Abstract
Background:
DNAJC12 (DnaJ heat shock protein family (Hsp40) member C12) encodes a member of the molecular chaperone Hsp40/DnaJ family, which are important protein folding and proteostasis regulators. Its role as a biomarker has been studied for a limited number of cancer types. Objectives: Here, we sought to investigate the potential of DNAJC12 mRNA and protein expression as a prognostic and predictive biomarker for breast cancer (BC).
Methods:
Using in silico analysis and data from immunohistochemistry analysis (IHC) of 292 samples from patients with primary BC, we determined the expression pattern and prognostic value of DNAJC12 mRNA and protein expression.
Results:
From online publicly available data, we were able to identify the transcripts of DNAJC12 as differentially expressed in patients with different clinicopathological characteristics, such as ER status (
Conclusion:
Our findings suggested a potential role for DNAJC12 as a prognostic and predictive biomarker for BC.
Introduction
Breast cancer (BC) is the most common malignant disease among women worldwide, with high mortality rates. 1 The disease is complex, and its occurrence and progression are linked to both environmental and genetic factors. 2 Molecular profiling has proven effective in assessing risk, prognosis, and managing BC patients.3 -5 Treatment for BC involves evaluating the expression of molecular markers such as estrogen, progesterone, and human epidermal growth factor receptors (ER/PR/HER2), 6 with several multigene assays now being used in clinical practice, 7 indicating progress in the use of BC biomarkers. However, due to the diverse nature of BC, understanding its biology continues to be a challenge for the scientific community. Therefore, identifying and validating new biomarkers could lead to the development of more precise tools for clinicians. This study aims to assess the prognostic and predictive potential of DNAJC12 for BC patients.
Heat-shock proteins (HSPs) are molecular chaperones involved in many cellular processes and help maintain proteostasis by preventing protein misfolding and aggregation. 8 They play crucial roles in tumor development, survival, and drug resistance and can be biomarkers. 9 In breast cancer, the expression of several HSP genes is dysregulated, and some are linked to patient prognosis. 10
The HSP40 family, also known as DnaJ or J-domain proteins (JDPs), is the most prominent family of HSPs and is characterized by the highly conserved J-domain region. It interacts with proteins from another chaperone family, the HSP70 family, by stimulating their ATPase activity and controlling their function in protein folding, transportation, and degradation. 11
DNAJC12 is located on chromosome 10q21.3 and encodes 2 isoforms of a protein member of the HSP40 family. The protein consists of either 198 or 107 amino acids. Mutations in this gene have been associated with specific disorders such as hyperphenylalaninemia 12 and Parkinsonism.13,14 Additionally, studies have investigated its association with cancer, including lung cancer, gastric cancer, rectal cancer, and breast cancer.15 -18
In previous research, our team found that DNAJC12 was expressed differently in breast cancer and showed a connection between DNAJC12 mRNA expression and ER status in primary breast tumors. 19 We also demonstrated that treatment with 17β-estradiol increased DNAJC12 expression in MCF-7 cells through ER. 18 In this study, we aim to investigate further how DNAJC12 expression correlates with ER status and other clinicopathological characteristics in a more significant number of primary breast tumors to gain new insights and a better understanding of the potential prognostic and predictive values of DNAJC12 in breast cancer.
Materials and Methods
Data mining
We accessed the expression profiles of DNAJC12 mRNA in samples from 1108 patients with breast cancer through the Firehose Legacy TCGA dataset downloaded from cBioPortal (https://www.cbioportal.org/).20 -23 We included only samples with information regarding the PAM50 intrinsic subtype and DNAJC12 expression. Tumors classified as normal-like were excluded, and the resulting sample size was n = 808.
The Cancer Genome Atlas (TCGA) data was also accessed through the UALCAN platform to compare mRNA expression between normal and tumor samples.22 -25
We assessed the prognostic value of DNAJC12 mRNA expression using the Kaplan-Meier Plotter tool (https://kmplot.com/analysis/).24 -27 Overall survival (OS) and Relapse-free survival (RFS) times were plotted, with a follow-up period of 120 months. Patient groups were split using the best cutoff value, and we used JetSet best probe sets only.
ROC Plotter (https://www.rocplot.org/)25 -28 tool was used to investigate the predictive potential of the DNAJC12 gene, comparing mRNA expression and response to treatment. Pathological complete response and 5-year relapse-free survival were analyzed for hormonal therapy, anti-HER2 therapy, and chemotherapy.
A flowchart illustrating the analysis of data mining and the distribution of cohorts is depicted on the left side of Figure 1.
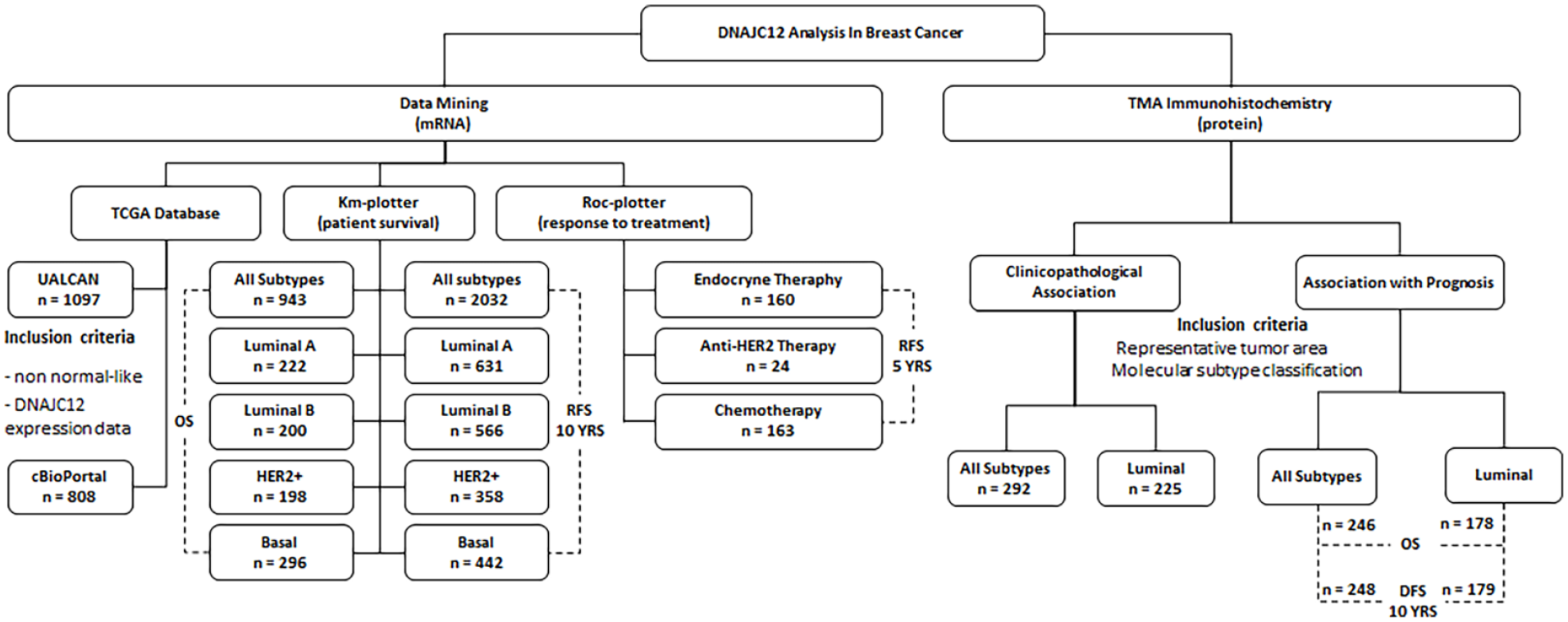
Study flowchart. The analysis of DNAJC12 expression was performed through data mining using public databases containing mRNA expression data (https://ualcan.path.uab.edu/; https://www.cbioportal.org/; https://www.kmplot.com/; https://rocplot.org/) and by immunohistochemistry (IHC) on tissue microarrays (TMA) containing 292 primary breast tumors.
Patients and tumor specimens
A total of 292 tumor tissues from BC patients were obtained retrospectively from the A. C. Camargo Cancer Center and Cancer Institute of Sao Paulo State (ICESP), both in São Paulo, Brazil. The archives were evaluated in duplicate by immunohistochemistry being included in this study only those of female patients with confirmed histopathological diagnosis of breast invasive carcinoma and with available demographic, clinicopathological, and survival data. All 292 patients were diagnosed with invasive ductal carcinoma of the breast. The follow-up period was corrected for long-term survival (range 0.23-120, median 91.73 months).
The study design followed the main criteria defined by REMARK, 29 and the flowchart describing the IHQ cohort distribution is presented on the right side of Figure 1.
Tissue microarray construction and immunohistochemical staining
The tissue microarrays (TMAs) containing breast tissue samples were constructed, and the IHC reactions were performed as previously described. 27 For IHC, the primary antibody used was affinity purified goat polyclonal antibody anti-DNAJC12 1:50 (Santa Cruz, sc104901). The optimal dilution was defined using well-characterized positive samples and tested before staining the TMA slides.
Immunostaining evaluation
QuPath for digital pathology (Quantitative Pathology Bioimage Analysis, v0.3.2-m1, University of Edinburgh, UK)
26
was used to analyze DNAJC12 protein expression on TMA slides quantitatively. Cells were counted and classified according to their average intensity of DAB staining as 0—negative, 1—weak, 2—medium, and 3—strong staining. The
Statistical analysis
IBM SPSS Statistics (version 25.0; SPSS Inc., IL, USA) was used for all statistical analyses. For TCGA mRNA expression data and IHC staining data collected from the TMA slides, contingency tables were assembled, and frequencies were compared using Pearson’s chi-square test. Median expression values for the TCGA data were compared using the Mann–Whitney
Ethical considerations
Following the ethical standards of the 1964 Declaration of Helsinki and its later amendments that are included in the Brazilian national resolution 466/12 (Resolução do Conselho Nacional de Saúde no. 466/12) the Institutional Ethics Committees (Faculdade de Medicina da Universidade de São Paulo, Protocol report CAAE: 35295514.1.0000.0065; and Fundação Antonio Prudente - Hospital AC Camargo Protocol report CAAE: 35295514.1.3001.5432) approved this tumor study, and all subjects provided informed consent.
Results
Data mining
We used the UALCAN tool to assess the expression pattern of DNAJC12 mRNA based on the TCGA dataset. The TCGA dataset analysis revealed that breast tumor samples exhibited elevated levels of DNAJC12 mRNA expression compared to normal tissue samples (Figure 2Aa). Furthermore, luminal breast cancer demonstrated significantly higher DNAJC12 expression compared to both normal tissue samples and other molecular subtypes. In contrast, triple-negative breast cancer displayed lower levels of DNAJC12 expression compared to normal tissue samples (Figure 2Ab).

DNAJC12 expression pattern and association with prognosis and treatment response. (A) TCGA data analysis in breast cancer samples in comparison to normal samples (a) or among molecular subtypes (b) using the UALCAN portal (https://ualcan.path.uab.edu/). (B) Impact of DNAJC12 expression in long term survival of breast cancer patients (a, b) and in each intrinsic molecular subtypes luminal A (c, d), luminal B (e, f), HER 2 (g, h) and basal-like (i, j). Kaplan-Meier curves of relapse-free survival (a, c, e, g, i) and overall survival (b, d, f, h, j) according to DNAJC12 expression using probe set 223722_at at KM Plotter online tool (https://www.kmplot.com/). (C) Association of DNAJC12 expression with response to endocrine therapy (a) anti-HER2 therapy (b) chemotherapy (c) (https://rocplot.org/).
Additionally, we downloaded the TCGA dataset from cBioPortal to explore the associations between DNAJC12 expression and clinicopathological characteristics of BC patients (Table 1). We found that the distribution of DNAJC12 expression levels significantly differs among groups when considering all 3 classical BC markers (ER
Associations between clinicopathological factors and DNAJC12 expression in breast cancer patients using data collected from TCGA public database.

Abbreviations: ER, estrogen receptor; PR, progesterone receptor.
Statistically significant (
For survival analysis, we used the KM-Plotter tool, and Kaplan-Meier curves were plotted to evaluate relapse-free survival (RFS) and overall survival (OS) when comparing groups with high- and low-expression of DNAJC12 (Figure 2B). When considering all BC patients, the expression level of DNAJC12 was able to distinguish groups of different prognoses for both RFS and OS, with the lower DNAJC12 expression associated with a worse prognosis (RFS
We used the ROC-Plotter tool to evaluate the predictive potential of DNAJC12 for response to treatment. Three different treatment regimens were considered in the analysis: neoadjuvant endocrine, anti-HER2 therapies, and adjuvant chemotherapy. For all 3 therapies, tumors of non-responder patients presented lower DNAJC12 expression. This biomarker was shown to be an effective classifier with high sensitivity for endocrine (
TMA immunohistochemistry
To further validate the expression pattern and prognostic value of DNAJC12, we assessed DNAJC12 protein expression by IHC on TMAs containing 292 primary BC samples. DNAJC12 protein was predominantly found in the cytoplasm of the tumor cells and was effectively scored in all 292 analyzable cases, which showed H-scores ranging from 100 to 340 (median, 51.5; Figure 3). This cohort comprised of 292 female patients with BC aged 26 to 91 years (median 53 years). Most of the patients were classified as ERpos (76.0%, n = 222), PRpos (69.5%, n = 203), HER2- (80.5%, n = 135), and early TNM stage (63.4%, n = 185). The molecular classification based on IHC expression data of ER, PR, and HER2 included 169 luminal A (57.9%), 56 (19.2%) luminal B, 15 (5.1%) HER2, 47 (16.1%) triple-negative subtypes, and 5 not classified (1.7%). The distribution of the patient’s demographic and clinicopathological parameters is presented in Supplemental Table 1.

DNAJC12 immunostaining in breast cancer primary tumor samples. Representation of the 4 DNAJC12 cytoplasmic staining intensities (A) low and (B) high. Image magnification is described in the figure.
As shown in Table 2, considering all valid cases, DNAJC12 protein expression was significantly different when comparing patients by ER tumor status (
Associations between clinical factors and DNAJC12 protein expression in breast cancer patients using data collected from immunohistochemistry-stained samples.

Abbreviations: ER, estrogen receptor; SBR, Scarff-Bloom-Richardson; HT, hormone therapy.
Considered statistically significant (
Survival analysis was also performed, considering patients’ disease-free survival (DSF) and overall survival (OS). We found that low DNAJC12 protein expression was associated with an increased risk of relapse and reduced rates of overall survival for all BC patients. (DFS:
Kaplan-Meier survival plots were also assembled, and in agreement with Cox Regression analysis, lower DNAJC12 expression was also associated with worse prognosis with all cases considered (DSF Logrank

Breast cancer survival analysis according to DNAJC12 protein expression. Patients were categorized in accordance with the median

Breast cancer survival analysis according to estrogen receptor status alone or in combination with DNAJC12 protein expression. Patients were categorized in accordance with estrogen receptor (ER, A, and B) the median
Discussion
In our previous study, we identified a correlation between the mRNA expression level of DNAJC12 and the estrogen receptor (ER) status in breast cancer and demonstrated its transcriptional regulation by ER. 18 Here, we performed in silico analysis using publicly available datasets and IHC on TMAs to better evaluate the potential role of DNAJC12 in breast cancer. We found that both DNAJC12 mRNA and protein down-regulation were associated with BC progression. DNAJC12 (DnaJ heat shock protein family (Hsp40) member C12) encodes a member of the heat shock protein family that constitutes a large family of proteins evolutive conserved and often classified based on their molecular weight into HSP10, HSP40, HSP60, HSP70, HSP90, etc. 8 DNAJC12 is widely expressed in cells and tissues and has been associated with hyperphenylalaninemia, dystonia, intellectual disability, and Parkinson’s disease.12,30,31 However, the role played by DNAJC12 in the tumorigenic process still remains unclear. Altered expression of DNAJC12 has been documented in lung cancer, rectal cancer, gastric cancer, and breast cancer.15 -18 Overexpression of DNAJC12 has been reported in lung cancer, and knockdown of DNAJC12 was able to inhibit cell proliferation, colony formation, migration, and invasion and induced apoptosis in lung cancer cells. 15 These authors also demonstrated that DNAJC12 knockdown reduced lung cancer cell growth in vivo through the suppression of p65 NF-κB phosphorylation, downregulating the expression levels and inhibiting the subsequent activation of β-catenin. 15 In rectal cancers, high expression of DNAJC12 protein was found to be associated with poor prognosis and poor response to neoadjuvant chemotherapy. 17 High levels of DNAJC12 mRNA expression were significantly associated with shorter overall survival in gastric cancer, and in vitro experiments showed that DNAJC12 knockdown decreases cell proliferation and invasion abilities of gastric cancer cells. 16 Different from our findings suggesting that DNAJC12 might be acting as a tumor suppressor gene in breast cancer, those that reported in lung, rectal, and gastric cancer suggest that DNAJC12 could act as an oncogene, implicating that DNAJC12 may have a dual effect as an oncogene or a tumor suppressor gene in a tissue-dependent manner.
The HSP40 family is the most prominent family of DNAJ proteins, encompassing 49 members, that acts as a cochaperone of HSP70. 32 All members of the HSP40 family possess 1 or more J domains, which are composed of 70 amino acids forming 4 α-helices containing the highly conserved HPD (histidine–proline–aspartate) motif, that facilitate the interaction with HSP70 and stimulate the ATPase domain in HSP70. 33 Therefore, the members of the HSP40 family, such as DNAJC12, act as cochaperones of Hsp70 and Hsp90 that participate in the steroid hormone receptor folding and activation.34,35 In yeast, Ydj1/Hsp40 mutants increase basal levels of ER activity in the absence of estrogen, indicating that the function of HSP40 is important for directing the ER to the HSP90 pathway. 36 Progesterone receptor (PR) chaperoning has been shown to start with the binding of a Hsp40 to a specific site on the PR, which is followed by the recruitment of Hsp70-ATP leading this complex to the HSP90 pathway through a dynamic association and re-association to the maintenance the pool of active PR. 37 Using an immunoprecipitation assay, Choi et al 38 demonstrated that the endogenous DNAJC12 and HSP70 proteins interact in LNCaP cells, confirming the role of DNAJC12 as a cofactor of the chaperone HSP70.
To date, little is known about the mechanisms of transcriptional regulation of DNAC12. In LNCaP prostate cancer cells, DNAJC12 expression is predominantly found in the cytoplasm and was up-regulated by the stress-inducing drug A23187 via the stress-regulated transcription factor AIbZIP/CREB3L4. 38 Previously, we were able to identify potential estrogen response elements (EREs) in the promoter region of the DNAJC12 gene and have demonstrated that in MCF-7 breast cancer cells, the DNAJC12 transcripts were up-regulated in a time-dependent fashion by 17ß-estradiol via ER. 18 More recently, using functional studies with MCF7 cells with overexpression or with knockdown of ER, Lin et al 39 identified that ER increases the expression of DNAJC12 and ERBB4 and DNAJC12 increases the expression of ERBB4, suggesting a regulatory relationship between ER, DNAJC12, and ERBB4 in breast cancer cells.
Conforming our previous results,18,19 we found a direct association between DNAJC12 mRNA and protein expression and the ER status in primary breast tumors. Corroborating our previous results and the findings presented here, Lin et al 39 also identified a positive correlation between ER status and the expression of DNAJC12. Moreover, in the present study we found that low DNAJC12 expression levels can stratify patients with ER-positive tumors with short RFS rates and patients with ER-positive and ER-negative tumors with poor overall survival, suggesting that the mRNA and protein expression of DNAJC12 would be helpful tool for identifying patients who need a more aggressive therapy and those with low relapse risk. In conclusion, the detection of DNAJC12 expression could be a promising biomarker for predicting the prognosis and hormone therapy for breast cancer patients. Here, we provide new information on the role of DNAJC12 as a prognostic factor in breast cancer that warrants new clinical and experimental studies better to define its importance as a breast cancer biomarker.
Conclusions
In conclusion, based on our current findings and previous research showing that DNAJC12 is up-regulated by estrogen through ER, we suggest that there is a feedback loop involving the DNAJC12/ER signaling pathway. This loop is linked to the advancement of breast cancer, holds prognostic potential, and could serve as a predictive factor for hormone therapy response in breast cancer.
Supplemental Material
sj-tif-1-bmi-10.1177_11772719251323095 – Supplemental material for Downregulation of DNAJC12 Expression Predicts Worse Survival for ER-Positive Breast Cancer Patients
Supplemental material, sj-tif-1-bmi-10.1177_11772719251323095 for Downregulation of DNAJC12 Expression Predicts Worse Survival for ER-Positive Breast Cancer Patients by Pedro Henrique Fernandes Gatti, Flavia Regina Rotea Mangone, Ana Carolina Pavanelli, Suely Nonogaki, Cynthia Aparecida Bueno de Toledo Osorio, Vera Luiza Capelozzi and Maria Aparecida Nagai in Biomarker Insights
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
