Abstract
Objective:
To elucidate mechanistic links among selenium measures, selenium-related genes, and immune traits in laryngeal carcinoma (LC) by integrating Mendelian randomization, bulk transcriptomic, and single-cell analyses.
Methods:
Two-sample Mendelian randomization (MR) was used to evaluate the causal effects of selenium measures and genetically predicted expression of selenium-related genes instrumented by eQTLs on LC risk, and immune traits were screened as candidate mediators via a directionality-based filter. We then examined PNPO expression in relation to pathway activity and immune infiltration in the TCGA and GEO cohorts. Single-cell RNA sequencing was further analyzed to characterize PNPO-associated programs in malignant epithelial and immune cells. The scPagwas and deconvolution algorithms were applied to prioritize disease-relevant T-cell subsets, and hdWGCNA was used to identify phenotype-associated gene modules. Intercellular communication, signaling activity, developmental trajectories, and transcription factor programs were evaluated in complementary analyses. The reporting of this study conforms to the STROBE-MR guideline for Mendelian randomization studies.
Results:
Genetically predicted selenium measures showed no evidence of a causal association with LC risk. In contrast, PNPO showed evidence consistent with involvement in LC susceptibility, potentially involving CD127− CD8+ T cells. In bulk transcriptomic cohorts, tumors with low PNPO expression were enriched for oncogenic pathways and were associated with poorer survival. PNPO expression correlated positively with inferred CD8+ T-cell infiltration. At single-cell resolution, PNPO− malignant epithelial cells displayed transcriptional features consistent with stemness, drug resistance, and immune-evasion programs. We further identified a pre-exhausted CD127− CD8+ T-cell subset with distinct molecular and functional characteristics.
Conclusion:
These analyses implicate PNPO-linked vitamin B6 metabolism and a pre-exhausted CD127− CD8+ T-cell state in LC, highlighting candidates for mechanistic validation and potential therapeutic exploration.
Introduction
Laryngeal carcinoma (LC) is a common head and neck malignancy, and approximately 98% of cases arise from the squamous epithelium of the laryngeal mucosa. 1 Laryngectomy remains a cornerstone treatment for selected patients with locally advanced, recurrent, or non-organ-preservation-eligible LC. 2 However, recurrence remains a major clinical challenge: postoperative relapse after primary total laryngectomy has been reported in ~27% to 31% of patients, 3 and the overall risk of recurrence within the first 3 years after treatment is ~20.5%. 4 Moreover, treatment decisions often require a trade-off between oncologic control and preservation of laryngeal function.5,6 Together, these issues underscore the need for biologically informed strategies that improve durable disease control while minimizing functional impairment.
Established risk factors—including tobacco use, alcohol consumption, and infection with human papillomavirus or Epstein–Barr virus—have been associated with LC etiology and progression.7,8 Yet LC remains clinically heterogeneous, with subsite-dependent behavior and outcomes, indicating that established exposures do not fully explain inter-individual variability. 9 Moreover, despite advances in multimodal therapy, outcomes remain poor for high-risk locally advanced disease and for recurrent/metastatic LC. 10 These gaps motivate efforts to identify actionable molecular pathways and immune mechanisms that can inform prevention and guide treatment.
Conceptual frameworks in cancer immunotherapy emphasize that immune cell states and signaling circuits can be leveraged for therapeutic targeting.11,12 CD8+ T cells are central mediators of tumor control, but chronic antigen exposure can drive a heterogeneous dysfunction program that spans precursor and terminally exhausted states.13,14 Importantly, marker-defined exhausted CD8+ T-cell subsets in epithelial tumors can represent biologically distinct states rather than a single terminal phenotype,15,16 with implications for biomarker development and target prioritization.
Against this background, selenium-related pathways have attracted interest; however, observational associations are difficult to interpret, and causal evidence in LC remains limited. We therefore used 2-sample Mendelian randomization to test causal links between selenium measures and selenium-related genes (focusing on PNPO) and LC risk. We further integrated bulk and single-cell datasets to characterize PNPO-associated epithelial programs and tumor-infiltrating CD127− CD8+ T-cell states.
Materials and Methods
An overview of the study workflow is shown in Figure 1; additional methodological details are provided in the Supplemental Materials and Supplemental Tables.

Flowchart of the study design and analytical workflow.
Reporting Guideline
The reporting of this study conforms to the STROBE-MR guideline, 17 and the completed checklist is provided in Supplemental Materials (STROBE-MR checklist).
Results
Causal Effects of Selenium Measures on LC Risk
We selected 12 single-nucleotide polymorphisms (SNPs) associated with selenium measures as instrumental variables (IVs; Supplemental Table S4), in line with a previous study. 18 Mendelian randomization (MR) analyses showed no evidence of a causal effect of genetically predicted selenium measures on LC risk (Supplemental Figure S1 and Table S15). Results of heterogeneity and horizontal pleiotropy assessments are provided in Supplemental Tables S19 to S20.
Causal Effects of Selenium-Related Genes on LC Risk
We included 109 selenium-related genes and 1171 corresponding expression quantitative trait loci (eQTLs; Supplemental Tables S2 and S5). MR screening across these genes nominated PNPO as a candidate associated with LC risk (Supplemental Table S16 and Figure S2). No evidence of heterogeneity or horizontal pleiotropy was observed (Supplemental Tables S19-S20). The inverse-variance weighted (IVW) analysis supported a statistically significant inverse association between genetically predicted PNPO expression and LC risk. In the FinnGen_R8_C3_LARYNX dataset, the odds ratio (OR) was 0.741 (95% CI, 0.569-0.963; P = .025; Figure 2A). This association was consistent in the independent GCST90041800 validation dataset (OR, 0.581; 95% CI, 0.435-0.776; P < .001; Figure 2B).

Forest plots showing an inverse causal association between genetically predicted PNPO expression and laryngeal carcinoma (LC) risk in 2 independent datasets: (A) FinnGen_R8_C3_LARYNX and (B) GCST90041800.
Associations of Immune Cell Traits with LC Risk and with Selenium-Related eQTLs
MR analyses identified immune cell traits showing evidence of association with LC risk (Supplemental Table S17). In the IVW analysis, CD127− CD8+ T cells were associated with LC risk in the FinnGen_R8_C3_LARYNX test set (OR, 0.498; 95% CI, 0.274-0.905; P = .022; Supplemental Figure S3). In the GCST90041800 validation cohort, CD20 on switched memory B cells was associated with LC risk (OR, 1.234; 95% CI, 1.006-1.514; P = .043; Supplemental Figure S4). We then evaluated associations between selenium-related eQTLs instruments and these immune phenotypes (Supplemental Table S18). In the IVW analysis, genetically predicted PNPO expression was significantly associated with CD127− CD8+ T cells in the test set and with CD20 on switched memory B cells in the validation cohort (Supplemental Figure S5).
Immune Mediation Screening and MR Sensitivity Analyses
Using a directionality-based filter, we identified immune traits showing a pattern consistent with potential mediation of the PNPO–LC association (Supplemental Table S21). Candidate mediators included CD127− CD8+ T cells, CD27 on IgD+ CD24+ B cells, CD20 on switched memory B cells, and CD33 on CD33+ HLA-DR+ CD14^dim cells, among others. Given the convergence of signals across the mediation screening, we prioritized the PNPO–CD127− CD8+ T-cell axis for downstream analyses (Figure 3). Leave-one-out analyses showed that the overall MR estimates were robust to removal of individual SNPs, with no single variant materially influencing the results (Supplemental Figures S2 and S6). Steiger directionality tests supported the assumed direction of effect, with no evidence favoring reverse directionality across instruments (Supplemental Tables S9-S14). Heterogeneity and horizontal pleiotropy assessment results are summarized in Supplemental Tables S19 to S20.

Schematic overview of the potential mediating role of CD127− CD8+ T cells in the association between PNPO expression and laryngeal carcinoma (LC).
Prognostic Relevance of PNPO and Association with Clinicopathologic Features
Kaplan–Meier analyses showed that survival differed significantly between PNPO-high and PNPO-low groups in both the GEO (P = .003) and TCGA (P = .033) cohorts. Higher PNPO expression was associated with more favorable outcomes—longer DFS in the GEO cohort and longer PFI in the TCGA cohort (Figure 4A). In the prognostic scoring framework, PNPO expression and clinicopathologic variables contributed to risk estimation to differing extents across stages. The integrated model showed that higher total scores corresponded to lower predicted 1 to 3-year survival probabilities (Figure 4B). Calibration curves showed good agreement between nomogram-predicted and observed outcomes (Supplemental Figure 7A). Notably, time-dependent receiver operating characteristic (ROC) analyses suggested that PNPO expression alone had limited prognostic discrimination in the GEO cohort, with a similar pattern observed in TCGA (Supplemental Figure 7B). PNPO expression was significantly higher in tumor than normal tissues (Supplemental Figure 7C). In contrast, PNPO expression showed no significant association with the evaluated clinicopathologic parameters (Supplemental Figure 7C).

Association between PNPO expression and prognosis in laryngeal carcinoma (LC): (A) Kaplan–Meier curves comparing survival between PNPO-high and PNPO-low groups, (B) Nomogram integrating PNPO expression and clinical variables for outcome prediction, and (C) Scatter plot of PNPO expression.
Single-Cell Transcriptomic Identification and Annotation of Cell Clusters
scRNA-seq datasets GSE206332 and GSE252490 were integrated and batch-corrected to generate a single Seurat object for downstream analyses. After quality control and normalization, and after removing 8608 doublets, 47 624 cells were retained; 30 principal components (PCs) were used for downstream analyses (Supplemental Figure 8A-C). PNPO expression was detected in 6.44% of cells (Figure 5B), consistent with sparse detection in scRNA-seq and potential dropout effects. Unsupervised clustering using the first 30 PCs (resolution = 0.7) identified 27 clusters, which were annotated into 9 major cell types based on canonical marker genes: epithelial cells, T/NK cells, neutrophils, myeloid cells, B cells, fibroblasts, plasma cells, endothelial cells, and mast cells (Figure 5A-C). Using CopyKAT, epithelial cells from tumor samples were classified as aneuploid (n = 18 093) or diploid (n = 3244; Figure 6A). Malignant epithelial cells accounted for >50% of the epithelial compartment and showed higher PNPO expression than non-malignant epithelial cells (Figure 6B and C). T/NK cells were then extracted and re-clustered, yielding 3262 CD127− CD8+ T cells and 928 CD127+ CD8+ T cells (Supplemental Figure 9A and B). Based on well-established marker genes and integrated T-cell functional scoring, CD127− CD8+ T cells were further subdivided into 4 states: cytotoxic, precursor-like exhausted (hereafter, “pre-exhausted”), cycling-exhausted, and dysfunctional exhausted (hereafter, “dysf-exhausted”; Figure 7).

Identification of 9 major cell types and the distribution pattern of PNPO expression in single-cell RNA data: (A) t-SNE plot displaying 27 distinct clusters annotated into 9 primary cell types, (B) scatter plot illustrating the expression pattern of PNPO across the identified cell types, and (C) canonical marker genes used for cell type annotation.

Identification of malignant epithelial cells and comparison of cell proportions and functional enrichment between malignant and non-malignant epithelial subpopulations: (A) t-SNE plot illustrating the identification and distribution of malignant epithelial cells, (B) comparison of PNPO expression levels between malignant and non-malignant epithelial cells, (C) relative proportions of different epithelial subpopulations, and (D) functional differences between malignant and non-malignant epithelial cells based on enrichment analysis.

Identification and functional characterization of CD127− CD8+ T cell subsets: (A) workflow and UMAP visualization of CD127− CD8+ T-cell subsets, (B) evaluation of cytotoxic and exhaustion-related functional states among CD127− CD8+ T-cell subsets, and (C) UMAP plot illustrating the distribution of pre-exhausted and proliferative states within CD127− CD8+ T cells.
Pathway and Functional Enrichment Analyses Based on Transcriptomic Data
In bulk transcriptomic analyses, GSEA and GSVA consistently indicated that low PNPO expression was associated with enrichment of angiogenesis, epithelial–mesenchymal transition (EMT), and Hedgehog signaling (Figure 8A and C). In the TCGA cohort, differential expression analysis identified 4003 DEGs between tumor and adjacent normal tissues (3031 upregulated and 972 downregulated; Supplemental Table S22 and Supplemental Figure 10A). Pearson correlation analysis (adjusted P < .05, r > .3) identified 444 PNPO-correlated genes in GEO and 1237 in TCGA. Intersecting the 2 co-expression gene sets with the TCGA DEGs yielded 10 hub genes (Supplemental Figure 10B and C; Figure 8E): RUVBL1, AHSA1, GNPDA1, NELFCD, DHX58, RAB7A, NMT1, ERAL1, NPEPPS, and COX7A1. PNPO showed a negative association with COX7A1 and positive correlations with the other 9 hub genes (Figure 8E). GO and KEGG enrichment analyses of genes positively correlated with PNPO across both cohorts highlighted pathways related to ubiquitin-mediated proteolysis, RNA degradation, valine/leucine/isoleucine degradation, catalytic activity acting on RNA, and transcription coactivator activity (Figure 8B).

Functional enrichment analysis and the relationship between PNPO expression and immune checkpoint genes: (A) gene set enrichment analysis (GSEA) comparing functional pathways between high and low PNPO expression groups, (B) Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses of genes positively correlated with PNPO expression, (C) Gene Set Variation Analysis (GSVA) revealing differences in pathway enrichment between the 2 cohorts, (D) Correlation analysis between PNPO expression and the expression levels of immune checkpoint genes, and (E) Correlation between PNPO expression and that of the 10 hub genes.
We next examined immune-relevant transcripts in bulk data. In TCGA, tumors with high PNPO expression showed higher expression of several immunotherapy-associated targets (PDCD1, CD27, CD48, and CD40), suggesting a more immune-infiltrated or immunologically active context in this subgroup. This pattern was not observed in the GEO cohort, indicating cohort-dependent variability and the need for further validation (Figure 8D).
To complement the bulk findings, we compared tumor and non-tumor cells in the scRNA-seq cohort. PNPO was upregulated in tumor-derived cells, consistent with the TCGA results (Supplemental Figure 11A). Differential expression between tumor and normal cells is summarized in the heatmap (Supplemental Figure 11B), and enrichment analysis of these DEGs indicated significant overrepresentation of immune response-related pathways in tumor cells (Supplemental Figure 11C and D). At the epithelial compartment level, malignant epithelial cells displayed increased activity of multiple oncogenic hallmark programs, including PI3K–AKT–mTOR, mTORC1 signaling, MYC targets, and endoplasmic reticulum stress response pathways (Figure 6D). In contrast, non-malignant epithelial cells were relatively enriched for immune and inflammatory signatures, consistent with potential roles in epithelial barrier function and immune surveillance (Figure 6D).
Malignant epithelial cells were then stratified into PNPO+ (n = 1703) and PNPO− (n = 16 390) groups based on PNPO detection in the normalized expression matrix (PNPO+, expression > 0; PNPO−, expression = 0). The corresponding expression patterns are shown in Figure 9A. Inflammation-related pathways were predominantly enriched in the PNPO− cells, likely reflecting tumor-intrinsic stress/inflammatory signaling rather than effective anti-tumor immune infiltration, whereas metabolic reprogramming and cell proliferation pathways were significantly enriched in the PNPO+ cells (Figure 9B and C; Supplemental Figure S12A and B). Notably, PNPO− malignant cells also exhibited higher scores for immune suppression, immune evasion, tumor stemness, and drug resistance (Figure 9D; Supplemental Figure S12B), and showed a trend toward enrichment of angiogenesis and EMT pathways, consistent with the bulk RNA-seq analyses (Supplemental Figure 12C).

Functional and immunological differences between PNPO − and PNPO+ malignant epithelial cells: (A) differentially expressed genes (DEGs) between PNPO+ and PNPO− malignant epithelial cells, (B and C) functional enrichment analyses illustrating pathway differences between PNPO− and PNPO+ malignant epithelial cells, and (D) comparison of immunosuppressive and immune evasion features between the PNPO+ and PNPO− groups.
Because PNPO+ CD127− CD8+ T cells were rare (n = 50) and exhibited low expression, we did not detect robust transcriptional differences between PNPO+ and PNPO− CD127− CD8+ T cells. Therefore, subsequent analyses focused on precursor-like exhausted (pre-exhausted) and cycling-exhausted CD127− CD8+ T-cell subsets. Functional enrichment results are shown in Figure 10, and pseudo-bulk DEGs are summarized in Figure 10B.
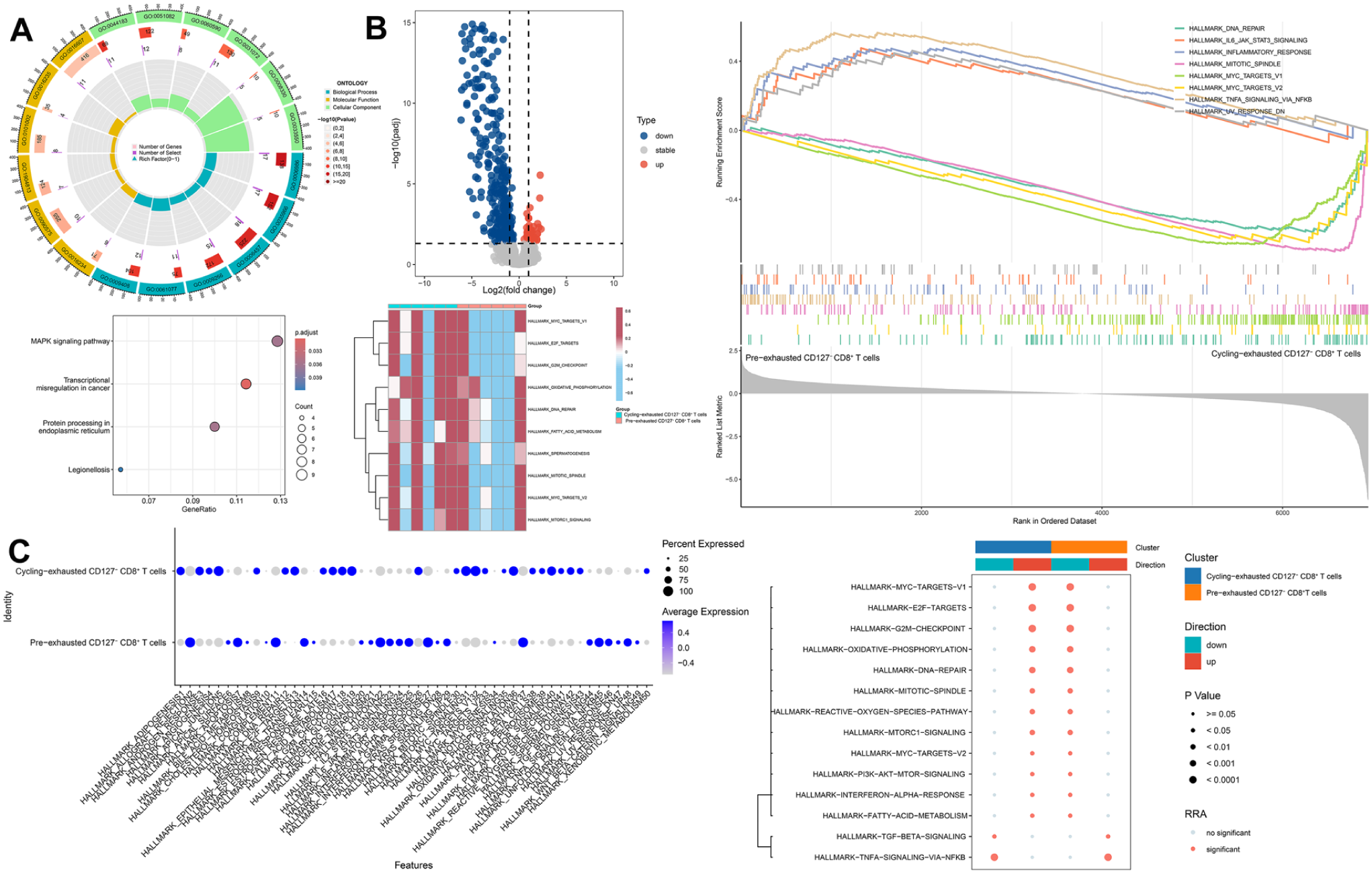
Functional differences between pre-exhausted and cycling-exhausted CD127− CD8+ T cells: (A) Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses of characteristic genes in pre-exhausted CD127− CD8+ T cells, (B) volcano plot showing differentially expressed genes (DEGs) between pre-exhausted and cycling-exhausted CD127− CD8+ T cells, generated using the DESeq2 package, followed by functional enrichment analyses based on Gene Set Enrichment Analysis (GSEA) and Gene Set Variation Analysis (GSVA), and (C) functional enrichment analyses highlighting the biological pathways and molecular processes associated with hallmark gene sets.
Metabolic Pathway Analyses and Identification of Cd127− Cd8+ T-Cell Subgroups Associated with LC
Metabolic pathway scoring indicated enrichment of multiple programs linked to immunosuppression and immune evasion in both PNPO− and PNPO+ malignant epithelial cells, including tryptophan metabolism, arachidonic acid metabolism, fatty acid metabolism, and steroid hormone biosynthesis (Figure 11A). Compared with PNPO+ cells, PNPO− malignant epithelial cells exhibited additional metabolic heterogeneity, with preferential enrichment of sphingolipid metabolism, retinoic acid metabolism, mucin-type O-glycan biosynthesis, and amino-sugar and nucleotide-sugar metabolism (Supplemental Figure 13A). PNPO− cells also showed stronger enrichment of drug metabolism–cytochrome P450 pathways, which may be consistent with their higher drug-resistance signatures (Figure 11A).

Metabolic differences among subtype-specific cells and the correlation of CD127− CD8+ T cells with laryngeal carcinoma (LC): (A and B) metabolic variations among subtypes of malignant epithelial cells and CD127− CD8+ T cells and (C) trait-relevant score (TRS) of CD127− CD8+ T-cell subtypes, highlighting their potential relevance to LC.
Across CD127− CD8+ T-cell subsets, we observed marked metabolic differences (Figure 11B and Supplemental Figure S13B). Overall, pathway activity tended to be higher in the cycling-exhausted subset, with the strongest activation in dysfunctional exhausted (dysf-exhausted) CD127− CD8+ T cells, suggesting metabolic remodeling that accompanies functional dysregulation.
To prioritize immune subpopulations enriched for LC-associated genetic signals, we applied scPagwas using the Reactome gene set (c2.cp.reactome.v2025.1.Hs.symbols). Among T-cell subsets, pre-exhausted CD127− CD8+ T cells showed relatively higher trait-relevant scores (TRS) than other cell types (Figure 11C and Supplemental Figure S13C), indicating that this state is more strongly enriched for LC-associated genetic relevance and was therefore prioritized for downstream analyses.
Classification of the Immune Infiltration of LC and Drug Response Prediction
To assess the relationship between PNPO expression and tumor immune contexture, bulk RNA-seq deconvolution indicated a significant positive association between PNPO expression and inferred CD8+ T-cell infiltration (Figure 12A). Estimated fractions of 22 immune cell types and their correlation are shown in Supplemental Figure 14A and B. In a targeted analysis of 4 immune subsets, CIBERSORTx did not identify a comparable association between PNPO expression and CD127− CD8+ T cells (Figure 12B). This likely reflects the limited resolution of bulk deconvolution for CD127-defined CD8+ sub-states and differences between GWAS immune traits and transcriptome-based estimators. In the GEO cohort, stromal and immune components differed between PNPO expression subgroups; however, these differences were not reproduced in TCGA, indicating cohort-dependent effects that require additional validation (Supplemental Figure 14C). Consistently, Immunophenoscore (IPS) values did not differ across PNPO subgroups (Supplemental Figure 14C).
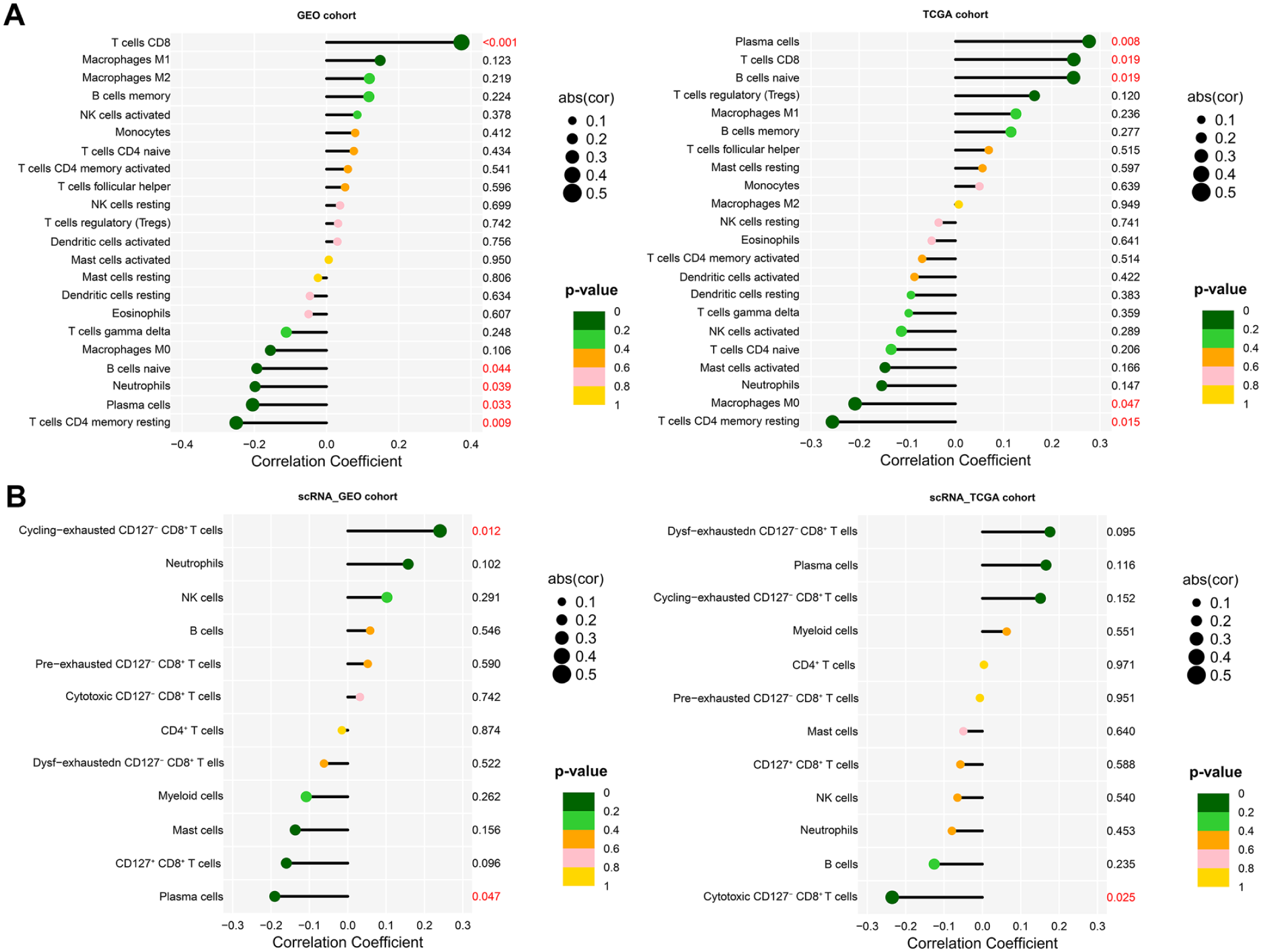
Comparison of immune cell infiltration between the Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA) cohorts using LM22 (A) and a single-cell RNA-derived signature gene matrix (B).
We next explored whether PNPO expression relates to predicted drug response. Using oncoPredict trained on the Genomics of Drug Sensitivity in Cancer (GDSC) resource, we estimated drug sensitivity as predicted IC50 values. PNPO-high tumors showed differential predicted sensitivity to gefitinib, erlotinib, osimertinib, sapitinib, and OF-1, agents largely linked to the EGFR pathway (Figure 13A and B). Predicted sensitivity profiles for CD127− CD8+ T-cell subtypes and malignant epithelial subtypes are provided in Supplemental Figure 15A to C. Notably, SB505124 (an RTK signaling inhibitor) showed lower predicted IC50 in malignant epithelial cells (Supplemental Figure 15C).

Prediction of drug sensitivity based on PNPO expression in the GEO cohort (A) and the TCGA cohort (B).
Identification of Differentiation Trajectories and Co-expression Gene Modules of Cd127− Cd8+ T Cells
We reconstructed CD127− CD8+ T-cell differentiation trajectories using Monocle3, setting cluster 7 (cytotoxic CD127− CD8+ T cells) as the pseudotime root (Figure 14A). The inferred trajectory indicated a progression from cytotoxic cells toward a precursor -like exhausted state, followed by divergence into terminal fates, including cycling-exhausted and dysfunctional-exhausted (dysf-exhausted) CD127− CD8+ T cells (Figure 14A and Supplemental Figure S16A).

Pseudotime trajectory and identification of key gene modules in CD127− CD8+ T cells: (A) Pseudotime analysis shows the developmental trajectory of CD127− CD8+ T cells originating from cluster 7, with PDCD1, TOX, MKI67, and GZMB expression illustrating key functional states. (B) Identification of the M2 module as a key independent gene module significantly associated with CD127− CD8+ T cells. (C) Visualization of the top 10 hub genes within the M2 module along the pseudotime trajectory.
To characterize coordinated transcriptional programs, we performed hdWGCNA in LC using a soft-thresholding power of β = 14, which identified the M2 module as specifically associated with pre-exhausted CD127− CD8+ T cells (Figure 14B). Functional enrichment analyses showed that M2 module genes were significantly enriched for the MAPK signaling pathway. Consistently, pseudotime mapping of the top 10 hub genes from this module suggested increasing activity along MAPK signaling and lipid metabolism-related programs (Figure 14C and Supplemental Figure S16C).
Finally, CytoTRACE analysis of epithelial cells suggested that PNPO− malignant epithelial cells occupy a more differentiated, later-stage state, whereas PNPO+ malignant epithelial cells exhibit higher developmental potential, consistent with a less differentiated phenotype (Supplemental Figure S17), noting that CytoTRACE scores can be influenced by transcriptional diversity and cycling-related signals and may not map one-to-one onto EMT/stemness signatures.
Ligand-Receptor Communication Network Construction and Transcription Factor Prediction
CellChat analysis indicated that both PNPO+ and PNPO− malignant epithelial cells participate broadly in intercellular communication, acting as both signal senders and receivers across multiple pathways. Although their overall outgoing signaling strength was similar, PNPO− malignant epithelial cells exhibited markedly higher incoming signaling strength, consistent with a stronger role as signal recipients (Figure 15 and Supplemental Figure S18). Across the network, the predominant pathways were enriched for immunosuppressive and immune-evasion-related signaling (Figure 15).

Cell–cell communication relationships among different cell types in the cell–cell contact category (A) and the secreted signaling category (B).
Notably, CD39 signaling was primarily initiated by pre-exhausted and cycling-exhausted CD127− CD8+ T cells and was directed toward PNPO+ malignant epithelial cells (Figure 15A). Because CD39 (ENTPD1) promotes the conversion of extracellular ATP to AMP and supports downstream adenosine production, this axis is widely linked to an immunosuppressive milieu that can dampen effector T-cell function and facilitate immune evasion. In addition, pre-exhausted CD127− CD8+ T cells showed active SEMA4 and type II interferon (IFN-γ) signaling and expressed PAR receptors (Figure 15A and B and Supplemental Figure S19). Together, these communication patterns suggest that the pre-exhausted subset may retain partial effector-like activity while engaging in immunomodulatory interactions within the tumor microenvironment: IFN-γ signaling is compatible with residual cytotoxic potential, whereas SEMA4 signaling and PAR receptor activity may reflect programs related to tissue remodeling and immune regulation.
We next used SCENIC together with decoupleR to infer transcription factor (TF) regulatory programs associated with LC. In both the GEO and TCGA cohorts, the PNPO-low group showed broader activation of TFs previously linked to immunosuppression, immune evasion, angiogenesis, and EMT (Figure 16). At single-cell resolution, inferred TF activity varied across cell populations (Figure 17 and Supplemental Figure S20). For example, BCL3 (+), KLF16 (+), and NFYA (+) were preferentially upregulated in pre-exhausted CD127− CD8+ T cells (Figure 17A), consistent with transcriptional programs that may influence cellular metabolism, cell-cycle regulation, and NF-κB-related signaling, thereby shaping T-cell persistence and exhaustion trajectories. Conversely, JUND (+), TFAP2A (+), XBP1 (+), FOSB (+), and KLF5 (+) were predominantly upregulated in PNPO− malignant epithelial cells (Figure 17B). Given their reported roles in tumor progression and immune modulation, these TFs may support checkpoint-associated and other immunoregulatory programs that contribute to an immune-evasive phenotype. The inferred regulatory networks and predicted targets for key TFs in CD127− CD8+ T cells and malignant epithelial cells are shown in Supplemental Figures S21 and S22, respectively.

Comparison of the top 20 differentially expressed transcription factors between PNPO+ and PNPO− groups in the GEO cohort (A) and the TCGA cohort (B). *P < .05. **P < .01. ***P < .001.

Top 5 cell type-specific transcription factors identified in CD127− CD8+ T-cell subpopulations (A) and epithelial cell subpopulations (B).
Discussion
Laryngeal carcinoma (LC) reflects the combined influence of environmental exposures, host genetics, and the tumor immune microenvironment.7,8 Building on immunotherapy frameworks that link immune cell states and signaling programs to therapeutic opportunities,11,12 and on evidence that exhausted CD8+ T cells comprise heterogeneous, marker-defined states rather than a single terminal endpoint,15,16 we integrated Mendelian randomization (MR) with bulk and single-cell transcriptomics to interrogate selenium-related biology in LC. Three findings emerged. First, genetically predicted selenium measures showed no evidence of a causal association with LC risk in our MR analyses. Second, PNPO—a selenium-related gene encoding pyridoxal-5′-phosphate oxidase, which generates pyridoxal-5′-phosphate (PLP), the active form of vitamin B6—showed evidence consistent with a protective association with LC susceptibility. Third, multi-omics profiling linked PNPO to malignant epithelial programs associated with aggressive behavior and to CD127− CD8+ T-cell states, including a pre-exhausted subset prioritized by enrichment of LC-associated genetic signals.
Selenium contributes to redox homeostasis and immune regulation and has been implicated in cancer-relevant pathways in experimental systems. 19 However, our MR analyses did not support a causal association between selenium measures and LC risk. This does not negate selenium’s biological relevance; rather, it indicates that mechanistic plausibility may not translate into population-level causal effects for LC. Selenium-cancer associations also appear context dependent (eg, baseline status, exposure timing, and population background), and high-quality human evidence has not consistently shown a universal cancer-preventive benefit of supplementation. 20 Even so, experimental work supports several plausible routes by which selenium could influence tumor biology. For example, selenium has been reported to enhance antioxidant defenses via Keap1–Nrf2 signaling, including increased GPx and TrxR activity, thereby limiting ROS-driven inflammatory signaling (eg, NF-κB/AP-1) and downstream genomic damage.21 -24 Selenium compounds have also been linked to apoptosis-related programs through AMPK/mTOR/FOXO3a and p53 signaling.25,26 Additional effects have been described on cell-cycle control,27,28 angiogenesis,28,29 immune effector function,30,31 and epigenetic regulation.32,33 Taken together, the null MR results may reflect heterogeneity in exposure context and study populations, as well as limitations of the available genetic instruments.
Our MR analyses did not support a causal association between selenium measures and LC risk, whereas PNPO showed evidence consistent with a protective association. PNPO encodes pyridoxal-5′-phosphate oxidase, which generates pyridoxal-5′-phosphate (PLP), the active form of vitamin B6. Together, these findings support a PNPO–PLP-linked hypothesis for laryngeal carcinogenesis. Notably, in our data the signal was driven by PNPO-related genetic instruments rather than selenium measures, suggesting that any effect—if causal—may reflect vitamin B6/PLP-related biology rather than selenium exposure per se. Epidemiologic studies have reported that higher circulating vitamin B6/PLP levels are associated with a lower risk of several cancers and with improved prognosis in colorectal cancer.34 -37 Immune-related mechanisms have been proposed as 1 potential contributor to these associations.38 -40 Vitamin B6 status has also been linked to broader health outcomes, including all-cause mortality and cardiovascular endpoints,41,42 and lower vitamin B6/PLP levels have been reported in smokers and in older individuals with polypharmacy. 43 Although these observational findings do not establish causality for LC, they provide context for prioritizing PNPO as a biologically credible candidate and motivate mechanistic studies of the PNPO–PLP axis in both tumor-intrinsic programs and anti-tumor immunity.
PLP is a cofactor for more than 150 enzymatic reactions, 44 and experimental studies suggest that PLP insufficiency can promote genomic instability and tumor-promoting phenotypes.45 -49 In line with these observations, our data associate low PNPO expression in malignant epithelial cells with poorer outcomes (shorter DFS/PFI) and with transcriptional programs related to EMT, angiogenesis, stemness, drug resistance, immunosuppression, and immune evasion. By contrast, higher PNPO expression was associated with increased proliferative activity in malignant epithelial cells. This pattern is consistent with prior reports that PNPO can promote proliferation through Cyclin B1/CDK1 phosphorylation, with downstream effects on lysosomal biogenesis and autophagosome turnover. 50 In addition, PNPO overexpression has been linked to activation of TGF-β and Wnt/β-catenin signaling, together with increased proliferation, migration, and invasion, in other cancer types.51,52 Although these mechanisms have not been demonstrated in LC, they provide a reasonable context for our observation that PNPO+ malignant epithelial cells exhibit a proliferation-skewed transcriptional profile, whereas PNPO− malignant epithelial cells are enriched for EMT- and stress-related programs. Direct evidence linking PNPO to EMT remains limited. However, ROS-related stress has been implicated in EMT induction and maintenance of stem-like states.53,54 One plausible interpretation is that reduced PNPO/PLP activity may indirectly favor EMT-like programs by exacerbating redox imbalance and genomic stress. This interpretation is compatible with the pathway patterns observed in our transcriptomic analyses, but it remains hypothesis-generating and will require experimental testing in LC models.
Vitamin B6 catabolism can be indexed by 4-pyridoxic acid (4-PA), and a higher 4-PA/PLP ratio has been associated with cancer risk in prior work. 42 In addition, negative feedback regulation of PNPO by PLP has been proposed in ovarian cancer. 51 Although the mechanisms by which PLP or 4-PA regulate PNPO expression remain unclear, these observations suggest that PNPO abundance may reflect both inherited regulation and metabolic context. In ovarian cancer models, targeting the PNPO–LAMP2–autophagy axis has shown anti-tumor activity, 50 supporting the broader view that PNPO-linked pathways may be therapeutically actionable, albeit in a tumor- and context-dependent manner.
Importantly, whereas many prior studies focus on PNPO at the level of tumor protein expression, our MR framework interrogates genetically predicted PNPO expression using eQTL instruments. This difference in biological level and tissue context may help explain why reported directions of association vary across tumor types. In our study, the genetic evidence is consistent with a protective association of PNPO with LC susceptibility; however, the extent to which circulating-cell eQTLs capture PNPO regulation in tumor tissue remains an important consideration. Several studies38 -40,55,56 link vitamin B6/PLP status and PNPO-related biology to anti-tumor immunity and immune-cell infiltration. In our single-cell analyses, PNPO− malignant epithelial cells exhibited transcriptional programs consistent with a more immune-suppressive/immune-evasive milieu (ie, reduced effector-supporting signals despite active stress/inflammatory programs), which may contribute to poorer clinical outcomes. Prior work suggests that tumor growth can induce local vitamin B6 deficiency and dampen NK-cell activation, and that vitamin B6 supplementation—particularly when combined with one-carbon metabolism blockade—can reduce tumor burden in vivo. 56
Mechanistically, reduced PNPO activity may lower PLP availability, with downstream effects on lysosomal acidification, macrophage function, iron homeostasis, and TET2 stability that collectively favor an immunosuppressive microenvironment. 57 Although these mechanisms have not been established in LC, they suggest testable links between the PNPO–PLP axis and immune regulation and are consistent with the immune-cold programs observed in PNPO− malignant epithelial cells in our dataset.
Our directionality-based screening identified a mediation-consistent pattern linking genetically predicted PNPO expression, CD127− CD8+ T-cell traits, and LC risk, suggesting that CD127− CD8+ T cells may be 1 immune route through which PNPO relates to LC susceptibility. This is consistent with immunotherapy-oriented target discovery concepts in which therapeutic and biomarker opportunities can emerge when specific signaling programs are mapped onto defined immune cell states within the tumor microenvironment. 11 Accordingly, integrating MR with bulk and single-cell profiling provided an immune-state context for interpreting PNPO-linked genetic signals and highlighted a CD127− CD8+ T-cell dysfunction axis. Consistent with this, scPagwas ranked pre-exhausted CD127− CD8+ T cells among the highest TRS-scoring T-cell states, indicating enrichment of LC-associated genetic signals in this population. This focus is not merely descriptive: in epithelial tumors, “exhausted” CD8+ T cells comprise multiple transcriptionally and functionally distinct states rather than a single terminal endpoint, and marker-defined subsets can differ in functional potential and therapeutic relevance. 15 In LC, resolving CD127− CD8+ T-cell heterogeneity helps specify which dysfunction-associated programs warrant mechanistic follow-up and which states are most informative for biomarker development or therapeutic exploration.
In this framework, checkpoint- and stress/hypoxia-related programs form the pathway/target layer, whereas the genetically prioritized pre-exhausted CD127− CD8+ T-cell population captures the state layer (higher TRS than other T-cell states). Although this does not establish mechanism, it motivates focused testing of whether PNPO/PLP-linked regulation of these programs influences pre-exhausted-state stability or state transitions, with implications for target and biomarker prioritization.
Vitamin B6 is required for normal T-cell function, and pharmacologic inhibition of PLP synthesis limits CD8+ T-cell proliferation and differentiation and weakens anti-tumor immunity in mice. 38 To place these findings in an immunologic context, we briefly outline how PLP-linked pathways and CD127/KLRG1-defined effector differentiation relate to the emergence and evolution of exhausted CD8+ T-cell states in tumors. PLP has also been reported to promote PD-L1 degradation, which may enhance T-cell cytotoxicity by modulating the programmed cell death protein 1 (PD-1)/programmed death-ligand 1 (PD-L1) axis. 58 During effector differentiation, CD127 and KLRG1 are commonly used to distinguish memory-precursor effector cells (KLRG1low CD127high) from short-lived effector cells (KLRG1high CD127low). After antigen clearance, CD127− CD8+ T cells typically undergo contraction and apoptosis. By contrast, sustained AhR signaling has been reported to antagonize HIF-1α activity and support memory formation and long-term survival, although the contribution of this axis likely depends on context. 59 Within the tumor microenvironment, CD127− CD8+ T cells commonly exhibit exhaustion features. 60 Pathways linking hypoxia and immune regulation may shape how these cells move across exhaustion states. Vitamin B6 has been described as a hypoxia-inducible factor (HIF) inhibitor, 61 and has been linked to memory-leaning CD8+ T-cell differentiation. 59 Vitamin B6 has also been reported to enhance CXCL13 secretion by CD160+ CD8+ T cells, which may support tertiary lymphoid structure formation, potentially by limiting HIF-1α degradation. 62 Along the same axis, PNPO-linked PLP metabolism has been implicated in PD-L1 turnover, raising the possibility that it modulates PD-1/PD-L1 signaling and, in turn, exhaustion dynamic. 58 Notably, exhausted CD8+ T cells span a continuum of functionally distinct states rather than a single terminal phenotype, with clear implications for immunotherapy target selection and biomarker development. In head and neck squamous cell carcinoma (HNSCC), Wang et al 16 reported that TCF1+ terminally exhausted cells represent a predominant subset among tumor-infiltrating CD8+PD-1+ T cells and that their presence correlates with improved survival. Similarly, proliferative exhausted CD8+ T cells have been associated with durable anti-tumor responses, particularly in human papillomavirus-positive HNSCC treated with immune checkpoint inhibitors (ICIs). 63 Conversely, other studies have linked deeper CD8+ T-cell exhaustion within the tumor microenvironment to poorer prognosis. 64
Taken together, these reports provide clinically relevant context for interpreting our findings, but they do not, on their own, establish a mechanism in LC. In our study, the data-supported observations are that PNPO shows genetic evidence consistent with a protective association and that we identify and prioritize a pre-exhausted CD127− CD8+ T-cell state in LC. A plausible interpretation is that the PNPO–vitamin B6/PLP axis intersects with immune checkpoint and hypoxia-related programs to shape CD8+ T-cell states, potentially contributing to the more favorable outcomes observed in PNPO-high tumors. Whether PNPO—alone or combined with immune checkpoint inhibitors—can shift pre-exhausted CD127− CD8+ T cells toward a more functional, therapy-responsive state remains a testable hypothesis requiring mechanistic and translational validation.
Although integrating multi-omics datasets can clarify links among genetic variation, immune cell states, and cancer phenotypes, our study has several limitations. First, we did not perform experimental validation; therefore, mechanistic interpretations should be regarded as hypothesis-generating. Second, the MR analyses were conducted in European-ancestry datasets, whereas the scRNA-seq data were derived from Asian cohorts, which may limit generalizability across ancestries. Third, immune-mediated effects were not consistently replicated in the validation dataset, and no associations remained significant after Bonferroni correction, which limits the strength of inference. In addition, relatively lenient instrumental-variable selection thresholds may increase the risk of false-positive findings. Fourth, PNPO instruments were derived from circulating-cell eQTLs and may not fully capture PNPO regulation within tumor tissue. Moreover, PNPO+ cells were rare within the CD127− CD8+ T-cell compartment in our integrated scRNA-seq dataset, limiting state-specific single-cell comparisons. Finally, despite the Steiger directionality test, the limited number of available instruments restricted sensitivity analyses for reverse causality. Accordingly, our findings should be interpreted cautiously and validated in larger, ancestry-matched cohorts, alongside functional experiments to test whether PNPO/PLP perturbation alters malignant epithelial programs and the differentiation and function of pre-exhausted CD127− CD8+ T cells in LC.
Conclusion
In conclusion, our MR analyses did not support a causal effect of selenium measures on LC risk, whereas PNPO showed evidence consistent with involvement in LC susceptibility. The association of PNPO− malignant epithelial cells with stemness, drug-resistance, and immune-evasion programs is consistent with a poorer prognostic profile. Together, these findings highlight the PNPO–vitamin B6/PLP axis and a pre-exhausted CD127− CD8+ T-cell state as priorities for mechanistic validation and potential therapeutic exploration in LC.
Supplemental Material
sj-docx-1-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-docx-1-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-docx-2-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-docx-2-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-10-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-10-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-11-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-11-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-12-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-12-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-13-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-13-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-14-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-14-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-15-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-15-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-16-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-16-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-17-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-17-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-18-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-18-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-19-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-19-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-20-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-20-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-21-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-21-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-22-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-22-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-23-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-23-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-24-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-24-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-3-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-3-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-4-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-4-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-5-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-5-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-6-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-6-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-7-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-7-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-8-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-8-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Supplemental Material
sj-tif-9-cix-10.1177_11769351261438477 – Supplemental material for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma
Supplemental material, sj-tif-9-cix-10.1177_11769351261438477 for Integration of Multi-Omics Data Identifies the Role of the Selenium-Related Gene PNPO and Pre-exhausted CD127− CD8+ T Cells in Laryngeal Carcinoma by Lin Xiang, Lin Chen, Cheng Gong and Yao Tang in Cancer Informatics
Footnotes
Acknowledgements
We thank the participants and investigators of FinnGen and the IEU OpenGWAS Project, as well as all researchers who made their data publicly available.
Ethical Considerations
This study used only publicly available, de-identified datasets; therefore, ethical approval was not required.
Consent for Publication
All authors have read and approved the final manuscript and agree to its publication.
Author Contributions
Lin Xiang and Lin Chen performed the research. Cheng Gong and Yao Tang designed the study. Lin Chen analyzed the data. Lin Xiang, Yao Tang, and Lin Chen drafted the manuscript. All authors reviewed the manuscript and approved the final version.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Science and Technology Project of Enshi Tujia and Miao Autonomous Prefecture (Grant Nos. D20220022 and D20230055). The funder had no role in the study design; data collection, analysis, or interpretation; manuscript preparation; or the decision to submit for publication.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data analyzed in this study were obtained from publicly available resources and previously published studies. Accession numbers and dataset details are provided in the Supplementary Materials. The supplementary dataset supporting this study is available via Zenodo (DOI: 10.5281/zenodo.18690888).
Supplemental Material
Supplemental material for this article is available online.
Code Availability
Custom R scripts used for data processing and analysis are available from the corresponding author upon reasonable request.
AI-assisted Writing Disclosure
During manuscript preparation, the authors used ChatGPT to assist with language editing and readability. The authors reviewed and edited the output and take full responsibility for the final content. No scientific data were generated, modified, or interpreted using AI.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
