Abstract
Background:
Bacterial infections are often an overlooked factor in female infertility. Staphylococcus aureus has been identified as the predominant vaginal pathogen in infertile women, with a prevalence of 57.33%. Previous studies showed that S. aureus induces infertility in mice by sperm impairment, suggesting its asymptomatic vaginal colonization creates a hostile environment for sperm. While sperm immobilization factor (SIF) from culture supernatant has been identified, its production within host’s environment remained unexplored.
Objective:
To unveil S. aureus-derived signature protein(s) in vaginal lavage fluid (VLf).
Design:
Mass spectrometry combined with experimental studies.
Methods:
VLf was obtained from female mice administered either with sperm immobilizing S. aureus (test group) or PBS alone (control group) and analyzed using nano-LC-MS/MS, gel filtration chromatography, SDS-PAGE, functional assays, and in silico studies.
Results:
Nano-LC-MS/MS yielded 5 distinct bacterial proteins in test group and no bacterial protein in control. Elution profile of test VLf revealed a single peak and indicated 1 protein band (~36 kDa) using SDS-PAGE that aligned with GMP reductase. VLf-protein showed impairment of sperm motility and viability in concentration-dependent manner and disrupted sperm morphology. Binding studies using FITC-labeled VLf-protein depicted presence of green fluorescence over entire surface of mouse spermatozoa. These results were akin to SIF, already isolated and characterized in our laboratory, from culture supernatant of S. aureus, causing sperm impairment and hence, designated as vaginal lavage fluid-derived sperm immobilization factor (VLf-SIF). Through in silico analysis, superimposition of VLf-SIF and SIF, already known to show sequence homology to cysteine-tRNA ligase, revealed close structural alignment. Molecular docking analysis depicted energetically favorable binding between VLf-SIF and spermatozoa surface protein (Heat shock-related 70 kDa protein 2).
Conclusion:
This study provides novel evidence of sperm-impairing S. aureus signature proteins as key mediators of bacterial-induced infertility, paving way for diagnostic, and therapeutic advancements.
Introduction
Amidst the backdrop of a burgeoning global population, the plight and anguish experienced by infertile couples cannot be overlooked, thus emphasizing the paradoxical trend of diminishing concerns regarding fertility. Infertility, touching 1 in 6 people globally, reverberates beyond individuals to impact families and communities worldwide. Among the myriad factors implicated in infertility, the role of bacterial infections has ignited considerable debate. While some bacteria, notably Mycoplasma genitalium, Neisseria gonorrhoeae, Ureaplasma urealyticum, and Chlamydia trachomatis have been extensively studied and firmly associated with infertility,1,2 the links between various other bacteria and infertility remain elusive. According to Ghiasi et al, 3 Staphylococcus aureus emerges as the predominant pathogen in the vagina of infertile women, with a prevalence of 57.33%, followed by Escherichia coli at 25.33%, indicating S. aureus as the most frequently encountered invading vaginal bacteria among this demographic.
Building upon this groundwork, earlier in our laboratory, infertility as a result of asymptomatic vaginal colonization with sperm impairing S. aureus (a clinical isolate from cervix of infertile woman) has been observed. Of note, no histopathological changes were detected in the reproductive tract, which ruled out the possibility of infertility due to tubal blockage or obstruction from inflammation caused by the organism.4,5 This indicated that presence of sperm-impairing microbes in the vagina may have transformed vaginal environment into one that is unfavorable for sperm survival, potentially resulting in infertility in mice. Further supporting this, the introduction of a non-sperm agglutinating and non-sperm immobilizing standard strain into the vagina did not negatively affect fertility outcomes. 5 Identifying these proteins and their mechanisms of action is crucial for understanding S. aureus-mediated reproductive dysfunction and developing targeted diagnostic and therapeutic strategies for infertility linked to asymptomatic bacterial colonization.
Thaper et al 6 delved deeper into the ramifications of S. aureus and depicted that it was capable of 100% immobilization of mouse spermatozoa in vitro. Extraction and purification of an active component or precipitating factor that endowed S. aureus with this sperm immobilizing property viz., sperm immobilization factor (SIF; ~59 kilodaltons (kDa)) was carried out. Their findings unveiled SIF’s capacity to compromise the motility, viability, and morphology of mouse spermatozoa in vitro and ultimately rendering female mice infertile upon intravaginal inoculation. 6 The concentration-dependent immobilization of spermatozoa suggests that SIF directly interacts with the sperm membrane. This effect is unlikely due to non-specific disruption of the membrane and is more probably a result of receptor-mediated action. This meant that SIF-induced immobilization of spermatozoa might have been resulted from binding with distinct surface receptors on mouse spermatozoa. In order to observe the localization of receptor, SIF was labeled with Fluorescein isothiocyanate (FITC) dye and interestingly, studies revealed green fluorescence distributed uniformly over the entire surface of spermatozoa, indicating the presence of SIF-binding receptor. 6
Despite identifying the sperm immobilization factor (SIF) from culture supernatant of S. aureus as a contributor to infertility, significant gaps remain in understanding whether the specific protein signatures are also produced in vaginal lavage fluid post S. aureus-administration in the host environment. Also, a constant quest for answers still exists in regards to infertility in females due to asymptomatic microbial colonization of the genital tract. Little attention has been paid toward the early diagnosis of these asymptomatic colonizers of genital tract leading to female infertility.
As proteomics is a powerful tool in biomarker identification for various diseases, 7 it offers a promising avenue for exploring the interaction between host and pathogen during infection.8,9 Leveraging vaginal lavage fluid proteome by combining mass spectrometry and bioinformatics, our study endeavors to unveil differential proteins in lavage fluid post-bacteria administration; offering to unveil S. aureus-derived signature protein(s) underlying bacterial-induced infertility. Given that no reliable biomarkers currently exist for asymptomatic bacterial colonization leading to infertility, this study presents a novel approach to understanding and potentially mitigating the reproductive consequences of S. aureus infections.
Materials and Methods
Animals
In this study, sexually mature BALB/c mice were used: females aged 4 to 5 weeks (22 ± 2 g; n = 20) and males aged 5 to 6 weeks (25 ± 2 g; n = 5). The mice were housed individually in polypropylene cages in the animal house of the Department of Microbiology at Panjab University, Chandigarh, India. They were given standard pellet food and water ad libitum, and maintained under standard laboratory conditions with a 12:12 hour light-dark cycle. All experimental procedures adhered to the approved protocols of the Institutional Animal Ethics Committee, Panjab University (reference no. PU/45/99/CPCSEA/IAEC/2023/820). The reporting of this study conforms to the ARRIVE 2.0 guidelines 10 (see Supplemental File for the completed checklist) to ensure transparency and reproducibility in animal research.
Microorganism and Preparation of Inoculum
The sperm immobilizing strain of Staphylococcus aureus, a clinical isolate from cervix of woman, already present in our laboratory was used. 5 The strain was inoculated into Nutrient broth and incubated under shaking conditions at 37°C for 24 hours. After incubation, the culture was centrifuged at 10 000 rpm for 20 minutes. The resulting pellet was washed twice with PBS (50 mM, pH 7.2), and the cells were resuspended in the same buffer. The serial dilution method was then employed to achieve a cell count of 108 CFU/20 µl.
Experimental Groups and Intravaginal Administration
Twenty female BALB/c mice were divided into 2 groups with 10 mice in the control group and 10 mice in the experimental group following the randomization technique. Only healthy, sexually mature mice with no visible signs of infection, illness, or reproductive abnormalities were included. For intravaginal administration, mice were placed on a flat grated surface (cage top), held from the base of the tail with 2 fingers and lifted upwards so that the mice provided resistance against the tail restraint. With the help of a sterile pipette, Group I was inoculated with 20 µl of PBS alone (Control group) and Group II with 108 CFU of sperm immobilizing S. aureus per mouse in 20 µl PBS (Test group) for 10 consecutive days by inserting the pipette tip about 2 to 3 mm deep into the vaginal lumen. After administration, mice were retained in this position for 5 minutes to avoid gravity-induced leakage of the PBS/S. aureus. Every third day, vaginal swabs were taken to ensure that the administered microorganism had colonized the vagina and the re-isolates were identified to be the same by Gram reactions and colony morphology Mannitol Salt Agar (results shown in Supplemental File; Figure S1). Also, the re-isolates were checked for presence of sperm-immobilization of mouse spermatozoa in vitro.
Sample Collection
To follow the itinerary of the experiment, on Day 12, vaginal lavages were obtained by introducing 100 μl of sterile PBS (50 mM, pH 7.2) into the vaginal canal. This was done using a pipette tip inserted 2 to 3 mm into the vaginal orifice, with the fluid being repeatedly aspirated and agitated. The lavage fluids were collected and pooled into a microcentrifuge tube followed by supplementation with 1% v/v protease inhibitor cocktail (Sigma-Aldrich, USA) and protein concentrations for the control and test group vaginal lavage fluid were measured using the Bradford assay. 11
Identification of S. aureus-Derived Proteins in the Host Vaginal Lavage Fluid by Gel-LC-MS/MS
1D-Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis (SDS) Gel Cleanup
Prior to use, the gel running apparatus, glass plates, cutting tools and the working surface were thoroughly cleaned with methanol, followed by Milli-Q water to avoid any contamination. Control and test group vaginal lavage fluid were run using 1D-SDS gels (12%) for 0.5 cm at a constant voltage of 100 V and the run was stopped when the dye front had migrated 0.5 cm, 12 followed by staining with 0.25% Coomassie brilliant blue (R-250). Using a clean razor blade, the entire stained region of each gel lane was excised. Further, the excised band was chopped into small cubes of 1 mm × 1 mm size, and pieces were duly transferred into labeled 1.5 ml centrifuge tubes.
In-Gel Trypsin Digestion and Sample Preparation for Label-Free Liquid Chromatography Tandem Mass Spectrometry
The gel bands were washed with MilliQ water for 1 hour while vortexing at 1500 rpm. After discarding the water, this washing step was repeated twice. The gel cubes were then vortexed at 3000 rpm for 20 seconds to ensure complete removal of the water. About 300 µl of de-staining solution [100 mM ammonium bicarbonate/ acetonitrile (1:1, v/v)] was added and vortexed for 30 minutes @1500 rpm. Next, 500 µl of 100% acetonitrile was added to the gel pieces, which were then incubated at room temperature with vortexing until the gel pieces turned white and shrank. The acetonitrile was completely removed, and the vials were left open for 20 minutes to allow for the complete evaporation of the solvent. After ensuring complete drying of gel pieces, 100 µl of 10 mM Dithiothreitol was added to the gel pieces for reduction and incubated at 55°C for 45 minutes. Next, 100 µl of 55 mM Iodoacetamide was added for alkylation and incubated for 30 minutes at room temperature in the dark. 30 to 50 µl of MS-grade trypsin buffer (13 ng/µl; Sigma-Aldrich, USA) was added so that it covered the dry gel pieces and was left in an ice bucket for saturation of gel. The trypsin buffer level was checked and was added in vials again to keep the gel submerged. The vials were incubated at 37°C at 300 rpm/overnight. The supernatant containing the peptides was transferred from the sample vials to freshly labeled 0.5 ml microcentrifuge tubes. Then, 100 µl of extraction buffer [5% formic acid/acetonitrile (1:2, v/v)] was added to the gel pieces in the sample tubes and vortexed vigorously for 20 minutes. The extraction buffer was then aspirated and combined with the supernatant containing the peptides in the 0.5 ml microcentrifuge tubes. The peptides were purified by C-18 spin columns, dried using a Speed Vacuum concentrator, and stored at −20°C until used. Before LC/MS analysis, samples were resuspended in 30 µl of 3% acetonitrile.
ESI-nano-LC-MS/MS
LC/MS/MS analysis was carried out using a nano ACQUITY UPLC System (Waters, Manchester, UK) paired with a Quadrupole Time-of-Flight (Q-TOF) mass spectrometer (SYNAPT-G2, Waters). Each sample, including both the Control and Test groups, was analyzed in technical replicates. The Mass Lynx 4.1 SCN781 software was used for instrument control and operation. A 3 µl aliquot of each sample was injected for analysis. To remove salts, a trap column (Symmetry® 180 µm × 20 mm, C18, 5 µm, Waters) was utilized. Peptide separation was achieved using a 75 µm × 200 mm, HSS T3 C18 column (Waters) with a particle size of 1.8 µm, maintained at 35°C. The mobile phases were water (Phase A) and acetonitrile (Phase B), both containing 0.1% formic acid. The total analysis time was 60 minutes, and the chromatography gradient was set as follows:
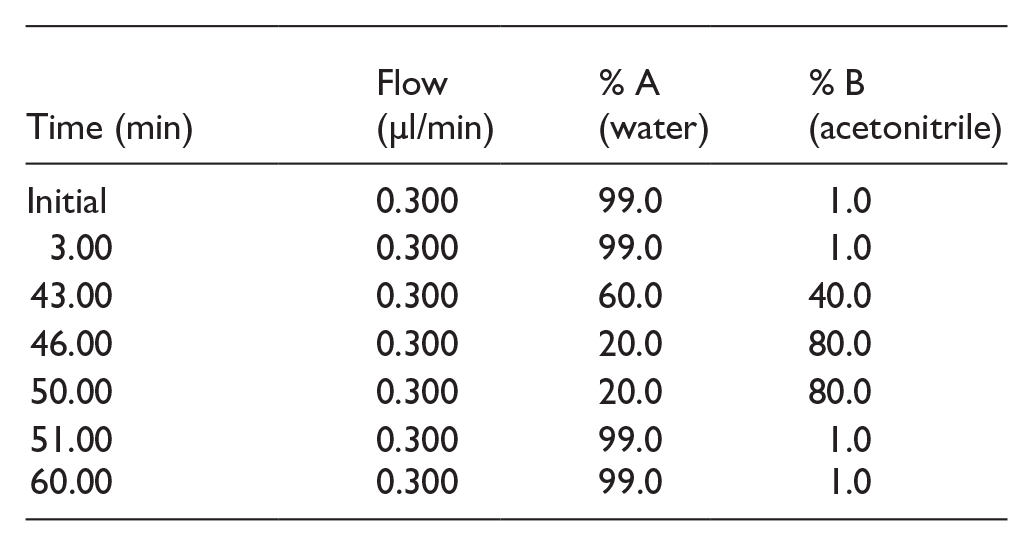
Mass Analysis and Database Search
After data acquisition, the raw files were processed for protein identification using Progenesis QI for Proteomics v4.2 (Non-Linear Dynamics, Waters). Protein identification was performed against the UniProt/Swiss-Prot “Staphylococcus” database (https://www.uniprot.org/), applying fixed carbamidomethylation of cysteine (C) and variable oxidation of methionine (M) as modifications. An MS/MS tolerance of 10 ppm was used for identification. A protein was deemed identified if it had at least 1 unique peptide; proteins not meeting this criterion were excluded. The data has been deposited in the MassIVE repository under ID: MSV000095516.
Elution Profile of Vaginal Lavage Fluid on Sephadex G-100 Column: Molecular Sieving
Gel-filtration chromatography was performed to analyze the elution pattern of proteins in the vaginal lavage fluid. The Sephadex G-100 column was equilibrated and eluted with PBS. About 3 ml fractions were collected, and their absorbance at 280 nm was recorded using a UV spectrophotometer. These fractions were then pooled and concentrated using polyethylene glycol at 4°C and designated as vaginal lavage fluid-protein (VLf-protein). The molecular mass of VLf-protein was assessed by SDS-PAGE, following the procedure described by Laemmli. 13
Validating VLf-Protein-Induced Sperm Impairment Through In vitro Analysis
Sperm Preparation
The mice (n = 5) were humanely euthanized by cervical dislocation, and their abdomens were then opened to access the vas deferens. The vas deferens was meticulously isolated and placed into 200 μl of phosphate-buffered saline (PBS, pH 7.2, 50 mM) on a glass surface. Spermatozoa were extracted by applying pressure to one end of each vas deferens and gently moving down the length with forceps to transfer the sperm into the buffered solution. The goal was to achieve a final concentration of 40 × 106 spermatozoa/ml.
Motility
The effect of VLf-protein on motility of mouse spermatozoa was determined by Sander–Cramer assay. 14 The immobilization effect of various dilutions of VLf-protein, prepared in PBS, on spermatozoa was determined by combining equal volumes (100 μl) of VLf-protein (ranging from 25 to 175 μg/ml) and sperm suspension. At different time of exposure (20 seconds, 15, 30, 45 minutes), observations were made by placing 10 μl of each sperm-protein mixture on a clean glass slide. Also, SIF, already isolated and characterized in our lab served as a positive control. 15 Various dilutions of SIF were prepared in PBS (10-100 µg/ml). PBS was used as the negative control. A minimum of 10 fields were rapidly examined under high-power magnification (400×) with a bright field microscope. Motility of sperm was calculated as following. 16

Viability
In order to determine the effect of VLf-protein, SIF (positive control), PBS (negative control) on viability of spermatozoa at various time intervals, sperm viability testing by eosin staining method was performed. Different dilutions of VLf-protein (25–250 μg/ml) and SIF (50-200 µg/ml) were prepared in PBS. Aliquots (100 μl) were taken containing VLf-protein/SIF/PBS and mixed with 100 μl of sperm suspension. After mixing for 20 seconds, 15, 30, and 45 minutes, sperm viability was subjectively assessed by eosin staining. A 10 μl aliquot from each tube was added to 10 μl of Eosin Y solution. A portion of the suspension was introduced onto a clean glass slide and mounted with a coverslip (22 mm × 22 mm) and left for 30 seconds. This dye exclusion assay relied on the ability of live spermatozoa to resist absorption of Eosin Y and easy penetration of the same in case of non-viable/dead spermatozoa, thereby, staining them pink. Ten fields were rapidly examined under high-power magnification (400×) and the percentage of unstained and stained spermatozoa was calculated.
FE-SEM Studies to Determine the Morphological Alterations
Standard operating procedures were followed throughout the sample processing. 17 After centrifugation at 500 rpm for 10 minutes, the sperm samples (>80% with normal morphology) were treated for 30 minutes with either phosphate-buffered saline or VLf-protein (200 μg/ml)/SIF (150 μg/ml). Subsequently, 4 ml of a 2.5% phosphate-buffered glutaraldehyde solution was added as a fixative, and the samples were incubated at 4°C for 1 hour. The samples were then washed twice with PBS and centrifuged for 5 minutes at 500 rpm. Following fixation and cleaning, the spermatozoa were mounted on adhesive tape on brass stubs and coated with gold using a Jeol ion sputter (JFC-1100). The coated samples were then examined using a field emission scanning electron microscope (Hitachi, Japan, SU8010 Series).
Binding Studies Using Fluorescein Isothiocyanate (FITC)-Labelled VLf-Protein: Proof-of-Evidence Using Fluorescence Microscopy
To substantiate whether VLf-protein mimics the pattern of SIF in binding the mouse spermatozoa, binding studies were conducted. Binding of FITC tagged VLf-protein and SIF to mouse spermatozoa was analyzed. The control was set up using spermatozoa and untagged VLf-protein/SIF to rule out autofluorescence. VLf-protein (1 mg/ml) and SIF (1 mg/ml) were labeled using an FITC Protein Labeling Kit (Bangalore Genei, Pvt. Ltd, India). Various volumes of FITC (2.5, 5, and 10 μl) were used, and the fluorophore-to-protein (F:P) ratios were determined based on absorbance measurements at 280 and 495 nm, in accordance with the instructions provided with the labeling kit. The sperm preparation was rinsed with 2× PBS, and pellet was then reconstituted in 500 μl of PBS (50 mM, pH 7.2). To each sperm suspension, 100 μl of tagged VLf-protein/SIF at the same concentration was added and incubated at 37°C for 1 hour. After incubation, the mixture was treated with 100 μl of 3% formaldehyde and incubated at 37°C for 1 hour. Following this, the samples were washed 3 times with PBS. The final pellet was resuspended in 50 μl of PBS (50 mM, pH 7.2). Wet-mount analysis was performed using a fluorescent microscope (Nikon, Japan).
Based on the results, this protein (VLf) was designated as vaginal lavage fluid-derived sperm immobilization factor (VLf-SIF), as it mimicked the effects of sperm immobilization factor (SIF) isolated from culture supernatant of S. aureus.
In silico Studies: Assessing the Structural Similarity Between Cysteine-tRNA Ligase (SIF) and GMP Reductase (VLf-SIF)
As previously determined in our laboratory, SIF showed sequence homology to cysteine-tRNA ligase of Myco-plasma penetrans (strain HF-2) with a score of 60 by MALDI-TOF (protein scores greater than 59 are significant; P < .05). So, the amino acid sequences corresponding to cysteine-tRNA ligase from Mycoplasma penetrans (strain HF-2) were obtained from the UniProt database, accessible at ( https://www.uniprot.org/ ). The sequence was retrieved in FASTA format for further analysis. Similarly, the sequence for GMP reductase (guaC_2) was retrieved which was selected in the present study as the potential VLf-SIF from the convergence of data from SDS-PAGE and proteomic analysis.
Three-Dimensional Modeling and Qualitative Analysis
The 3-dimensional modeling of proteins viz., cysteine-tRNA ligase (SIF) and GMP reductase (VLf-SIF) were carried out using I-TASSER 18 and Phyre2 19 packages. The sequences of cysteine-tRNA ligase (UniProt ID: Q8EX28) and GMP reductase (UniProt ID: A0A7Z7VSM3) were submitted directly to the I-TASSER and Phyre2 servers. PyMOL Molecular Graphics System, Version 2.0 Schrodinger, LLC packages were utilized to illustrate the predicted protein models. Further, the predicted 3-dimensional models of respective proteins from I-TASSER and Phyre2 were proceeded for qualitative evaluation. The evaluation of a protein’s 3-dimensional model quality was carried out using various tools, including the PROCHECK module 20 of the PDBSum server, ERRAT, 21 and ProSA 22 servers.
Superimposition of Cysteine-tRNA Ligase (SIF) and GMP Reductase (VLf-SIF) Using UCSF Chimera
To ascertain the structural resemblance between the 3D models of cysteine-tRNA ligase (SIF) and GMP reductase (VLf-SIF) proteins, a structural alignment was carried out. This alignment analysis was executed utilizing the Match Maker extension integrated into UCSF Chimera. 23 The Root Mean Square Deviation (RMSD), representing the disparity between the structures of the 2 proteins, was computed as a quantifier of structural similarity. A lower RMSD value, specifically when below 2 Ångströms (Å), denotes a heightened degree of similarity in the protein structures. These analyses were undertaken to assess the degree of superimposition or alignment between the 2 proteins.
In silico Demonstration via Protein-Protein Docking of GMP Reductase (VLf-SIF) With Heat Shock-Related 70 kDa Protein 2 (Mouse Spermatozoa Surface Protein)
Three-Dimensional Modeling and Qualitative Analysis
As earlier described in our laboratory, sperm immobilization by S. aureus is probably associated with the interaction of SIF with a surface spermatozoa protein of mouse. 6 By bioinformatics approach, this mouse spermatozoa surface protein showed homology to Heat shock-related 70 kDa protein 2 (UniProt ID: P17156) and exhibited stable intermolecular interactions with cysteine-tRNA ligase (SIF; an unpublished study). In this study, we aimed to determine whether VLf-SIF (GMP reductase) exhibits a similar binding pattern with this surface spermatozoa protein (Heat shock-related 70 kDa protein 2). As described in Section 2.7, the 3-dimensional models of GMP reductase (VLf-SIF) and Heat shock-related 70 kDa protein 2 were generated through a comprehensive in silico approach.
Prediction of Binding Pockets
The determination of the active/binding sites of the protein models for GMP Reductase and Heat shock-related 70 kDa protein 2 was conducted by utilizing the CDD module of the NCBI database. Additionally, the respective protein models were also submitted to CASTp 24 and COACH 25 servers to find the binding pockets.
Protein-Protein Docking Using ClusPro 2.0
Protein-protein docking studies were conducted between GMP Reductase (ligand) and Heat Shock Protein 70 kDa (receptor) using the ClusPro 2.0 server. 26 The server performs 3 main computations: first, it conducts rigid body docking by sampling billions of possible conformations; second, it clusters the 1000 lowest energy structures based on root-mean-square deviation (RMSD) to identify the largest clusters, which represent the most probable complex models; and third, it refines these selected structures through energy minimization. The ClusPro 2.0 software employs scoring strategies to evaluate the quality of the predicted complex structures, considering factors such as van der Waals interactions, electrostatic forces, hydrogen bonding, desolvation energy, and shape complementarity.
Results
Proteomic Profile of Vaginal Lavage Fluid by LC–MS/MS Analysis to Identify S. aureus-Derived Signature Proteins
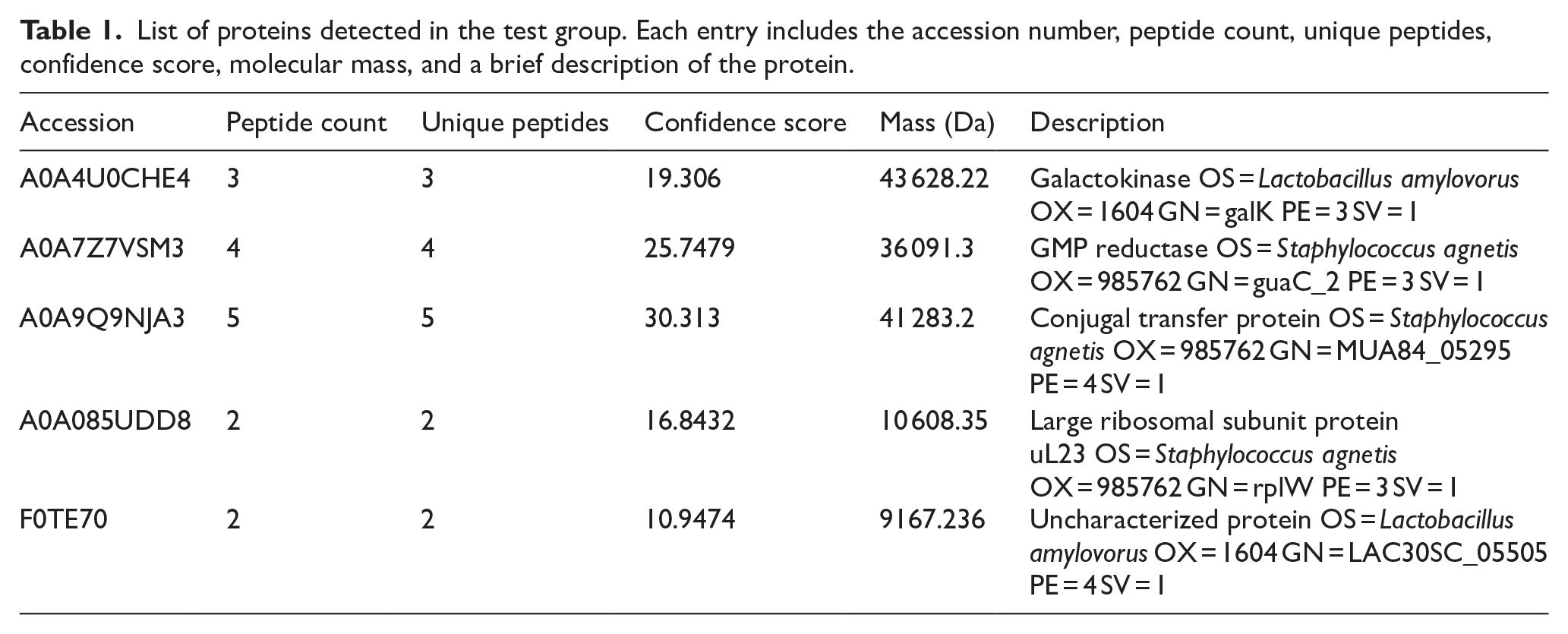
In the vaginal lavage fluid obtained from the test group samples, after administering sperm immobilizing S. aureus, the protein identification process yielded 5 distinct bacterial proteins. Each of these proteins contained at least 1 unique peptide. The analysis was conducted by searching the raw data against a comprehensive “Staphylococcus” database. In contrast, no bacterial protein was identified in the control group samples of mice administered with PBS. Details of the differentially expressed proteins detected in the test group are provided in Table 1.
List of proteins detected in the test group. Each entry includes the accession number, peptide count, unique peptides, confidence score, molecular mass, and a brief description of the protein.
Elution Profile of Vaginal Lavage Fluid on Sephadex G-100 Column: Molecular Sieving
After applying pooled vaginal lavage fluid of test group on a Sephadex G-100 column for purification, the elution profile revealed a single prominent protein peak. The active fractions (Fraction 12-18; Figure 1) were pooled and concentrated using polyethylene glycol at 4°C and designated VLf-protein, that were further tested for sperm immobilization activity. No significant peaks were detected in the lavage fluid of control group.

Elution profile of Vaginal Lavage fluid on Sephadex G-100 column: Bioactivity entity resides in Fractions 12-18.
Determination of the Molecular Mass of VLf-Protein Using SDS-PAGE Analysis
SDS-PAGE analysis was conducted to ascertain the molecular mass of VLf-protein following purification. The single protein band was obtained that corresponded to an apparent molecular weight of approximately 36 kDa when compared to molecular weight reference markers (Figure 2). Therefore, GMP reductase (guaC_2; molecular weight of 36,091.3 Da), identified with a significant confidence score and unique peptide matches from the proteomic profile of test group vaginal lavage fluid (Table 1), was selected as the potential VLf-protein. This convergence of data from SDS-PAGE and proteomic analysis provide preliminary support for the identification of the 36 kDa protein band as GMP reductase identified through mass spectrometry (OS = Staphylococcus agnetis; OX = 985762; Figure S2).

Molecular mass demonstrated through SDS-PAGE analysis: Lane 2 shows the results for pooled and concentrated fractions (12–18), while Lane 1 contains protein molecular weight markers for reference.
Effect of VLf-Protein on Various Sperm Parameters: Assessing Functional Congruence with SIF’s Pattern
Motility
When different concentrations of VLf-protein ranging from 25 to 175 µg/ml were incubated with mouse spermatozoa, it was observed that VLf-protein affected mouse spermatozoa in a dose dependent manner. With increasing concentration of VLf-protein, there was an increase in the percentage of sperm immobilization. Specifically, it was observed that a concentration of 100 µg/ml induced 100% sperm immobilization within a 30-minute incubation period. Interestingly, concentration of 150 µg/ml achieved the same effect almost instantly, within just 20 seconds, in comparison to the control group. In case of positive control, 50 µg/ml of SIF caused 100% immobilization of spermatozoa within a 30-minute incubation period and 70 µg/ml of SIF achieved the same effect within 20 seconds.
Viability
Eosin staining was employed to evaluate the impact of VLf-protein and SIF on sperm viability through light microscopy. At elevated concentrations, VLf-protein and SIF demonstrated spermicidal properties, with a concentration of 175 and 120 µg/ml, respectively, resulting in the complete elimination of viable spermatozoa within a 30-minute incubation period. Similarly, a concentration of 200 μg/ml of VLf-protein and 150 μg/ml SIF achieved a comparable effect within a rapid 20-second incubation period when compared to the control sample (Figure 3).
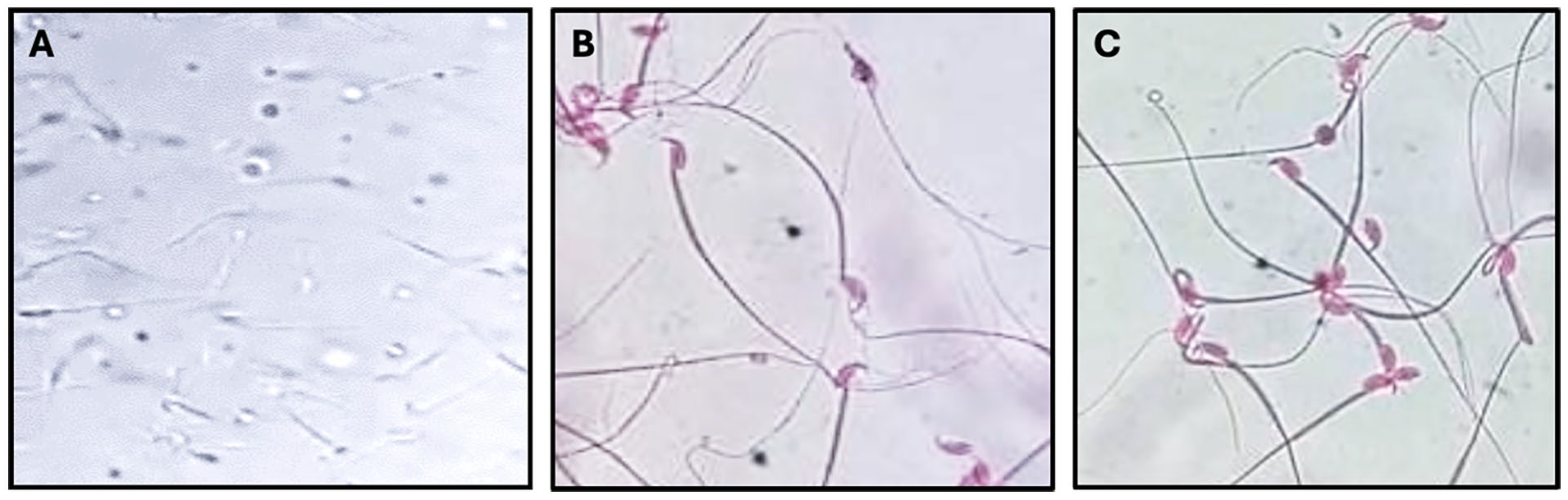
Photomicrographs illustrating the effect of VLf-protein on spermatozoa viability. Normal mouse spermatozoa incubated with PBS (A), Dead spermatozoa (pink-stained) after treatment with VLf-protein (200 μg/mL) (B), and Dead spermatozoa (pink-stained) treated with SIF (150 μg/mL) (C) (Original magnification X400).
Morphology
A scanning electron microscopy examination was performed to conduct an in-depth investigation of the topological structures of spermatozoa. The results clearly indicated that the control samples demonstrated typical characteristics, including the characteristic hook-shaped appearance of the head and an intact plasma membrane enveloping both the head and tail (Figure 4a). Conversely, upon exposure to a concentration of 200 μg/ml of VLf-protein (Figure 4b) and 150 μg/ml of SIF (Figure 4c), a notable modification in the morphology of sperm was noticed in both samples, with the most salient being head malformations.

Representative scanning electron micrographs illustrating normal mouse spermatozoa observed after incubation with PBS (A); Morphological defects evident upon incubation with VLf-protein (200 μg/mL) (B); and SIF (150 μg/mL) (C).
Binding Studies Carried Out with Fluorescein Isothiocyanate (FITC)-Labelled VLf-Protein/SIF
Fluorescence microscopy studies depicted no fluorescence in untagged VLf-protein/SIF incubated with spermatozoa. Whereas, bright green fluorescence was observed in the case of tagged VLf-protein/ SIF incubated with spermatozoa (Figure 5).

Fluorescent microscopy. Representative images of unlabelled VLf-protein/ SIF incubated with spermatozoa, serving as control (A); FITC labelled VLf-protein incubated with mouse spermatozoa (B); FITC labelled SIF incubated with mouse spermatozoa (C).
In silico Analysis to Assess the Structure Similarity by Superimposing Cysteine-tRNA Ligase (SIF) and GMP Reductase (VLf-SIF)
Three-Dimensional Modeling and Qualitative Analysis
For cysteine-tRNA ligase of Mycoplasma penetrans (strain HF-2; SIF), I-TASSER predicted 3D structure was selected based on the qualitative analysis. Only 1 model out of the 5 generated models of the query sequences was selected based on a requirement of significant alignment with the given template, as well as reasonable C-score, TM score, and RMSD values. It depicted RMSD of 4.1 ± 2.8 Å, a substantial C-score of 1.38, and a TM score of 0.91 ± 0.06. This was validated using PROCHECK, ERRAT, and ProSA. The PROCHECK analysis revealed that 91.8% were in most favored regions, 8.2% in additionally allowed regions (Figure S3a). ERRAT demonstrated the overall quality factor of 94.43 for SIF. The ProSA analysis revealed that the amino acid residues showed negative energy and had z-scores of −10.24 for SIF as depicted in Figure S3b.
For GMP Reductase (VLf-SIF), predicted 3D structure from Phyre2 was selected based on the qualitative analysis. This was validated using PROCHECK, ERRAT, and ProSA. The PROCHECK analysis revealed that 91.7% were in most favored regions, 7.9% in additionally allowed regions (Figure S4a). ERRAT demonstrated the overall quality factor of 77.14 for VLf-SIF. The ProSA analysis revealed that the amino acid residues showed negative energy and had z-scores of −8.43 for VLf-SIF as depicted in Figure S4b.
The qualitative study carried out suggested that the predicted models for cysteine-tRNA ligase and GMP Reductase have good quality and were proceeded for further analysis.
Evaluation Using UCSF Chimera
The structural concordance between the 3D models of cysteine-tRNA ligase (SIF) and GMP Reductase (VLf-SIF), a structural alignment was conducted. This alignment procedure was performed employing the Match Maker extension within the UCSF Chimera software platform. In this analysis, we computed an RMSD value of 1.411 Å, signifying the extent of superimposition between cysteine-tRNA ligase (SIF) and GMP Reductase (VLf-SIF). This observation substantiates the notion that proteins exhibiting pronounced structural similarity are also inclined to share functional congruence (Figure 6). This level of similarity could imply a conserved structural framework or potential functional similarities despite possible differences in primary sequences.

Superimposed image of cysteine-tRNA ligase (SIF) and GMP Reductase (VLf-SIF) depicting RMSD value of 1.411 angstroms (across all 257 pairs: 22.561) showing high structural similarity.
GMP Reductase (VLf-SIF) Binding with Heat Shock-Related 70 kDa Protein 2 (Mouse Spermatozoa Surface Protein) Through Protein-Protein Docking Analysis
For the structure of Heat shock-related 70 kDa protein 2 of mouse spermatozoa, predicted 3D structure from I-TASSER was selected based on the qualitative analysis. It depicted RMSD of 5.5 ± 3.5 Å, a substantial C-score of 1.07, and a TM score of 0.86 ± 0.07. The PROCHECK analysis revealed that 94.6% were in most favored regions, 5.0% in additionally allowed regions (Figure S5a). ERRAT demonstrated the overall quality factor of 88.5. The ProSA analysis revealed that the amino acid residues showed negative energy and had z-scores of −12.69 as depicted in Figure S5b.
The CDD, CASTp and COACH analysis revealed the binding pockets consisting of GLN 331, ASN107, SER106, GLU85, SER88, GLN331, ASN336, PHE110 in Heat shock-related 70 kDa protein 2 of mouse spermatozoa and LYS135, GLY183, MET1, ARG316, ASN168 in GMP Reductase. Subsequently, protein-protein complexes of Heat shock-related 70 kDa protein 2 with GMP Reductase was generated and submitted to ClusPro 2.0, revealing interactions among binding pocket residues. Docking results indicated confident docking scores using PyMOL, results are depicted in Figure 7. The Heat shock-related 70 kDa protein 2 with GMP Reductase complex displayed a weighted score of −823.2 and a lowest energy of −1103.9 kcal/mol. Consequently, the protein complexes assumed energetically favorable and stable conformations, as indicated by the low energy scores.

Representation of Protein-Protein interaction between GMP Reductase (VLf-SIF) & Heat shock-related 70 kDa protein 2 of mouse spermatozoa surface protein at various binding sites.
Discussion
The impetus for this study stems from the significant yet often overlooked impact of bacterial diseases on female infertility. Previous findings from our laboratory have indicated that Staphylococcus aureus, a predominant pathogen in the vagina of infertile women, can impair sperm function and render mice infertile without causing histopathological changes.4,5 This highlights the potential for asymptoma-tic vaginal colonization by S. aureus to create a hostile environment for spermatozoa, yet the precise factors involved remain unclear. Notably, the sperm immobilization factor (SIF) from culture supernatant of S. aureus has been identified as a key player, 6 but, to our knowledge, its production within the host’s internal environment have not been investigated. This knowledge could lead to the development of new treatments (therapeutic biomarkers) or preventive (diagnostic/prognostic biomarkers) measures for infertility caused by bacteria.
Traditional liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS) of peptides resultant from proteolytic digestion offers the greatest proteomic sensitivity 27 and aids to quickly attain pathological information by evaluating biomarkers that distinguish diseases and determine the rules of pathological changes.28,29 Bekmurzayeva et al 30 states that for pathogen-derived biomarkers to be viable targets for antigen detection assays, they must be present at detectable levels within sample matrices. In the current research, we have therefore considered vaginal lavage fluid as 1 such matrix. The existing work intended to determine antifertility signature proteins using LC-MS/MS in the proteome of vaginal lavage fluid of sperm immobilizing S. aureus-administered female BALB/c mice (test group) in comparison to the control group, intravaginally inoculated with PBS alone.
The proteomic analysis of pooled vaginal lavage fluid identified 5 distinct S. aureus-derived proteins exclusively in the test group, with no bacterial proteins detected in the control group treated with PBS. This differential expression suggested a significant proteomic response to S. aureus exposure, highlighting the impact of the bacterial interaction on the vaginal environment. The presence of S. aureus in the vaginal milieu likely have altered the biochemical landscape, leading to the synthesis and release of specific bacterial proteins as part of the bacterial colonization and pathogenesis process. These proteins might have interfered with various functions, including sperm motility and viability, which are consistent with the known effects of S. aureus on sperm function. On the same note, Bi et al 7 conducted a label-free proteomics study to investigate protein alterations in sera of Schistosoma japonicum infected rabbits to understand the pathogenesis of schistosomiasis and identified 10 S. japonicum-derived proteins that significantly changed across infection stages.
Given the identification of these 5 proteins as potential contributors to sperm impairment, we sought to isolate the particular protein responsible for infertility. We purified the vaginal lavage fluid using gel filtration chromatography, resulting in an elution profile with a single prominent protein peak in the test group, while no significant peak was observed in the control group’s elution profile. This suggests that the prominent peak in the test group was not composed of host proteins. Active fractions from the test group were pooled, concentrated, and subsequently tested for sperm immobilization activity. The presence of bioactivity in these fractions confirmed the existence of a functional protein entity responsible for the observed effect. Further assessment for purity using SDS-PAGE revealed a single unique protein band with an apparent molecular mass of 36 kDa, which aligned with the GMP reductase (guaC_2) identified through proteomic profiling. This vaginal lavage fluid-derived protein (VLf-protein) was proposed as the mediator of the observed biological activity. Moreover, the detection of the bacterial protein in the test lavage fluid, despite the presence of host proteins, indicates a significant threshold of bacterial protein presence, reinforcing the idea that this specific protein is involved in the impairment of sperm function.
As morphometric parameters including motility, viability, and morphology are perilous indicators in fertility studies, 31 above findings were validated through spermiogram analysis, which unequivocally demonstrated that VLf-protein plays a pivotal role in sperm immobilization and also, corroborated closely with the findings of SIF. This analysis revealed a clear, dose-dependent impact of VLf-protein on sperm motility and viability where higher concentrations of VLf-protein led to pronounced reductions. This highlighted the comprehensive effect of VLf-protein which obliterates not only motility, but also vitality of mouse spermatozoa. The parallel observed effects with SIF, as previously proposed by Thaper et al 6 suggested a concentration-dependent pattern of sperm immobilization mediated possibly through binding to a membrane-bound receptor present on mouse spermatozoa. A similar mechanism of action leading to compromised sperm function was notable in both VLf-protein and SIF treated spermatozoa as FE-SEM revealed significant morphological changes in spermatozoa, reinforcing the comparable impact that were absent in untreated spermatozoa. These morphological changes due to 36 kDa antigen indicates a disruption of sperm structural integrity, potentially contributing to negative impacts on various sperm properties and fertilization potential. Similarly, functional studies have revealed that various bacterial components, including immune-dominant antigen A, alkaline shock proteins, and thermonuclease, negatively impact sperm motility. 32 Li et al 33 have shown that S. aureus strains isolated from infertile males impair sperm motility and morphology through their bacterial virulence factors.
Next, we conducted binding studies using fluorescence microscopy with FITC-labeled VLf-protein to determine the distribution of VLf-protein-binding receptors on the surface of mouse spermatozoa to investigate whether it parades an action corresponding to SIF. The results revealed bright green fluorescence across the entire surface of mouse spermatozoa, suggesting that the biological effects of VLf-protein on sperm cells could be due to interactions with specific cellular receptors on the spermatozoa.
Taking into account all these observations, VLf protein was named as VLf-SIF because it exhibited similar effects to that of the sperm immobilization factor (SIF) isolated from the culture supernatant of S. aureus.
Having identified VLf-SIF as GMP reductase from the above experiments, structural similarity with the previously known and characterized SIF protein (cysteine-tRNA ligase) was also carried out. In silico validation was conducted by superimposing their predicted 3D structures using the Match Maker extension in UCSF Chimera. The structural alignment indicated a high degree of superimposition between the 2 proteins. According to Kirchmair et al, 34 RMSD values below 5 Å are generally regarded as indicative of good alignment, with lower values indicating greater structural similarity. Our findings suggest that VLf-SIF and SIF share a substantial degree of structural overlap, supporting the hypothesis that VLf-SIF, identified as GMP reductase, possesses functional characteristics akin to SIF in sperm immobilization. Also, the earlier findings strongly pointed toward the hypothesis of SIF utilizing a mouse spermatozoa surface protein as receptor for binding to spermatozoa, 6 thereby impairing their function. Subsequent molecular docking analysis in the present study confirmed this interaction with spermatozoa surface protein, as indicated by the energetically favorable and stable conformations of the protein complexes.
Our study provides comprehensive evidence regarding the role of VLf-SIF, identified as GMP reductase, in impairing sperm function through functional and structural analyses. The similarity in action between VLf-SIF and SIF emphasizes their potential functional congruence, offering insights into the pathophysiology of infertility associated with asymptomatic bacterial infections. Of note, the discrepancy in protein identification between in vitro and in vivo isolation scenarios can be attributed to several factors related to the complex biological environments and conditions involved such as host organism’s immune response, tissue environments, and physiological conditions. However, our study has addressed this limitation by isolating and identifying GMP reductase directly from in vivo conditions, ensuring that the interactions with the host’s internal environment, which can significantly affect the expression and function of bacterial proteins, are accurately accounted for. This approach enhances the relevance and applicability of our findings to real biological scenarios. The identification of VLf-SIF (GMP reductase) as a sperm-impairing bacterial protein suggests a broader role for bacterial infections in infertility, particularly in humans. If similar bacterial proteins interact with sperm in the human reproductive tract, they could impair motility, viability, and structural integrity, thus reducing fertilization potential. Given that asymptomatic bacterial colonization is often overlooked in infertility diagnoses, the presence of sperm-agglutinating or immobilizing bacterial factors may contribute to unexplained infertility. To validate VLf-SIF as a biomarker for bacterial-induced infertility, clinical studies should assess its presence in vaginal lavage fluid from infertile women with S. aureus colonization using LC-MS/MS. Additionally, developing ELISA or lateral flow immunoassays could enable non-invasive detection, facilitating its potential use as a diagnostic tool.
Conclusion
Based on the findings of this study, it is evident that the vaginal lavage fluid-derived sperm immobilization factor (VLf-SIF), identified as GMP reductase, plays a significant role in impairing sperm function through structural and receptor-mediated mechanisms. This research underscores the potential impact of asymptomatic bacterial infections on female infertility, providing insights into the sperm-impairing factor involved. Further investigation and validation studies could also serve VLf-SIF as potential diagnostic biomarker.
Supplemental Material
sj-docx-1-bmi-10.1177_11772719251340518 – Supplemental material for Staphylococcus aureus-derived Signature Protein(s) in Murine Vaginal Lavage Fluid: Investigating the Underlying factors of Infertility as a Consequence of Sperm Impairing Bacterium
Supplemental material, sj-docx-1-bmi-10.1177_11772719251340518 for Staphylococcus aureus-derived Signature Protein(s) in Murine Vaginal Lavage Fluid: Investigating the Underlying factors of Infertility as a Consequence of Sperm Impairing Bacterium by Ishwerpreet Kaur Jawanda, Thomson Soni, Seema Kumari and Vijay Prabha in Biomarker Insights
Footnotes
Acknowledgements
Authors would like to acknowledge SAIF/CIL, Panjab University, Chandigarh, India and DBT-SAHAJ National Facility for Mass Spectrometry-based Proteomics, Metabolomics and Lipidomics at RGCB, India.
Ethical Considerations
All experimental procedures adhered to the approved protocols of the Institutional Animal Ethics Committee, Panjab University (reference no. PU/45/99/CPCSEA/IAEC/2023/820). The reporting of this study also conforms to the ARRIVE 2.0 guidelines. This study was carried out in strict accordance with the recommendations in the guidelines for reporting experiments involving animals. The full ARRIVE 2.0 guidelines checklist has been reviewed, and the methods and results are presented in accordance with these guidelines.
Consent for Publication
Not applicable.
Authorship Contributions
Ishwerpreet Kaur Jawanda: Writing – original draft, Data curation, Conceptualization, Methodology, Investigation. Thomson Soni: Conceptualization, Data curation, Investigation, Methodology, Writing-Reviewing and Editing. Seema Kumari: Investigation, Data curation, Project Administration, Formal Analysis. Vijay Prabha: Writing – review & editing, Validation, Supervision, Resources, Project administration, Formal Analysis, Methodology, Conceptualization.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding was provided from Panjab University, Chandigarh, India to I.K.J. (D-DRES/22/1898-1917; Dated 30-06-2022). The funders had no role in the study design, data collection and analysis, decision to publish or prepare the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Informed Consent Statement
Not applicable.
Institutional Review Board Statement
Not applicable.
Data Availability Statement
The proteomics data generated and analyzed during this study are available in the repository mentioned within the article. Additional information supporting the findings of this study is included in the Supplemental File. Any other data related to this study will be made available upon reasonable request to the corresponding author.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
