Abstract
Background:
Hypoxia alters skeletal muscle metabolism and function through complex regulatory mechanisms, including exosome-mediated microRNA (miRNA) signaling.
Objectives:
This study profiled exosomal miRNAs from hypoxic human skeletal muscle cells (HSMCs) to explore their roles in hypoxic adaptation.
Design:
Human skeletal muscle cells were cultured under normoxic or hypoxic conditions, and secreted exosomes were isolated for comprehensive molecular profiling. High-throughput miRNA sequencing combined with integrative bioinformatic analyses was used to uncover hypoxia-responsive regulatory networks and key miRNA hubs involved in skeletal muscle adaptation.
Methods:
HSMCs were cultured under normoxic or hypoxic conditions for 24 hours. Exosomes were isolated and characterized by transmission electron microscopy, nanoparticle tracking analysis, and immunoblotting. Exosomal miRNAs (n = 3 per group) were profiled using high-throughput sequencing, followed by differential expression, target prediction, enrichment, and network analyses.
Results:
Isolated exosomes displayed typical morphology (mean size: 82.4 ± 3.2 nm) and expressed markers CD9, CD63, and TSG101. Seventy-four miRNAs were significantly dysregulated under hypoxia (23 upregulated, 51 downregulated; FDR < 0.05, |log2FC| ⩾ 1), including upregulated hsa-miR-210-3p and downregulated hsa-miR-486-5p, hsa-miR-127-3p, and hsa-miR-126-3p. Predicted targets (~2000 genes) included 451 genes differentially expressed in hypoxic versus normoxic skeletal muscle cells. Functional enrichment highlighted cancer-related, MAPK, PI3K-Akt, and HIF-1 signaling pathways, along with muscle differentiation processes. Network analysis identified hsa-miR-20a-5p as a central regulatory hub (46 targets), followed by hsa-miR-24-3p and hsa-miR-152-3p. hsa-miR-24-3p showed the strongest disease associations in the miRNA-disease network.
Conclusions:
Hypoxia induces distinct exosomal miRNA signatures in skeletal muscle, regulating genes involved in differentiation, migration, and stress response. These findings suggest that exosome-mediated miRNA signaling contributes to hypoxia-driven muscle adaptation and intercellular communication.
Plain Language Summary
When muscle cells do not get enough oxygen (a condition called hypoxia), they change how they work and communicate. One way they send signals is through exosomes—tiny packages that carry microRNAs (miRNAs), which help control gene activity. In this study, we compared exosomes released by human muscle cells under normal oxygen and low-oxygen conditions. We looked at what miRNAs these exosomes carried and what genes those miRNAs might affect. We found that hypoxia caused major shifts in the exosomal miRNA profile: 74 miRNAs changed, with some increasing and others decreasing in low-oxygen condition. Important changes included increases in miR-210-3p and decreases in miR-486-5p, miR-127-3p, and miR-126-3p levels. These miRNAs are predicted to influence hundreds of genes involved in muscle function and stress responses. Further analysis showed that these miRNAs are linked to key pathways that help cells adapt to low oxygen, including HIF-1, MAPK, and PI3K-Akt signaling. One miRNA, miR-20a-5p, appeared to be a major regulator affecting many genes. Overall, our results suggest that when muscle cells experience low oxygen, they release exosomes with a unique set of miRNAs that help coordinate how cells adapt, grow, and respond to stress.
Introduction
Lower limb ischemia (LLI), a severe manifestation of peripheral artery disease (PAD), is characterized by reduced blood flow and oxygen supply to the lower extremities, often leading to pain, non-healing ulcers, and even limb amputation.1,2 Affecting approximately 236 million people worldwide as of 2015, with numbers continuing to rise, PAD represents a growing global health burden. 3 Modifiable risk factors account for approximately 70% of the global PAD burden, 4 underscoring the urgent need for effective therapeutic interventions. Therapeutic angiogenesis – the stimulation of new blood vessel formation – has emerged as a promising strategy to restore perfusion and promote tissue repair in ischemic limbs. 5 However, current therapeutic approaches, including gene therapy and recombinant growth factors, have shown limited efficacy and safety in clinical settings. Hence, novel therapeutic strategies are urgently needed.
Although hypoxia is recognized as a key driver of skeletal muscle adaptation, the precise regulatory pathways that coordinate these responses at the intercellular level remain incompletely understood. Most existing studies have focused on intracellular hypoxia-inducible factor (HIF)-dependent transcriptional programs regulating metabolism, angiogenesis, and stress responses within individual muscle fibers. 6 However, skeletal muscle functions as a highly integrated tissue, requiring coordinated communication among myofibers, endothelial cells, fibroblasts, and immune cells during hypoxic stress. Recent advances in extracellular vesicle biology have revealed that exosomes – nanosized vesicles secreted by virtually all cell types – play critical roles in intercellular communication.7,8 Notably, exosomes derived from skeletal muscle under hypoxic stress are increasingly recognized as potent mediators of tissue adaptation and regeneration.9,10 These exosomes encapsulate various bioactive molecules, including proteins, lipids, and non-coding RNAs, particularly microRNAs (miRNAs), which are known to post-transcriptionally regulate gene expression in recipient cells. 11 Nevertheless, the specific miRNA regulatory networks and key nodal miRNAs packaged into muscle-derived exosomes under hypoxic conditions remain largely undefined.7,12 Defining these regulatory pathways is essential for understanding how local hypoxic stress is translated into coordinated tissue-level adaptation.
Several hypoxia-responsive miRNAs (hypoxamiRs), such as hsa-miR-210, hsa-miR-126, and hsa-miR-23a, have been implicated in promoting endothelial cell survival, migration, and angiogenesis.13 -15 These miRNAs are often enriched in exosomes secreted under hypoxic conditions, enabling them to act as important mediators of intercellular signaling in ischemic tissues.16,17 Previous studies suggest that exosomes derived from hypoxia-conditioned human skeletal muscle cells (HSMCs) may contribute to vascular remodeling by delivering pro-angiogenic miRNAs to endothelial cells.18,19
Despite these promising preclinical findings, the translation of exosome-based therapies to clinical applications remains in its early stages. Accumulating evidence demonstrates that exosome-based therapies can modulate angiogenesis, inflammation, and tissue regeneration in hypoxia- and ischemia-related conditions, including cardiovascular and musculoskeletal disorders.20,21 Importantly, the therapeutic efficacy of exosomes appears to be highly dependent on their molecular cargo, particularly miRNAs that regulate key adaptive and repair pathways. 7 However, rational development of exosome-based interventions for skeletal muscle disorders is currently limited by an incomplete understanding of endogenous hypoxia-driven exosomal signaling mechanisms. Elucidating these pathways in physiological settings may therefore inform the design of more targeted, mechanism-based exosome therapies for muscle regeneration and hypoxia-associated diseases. Moreover, the specific alterations in exosomal miRNA cargo under hypoxia and the regulatory mechanisms through which these miRNAs influence angiogenesis in the context of lower limb ischemia are not fully elucidated.
Given the current limited clinical experience with exosome therapeutics, understanding the precise molecular mechanisms underlying skeletal muscle exosome-mediated angiogenesis is essential for advancing this therapeutic approach from bench to bedside. In this study, we aim to address this knowledge gap by systematically profiling exosomal miRNAs derived from HSMCs cultured under hypoxic versus normoxic conditions and performing downstream bioinformatic analyses to identify candidate miRNAs and pathways that may underlie their angiogenic effects. Our work provides novel insights into the role of skeletal muscle-derived exosomal miRNAs as endogenous modulators of angiogenesis in ischemic conditions and highlights their therapeutic potential in treating peripheral vascular diseases.
Methods
Human Skeletal Muscle Cells (HSMCs) Culture
The human skeletal muscle cell line (CP-H095, Wuhan Procell life science & technology Co., Ltd.) was donated by the Endocrinology Department of Shandong Provincial Hospital. The HSMCs were cultured for 24 hours under either hypoxic (1% O2) or normoxic (21% O2) conditions. For hypoxia induction, cells were maintained in a tri-gas incubator with a controlled atmosphere of 95% N2, 4% CO2, and 1% O2, simulating physiological low-oxygen conditions. Normoxic controls were cultured in a standard humidified incubator (37°C, 5% CO2, 21% O2). Both groups were grown in DMEM/F-12 medium supplemented with 10% exosome-depleted FBS (to prevent exogenous exosome interference) and 1% penicillin-streptomycin. After 24 hours, conditioned media were collected for subsequent exosome isolation.
Exosome Isolation
Following collection of conditioned media, exosomes were isolated through sequential centrifugation. First, cellular debris was removed by centrifugation at 3000g (4°C, 10 minutes), with repeated cycles if significant pellets were observed. The supernatant was then cleared of contaminating particles by centrifugation at 10 000g (4°C, 10 minutes) and transferred to fresh tubes. Exosome Concentration Solution (ECS) was added proportionally to the clarified supernatant, followed by vigorous vortexing (1 minute) and overnight incubation at 4°C (8-24 hours). Exosomes were pelleted by centrifugation (10 000g, 4°C, 60 minutes), with the supernatant carefully aspirated. The pellet was washed (10 000g, 4°C, 2 minutes) and resuspended in 1× PBS (200 μL per 20 mL initial media) through gentle pipetting. A final clarification step (12 000g, 4°C, 2 minutes) yielded purified exosomes in the supernatant, which were stored at −80°C for downstream analysis.
Exosome Characterization
Isolated exosomes were characterized using standardized techniques. For morphological analysis, exosomes were fixed in 2% paraformaldehyde, adsorbed onto 200-mesh copper grids, and negatively stained with 2% uranyl acetate before imaging with a transmission electron microscope (TEM; Hitachi HT7800, 80 kV) to verify cup-shaped ultrastructure. Particle size distribution and concentration were quantified by nanoparticle tracking analysis (NTA; Malvern NanoSight NS300) following dilution in PBS to achieve optimal particle counts (20-100 particles/frame). For molecular characterization, exosomal proteins were lysed in RIPA buffer, separated by 10% SDS-PAGE, and probed with antibodies against canonical exosomal markers (anti-CD9 [1:1000, Abcam ab92726], anti-CD63 [1:500, Santa Cruz sc-15363], and anti-TSG101 [1:1000, Invitrogen MA1-23 296]). Cellular contamination was assessed using anti-calnexin (1:1000, Cell Signaling #2679) as a negative control, with β-actin serving as a loading control.
Exosomal miRNA Sequencing
To investigate hypoxia-induced alterations in exosomal miRNA profiles, we performed comprehensive small RNA sequencing on exosomes isolated from hypoxic (1% O2) and normoxic (21% O2) HSMCs. Library preparation was conducted using the NEBNext® Multiplex Small RNA Library Prep Set for Illumina® (NEB #E7300S, USA) following manufacturer protocols with modifications for exosomal RNA input. Briefly, 3 μg of exosomal RNA per sample underwent 3′- and 5′-adapter ligation, first-strand cDNA synthesis, and PCR amplification (12 cycles) with unique barcodes for multiplexing. Library quality was rigorously assessed using the Agilent Bioanalyzer 2100 system with High Sensitivity DNA chips (Agilent #5067-4626), confirming optimal size distributions (140-160 bp) and absence of adapter dimers. Quantification was performed via qPCR using the KAPA Library Quantification Kit (Roche #KK4824). Pooled libraries were sequenced on an Illumina NovaSeq 6000 platform (50 bp read) to achieve >10 million reads per sample, ensuring statistical power for differential expression analysis.
Exosomal miRNA Sequencing Data Processing
The miRNA sequencing data was processed using sRNAtoolbox version 2022, 22 a comprehensive platform for small RNA analysis. Raw FASTQ files were uploaded to the sRNAbench server and processed through its integrated pipeline. Adapter trimming was performed with a minimum adapter length of 10 nucleotides allowing a mismatch of 1 base between the adapter and reference sequences. The quality filtering retained reads with a mean phred score above 20 and minimum length of 15 nucleotides. The filtered reads were aligned to the reference genome and miRBase database version 22.1 23 using Bowtie aligner 24 with the following parameters: maximum of 1 mismatch permitted per alignment, seed length of 20 nucleotides for initial mapping, and maximum of 10 multiple mapping locations allowed per read to balance sensitivity and specificity. miRNA abundance quantification was performed using raw read counts, followed by normalization to reads per million (RPM) to account for differences in sequencing depth between samples.
Differential Expression Analysis
Differential expression analysis of miRNAs was conducted using sRNAde, an integrated module within the sRNAtoolbox platform that implements multiple statistical methods for robust differential expression detection. The module encompasses several established algorithms including DESeq, DESeq2, NOISeq, paired/unpaired t-tests, and edgeR, each with distinct statistical assumptions and normalization approaches. We selected DESeq2 25 as the primary method due to its superior performance with count-based RNA-seq data, utilizing negative binomial distribution modeling to account for overdispersion, commonly observed in small RNA sequencing datasets and batch function to remove the batch effects. Multiple testing correction was applied using the Benjamini-Hochberg false discovery rate (FDR) method. miRNAs were considered differentially expressed (DEMs) if they met the following criteria: FDR-adjusted P-value < .05 and log2-fold change ⩾ 1. To enhance the robustness and reliability of our findings, we implemented a consensus approach by retaining only those DEMs that were independently identified as significantly differentially expressed by at least 1 additional method among the available algorithms, thereby reducing method-specific false discoveries and increasing confidence in the biological relevance of the identified miRNAs.
miRNA Target Prediction
The miRNA target prediction for DEMs was performed using miRWalk database version 3.0, 26 which provides comprehensive target prediction through multiple complementary algorithms. The primary prediction method employed TarPmiR, a machine learning-based algorithm utilizing random forest classification that incorporates sequence complementarity, thermodynamic stability, and evolutionary conservation features. 27 To enhance prediction reliability, miRWalk integrates predictions from 2 additional well-established algorithms: TargetScan version 7.2, 28 which focuses on conserved seed matches and context scores, and miRDB version 6.0, 29 which employs support vector machine-based prediction with experimental training data. Experimentally validated miRNA-target interactions were incorporated from miRTarBase version 9.0, 30 which contains manually curated interactions supported by reporter assays, Western blot, qPCR, and other experimental evidence. To ensure high-confidence predictions and minimize false positives, only targets that were consistently predicted by all the 3 algorithms (TarPmiR, TargetScan, and miRDB) were retained.
miRNAs mainly affect cellular processes by regulating the expression of their target genes. Although the predicted targets obtained by us contain miRNA binding sites, all the targets may not be functionally important in the skeletal muscle and/or under hypoxic conditions. Hence, we filtered the targets that exhibited a differential expression (FDR P < .05 and |logFC ⩾ 1|) between hypoxia and normoxia in mouse C2C12 skeletal muscle cell line (dataset GSE114086), a model that has been extensively used in muscle research. The differentially expressed targets were obtained from the iHypoxia database 31 and were used for further analysis.
Functional Enrichment Analysis of Target Genes
The predicted miRNA gene targets that were also differentially expressed in hypoxic versus normoxic condition in skeletal muscle cell line were subjected to functional enrichment analysis using WebGestalt (WEB-based GEne SeT AnaLysis Toolkit) 32 to identify significantly enriched biological pathways, and Gene Ontology (GO) terms. The analysis utilized the Over-Representation Analysis (ORA) method with protein-coding genes serving as the reference gene set for background comparison. Multiple testing correction was applied using the Benjamini-Hochberg false discovery rate (FDR) method, with enriched terms considered statistically significant at FDR-adjusted P < .1. Targets of significantly up- and down-regulated miRNAs were analyzed separately to capture direction-specific functional impacts.
miRNA-Target Network Construction
The DEMs along with their differentially expressed targets were used for network construction and analysis. The miRNA-target network was constructed and analyzed using Cytoscape version 3.10.3. 33 The network statistics including the degree and betweenness of the nodes were obtained using the CytoHubba plugin integrated within Cytoscape.
miRNA-Disease Network Construction
The DEMs were submitted to miRNet 34 for constructing miRNA-disease networks. By integrating multiple high-quality miRNA-target data sources and advanced statistical methods, miRNet allows to easily navigate the complex landscape of miRNA-target interactions to obtain deep biological insights. The miRNA-disease interactions have been collected from literature along with experimental and study details. Currently the database contains ~32 000 miRNA-disease interactions.
Result
Characterization of Hypoxic Skeletal Muscle-Derived Exosomes
Hypoxic exposure duration was first optimized to ensure cell viability while maximizing exosome yield. HSMCs maintained normal morphology during hypoxia for <24 hours, whereas extended exposure (36-48 hours) triggered apoptotic changes (Figure 1A). TEM imaging of isolated vesicles confirmed classical exosomal ultrastructure, with 92.3% of particles exhibiting cup-shaped morphology (n = 427 counted vesicles) and diameters conforming to the expected 30 to 150 nm range (Figure 1B). NTA quantification revealed a mean particle size of 82.4 ± 3.2 nm (mode: 78.1 nm) with a peak concentration of 3.8 × 1010 particles/mL (Figure 1C). Immunoblotting demonstrated strong enrichment of exosomal markers CD9 (24 kDa), CD63 (53 kDa), and TSG101 (44 kDa), while calnexin (90 kDa) was undetectable – confirming the absence of endoplasmic reticulum contaminants (Figure 1D). The combined datasets established that our isolation protocol reliably yielded high-purity exosomes from hypoxic HSMCs, with physical and molecular characteristics matching established exosome criteria.
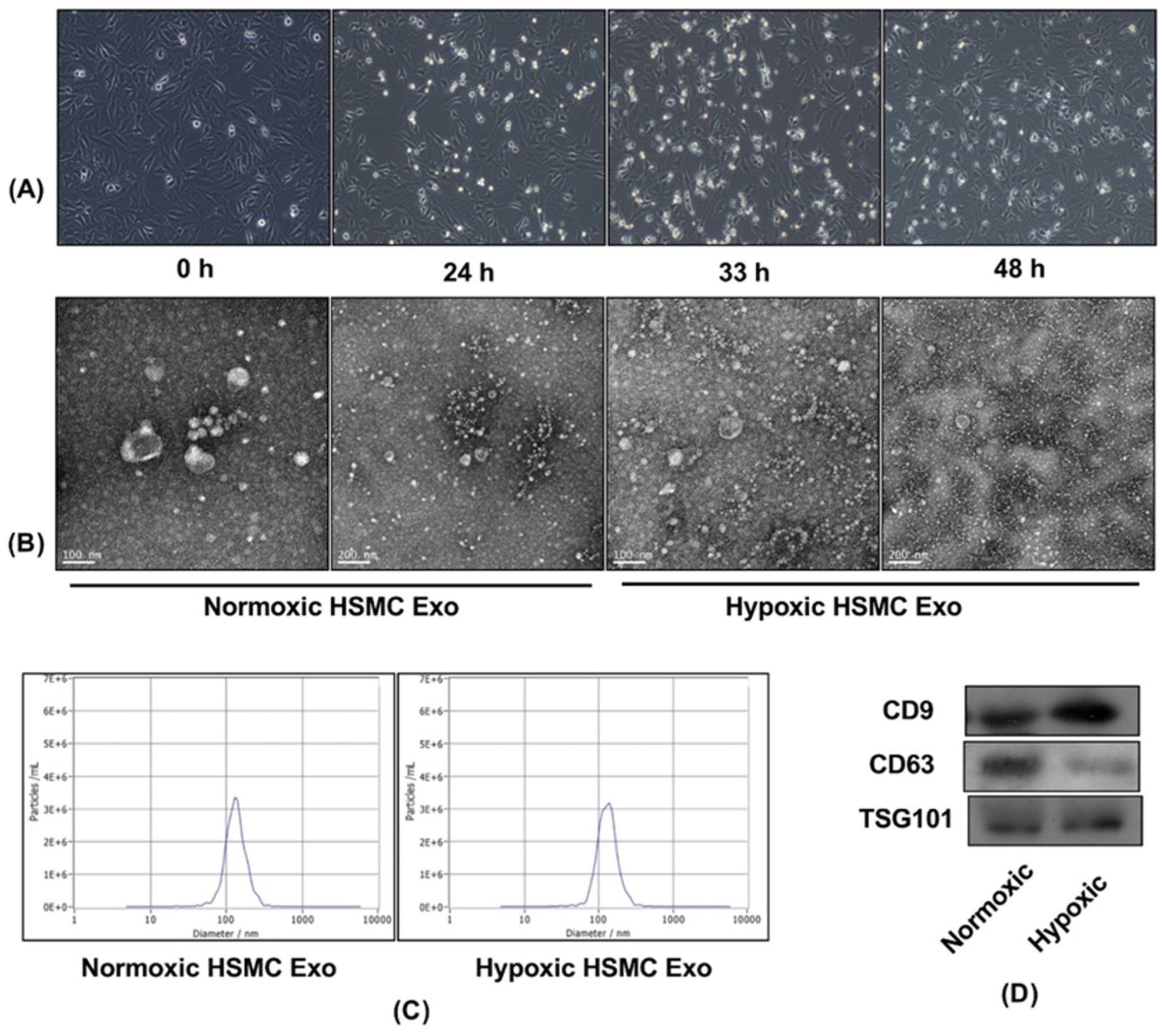
Characteristics of exosomes from skeletal muscle cells: (A) the 24-hour hypoxic condition was selected for the follow-up experiment because the hypoxic cells showed no morphological change at a duration shorter than 24 hours, (B) the morphological characteristics of exosomes under an electron microscope, (C) the distribution of particle size and concentration, and (D) the Western Blot revealed the presence of exosome-specific proteins CD9, CD63, and TSG101 in both normoxic and hypoxic exosomes from HSMC cells.
miRNA Sequencing and Analysis of Hypoxic Muscle Cell Derived Exosomes
High-throughput miRNA sequencing was performed on exosomes isolated from muscle cells under normoxic (n = 3, C1-3) and hypoxic (n = 3, Q1-3) conditions using the Illumina platform. Three replicates provide enough data to assess variability, perform proper normalization, and conduct meaningful statistical analysis, particularly when combined with strict criteria for identifying significant changes. A total of ~75 million raw reads were generated across all samples, with an average of 12.4 million reads per sample. As shown in Table 1, following quality control and adapter trimming, 93.5% of reads passed filtering criteria, yielding an average of ~11.6 million high-quality reads per sample (range: 10.7-13.5 million). Alignment to the reference genome and miRBase database (v22.1) achieved an average mapping rate of 65%, with 14% of mapped reads corresponding to known miRNAs (Supplemental File 1). A total of 295 unique miRNAs (average of ~275 miRNAs) were detected across all samples (Figure 2A). Supplemental File 2 contains the list of miRNAs detected per sample. Principal component analysis (PCA) demonstrated clear separation between normoxic and hypoxic samples, with PC1 and PC2 explaining 52.6% and 19.4% of the total variance, respectively based on the expression of these miRNAs (Figure 2B).
Raw read trimming and filtering.

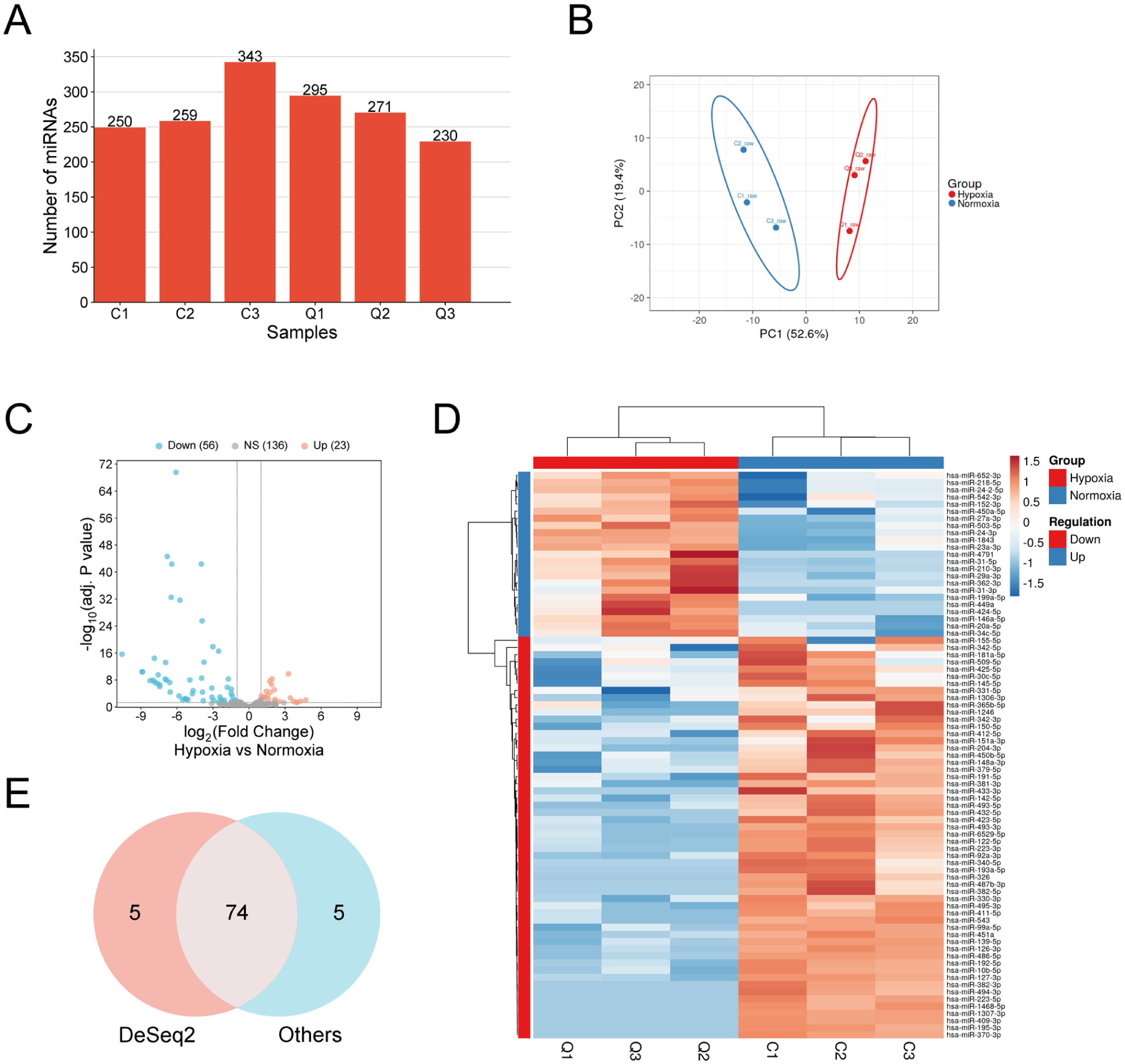
miRNA-seq analysis of skeletal muscle derived exosomes under hypoxic conditions. (A) Number of miRNAs detected per sample. C1 to C3: normoxic controls; Q1 to Q3: hypoxic samples. (B) Principal Component Analysis showing clear separation between normoxic (blue) and hypoxic (red) samples based on miRNA expression profiles. PC1 and PC2 explain 52.6% and 19.4% of variance, respectively. (C) Volcano plot showing differentially expressed miRNAs between hypoxia and normoxia conditions. The x-axis represents log2(fold change) and y-axis shows −log10(adjusted P-value). Significantly upregulated miRNAs (n = 23) are shown in red, downregulated miRNAs (n = 56) in blue, and non-significant miRNAs (n = 136) in gray. (D) Hierarchical clustering heatmap of differentially expressed miRNAs across samples. Rows represent individual miRNAs and columns represent samples grouped by condition (hypoxia in red, normoxia in blue). The color scale indicates expression levels with red representing high expression and blue representing low expression. Dendrograms show clustering relationships between miRNAs (left) and samples (top). (E) Venn diagram comparing differentially expressed miRNAs identified by DESeq2 analysis with those identified by other methods, showing 74 miRNAs commonly identified by both approaches.
Differentially Expressed miRNAs (DEMs)
Differential expression analysis identified 79 significantly dysregulated miRNAs with FDR-adjusted P < .05, |log2FC| ⩾ 1 (Figure 2C and D and Supplemental File 3). Among these, 74 were significantly differentially expressed by at least 1 other algorithm (Figure 2E) and comprised of 23 up-regulated and 51 down-regulated miRNAs in hypoxic conditions compared to normoxic controls. Top 5 up- and down-regulated miRNAs are provided in Table 2. Hierarchical clustering analysis of differentially expressed miRNAs showed distinct expression patterns that clearly distinguished hypoxic from normoxic exosome samples.
Top 10 differentially expressed miRNAs between hypoxic and normoxic samples.

miRNA Target Prediction
Target prediction was performed for the differentially expressed miRNAs using miRWalk database. The analysis employed 3 complementary algorithms: TarPmiR (miRWalk’s native random forest-based method), TargetScan, and miRDB, each utilizing distinct computational approaches for target prediction. To ensure high-confidence predictions and minimize false positives, only targets that were consistently predicted by all 3 algorithms (TarPmiR, TargetScan, and miRDB) were retained for downstream analysis. This stringent consensus approach identified 1974 putative target genes (with ~2678 miRNA-gene interactions) for the 53 (among the 74) differentially expressed miRNAs, with an average of 50 targets per miRNA (Supplemental File 4). The up-regulated miRNAs (n = 17) were predicted to target 893 genes, while down-regulated miRNAs (n = 36) showed predicted interactions with 797 genes, with 284 genes being common targets of both up- and down-regulated miRNAs. Among the most highly connected miRNAs, hsa-miR-20a-5p showed the highest number of predicted targets (n = 209), followed by hsa-miR-92a-3p (n = 155) and hsa-miR-27a-3p (n = 148). Cross-validation with experimentally validated interactions from miRTarBase revealed that 19% of our predicted targets had prior experimental support, providing confidence in the computational predictions. Furthermore, we filtered the targets that showed a differential expression (FDR < 0.05 and |logFC ⩾ 1|) between hypoxia and normoxia in mouse C2C12 skeletal muscle cell line (dataset GSE114086), a model that has been extensively used in muscle research. This resulted in a total of 270 genes targeted by upregulated miRNAs and 246 genes targeted by downregulated miRNAs, with 65 being common by both miRNA types (Supplemental File 4).
Functional Enrichment Analysis of Predicted Targets
miRNAs primarily negatively and sometimes positively regulate the expression of mRNAs. Hence, we analyzed the functional roles of the genes that showed differential expression in hypoxic versus normoxic skeletal muscle murine cell line. Based on the enrichment analysis, up- and down-regulated miRNA targets showed significant enrichment in multiple cancer-related KEGG pathways (Figure 3A and Supplemental File 5), with ‘Pathways in cancer’ demonstrating the highest significance (n gene: 27; FDR < 0.001). Additionally, the enriched pathways included cardiac related pathways and multiple signaling pathways. The signaling pathways that were enriched by the upregulated miRNA targets include MAPK signaling, PI3K-Akt signaling, HIF-1 signaling and mTOR signaling, all of which have been shown to be important in regulating gene expression under hypoxic conditions. The downregulated miRNA targets were enriched for FoxO signaling pathway.

Enrichment analysis of upregulated and downregulated miRNA targets. (A) KEGG pathway and (B) Gene Ontology Biological Processes (GO-BP) enrichment analysis for differentially expressed miRNA targets. X-axis indicates ratio, whereas y-axis shows the pathway or GO-BP terms. The color gradient indicates the P-value significance. Number of genes associated with each term are indicated by dot size. Top 20 significant pathways/processes are shown.
The corresponding Gene Ontology Biological Process analysis (Figure 3B and Supplemental File 5) revealed highest enrichment in Muscle cell differentiation (−log10(FDR) ~3.0), followed by Ameboidal-type cell migration, Fat cell differentiation, and Tissue migration, with enrichment ratios spanning 4 to 10 and gene counts of 5 to 20. Other significantly enriched biological processes included muscle organ development, response to oxygen levels, regulation of vascular development, cell-cell signaling, mesenchyme development, cellular response to vascular endothelial growth factor stimulus, and various developmental and stress-response processes including fibroblast proliferation, and response to oxidative stress. Interestingly, the muscle-related processes were enriched by both up- and down-regulated miRNA targets. These results indicate that miRNAs under hypoxic conditions target genes involved in cellular differentiation, migration, and developmental processes, suggesting their role in hypoxia-induced cellular reprograming and homeostasis.
miRNA-Target Network
The miRNA-gene network was constructed using differentially expressed miRNAs and genes in the skeletal muscle under hypoxic conditions. The core network consisted of 479 nodes, where 41 miRNAs were connected with 438 target genes with 610 edges (Figure 4 and Supplemental File 6). The network is highly dense with an average node degree of ~15 for the miRNAs. The network analysis reveals a miRNA-target gene regulatory network dominated by a few highly connected microRNA hubs alongside important protein-coding genes. hsa-miR-20a-5p emerges as the central regulatory node with 46 connections and the highest betweenness centrality (50 496), followed by hsa-miR-24-3p and hsa-miR-152-3p with 34 connections each, and hsa-miR-449a with 33 connections. Among the protein-coding genes, RORA stands out with 6 connections and high betweenness centrality (15 587), while genes like CADM1, PPM1A, SH3PXD2A, UHMK1, and AGO1 also show moderate connectivity (5 connections each) with varying regulatory importance. The network displays a typical scale-free topology where most nodes have low connectivity (degree 1-2), including many single-connection genes like VEGFA, PTEN, FOXO1, and KLF4 that represent specialized regulatory targets. The high bottleneck scores for hub miRNAs and key genes indicate they are critical control points for information flow through the network, suggesting their potential importance as therapeutic targets or disease biomarkers. This hierarchical structure reflects the regulatory architecture of biological systems where key miRNAs can simultaneously modulate numerous downstream genes, while certain protein-coding genes serve as important intermediate regulators, amplifying regulatory impact across cellular pathways.
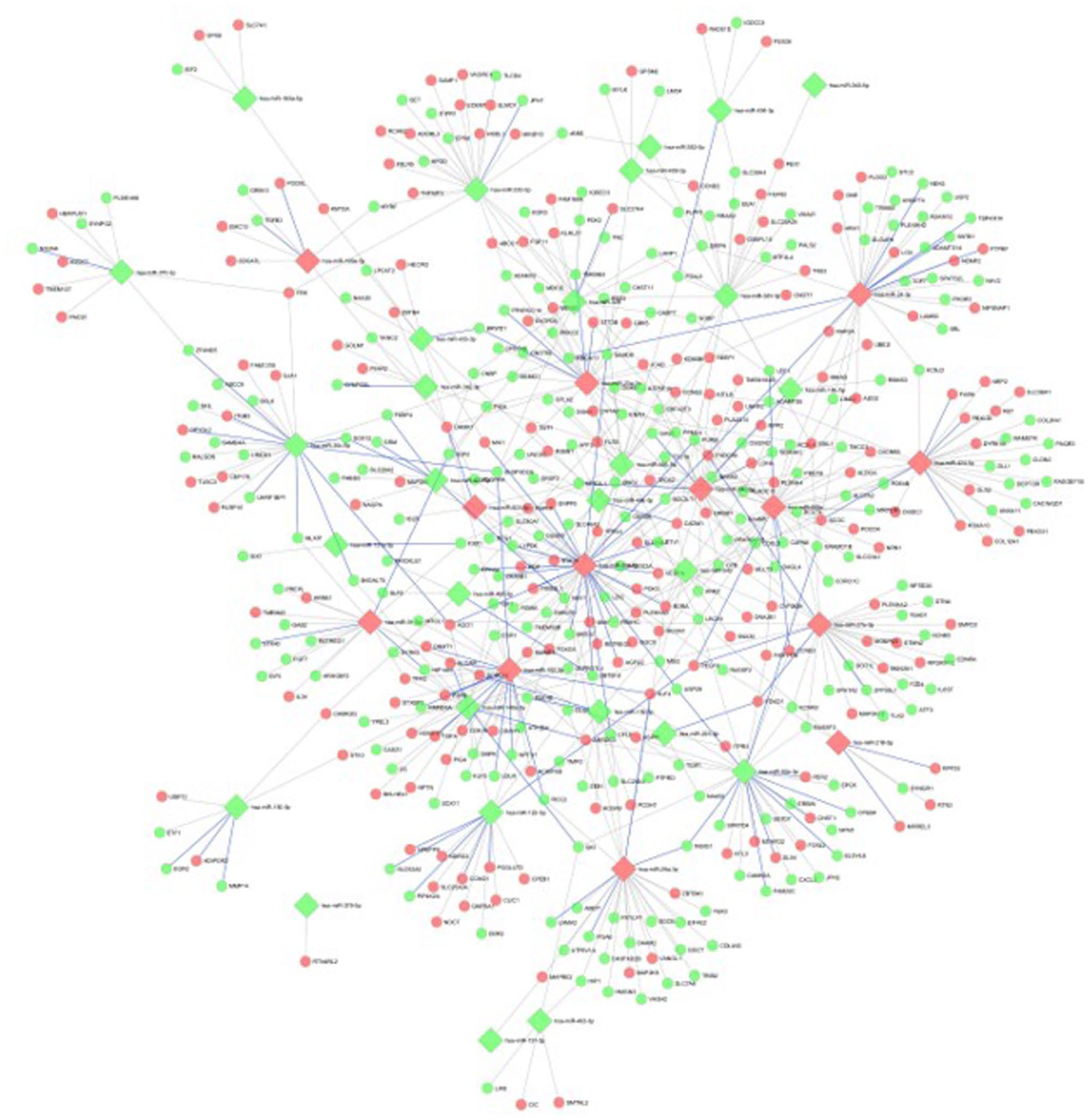
miRNA-target gene regulatory network under hypoxic conditions. The network illustrates interactions between differentially expressed exosomal miRNAs and their predicted or experimentally validated target genes identified under hypoxic conditions. Nodes represent miRNAs or target genes (mRNAs), while edges denote miRNA–mRNA regulatory interactions. Diamond-shaped nodes indicate miRNAs, and circular nodes indicate target genes. Node color reflects differential expression status, with red denoting upregulated miRNAs or mRNAs and green denoting downregulated miRNAs or mRNAs under hypoxia. Blue, thick edges represent experimentally validated miRNA-target interactions, whereas gray edges indicate computationally predicted interactions. The dense connectivity highlights the complexity of hypoxia-responsive post-transcriptional regulation, where individual miRNAs can regulate multiple target genes and single genes may be co-regulated by multiple miRNAs.
miRNA-Disease Network
The DEMs were used for constructing a miRNA-disease network using miRNet. Of the 74 DEMs, 20 were found to be linked to 473 different diseases (Figure 5). hsa-miR-24-3p was found to be linked with most number of diseases (n = 100) followed by hsa-miR-92a-3p (n = 99) and hsa-miR-451a (n = 98). Interestingly, hsa-miR-24-3p was among the top 2 in terms of degree when differentially expressed targets were considered, whereas hsa-miR-92a-3p was in the top 2 when all the targets were considered. This supports the importance of these miRNAs in hypoxic skeletal muscle cells. Furthermore, both these miRNAs were associated with ischemia, whereas hsa-miR-24-3p and hsa-miR-451a were also associated with muscle diseases. hsa-miR-92a-3p, hsa-miR-199a-5p, hsa-miR-409-3p, hsa-miR-486-5p, hsa-miR-493-3p were found to be associated with muscular dystrophy. Among the diseases involved, cancer was linked to most of the miRNAs (n = 7) followed by heart failure (n = 6) and Muscular Dystrophy, Duchenne (n = 5). Details of miRNA-disease associations can be found in Supplemental File 6.

miRNA-Disease Association Network. Network visualization of miRNA-disease associations generated using the miRNet platform. Blue rectangular nodes represent individual miRNAs, while pink nodes represent associated diseases or pathological conditions. Edges indicate reported miRNA-disease associations curated within the miRNet database. The size of the miRNA nodes corresponds to their degree of connectivity, with larger nodes indicating miRNAs associated with a higher number of diseases. Highly connected miRNAs may represent key regulatory hubs with broad relevance to disease pathogenesis.
Discussion
In the current study, we performed miRNA expression analysis of hypoxic skeletal-muscle cell derived exosomes. Differential analysis identified a total of 79 miRNAs with a significant difference between hypoxic and normoxic conditions, of which 74 were reported to be differentially expressed by at least 2 different methods (including DESeq2). The expression of these miRNAs was able to clearly segregate the hypoxic and normoxic samples. hsa-miR-210-3p was found to be the most significantly upregulated miRNA, whereas hsa-miR-486-5p was found to be the most significantly downregulated miRNA. hsa-miR-210-3p, a hypoxia-inducible microRNA, is known to play a central role in skeletal muscle during hypoxia. 35 It is upregulated in response to low oxygen levels and is associated with changes in mitochondrial function, muscle atrophy, and fibrosis. 36 It has been demonstrated that hsa-miR-210-3p has significant protective effects in muscle tissue during hypoxic conditions. In the ischemic skeletal muscle, it seems to be cytoprotective, regulating oxidative metabolism and oxidative stress. 37 The protective effects of hsa-miR-210-3p in muscle tissue involve several pathways including angiogenesis, metabolic reprograming and anti-apoptotic signaling. Furthermore, hsa-miR-210 is known to be a negative regulator of mitochondrial respiration, 38 suggesting its role in metabolic adaptation during hypoxia. hsa-miR-486 is a muscle-enriched microRNA, or ‘myomiR’, that is particularly important for muscle function and maintenance. It is a useful biomarker for dystrophic disease progression and its reduced expression is correlated with Duchenne muscular dystrophy (DMD). 39 Exosomal hsa-miR-486 has been shown to regulate hypoxia-induced erythroid differentiation of erythroleukemia cells by targeting Sirt1. 40 The other top miRNAs that showed strong differential expression between hypoxic and normoxic condition were hsa-miR-127-3p, hsa-miR-126-3p, hsa-miR-122-5p, and hsa-miR-450a-5p. hsa-miR-127-3p plays an important role in skeletal muscle development and myoblast proliferation and differentiation, potentially affecting the growth and development of muscle cells. 41 hsa-miR-126-3p also plays a crucial role in muscle function by modulating the levels of transcripts that are involved in myogenic differentiation. 42 hsa-miR-122-5p has been implicated in regulating fatty acid metabolism in skeletal muscle, potentially impacting muscle function during hypoxic conditions. 43 hsa-miR-450a-5p is significantly dysregulated in skeletal muscles of cachectic individuals suffering from tumors. 44 It is noteworthy that all the top miRNAs were significantly differentially expressed by multiple algorithms, confirming their strong association with hypoxia in skeletal muscle.
Next, to understand the role of these miRNAs with various cellular processes, we identified the corresponding target genes. The target prediction was performed using 3 different algorithms, TarPmiR, TargetScan, and miRDB, and those that were identified by all 3 algorithms were considered for further analysis. The interaction of the DEMs with approximately 2000 targets suggested the functional importance of these miRNAs in hypoxia and skeletal muscle. hsa-miR-20a-5p targeted the most number of genes followed by hsa-miR-27a-3p and hsa-miR-92a-3p. While the first 2 are upregulated, the later is downregulated under hypoxia. However, all 3 play an active role in regulating muscle function/differentiation under hypoxic conditions. hsa-miR-20-E2F1 axis has been linked to auto-regulatory feedback loop involved in myoblast proliferation and differentiation. 45 miRNA-92a regulates hypoxic stimulation of myogenic lineage differentiation in mouse embryonic stem cells. 46 Furthermore, decreased hsa-miR-92a-3p expression has been shown to potentially mediate the pro-angiogenic effects of oxidative stress-activated endothelial cell-derived exosomes by targeting tissue factor involved in angiogenesis. 47 hsa-miR-27a-3p targets ATF3 and reduces calcium deposition in vascular smooth muscle cells. 48 It plays a role in hypoxia-induced muscle dysfunction, affects muscle fiber size and arrangement by modulating insulin signaling. Furthermore, it is deregulated in chronic intermittent hypoxia (CIH), leading to reduced expression in skeletal muscle. 49 While we identified ~2000 miRNA targets, all of them may not be functionally significant. MiRNAs affect the cellular processes by deregulating the expression of their target genes, primarily by suppressing, but rarely by elevating the expression of their target genes through different mechanisms. Therefore, we shortlisted the targets that were differentially expressed under hypoxia in mouse skeletal muscle cell line C2C12, a well-known model for muscle research. The mouse cell line was selected due to the unavailability of a relevant high-throughput human study for comparison; nonetheless, it provides an opportunity for the cross-species validation of our targets. Among the differentially expressed (DE) targets, RORA, CADM1, PPM1A, SH3PXD2A, UHMK1, and AGO1 were among the top. Kim et al, 50 provide novel insights into the function of retinoic acid-receptor-related orphan receptor (RORA) in amplification of hypoxia signaling and suggest it as a potential therapeutic target for hypoxia-associated vascular diseases. Argonaute 1 (AGO1), a key component of the microRNA-induced silencing complex, plays a role in hypoxic response, mainly in muscle and endothelial cells. It is linked to hypoxia-induced angiogenesis and the regulation of various genes involved in muscle metabolism and function. 51
The differential expression of exosomal miRNAs observed under hypoxic conditions provides important insights into the regulatory mechanisms that support skeletal muscle function during oxygen deprivation. Hypoxia imposes substantial metabolic and oxidative stress on muscle tissue, necessitating coordinated adaptations involving energy metabolism, angiogenesis, cellular survival, and regeneration. The hypoxia-responsive miRNAs identified in this study are therefore likely to contribute to muscle adaptation by modulating pathways involved in mitochondrial function, stress tolerance, and vascular remodeling. Functional enrichment analysis of the targets of differential miRNAs identified pathways and processes potentially modulated by the DEMs. Both up- and down-regulated miRNA targets were enriched for multiple cancer related pathways. This could be because hypoxia modulates several molecules associated with cancer related processes. 52 We also identified a few heart-related pathways, which could be attributed to the muscular property of the organ. In addition, several signaling pathways, including HIF-1 signaling, PI3K-Akt signaling, MAPK signaling, mTOR signaling were enriched by the targets of upregulated miRNAs, whereas FoXO signaling was enriched by the targets of downregulated genes. All these signaling pathways have been shown to be associated with hypoxia53 -55 with HIF-1 signaling being prominently involved in skeletal muscle’s response to hypoxia. 56 Under low oxygen, HIF-1α accumulates and activates genes involved in energy metabolism and angiogenesis, which is important for both physiological conditions like exercise and pathological conditions like muscle fibrosis.56 -58 Among the enriched biological processes, muscle cell differentiation was found to be the most enriched. The top 20 GO-BP terms contained several muscle development-related biological processes. The other key processes enriched were response to oxygen levels, Wnt signaling, and vasculature development.
The miRNA-target network constructed using differentially expressed miRNAs as well as their targets revealed interesting findings. This network revealed hsa-miR-20a-5p, hsa-miR-24-3p, and hsa-miR-152-3p to be top 3 based on the degree (number of differentially expressed targets). Interestingly, hsa-miR-24-3p was also found to be linked to most number of diseases in the miRNA-diseases network, including muscle disease and ischemia, which was in agreement with the published literature.59,60 Hub miRNAs are often functionally critical because they integrate and coordinate multiple downstream pathways rather than acting as isolated regulators. 61 miR-20a-5p is a member of the miR-17-92 cluster, a well-established regulator of hypoxia-responsive processes, including cell-cycle control, angiogenesis, metabolic adaptation, and stress signaling. 62 In the context of skeletal muscle, such regulatory functions are particularly relevant during ischemic or hypoxic stress, where rapid restoration of perfusion and controlled myogenic proliferation are required for effective repair. The identification of miR-20a-5p as a hub miRNA in our exosomal network therefore suggests that it may function as a master coordinator of hypoxia-driven signaling cascades in recipient cells following exosome uptake. In contrast, miR-24-3p, exhibited extensive disease associations and substantial target breadth, which is consistent with its reported involvement in muscle biology, inflammation, oxidative stress regulation, and tissue remodeling. 63 miR-24-3p has also been reported to respond to hypoxic stress and to influence gene expression programs across multiple cell types, supporting its role as a broadly acting effector miRNA in intercellular communication. 64 Its presence in hypoxic muscle-derived exosomes suggests a role in disseminating stress-adaptive signals that influence both local and systemic responses during muscle injury or regeneration. While many miRNAs targeted both up- and down-regulated genes, some of the miRNAs targeted the genes that were specifically up- or down-regulated under hypoxia in skeletal muscle. For instance, hsa-miR-122-5p is downregulated under hypoxia, while most of its targets were up-regulated under hypoxic condition in skeletal muscle cell line. Similarly, hsa-miR-29a-3p was up-regulated, while most of its targeted genes were found to be downregulated under hypoxia in skeletal muscle. Both these miRNAs play critical roles in hypoxia and/or muscle tissue.43,65 hsa-miR-29a-3p could inhibit PI3K protein expression, resulting in a reduction in AKT phosphorylation levels and HIF-1α protein expression levels. 66 A recent study has shown that hsa-miR-29a-3p can be released by skeletal muscle as a myokine (a muscle-derived signaling molecule) after resistance exercise. 67 hsa-miR-122-5p plays a complex role in hypoxia and muscle, influencing various processes like fat metabolism, inflammation, and fibrosis. Furthermore, hypoxia can modulate hsa-miR-122 expression, potentially affecting skeletal muscle function and contributing to fat metabolism related diseases. 43
Although our study sheds light on the miRNA-target network and reveals several pathways associated with miRNAs underlying hypoxia in skeletal muscle derived exosomes, there are a few limitations. First, the differential expression of the target genes needs to be studied in human skeletal muscle tissue to reveal the mechanisms associated with hypoxia in detail. Second, the sequencing result needs to be validated using low-throughput techniques. Third, the current study used 3 samples each for hypoxic and normoxic conditions, limiting the statistical power. Increasing the sample size may result in slight variation in our results including higher number of differentially expressed miRNAs, but we believe that this will not drastically change the core findings including the pathway and functional analysis results. Nevertheless, a higher number of samples will improve statistical power and reduce inter-sample variability, thereby strengthening the generalizability and reproducibility of our findings.
Conclusion
This study demonstrates that hypoxic skeletal muscle-derived exosomes carry a reprogramed miRNA cargo that orchestrates cellular adaptation to oxygen deprivation. The identification of 74 differentially expressed miRNAs, including downregulation of muscle-specific hsa-miR-486-5p and upregulation of the hypoxia-responsive hsa-miR-210-3p, highlights coordinated hypoxic remodeling of exosomal signaling. Enrichment of target genes in HIF-1, PI3K-Akt, MAPK, and muscle differentiation pathways underscores the capacity of these miRNAs to regulate metabolism, survival, and myogenic responses. Network analysis further identifies hsa-miR-20a-5p as a central regulatory hub targeting key genes such as VEGFA, suggesting that hypoxic muscle-derived exosomes act as adaptive signals promoting angiogenesis and cellular resilience under oxygen-limited conditions.
Future studies integrating functional validation, in vivo ischemia models, and exosome-based intervention strategies will be essential to establish causality and translational relevance of the miRNAs identified in the current study. Evaluating the effects of hypoxia-conditioned or miRNA-enriched exosomes on angiogenesis, inflammation, metabolic recovery, and muscle regeneration will be essential to establish causality and therapeutic efficacy. In parallel, systematic assessment of exosome biodistribution, dosing, and safety in preclinical models will be critical for translational development. These efforts could ultimately inform early-phase clinical trials exploring exosome-based miRNA therapies for muscle-related diseases characterized by hypoxia or ischemia, such as peripheral artery disease, and ischemia-reperfusion injury, thereby bridging mechanistic discovery and clinical application.
Supplemental Material
sj-xlsx-1-bmi-10.1177_11772719261427170 – Supplemental material for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs
Supplemental material, sj-xlsx-1-bmi-10.1177_11772719261427170 for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs by Haijiao Wang, Baochang Shi, Wenting Song, Lu Lu, Akhilesh K. Bajpai and Qiu Li in Biomarker Insights
Supplemental Material
sj-xlsx-2-bmi-10.1177_11772719261427170 – Supplemental material for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs
Supplemental material, sj-xlsx-2-bmi-10.1177_11772719261427170 for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs by Haijiao Wang, Baochang Shi, Wenting Song, Lu Lu, Akhilesh K. Bajpai and Qiu Li in Biomarker Insights
Supplemental Material
sj-xlsx-3-bmi-10.1177_11772719261427170 – Supplemental material for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs
Supplemental material, sj-xlsx-3-bmi-10.1177_11772719261427170 for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs by Haijiao Wang, Baochang Shi, Wenting Song, Lu Lu, Akhilesh K. Bajpai and Qiu Li in Biomarker Insights
Supplemental Material
sj-xlsx-4-bmi-10.1177_11772719261427170 – Supplemental material for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs
Supplemental material, sj-xlsx-4-bmi-10.1177_11772719261427170 for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs by Haijiao Wang, Baochang Shi, Wenting Song, Lu Lu, Akhilesh K. Bajpai and Qiu Li in Biomarker Insights
Supplemental Material
sj-xlsx-5-bmi-10.1177_11772719261427170 – Supplemental material for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs
Supplemental material, sj-xlsx-5-bmi-10.1177_11772719261427170 for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs by Haijiao Wang, Baochang Shi, Wenting Song, Lu Lu, Akhilesh K. Bajpai and Qiu Li in Biomarker Insights
Supplemental Material
sj-xlsx-6-bmi-10.1177_11772719261427170 – Supplemental material for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs
Supplemental material, sj-xlsx-6-bmi-10.1177_11772719261427170 for Molecular Insights into the Regulatory Mechanisms Mediated by Hypoxia-Conditioned Skeletal Muscle Exosomal miRNAs by Haijiao Wang, Baochang Shi, Wenting Song, Lu Lu, Akhilesh K. Bajpai and Qiu Li in Biomarker Insights
Footnotes
Acknowledgements
None.
Ethical Considerations
None.
Consent to Participate
None.
Author Contributions
Conceptualization, Qiu Li; Investigation, Haijiao Wang and Baochang Shi; Data curation, Baochang Shi; Formal analysis, Haijiao Wang, Baochang Shi, Wenting Song and Akhilesh Bajpai; Methodology, Haijiao Wang, Lu Lu, Akhilesh Bajpai and Qiu Li; Project administration, Qiu Li; Supervision, Akhilesh Bajpai and Qiu Li; Validation, Haijiao Wang, Baochang Shi, Wenting Song and Lu Lu; Visualization, Akhilesh Bajpai; Writing – original draft, Haijiao Wang, Baochang Shi and Akhilesh Bajpai; Writing – review & editing, Wenting Song, Lu Lu and Qiu Li.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by Shandong Provincial Medical and Health Science and Technology Project (202303061275).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data that support the findings of this study are available from the corresponding author, Q. L., upon reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
