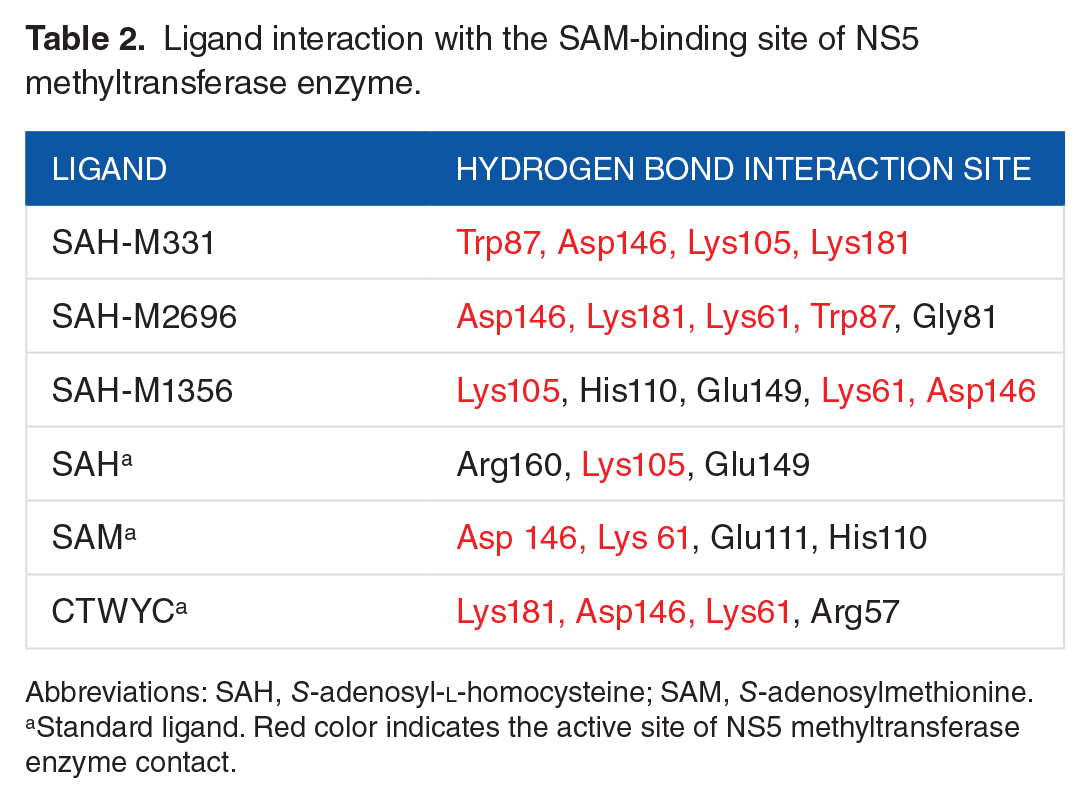
Abbreviations: SAH,
Standard ligand. Red color indicates the active site of NS5 methyltransferase enzyme contact.
In Table 2, the red-colored letters show the SAM-binding site residues. It can be seen that ligand SAH-M331 has formed 4 hydrogen bonding interactions with the active site of NS5 methyltransferase enzyme, of which 1 interaction occurs in the pi-pi cation interactions with Trp87. Moreover, 6 hydrogen bonds, with 4 of them occurring in the active site of NS5 methyltransferase enzyme, occur in the ligand SAH-2696. Finally, the interaction between SAH-1356 and NS5 methyltransferase enzyme has generated 4 hydrogen bonds, with 3 of them occurring in the active site (Figure 4). This result shows that the modified SAH-based ligands have a better binding affinity regarding the hydrogen bonding than the SAH, SAM, and CTWYC ligands, which has 1, 2, and 3 hydrogen bonds in the active site, respectively. The molecular interaction between NS5 methyltransferase enzyme and the modified SAH-based ligands, as well as the standard ligands, is shown in Figures 4 and 5, respectively.
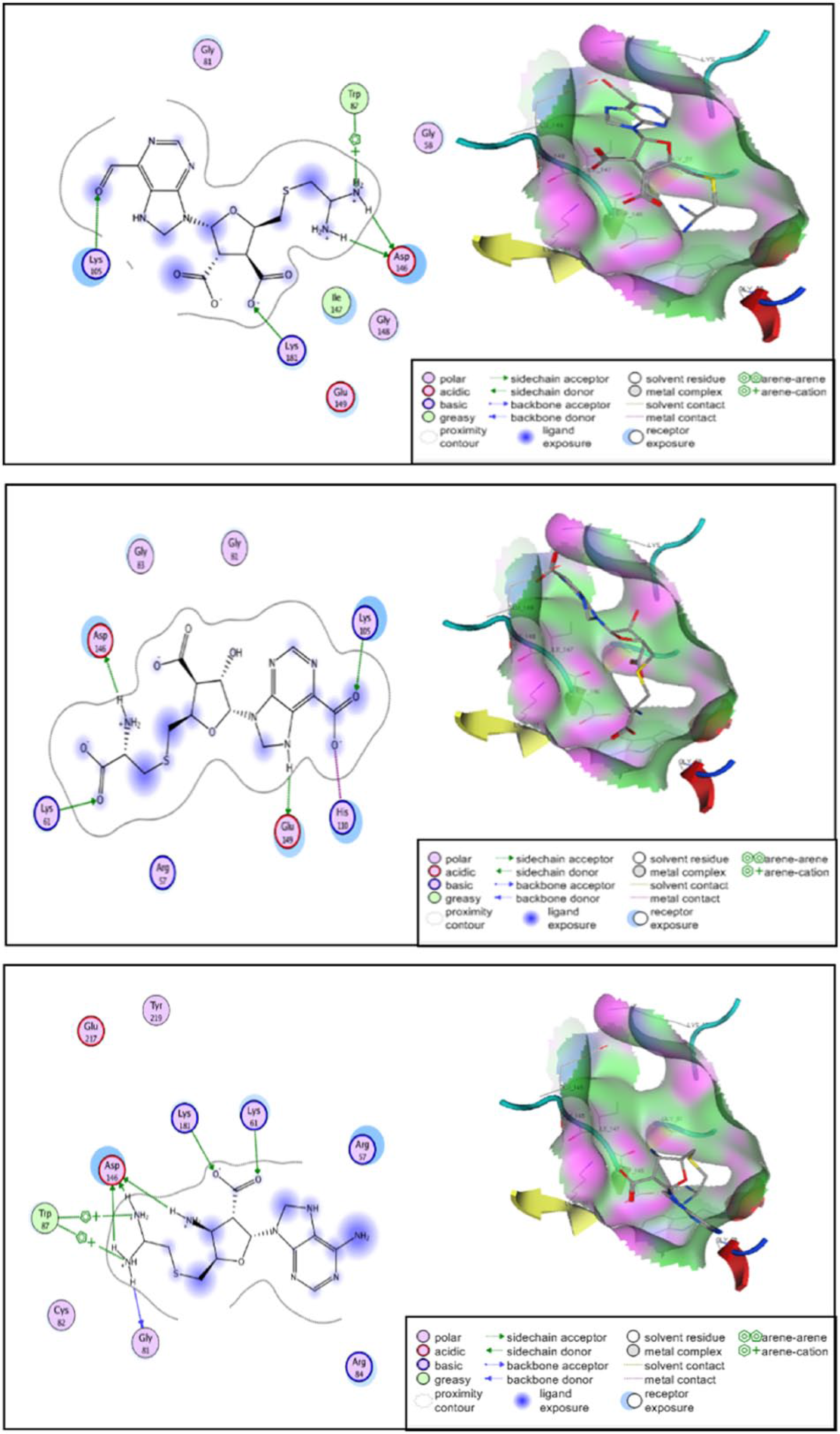
Molecular interaction and 3-dimensional visualization of binding site of SAH-based ligands, namely, SAH-M331 (top), SAH-M2696 (middle), and SAH-M1356 (bottom). SAH indicates
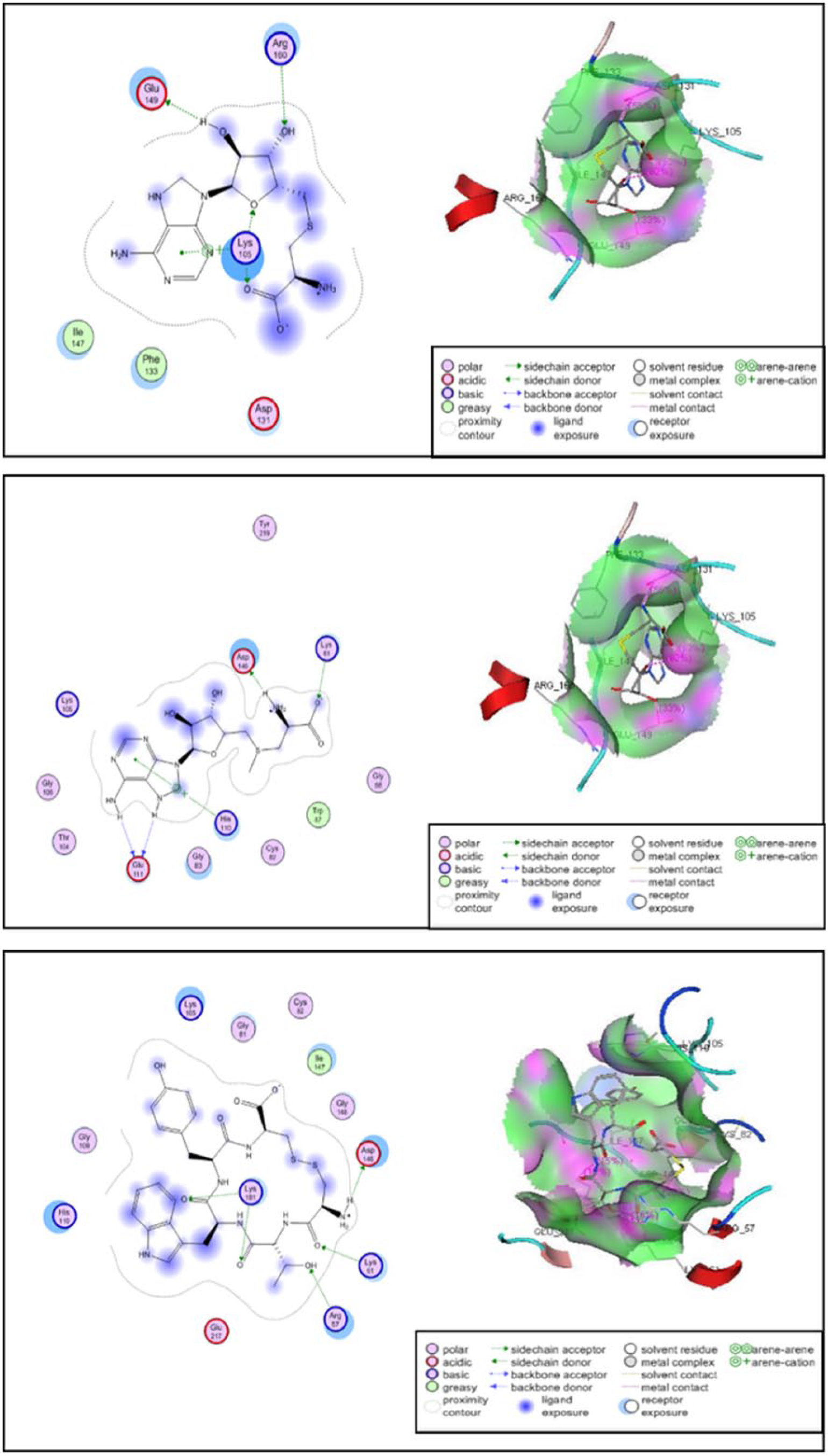
Molecular interaction and 3-dimensional visualization of binding site of standard ligands, namely, SAH (top), SAM (middle), and CTWYC (bottom). SAH indicates
In silico ADME-Tox prediction
Predicting ADME-Tox properties by ADME-Tox software
The process of finding a suitable, yet effective, drug for a disease can be considered by their ADME-Tox (Absorption, Distribution, Metabolism, Excretion, and Toxicity) properties. Any drug candidates that do not meet the ADME-Tox criteria will be eliminated to suppress the failure possibility in clinical trials. Nowadays, the ADME-Tox prediction can be easily made with the help of computers using in silico method; one of the software that is frequently used in the analysis is ADME-Tox, which is an online software available in ACD/Labs (via https://ilab.acdLabs.com/iLab2/). 59 Through this software, we can predict the toxicity properties of a compound by comparing them with each fragment’s properties that are stored in the database, including oral bioavailability, passive absorption, health effect, toxicity probability, and active transport. A bioactive molecule which has a high oral bioavailability is often considered and can be developed further as therapeutic agents for many diseases. 60 The ADME-Tox prediction using ADME-Tox software is shown in Table 3.
The absorption, distribution, metabolism, excretion, and toxicity (ADME-Tox) property prediction by ADME-Tox software.
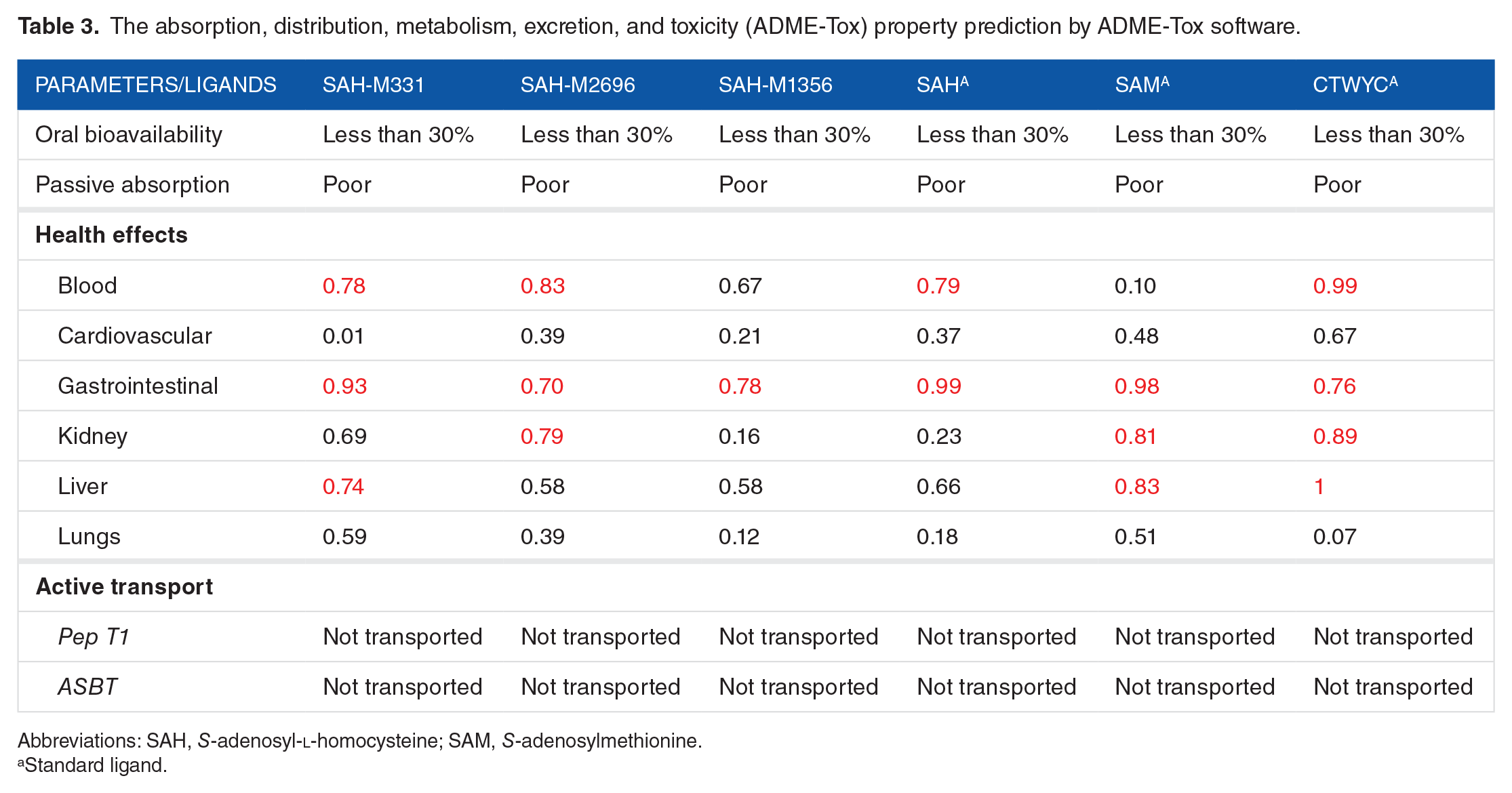
Abbreviations: SAH,
Standard ligand.
The numbers which are listed in the table show the toxicity probability; if the score is less than 0.7, then we assumed that the compound is safe for the selected organ/organ system. Based on Table 3, all of the 3 standard ligands have low oral bioavailability, yet they have weak active transport probability. However, the health effect prediction of these ligands shows a real promise, as they have great results compared with their standard ligands. Overall, this test revealed SAH-M1356 as the best ligand, whereas SAH-M331 and SAH-M2696 ligands show less favorable property than the former ligand but are still better than the standard ligands.
Carcinogenicity and mutagenicity predicting test by Toxtree software
The next in silico analysis is to determine the carcinogenicity and mutagenicity probabilities of a compound. Therefore, the Toxtree (v2.5.0) software was used.
42
This software predicts the probabilities based on the Benigni/Bossa rule for carcinogenicity and mutagenicity compounds, which involves searching the structural alerts of a fragment that could potentially contain either mutagenic or carcinogenic property in the compound.
61
The carcinogenic properties of a compound can fall into 3 categories: genotoxic, nongenotoxic, and quantitative structure activity relationship (QSAR)–based carcinogens, whereas the mutagenic feature of a compound can be determined by testing it against
Carcinogenicity and mutagenicity prediction by Toxtree software.
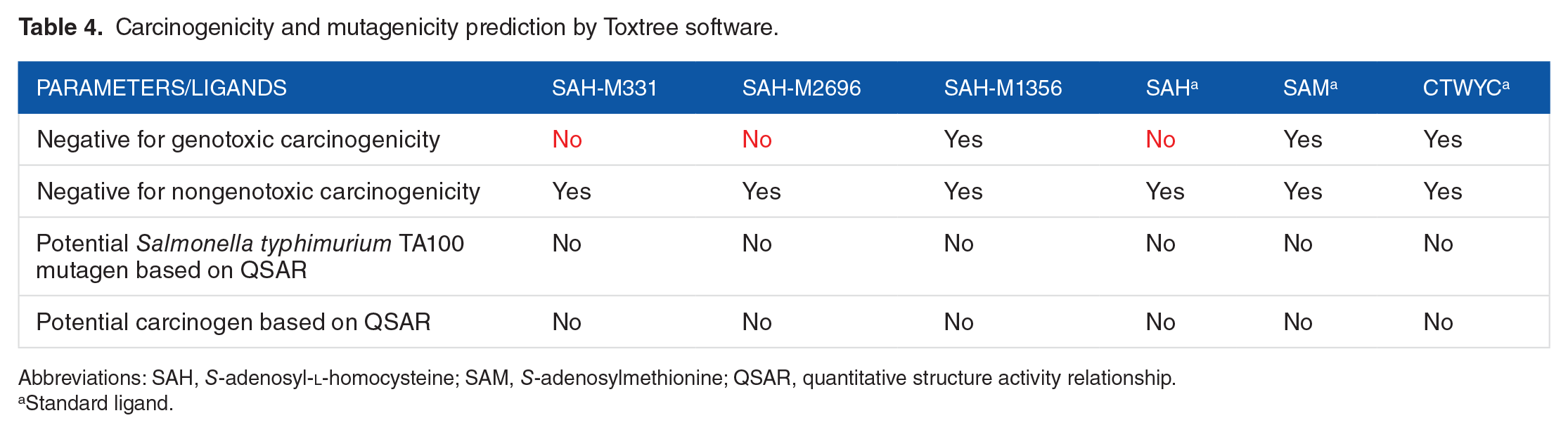

Abbreviations: SAH,
Standard ligand.
It can be seen from Table 4 that SAM and CTWYC peptide ligands have good properties, which did not show any carcinogenicity or mutagenicity probability. However, the SAH ligand showed positive results in genotoxic carcinogens. Interestingly, SAH-M331 and SAH-M2696 ligands are indicated to have same properties as well. In this test, once again, the SAH-M1356 ligand proves to be the best ligand for showing no carcinogenic or mutagenic possibility.
Druglikeness prediction by OSIRIS Property Explorer software
The toxicity and druglikeness analyses can be performed further using OSIRIS Property Explorer software (www.organic-chemistry.org).43,62 The analysis using this software is similar to Toxtree software, such as mutagenic and carcinogenic analysis. However, the OSIRIS Property Explorer also offers irritant, reproductive harmless, and druglikeness analysis, which is more useful and beneficial than the later software regarding drug design and discovery field. 63 Druglikeness is related to the Lipinski rule (also known as Lipinski rule of five, RO5). This rule helps us to distinguish between drug-like and non–drug-like molecules by predicting its oral bioavailability through its molecular properties. 64 In general, this rule states that the probability of a compound to be absorbed into the human body via lipid bilayer is high if it satisfies two or more criteria below:
The molecular weight of the compound is less than 500 g/mol;
Has high lipophilicity (logP less than 5.0);
Has less than 5 hydrogen bond donors;
Has no more than 10 hydrogen bond acceptors.
Furthermore, the determination of drug score parameter in the OSIRIS Property Explorer is based on the calculation of the Lipinski rule of five, the Veber rule, and the mutagenic, tumorigenic, irritant, and reproductive effect probabilities.60,65 The result of this analysis is shown in Table 5.
Druglikeness property prediction by OSIRIS Property Explorer software.
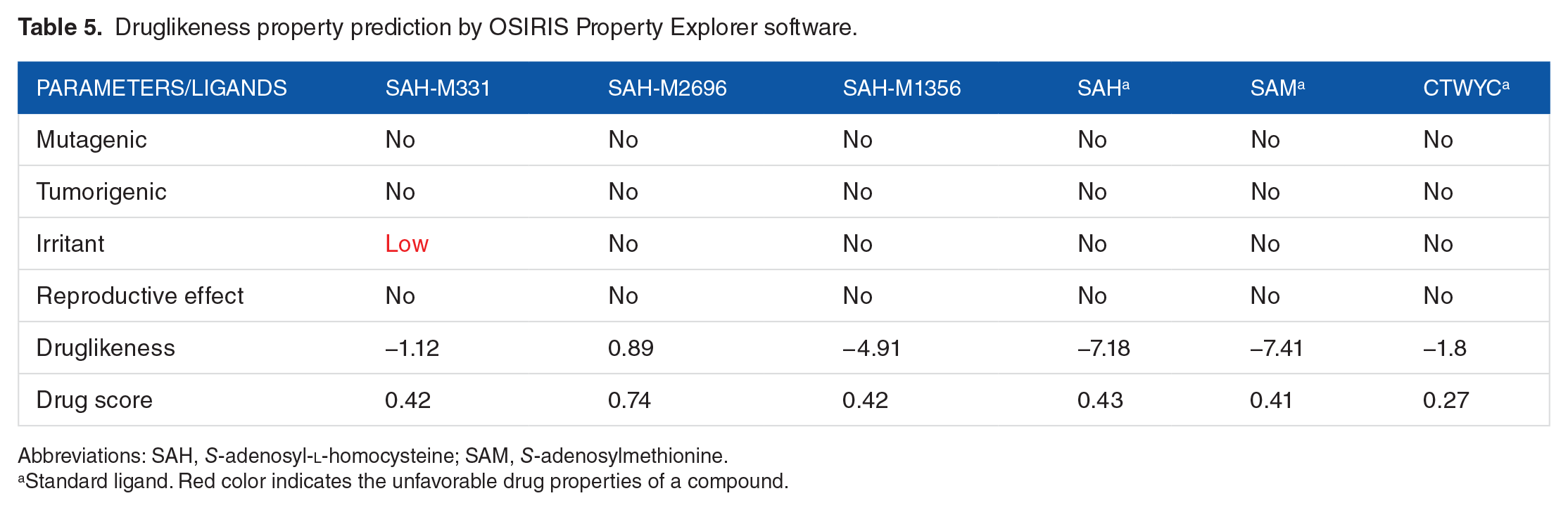

Abbreviations: SAH,
Standard ligand. Red color indicates the unfavorable drug properties of a compound.
The analysis shown in Table 5 indicates that the standard ligands have good drug results, such as no risk of being mutagenic, tumorigenic, and other health effects, although the druglikeness and the drug score of the standard ligands showed poor results. Furthermore, SAH-M2696 and SAH-M1356 ligands had no mutagenic, tumorigenic, irritant, and reproductive effect, although the former ligand has a good value of druglikeness and drug score than the latter. The SAH-M331, however, reveals to be an irritant, thus decreasing its druglikeness and drug score.
Synthetic accessibility prediction
The synthetic accessibility prediction is used to analyze whether any modification of a compound can be synthesized in a wet lab or not. This prediction can be made using several kinds of software. In this study, we used CAESA software to analyze the synthetic accessibility, residual complexity, and starting material of a compound based on databases such as Acros, Sigma-Aldrich, and Lancaster. The result of the analysis can be shown as a percentage; the greater the percentage, the bigger the chances for the compound to be synthesized. The result of synthetic accessibility prediction is shown in Table 6.
Synthetic accessibility prediction of the selected ligands.
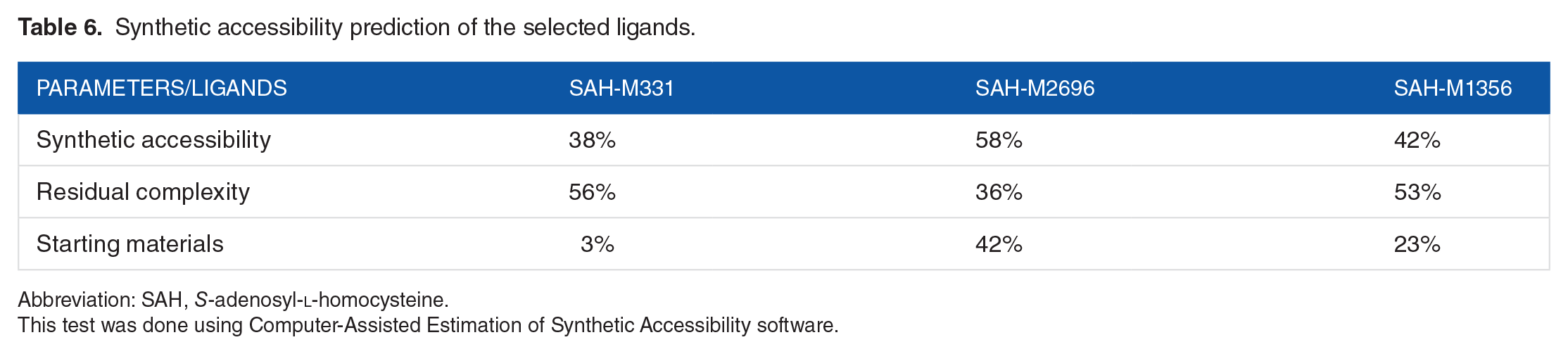

Abbreviation: SAH,
This test was done using Computer-Assisted Estimation of Synthetic Accessibility software.
The above data indicate that the SAH-M2696 ligand has the highest possibility to be synthesized rather than other ligands; this is because it has the highest percentage (at 58%) in synthetic accessibility, far greater than SAH-M331 and SAH-M1356 (at 38% and 42%, respectively).
Molecular dynamics simulation of NS5 methyltransferase enzyme and SAH-based ligands
Molecular dynamics simulation is used to predict the stability, movement, and conformation of protein-ligand complex that is observed over comparable time periods. 66 Thus, this simulation allows the enzyme-ligand complex to apply the induced-fit model and makes it more accurate and reliable than the docking simulation. 25 In this study, the molecular dynamics simulation was done using MOE 2008.10 software. The NPA algorithm was applied in this simulation, with the AMBER94 force field, NVT canonical ensemble, at a temperature of 310 and 312 K. Furthermore, this simulation was completely done using implicit solvent models.
In general, this simulation can be divided into 3 main phases: initialization, equilibration, and production. First, the initialization stage was conducted to determine the state of the system, such as atomic coordinates, velocity, and potential energy. Then, the equilibration stage was conducted to provide a relaxed state on the complex as a result of restraint when the system is heated. 58 Finally, the production stage was done to generate a trajectory of a simulation that showed the conformation changes of the complex in forms of atom coordinates in the simulation period. 67 The production stage in this study was conducted in 5 ns. The root-mean-square deviation (RMSD) value and molecular interaction of the complex from this simulation should be checked to determine the ligand-NS5 methyltransferase enzyme.
The purpose of RMSD value is to describe the conformational changes between 2 atomic coordinates during molecular dynamics simulation. Therefore, we can decide which ligand-enzyme complex has better stability. As the temperature increases, the molecular interaction between the enzyme and the ligand should change, but not in an insignificant manner. Hopefully, the presence of the modification of SAH ligand can inhibit the action of the active DENV at fever temperature (around 39°C or 312 K).
As shown in Figure 6, the ligand SAH-M1356 tends to have the most stable complex with NS5 methyltransferase enzyme; this is because the complex of SAH-M1356 ligand and NS5 methyltransferase enzyme has the smallest RMSD value at 312 K, indicating that the complex would be stably formed and the capping of new RNA of DENV would unlikely happen. In addition, the RMSD value of SAH-M331 complex at different temperatures was very different. Although at 310 K the complex seems to have low RMSD value (at 1.5-1.8 Å), the complex tends to have high RMSD value (at 3.5-4.5 Å) at 312 K. Moreover, the ligand SAH-M2696 has the highest RMSD value, compared with the other 2 ligands (3.5-4.0 Å and 4.0-5.0 Å at 310 and 312 K, respectively), suggesting that the ligand modification of SAH-M2696 is more active and forms less stable ligand-enzyme complex than the other ligands.
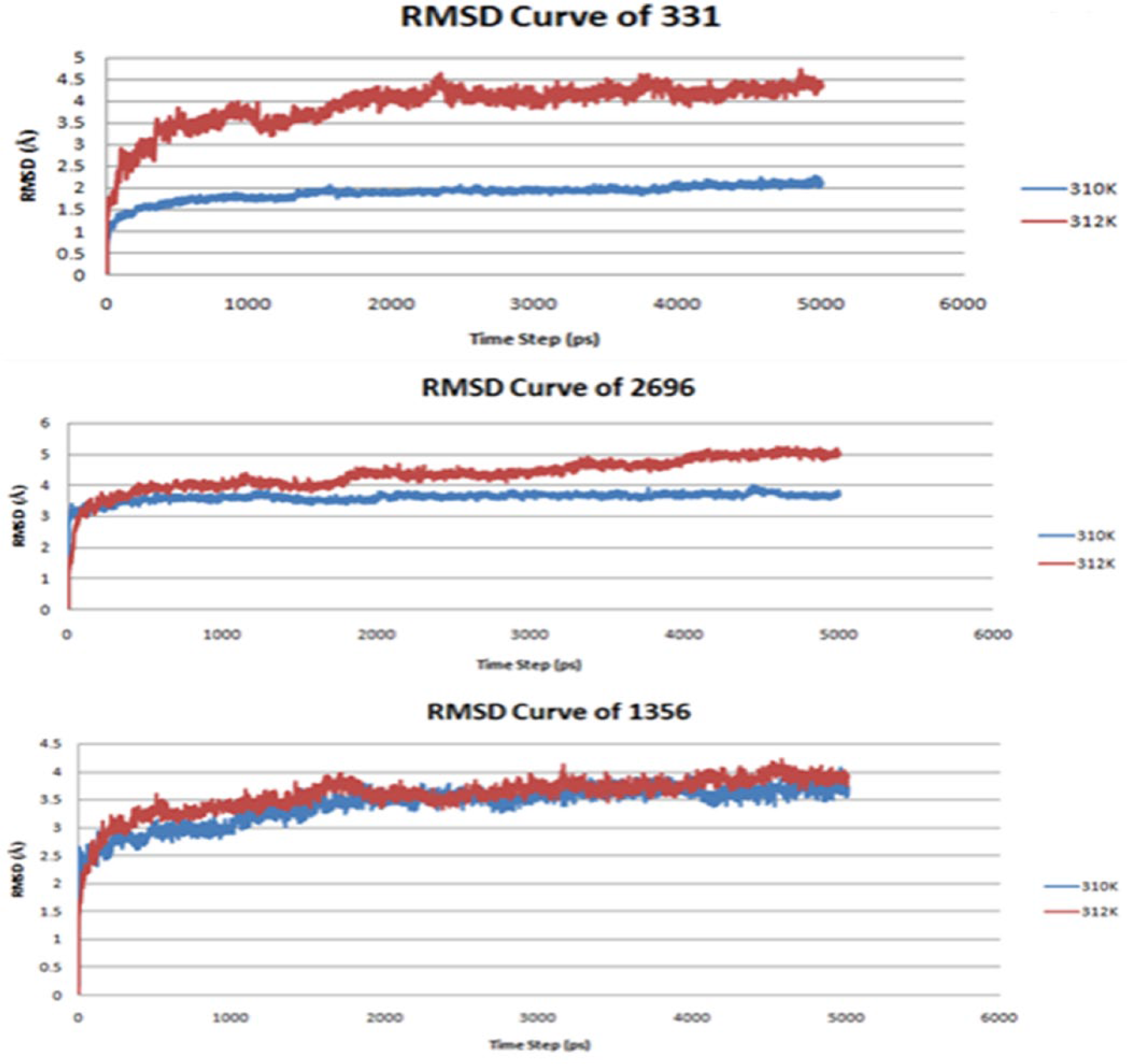
The RMSD curve of SAH-M331 ligand complex (top), SAH-M2696 (middle), and SAH-M1356 (bottom). RMSD indicates root-mean-square deviation; SAH,
Furthermore, the molecular interaction between the ligand-enzyme complexes between the different molecular dynamics simulation stages was also evaluated. In this study, we compared the molecular interaction, including hydrogen bond and residue contact, before and after molecular dynamics simulation was performed. As shown in Table 7, the molecular interaction between the ligand modification of SAH-M331, SAH-M2696, and SAH-M1356 still occurs with their respective enzymes at 2 different temperatures. However, we can also see that the molecular interaction of these ligands, at various temperatures and stages, has significantly changed as well. After molecular dynamics simulation at 310 K, the SAH-M331 complex tends to have better stability; this is shown from the 18 interactions that occur in the complex, including 5 interactions with Asp146 and 2 interactions with Lys181. Moreover, the arene-cation interaction between
Comparison of molecular interaction before (after docking) and after dynamics simulation.
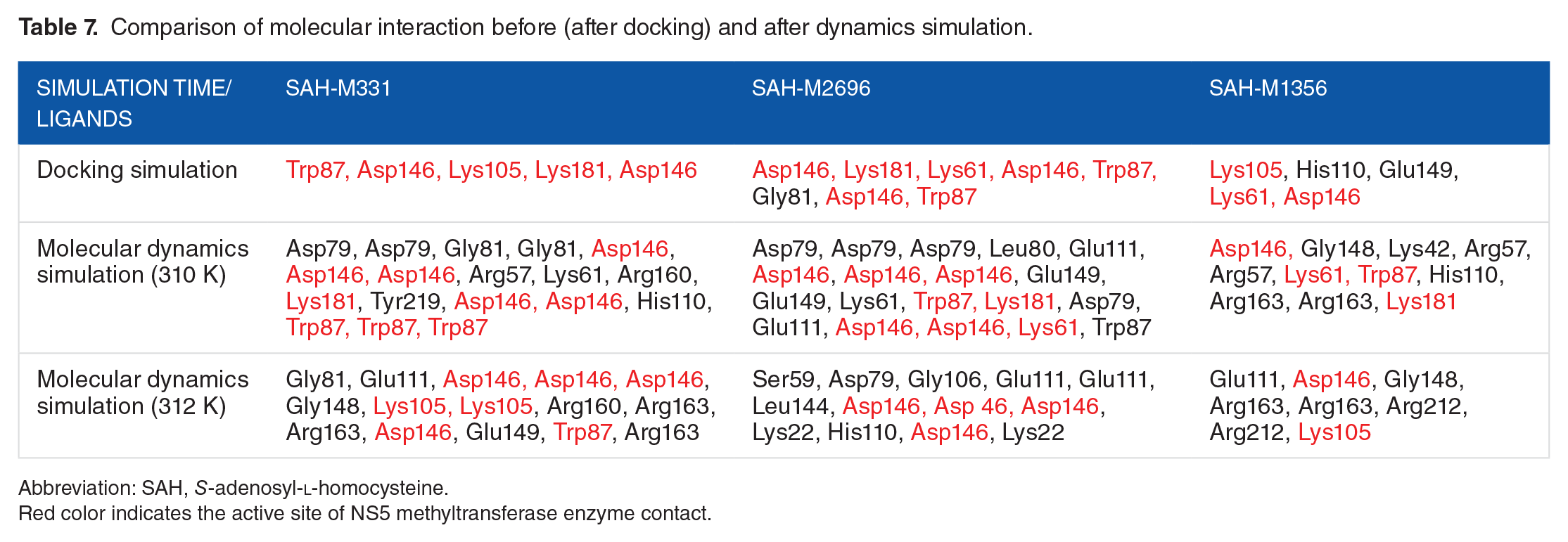
Abbreviation: SAH,
Red color indicates the active site of NS5 methyltransferase enzyme contact.
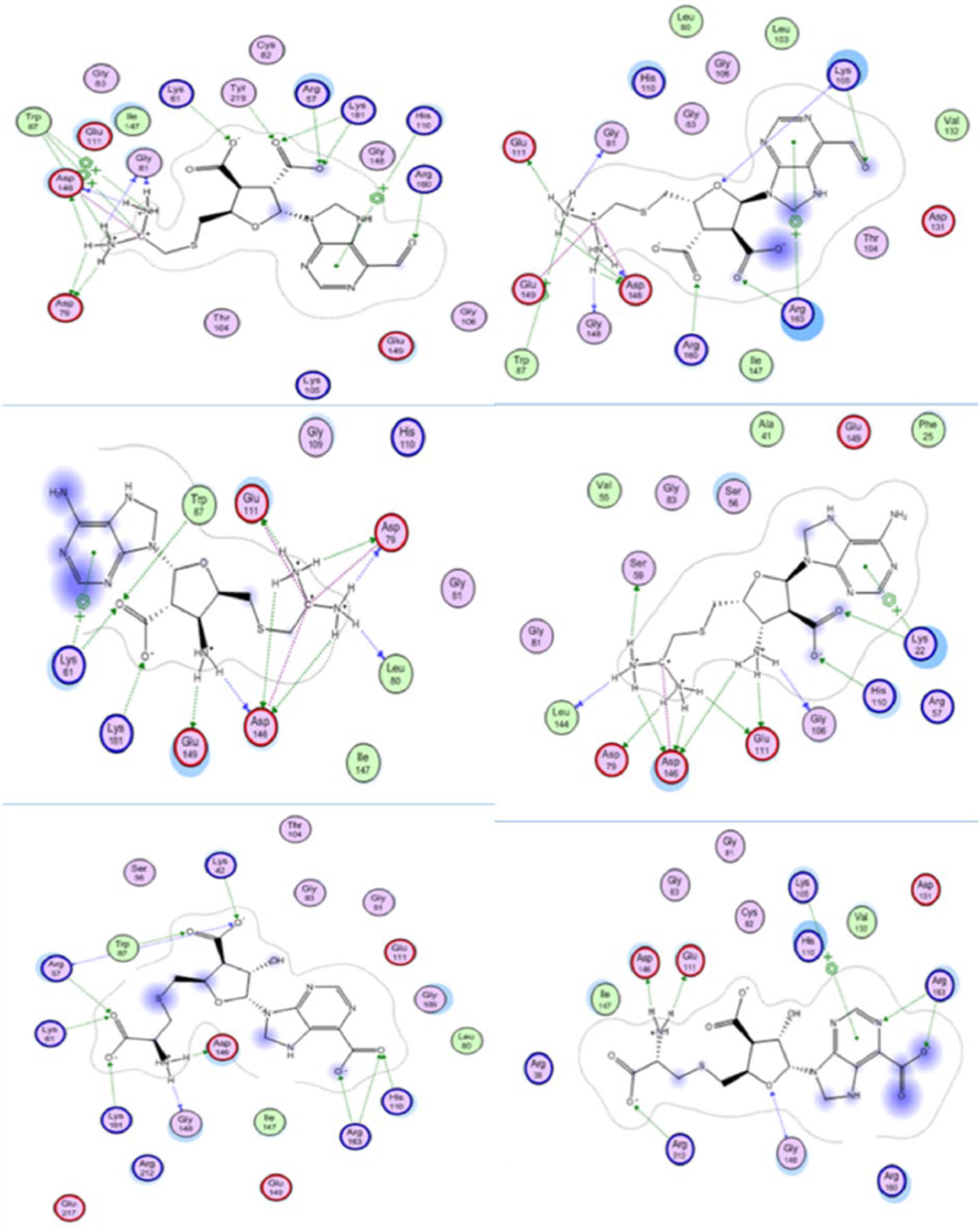
Molecular interaction and 2-dimensional visualization of SAH-based ligands after molecular dynamics simulation at 310 K (left side) and 312 K (right side), namely, SAH-M331 (top), SAH-M2696 (middle), and SAH-M1356 (bottom). SAH indicates
Conclusions
In this study, 3460 modified SAH-based ligands were created and tested against NS5 methyltransferase inhibitors through molecular docking and molecular dynamics simulation, along with computational ADME-Tox test. Through all the simulations that we have done, we concluded that SAH-M1356 is the most potential SAH-based ligand to inhibit NS5 methyltransferase. This ligand was chosen because it has lower Gibbs free binding energy and higher inhibition constant than our standard ligands (SAH, SAM, and CTWYC). Furthermore, this ligand also has good ADME-Tox properties, such as low health risk effect, no carcinogenicity and mutagenicity risk, and good drug-like properties, from the computational ADME-Tox results. The molecular dynamics results also confirmed that SAH-M1356 ligand has stable molecular interaction and RMSD score and indicated that the SAH-M1356 and NS5 methyltransferase complex can be likely formed. Therefore, to prove the computational results from our research, we highly recommended synthesizing this compound to further determine its bioactivity and inhibitory activity against DENV infection by wet lab experiments.
Footnotes
Acknowledgements
The authors would like to thank Ratih Dyah Puspitasari for helping us to proofread the final manuscript.
Peer review:
Six peer reviewers contributed to the peer review report. Reviewers’ reports totaled 1093 words, excluding any confidential comments to the academic editor.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Publication of this article was funded by the Directorate of Research and Community Engagement of Universitas Indonesia (DRPM UI) through Penelitian Unggulan Perguruan Tinggi (PUPT) 2016 (grant number: 1713/UN2.R12/HKP.05.00/2016).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
USFT and AAP designed the whole study and supervised this research. DK and SI created the general pipeline of this study and gave more relevant materials for the paper. MAFN and EPT helped in writing the paper. FA conducted the technical and experimental details. All authors approved the final manuscript.
