Abstract
Multiple sclerosis (MS) is a complex autoimmune disorder of the CNS that affects millions of people worldwide. The causes of the disease remain unknown despite extensive efforts to understand it. CircRNAs are a unique class of endogenous non-coding RNA that are abundant, stable, conserved, and specifically expressed molecules, making them a promising biomarker of diseases. This review investigates the role of circRNA in MS pathogenicity and their potential as a biomarker through a comprehensive literature search conducted in 8 scientific databases. The studies found that there are differentially expressed circRNAs in MS patients compared to healthy controls (HC), and this difference is even more pronounced in different MS subtypes. Enrichment of circRNAs in linkage disequilibrium (LD) blocks that harbor MS-associated SNPs suggests that these SNPs manipulate the levels of circRNAs in the surrounding area, contributing to disease pathogenicity. While circRNA shows promise as an indicator or biomarker for MS disease pathology, further research is needed to fully explore its potential and impact on human biology.
Keywords
Introduction
Multiple sclerosis (MS) is a complex, multifactorial autoimmune disorder of the CNS with no defined etiological factors and over 2.8 million diagnosed patients worldwide. 1 MS causes inflammation, demyelination and neurodegeneration and can result in the appearance of a wide spectrum of symptoms, including fatigue, cognitive dysfunction, paresthesia or numbness, motor weakness (mostly in the lower extremities), monocular visual disturbances (optic neuritis), ataxia and vertigo. Remyelination and neurogenesis may occur during the remission phase of the disease, whereupon patients show signs of recovery. In the chronic phase of MS, however, neurological disabilities are mostly irreversible due to axonal loss.2 -5 Four different subtypes of MS are clinically distinguishable: relapsing-remitting MS (RRMS), primary-progressive MS (PPMS), secondary-progressive MS (SPMS), and progressive-relapsing MS (PRMS). RRMS, which represents 85% of MS cases, is characterized by unpredictable relapsing episodes, alternating with periods of remission; in RRMS, cranial focal areas of inflammation and demyelination can be resolved over time, as the damage is reversible. PPMS comprises around 10% to 15% of MS cases, in which a steady accumulation of neurological disability, mainly progressive myelopathy, appears from disease onset. Some patients develop superimposed relapses; these amount to nearly 5% of MS cases, and are said to display PRMS, which, similarly to PPMS patients, have a progressive disability from the onset of symptoms, but with acute attacks or relapses. SPMS represents a stage subsequent to RRMS; patients progress from RRMS to SPMS with time and show progressive neurological decline, either with or without relapses after the initial relapsing-remitting pattern. 6
Efforts in understanding the disease has been extensively made with over than 106 121 published articles in PubMed database since the year 1879. However, the disease remains elusive, and no definite causes are known. Environmental, immunological, genetic, and epigenetic factors have all been implicated in the disease’s development, but none can fully explain it. Consolidating these factors may lead to a better understanding of the disease. Environmental factors such as vitamin D deficiency, Epstein-Barr virus (EBV), smoking, obesity, and alcohol consumption have been linked to the disease. Immunological dysregulation, where immune privilege attacks the brain, has been extensively studied. Genome-wide association (GWA) studies and association studies have identified several genetic variants, with HLA-DRB1*15:01 being one of the most prominent factors. Epigenetic modifications, including DNA methylation, histone modification, and mRNA silencing, have been implicated in the disease’s pathogenesis.2,5,7,8
Genetics has long been linked to the pathogenesis of MS, but it does not provide a complete explanation. Large-scale studies have identified several genomic locations containing genetic variants that appear to elevate the risk of MS. Currently, more than 236 independent genetic variants are known to increase MS risk, but their presence alone does not explain the disease pathology since they are not exclusive to MS. Most of these variants are non-coding and affect gene expression and silencing. They have been implicated in immunological, neurological, and hemostasis pathways, and have been found to impact innate and adaptive immune regulation. Each of these variants can increase MS risk, but none of them are essential for the development of MS. Environmental factors may also play a role in MS development, possibly through the effects of these variants (reviewed previously2,5,7 -9).
Non-coding RNA (ncRNA), specifically circular RNA (circRNA), is a new area of focus for understanding the disease’s pathogenicity. Of all transcribed RNA, only 3% produce mRNA, which results in protein production. The remaining are classified as ncRNA, which can be divided into 4 subtypes based on their shape, length, and location: microRNA (miRNA), long ncRNA (lncRNA), circRNA, and PIWI-interacting RNA (piRNA). miRNA are small RNA molecules that contain about 22 nucleotides and target mRNA to silence them by recruiting RNA-induced silencing complex (RISC). piRNA are 24 to 30 nucleotides in length and are involved in epigenetic regulation of chromatin with the help of PIWI family proteins. LncRNA and circRNA are longer than 200 nucleotides and regulate gene expression through different mechanisms, with lncRNA being linear and circRNA forming a circular structure. All of these RNA subtypes have been linked to the progression of diseases and cancer.10 -12 This review will focus on the role of circRNA in the pathogenesis of MS and its potential as MS biomarkers to further understanding the disease.
CircRNA molecules are a distinct type of ncRNA that are abundant, stable, conserved, and expressed specifically. They are more prevalent than linear transcripts and their abundance varies depending on the developmental stage or age. These molecules have a half-life of over 48 hours, and many are tissue-specific. CircRNAs are conserved throughout evolution and have been mapped in numerous species, including humans, mice, nematodes, zebrafish, Drosophila, protists, and plants.13 -16 According to their composition, circRNAs can be categorized as exonic circRNAs (ecircRNAs), circular intronic circRNAs (ciRNAs), and exonic-intronic circRNAs (EIciRNAs). 13 Other categories of circRNAs include fusion circRNA (f-circRNAs), read-through circRNAs (rt-circRNAs), and mitochondria-encoded circRNA (mecciRNAs). CircRNAs can be found in various cell compartments depending on their category. For instance, ciRNAs are located in the nucleus, while ecircRNAs are present in the cytoplasm and exosomes. EIciRNA is situated in the nucleus, f-circRNAs can be found in all cellular compartments, rt-circRNAs are located in the cytoplasm, and mecciRNAs are present in the mitochondrial environment. 17 CircRNAs have several functions, including acting as miRNA sponges, protein scaffolds, and protein translators. Their unique structure, which consists of covalently closed RNA circles without a 5′ cap or 3′ tail, makes them more resistant to endonucleases than linear RNA and, as a result, they are more suitable as biomarkers.13,14,18,19
Method
A comprehensive literature search was conducted to extract data regarding circRNA in MS and EAE, its effect on the pathogenicity of MS, and their possible role as MS biomarkers (Figure 1). This article represents a systematic review of the published literature. This review was performed by a single reviewer.

Depicts the methodology employed in this review to extract data on the role of circRNA in MS and summarize our current understanding.
Inclusion and exclusion criteria
The following inclusion criteria were developed and refined as needed during the search process. All the studies were required to be (1) published journal articles, or electronically published ahead of printing, (2) peer-reviewed articles, and (3) written in English. No restrictions were placed on the date of publication. There were no restrictions on the inclusion of human or animal species in the studies analyzed in this review.
Studies that (1) does not match the purpose of this article; (2) are written in languages other than English; (3) are not full-text articles, for example, an abstract or poster paper; (4) are not peer-reviewed papers; or (5) are duplicate papers were excluded.
Search strategy
In the initial stages of the search, the author’s focus was on establishing a precise definition of circRNA, encompassing its historical background, biogenesis, and function. To achieve this, relevant review papers were scrutinized, and additional resources were consulted, as needed, from the reference lists of the screened articles.
Subsequently, the author shifted their focus toward delineating the role of circRNA in MS, as well as exploring its potential as a biomarker for the disease. Additionally, a comprehensive list of search terms was developed, and keyword searches were conducted, which included:
- Multiple sclerosis or its variant terms (eg, multiple sclerosis, MS; relapsing-remitting multiple sclerosis, RRMS; secondary progressive multiple sclerosis, SPMS; primary progressive multiple sclerosis, PPMS; experimental autoimmune encephalomyelitis, EAE) combined with circular RNA or its variant terms (eg, circRNA, ciR).
Search source
Searches were conducted in NCBI PubMed, Googler Scholar, CNKI Scholar, WorldCat, Cochrane Library, WHO Global Index Medicus, SciELO, JSTOR databases.
• PubMed search was filtered to include only studies with (1) English language, (2) full-text articles, and (3) exclude preprints.
• Google Scholar search was filtered to include only papers with (1) English language (2) that contain the exact search terms in the title of the article (Advanced Search).
• The CNKI Scholar search was limited to (1) academic journals, (2) full-texted article, and (3) English language.
• The WorldCat search was limited to (1) articles, and (2) English language.
• The Cochrane Library search was limited to search terms found in title/ abstract/ keywords.
• The WHO Global Index Medicus search was filtered to include only papers with (1) English language, (2) full-text article, (3) that contain the search terms in the title, abstract and subject.
• No filters were applied to SciELO or JSTOR engines.
Additional references were identified via the reference list from the searched articles. All searches were performed before the end of July 2023.
Study selection
Titles and abstracts were screened for eligibility. Ineligible studies were excluded while the remaining studies underwent a full-text review. The studies were screened for mentions of circRNA and MS or its relevant terms.
Data extraction and summary measures
Data extracted included the following:
[1] History of circRNA.
[2] Biogenesis of circRNA.
[3] Function of circRNA.
[4] Significant dysregulation of circRNA in MS or its subtypes, supported by P-values (P) and fold change (FC).
[5] CircRNA as pathogenic molecules in MS disease development, supported by P and FC.
[6] CircRNA as biomarkers for MS or its subtypes, supported by P and FC.
Results
Study selection
Figure 2 illustrates the key stages that were followed to write and complete this review, in accordance with the PRISMA 2020 guidelines. 28 Database searches retrieved a total of 60 articles. The term “circRNA” or its variants yield 20 272 articles in PubMed that were published between the years 1964 and 2023. The term “Multiple Sclerosis” or its variants yield 90 924 articles in PubMed that were published between the years 1879 and 2023. The combined terms “circRNA” and “Multiple Sclerosis” or their variants have yield only 34 articles in PubMed database that were published between the years 2015 and 2023. The same terms yield only 3 articles in Google Scholar that were published between the years 2021 and 2023. The CNKI Scholar engine yield 1 articles that were published between the year 2019 and 2023. The WorldCat engine yield 20 articles that were published between the years 2019 and 2023. JSTOR search yielded irrelevant 2 articles. While in SciELO, Cochrane Library and WHO Global Index Medicus search engines, typing the same terms yield 0 matching articles.

Illustrates the key stages done for writing and completing this review as per PRISMA 2020 guidelines. 99
NCBI PubMed search
Out of the 34 articles reviewed, only 10 were deemed relevant and included. The remaining 24 were excluded for the following reasons: 16 were review papers and 8 were found to be irrelevant. These 8 papers included a study on the function of circ_0000518 that was associated with MS in the discovery experiment but ruled out in the validation experiment; an article describing a software model for predicting circRNA-disease association; a study on the role of MALAT1 dysregulation and alternative splicing alteration in MS pathogenesis; a study on the expression and interaction patterns of circRNA in Amyotrophic Lateral Sclerosis (ALS); a study on the expression and replication of circDNA in human cells; a study investigating the role of CRISPR/Cas9 system in eliminating Polyomavirus JC Infection; a study aimed at screening bioactive compounds for inhibition of Repeat-associated non-AUG (RAN) translation; and a study investigating the profile of circRNA in frail individuals.
Google Scholar search
One of the 3 articles obtained was included in this review, while the other 2 were found to be redundant and therefore excluded.
CNKI Scholar search
The article retrieved was already included in this review.
WorldCat search
Out of the 20 articles reviewed, 8 were duplicates and 12 were excluded for the following reasons: 8 were review papers and 4 were found to be irrelevant. These 4 papers included a study on the expression and interaction patterns of circRNA in ALS; a study focusing on the role of circ_HECW2 function in LPS-induced endothelial-mesenchymal transition; a study on the function of circ_0000518 that was associated with MS in the discovery experiment but ruled out in the validation experiment; an article describing a software model for predicting circRNA-disease association.
JSTOR search
The search yielded 2 irrelevant articles: one pertaining to miRNA and epilepsy, and another that was a review on the presence of a new class of competing endogenous RNA.
In total, 11 published articles were included in this review (see Table 1). Data collection for the review began in April 2023 and concluded in July of the same year.
The studies that were included in this review.

Abbreviations: circRNA, circular RNA; CIS, clinically isolated syndrome; CSF, cerebrospinal fluid; EAE, experimental autoimmune encephalomyelitis; EV, extracellular vehicle; HC, healthy control; IgG, immunoglobulin G; LD, linkage disequilibrium; LS-OCMB, anti-myelin lipid-specific oligoclonal IgM band; miRNA, microRNA; mRNA, messenger RNA; MS, multiple sclerosis; PBMC, peripheral blood mononuclear cell; qRT-PCR, quantitative reverse-transcriptase polymerase chain reaction; RRMS, relapsing-remitting MS; SNP, single nucleotide polymorphism; SPMS, secondary-progressive MS.
Data extraction
Data extracted were thoroughly reviewed under “a bit of history,” “circRNA biogenesis,” “circRNA functions,” “circRNA in MS,” and “circRNA as MS biomarkers” sections. This review only includes data related to circRNA in MS that have significant P-values (<.05).
A bit of history
CircRNA was discovered in 1976 by Kolakofsky, who identified defective interfering (DI)-RNA in 4 strains of Sendai virus. These circular structures had complementary endings and were described as unique segments of the viral genome. 20 Sanger et al discovered covalently closed circRNA molecules in plants in the same year, which they named Viroid. 21 CircRNA later was observed in viral genome and in several eukaryotic cells.22,23 In 1993, circular molecules that corresponded to gene transcripts were identified in human and mice testes.24,25 The origin of circRNAs has been debated, with some suggesting they were the result of nuclear spliceosomal mistakes, while others found they were nonrandom products of RNA splicing that could control gene expression.26,27 In 2012, Salzman et al reported that circRNAs were abundant, stable, conserved, and nonrandom products of RNA splicing that could be involved in controlling gene expression.15,28,29 Memczak et al provided evidence for the post-transcriptional regulatory function of circRNAs in 2013, 30 and in 2014 Ashwal-Fluss et al found that circRNAs functioned in gene regulation by competing with linear splicing. 31 By 2017, more functions of circRNAs were identified as protein-coding RNA or protein-translator32,33 and protein scaffold.34,35
CircRNAs biosynthesis
In 1979, Hsu and Coca-Prados observed circRNAs in eukaryotic cells and described them as non-random structures with high sequence homology that is rich in G-C content. The structures had pan-handles that were equidistant from each other, and their size varied. The circRNAs were observed in poly(A)+ form. 23 Over 2 decades later, the biogenesis of circRNAs began to be understood, with 3 main models being proposed (Figure 3).
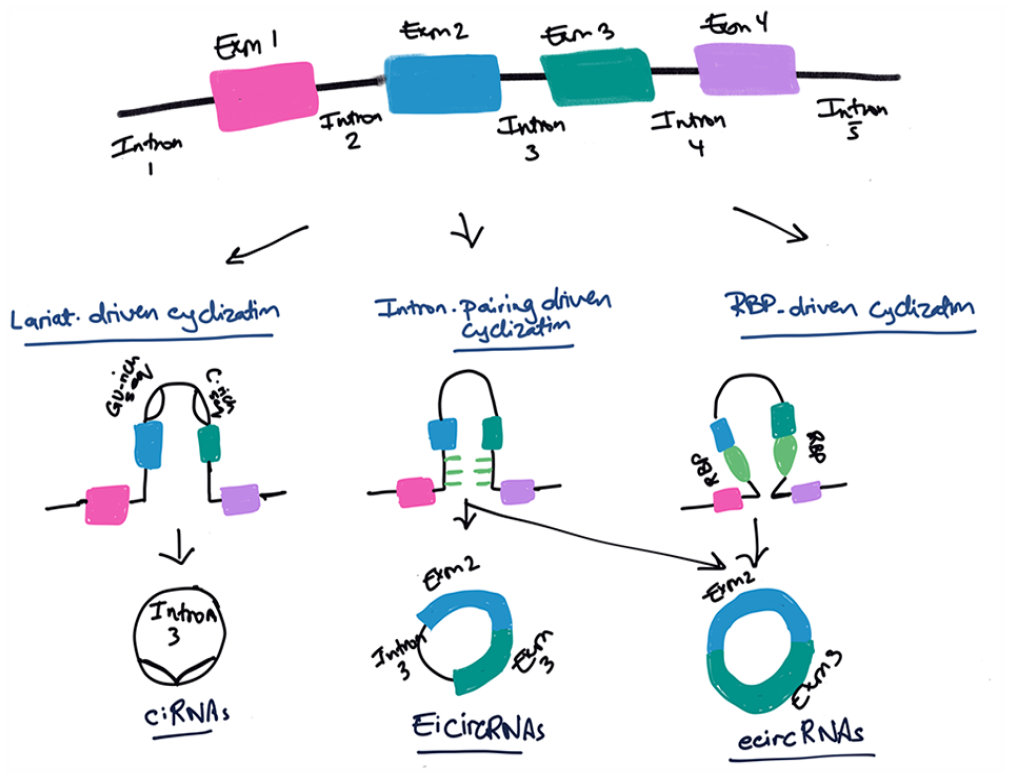
The 3 main models proposed for circRNA biogenesis.
Direct back-splicing
In back-splicing, a splice donor is jointed to an upstream splice acceptor (eg, 3′ end of exon 2 is joined to the 5′ end of exon 2). Most exons have splicing sites at both ends of the gene and can circularize in theory, but few were seen in circularization form and back-splicing seems to be tissue-specific. 36 Back-splicing machinery depends on the presence of the reverse complementary sequences or inverted repeats (IR) that can be repetitive complementary sequences (eg, ALU sequence) or non-repetitive complementary sequences. These sequences must be 30 to 40 nt or more and the longer the sequences are, the higher the production rate of circRNA.17,29,37 More so, the sequence and the length of the circularized exon determines the circularization rate and efficiency.31,36 Kramer et al suggested that the length of the IR dictates the timing of circularization, with long repeats (∼300 nt or longer) more likely promoting co-transcriptional back-splicing. 36 Upstream introns and downstream exons containing the IR elements will pair directly through base complementation. The splicing sites will be spatially close to each other, forming exon–intron circRNA without intron elimination or exon circRNA with intron elimination. The diversity of the complementary sequences allows selective and competitive pairing and generates multiple circRNAs in the same gene. 38
Exon-skipping or lariat-driven cyclization
Kelly et al suggest that the circularization of RNA is strongly associated with exon skipping, which means that exons that are not represented in mRNA are likely to be represented in circRNAs. 39 Zhang et al explained the process of circularizing long intronic sequences of pre-mRNA known as ciRNA. This process depends on the presence of a consensus motif that includes a 7 nt GU-rich element near the 5′ splice site and an 11 nt C-rich element adjacent to the branchpoint site. 40 Barrett et al stressed the significance of the 5′- and 3′-branch points of the circularized RNA in the lariat model; deleting either of these sites should disrupt circularization. In their model, they described the circularization of exon-containing lariats and found that exon skipping alone is insufficient for detectable levels of circRNAs. They also discovered that exon-containing lariats are not sufficient for RNA circularization and suggested that other regulatory mechanisms are responsible for producing circRNAs. They proposed that lariat production and back-splicing are not necessarily 2 distinct mechanisms in the production of circRNAs. Instead, lariat production could assist in back-splicing catalysis by positioning splice sites and preventing competing splice sites from functioning. Exon length was also found to be a significant factor in the circularization process, as small exons may restrict circularization due to physical limitations. 41
RNA-binding-protein-driven circularization
According to this model, RNA-binding proteins (RBPs) can identify and attach to specific gene sequences in introns, prompting the formation of circRNAs. RBPs will generate splicing sites at both ends of exons that are in close proximity to each other through protein interaction or dimer formation, creating covalent links between the splicing acceptor and splicing donor. This process results in circularization and the formation of circRNAs.38,42 RBPs, such as muscleblind protein and quaking protein, are known to facilitate the formation of circMbl and circQKI. 43
Biogenesis of other circRNA types
F-circRNA are produced from transcribed exons of distinct genes affected by the translocations and can contribute to disease development and survival. 44 Rt-circRNAs are fusion circles that include coding exons from 2 adjacent and similarly oriented genes, their function remains unknown. 45 MecciRNAs are encoded by both the light and heavy strands of the mtDNA, although the heavy strand of mtDNA encodes the majority. They appear to regulate essential physiological functions within the mitochondria. 46
Regulation of CircRNA biosynthesis
Ashwal-Fluss et al said that circRNAs are generated co-transcriptionally and their abundance is controlled at the biosynthesis level. They suggested that circRNA generation and pre-mRNA splicing are in competition. CircRNA production was regulated by the relevant gene expression, higher MBL gene expression resulted in higher circMbl. Knock-down of MBL resulted in 2- to 6-fold decrease of circMbl levels. The presence of putative MBL binding sites in the circMbl flanking introns was the reason that MBL overexpression regulated circMbl biosynthesis. MBL-induced circularization depended on the binding of MBL to both MBL intronic binding sites simultaneously, mutation of the MBL sites in only one of the flanking introns is sufficient to significantly affect the circMbl levels. 31
Kramer et al said that a set of different regulatory factors deduct the circularization of different RNAs. The elegant feedback loop seen in MBL example may not be relevant for all circRNAs. Knock-down of the Drosophila Laccase2 mRNA by targeting exon 3 of the gene did not affect Laccase2 circRNA levels. Rather, a specific set of heterogeneous nuclear ribonucleoprotein (hnRNP) and serine–arginine (SR) proteins regulated the expression levels of the Laccase2 circRNA. Testing 6 other genes indicated that different sets of hnRNP and SR proteins affect the splicing and biogenesis of different circRNAs. Likewise, not all circRNAs were controlled by hnRNP and SR proteins, circMbl expression was not affect by hnRNP and SR proteins knock-down. 36
CircRNA function
Despite the incomplete understanding of the function of circRNA, its role in controlling gene expression has been documented. CircRNA can function as competing endogenous RNA or miRNA sponge, translating proteins, and protein scaffold (interacting with RNA binding proteins, modulating the stability of mRNAs, regulating gene transcription, and protein transporters).19,47,48 A single circRNA molecule can neutralize several miRNAs simultaneously and thus influence post-transcriptional regulation. CircRNA can be translated into microprotein that can compete with their host encoded protein and act as a decoy molecule that can either silence the host encoded protein or prevent their degradation. Examples include CDR1as or ciRS-7, circ-SRY, circ-ITCH, circ- HIPK3, hsa_circ_0000615, and mm9_circ_012559. CircRNA can interact with RNA binding proteins, influencing their function. Examples include circ-Foxo3, and circ-MBL (muscleblind). Some circRNAs help stabilize the mRNAs; CDR1as and circ-RasGEF1B are example of which. Others regulate gene transcription and these always exist in the nucleus; circ-EIF3J, circ-PAIP2, ci-ankrd52 and ci-sirt7. They can influence DNA methylation of downstream genes; example of which is circRNA-ACR.47,48 They can act as reverse transposons that alter the structure of the genome and adjust gene expression.48,49 CircRNA can act as protein transporters that facilitate the translocation of functional proteins from the cytoplasm to nucleus.13,14,17,48
CircRNA role in immune regulation and disease progression
CircRNA plays a crucial role in regulating the immune system and has been linked to the development of cancer and disease progression. For instance, circNDUFB2 acts as a protein scaffold and enhances the interaction between TRIM25 and IGF2BPs to inhibit the growth and metastasis of non-small cell lung cancers (NSCLC). It also regulates RIG-I-MAVS signaling cascades and attracts immune cells to the tumor microenvironment. 50 Similarly, circIGF2BP3 is involved in NSCLC and negatively correlates with CD8+ T-cell infiltration. 51 In breast cancer patients, hsa_circ_001783 promotes tumor progression by sponging miR-200c-3p. 52 In rheumatoid arthritis and SLE, circRNA_09505 and circRACGAP1 regulate AKT1 and PTEN/AKT signaling, respectively, and circ-0008945 is associated with increased apoptosis of PBMCs.53 -55 There are multiple other documented examples of circRNAs that affect the regulation of the immune system and the progression of cancer and diseases.56 -61
CircRNA role in neurodegenerative diseases
CircRNAs have been implicated in the development of neurodegenerative diseases. In Parkinson’s disease, circEPS15 acts as a miR24-3p sponge to promote the stable expression of the target gene PINK1, enhancing the elimination of damaged mitochondria and maintaining mitochondrial homeostasis. 62 In Alzheimer’s disease, circ_0003012 affects the pathophysiology of the disease by manipulating cGMP-PKG signaling. 63 The hippocampus of Alzheimer’s patients has been observed to exhibit decreased levels of ciRS-7, leading to the reduced expression of UBE2A, a crucial enzyme for Aβ clearance. This circRNA acts on miR-7, which subsequently downregulates UBE2A. 64 In a study by Zhang et al, dysregulated circRNA m6A methylation was observed in Alzheimer’s mice models, implicating them in the disease pathology. 65 Additional examples of circRNAs that contribute to the development of neurodegenerative diseases exist.66 -69
CircRNA in MS
MS is a neurodegenerative disorder of the CNS that is mediated by the immune system. As circRNAs have been associated with immune regulation and the progression of neurodegenerative diseases, it is plausible that they may be involved in the development of MS. The search for scientific articles related to circRNA and MS only yielded 34 results. This indicates that our understanding of the role of circRNAs in MS pathogenesis is still limited. In fact, we are still in the process of comprehending the role of circRNAs in general. CircRNAs were only discovered in 1976 and new insights about the molecule continue to be published daily. Therefore, we should be cautious in drawing any conclusions about the role of circRNAs in MS. This paper may provide a preliminary understanding of the potential role of circRNAs in MS pathogenesis, if any.
Differences in circRNA expression between MS patients and healthy controls
Recent studies have shed light on the potential involvement of circRNAs in MS. Iparraguirre et al reported 406 out of 13 617 circRNA that were tested to be differentially expressed (FC ⩾ 1.5, P ⩽ .05) in RRMS-remission patients, compared to healthy controls (HC). 324 were downregulated and 82 upregulated. Of these 2 (circ_0005402, and circ_0035560) were confirmed to be downregulated in validation cohorts that included RRMS-remission, -relapse and CIS patients. Both circ_0005402, and circ_0035560 are located on chromosome 15 within the ANXA2 gene. 70 ANXA2 known to play a role in the traversal across the blood-brain barrier (BBB) 71 and promote miR-155 expression via STAT3 (Signal Transducer and Activator of Transcription 3) phosphorylation. 72 miR-155, interestingly, is a biomarker for MS that can be detected in patient body fluids before the onset of the disease and have been documented to increase in expression during relapses stages. 7 Cardamone et al identified a novel isoform of GSDMB circRNA (hsa_circ_0106803) (GSDMB gene encodes Gasdermin B) that was upregulated by 2.8-fold in RRMS patients when compared to HC (P = .0011). 73
Variations in circRNA expression among different MS subtypes
In another paper, Iparraguirre et al conducted a study that examined the differential expression of circRNAs collected from extracellular vehicles (EVs) in MS patients and HC. EVs are released by various cell types and play a role in numerous biological and pathological processes. The study identified 100 circRNAs that were differentially expressed between RRMS patients and HCs (FC > 1.5 and P < .05), with 47 being upregulated and 53 downregulated in RRMS patients. Additionally, 75 circRNAs were found to be differentially expressed between SPMS and HCs. Out of these, 32 were upregulated in SPMS patients (P-adj < .05), and 43 were downregulated (P-adj < .05). The study also revealed that the profile of circRNAs between RRMS and SPMS was different, where 2734 circRNAs were designated for RRMS and 658 circRNAs were designated for SPMS. Interestingly, the authors claimed that the function of circRNAs found in EVs and leukocytes is not primarily miRNA sponging, as the length and miRNA binding sites found in the sequence do not support such a function. 74
Enrichment of circRNAs in linkage disequilibrium blocks containing MS-associated SNPs
Paraboschi et al described a significant enrichment of circRNAs mapping in the 73-linkage disequilibrium (LD) blocks harboring MS-associated single nucleotide polymorphisms (SNPs) (a total of 482 circRNAs in MS-associated LD blocks vs 194 ± 65 in the random sets of LD blocks). To test whether the different genotypes of an MS-associated SNP could impact the expression levels of a circRNA mapping in the corresponding LD block, rs2293152 was genotyped and crossed against the expression levels of the circRNAs hsa_circ_0043813. 75 rs2293152 is located within the STAT3 gene, in particularly 50 nucleotides (nt) upstream of exon 14 and has been associated with MS in GWA studies. 76 Hsa_circ_0043813 is also located in the STAT3 gene and composed of exons 12, 13, and 14. The data showed significant different expression levels upon genotype stratification, with the homozygous CC healthy subjects showing the highest levels of expression (one-way ANOVA P = .023). They then analyzed RNA-seq data of 2 cell lines, SH-SY5Y and Jurkat cells, representing tissues relevant for MS, and found 18, 2 of which novel, circRNAs derived from MS-associated genes. As such they suggested MS-associated SNP exerts its functional effect by manipulating the levels of specific circRNAs in the near vicinity and, hence, modifying the ratio of the circular: linear RNA isoforms. 75 Cardamone et al proposed that MS-associated SNPs may influence the expression levels of circRNAs through exon-based DNA methylation. Their study analyzed 5663 circRNAs, of which 166 were found to be dysregulated in 10 RRMS samples compared to 10 HCs (P < .05), with 125 being downregulated. These circRNAs were associated with various immune functions, gene transcription, and chromatin regulation. During the validation step, the study identified PGAP3 hsa_circ_000799 as a top downregulated circRNA, with statistical significance found in both the US and Italian replication cohorts (FC = −0.44, and −0.40, P = .049 and .0075, respectively). The study also investigated the effect of MS-associated variants on circRNA expression levels and found that 89 out of 285 SNPs had a significant effect on circRNA expression. The rs7295386 polymorphism was found to affect the expression of 4 circRNAs, including ATF1 hsa_circ_0098746. The study observed a significant correlation between the genotype of rs7295386 and the expression of hsa_circ_0098746 in MS patients (US MS patients/ HCs ANOVA P = .0077; Italian MS-patients/HCs ANOVA P = .0027). Finally, the study found a significant correlation between exon methylation and circRNA expression levels. 77 These findings suggest a potential mechanism by which MS-associated SNPs may regulate circRNA expression levels and contribute to the pathogenesis of MS.
The circRNA/miRNA/mRNA networks
Huang et al conducted an intriguing analysis of circRNA/miRNA/mRNA networks using previously published differential expression results in MS patients. They utilized bioinformatics methods to examine the expression patterns of circRNAs, miRNAs, and mRNAs in MS patients from 3 previously published data sets (Mycko et al, 78 Keller et al, 79 Kemppinen et al 80 ). Starting with the top differentially expressed circRNA from the Mycko et al data set, 78 they identified the miRNAs that bind to it and their corresponding target mRNA. By analyzing the data presented by both Keller et al 79 and Kemppinen et al, 80 they were able to identify differentially expressed miRNAs and mRNAs that could be linked to the differentially expressed circRNA. This approach led to the discovery of 4 circRNA/miRNA/mRNA networks that were found to impact B-cell proliferation and B-cell receptor signaling pathways. Further analysis of these networks revealed their potential implications for MS pathology. 81
CircRNAs as biomarkers
Defining biomarkers for diseases is crucial for early discovery and treatment. Ideally, biomarkers that are stable, informative, and non-invasive are preferred. CircRNAs are stable molecules that resist several endonucleases, making them a promising biomarker material. If these molecules serve as biomarkers for MS, a simple non-invasive blood test could lead to early diagnosis, relapse discovery or treatment effect. In fact, several circRNA biomarkers have been identified for MS, including active disease stage, disability score and immune-mediated demyelination.
CircRNAs as biomarkers for active MS
Late in 2020, Iparraguirre et al proposed 6 circRNAs as global, specific, and sex-dependent MS-biomarkers; hsa_circ_0141241, hsa_circ_0058514, circPADI4, hsa_circ_0001947, hsa_circ_0001707, hsa_circ_0001459. All of which showed upregulation trends in female but not in male MS patients. CircRNA isoforms but not the linear isoforms had been upregulated (FC > 1.58, P < .0001). 82 They also defined 2 circRNAs (hsa_circ_0000478 and hsa_circ_0116639) and 2 linear RNAs (IRF5 and MTRNR2L8) as minimally invasive biomarkers of highly active MS that is characterized by the presence of anti-myelin lipid-specific oligoclonal IgM bands (LS-OCMBs) (performance accuracy of 70%). 83
Zurawska et al identified circRNA biomarkers for inflammatory activity in RRMS patients. The study analyzed the profile of circRNA in 104 participants (67 RRMS patients and 37 HC) using microarray and quantitative reverse-transcriptase polymerase chain reaction (qRT-PCR). Of the 13 617 human circRNAs included in the study, 914 transcripts were differentially expressed between RRMS-relapsed patients and HC (P < .05; FC for upregulated circRNAs > 2.0, and down-regulated circRNAs < 0.5) and 622 were differentially expressed between RRMS-remission patients and HC (P < .05; FC upregulated > 2.0, down-regulated < 0.5). The study identified 15 differentially expressed circRNAs, between RRMS-relapse and -remission patients (P < .05; FC > 2.0), with 2 of these circRNAs (hsa_circRNA_101348 and hsa_circRNA_104361) found to be associated with inflammatory activity measured by MRI in RRMS patients (P = .0039, FC = 2.4; P = .029, FC = 1.7, respectively). Additionally, the study found that the upregulation of these 2 circRNAs was linked to the upregulation of AK2 and IKZP3 mRNA expression, which are known to be involved in B-cell function. Upon analyzing the expression results of AK2 and IKZP3, an upregulation of gene transcription was found (P = 5.65E-22, FC = 3.1; P = 1.72E-17, FC = 3.0, respectively). 84
CircRNAs as biomarkers for disability score in MS
Zurawska et al after found a circRNA biomarker that was associated with disability score. The group analyzed the downregulated circRNAs in the RRMS-relapse versus the RRMS-remission group versus HC. Two hundred forty-six unique circRNAs were downregulated in either RRMS-relapse or RRMS-remission when compared to HC (P < .05, FC < 0.5). Expression levels of 3 of the downregulated circRNAs were further analyzed using qRT-PCR. Two of which hsa_circRNA_101145 (P = .0000332, FC = 0.385) and hsa_circRNA_001896 (P = .0455, FC = 0.591) were significantly lower in RRMS-remission patients versus HC. And hsa_circRNA_101145 showed a significantly lower expression in RRMS-remission versus RRMS-relapse (P = .026, FC = 0.589). None were significantly different between RRMS-relapse and HC. Hsa_circRNA_001896 decreased levels in patients was associated with a lower disability score (P = .0008) and none had association with disease duration or inflammatory activity. 78
CircRNAs as biomarkers of immune-mediated demyelinating diseases
He et al’s study demonstrated the presence of differential expression levels of circRNAs in the CSF exosomes of patients with immune-mediated demyelination diseases compared to controls. The study identified 5095 differentially expressed circRNAs, of which 26 reached significant levels (FC ⩾ 1.5 and P ⩽ .05). Hsa_circ_0087862 and hsa_circ_0012077 were further validated in an independent cohort and found to be significantly upregulated when compared to control samples (P < .05, FC = 3.55 and 1.45, respectively). The study also discovered a potential connection between exosomal hsa_circ_0012077 expression levels and immunoglobulin G (IgG) levels in CSF. Using hsa_circ_0087862 or hsa_circ_0012077 as biomarkers for immune-mediated demyelinating diseases showed a diagnostic accuracy of 100%. 85
These findings highlight the potential of circRNAs as novel biomarkers for MS, aiding in the early diagnosis and management of these diseases.
Discussion
MS is a multifactorial autoimmune disorder of the CNS with both immunological and neurological components. 1 The disease has been associated with various immunological, environmental, genetic, and epigenetic factors.2,5,7,8 Understanding the effect of each factor can aid in comprehending the disease’s pathophysiology. This paper provides initial insights into the possible involvement of circRNAs in MS pathogenesis based on recent research and explores the possibility of circRNAs serving as biomarkers for the disease.
Upon conducting a comprehensive literature search, it was found that circRNAs hold promise not only as a factor in understanding the disease but also as potential biomarkers. Iparraguirre et al 70,74,Cardamone et al 73 reported several differentially expressed circRNAs in MS patients compared to HC, including differential expression of several circRNAs in MS subtypes. Paraboschi et al observed a significant enrichment of circRNAs in the 73-LD blocks harboring MS-associated SNPs, suggesting that MS-associated SNPs manipulate the levels of specific circRNAs in the surrounding area, thereby modifying the ratio of circular: linear RNA isoforms and contributing to disease pathogenicity. 75 Cardamone et al added that MS-associated SNPs influence the expression levels of circRNAs through exon-based DNA methylation. 77 These findings suggest a potential mechanism by which MS-associated SNPs affect circRNA expression levels and contribute to disease pathogenicity. Huang et al analyzed circRNA/miRNA/mRNA networks in MS patients using bioinformatics methods and identified 4 networks that impact B-cell proliferation and B-cell receptor signaling pathways. These findings suggest potential implications for circRNAs in MS pathphysiology. 81
CircRNAs are stable molecules that are resistant to several endonucleases, making them an intriguing biomarker material. In fact, Iparraguirre et al identified 6 circRNAs as potential MS biomarkers that are sex-dependent and 2 as minimally invasive biomarkers for highly active MS.82,83 Zurawska et al found 2 circRNA biomarkers for inflammatory activity and disability score in RRMS patients.78,84 He et al identified 2 circRNAs as biomarkers for immune-mediated demyelinating diseases with a diagnostic accuracy of 100%. 85 These findings demonstrate the potential of circRNAs as novel biomarkers for immune-mediated demyelination diseases, which may facilitate early diagnosis and management of these diseases.
Despite the increasing awareness of circRNA in recent years, these molecules remain poorly understood. Further research is necessary before any definitive conclusions can be drawn about their functions, particularly in the context of complex diseases such as MS. While the data presented in this review suggest that circRNAs may be involved in the pathogenesis of MS, the limited available information on circRNA is insufficient to establish a definitive link between this molecule and MS pathology. Therefore, a greater amount of research is required to fully elucidate the role of circRNA in MS pathogenicity.
This review aims to highlight the contribution of circRNA to the pathogenesis of MS. While a search for circRNA in MS yielded 16 review papers, none of them discussed the role of circRNA in MS as comprehensively as this review. Some of these papers barely mentioned circRNA,64,86 -92 while others focused on other types of ncRNAs such as miRNA and LncRNA.93,94 A few papers discussed the role of circRNA in other types of diseases but did not mention MS.95 -98 The lack of comprehensive discussion on the role of circRNA in MS and other diseases may be attributed to the limited understanding of this molecule. Further research is needed to fully elucidate the functions of circRNA in health and disease, particularly in the context of complex diseases such as MS.
Several limitations to this review must be noted. Firstly, the search for scientific articles related to circRNA and MS produced a mere 34 results out of which only 11 qualified as research papers, indicating a limited understanding of the role of circRNAs in MS pathogenesis. Due to our limited knowledge of circRNA, as it has only been spotted in 1976, little have attempted to understand the role of circRNA in MS and tried to focus on their role as biomarkers for the disease. Secondly, data reproducibility can affect our understanding of the role of circRNA in MS. While some studies have suggested a potential role for circRNA as biomarkers for MS, further research is needed to establish this connection more definitively. Finally, this review has focused solely on papers published in English, which may exclude other languages and perspectives.
It is worth noting that data reproducibility has been a significant drawback in this area. This could be attributed to a variety of factors, such as: (1) previous research has mainly concentrated on other types of ncRNA in MS, such as miRNA; (2) our comprehension of the function or role of circRNA has only recently become more compelling; and (3) researchers are exploring divergent directions in their investigations, which makes data reproducibility more difficult.
Prospective
CircRNA is thought to potentially play a significant role in the pathogenicity of MS, possibly mediated by MS-associated SNPs. CircRNA is highly abundant, stable, conserved, and expressed specifically, making it a promising candidate as a disease biomarker. In fact, several studies have demonstrated the accuracy of certain circRNAs as disease biomarkers. However, as our understanding of circRNA is still limited, caution should be exercised in underestimating what is still unknown. Further research is necessary to explore the full potential of this molecule and its impact on human biology and disease pathology.
Multifactorial complex diseases like MS involve multiple factors such as environmental, viral, and genetic factors that can interact with each other in complex ways. Changes or modifications in one factor can have a cascading effect on other factors. In the case of MS, it has been found that certain genetic variations or SNPs can impact the expression of circRNA molecules which in turn can affect the miRNA/mRNA networks and other biological processes. Therefore, careful consideration and analysis of multiple factors are required when interpreting information related to such complex diseases.
In recent decades, significant progress has been made in our understanding of the biology of MS. However, there is still much to be done to fully elucidate the etiology of MS, which could lead to more effective therapeutic interventions and ultimately improve the quality of life for patients.
Footnotes
Acknowledgements
My sincere gratitude extends to those who inspire me to do more.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
The author alone is responsible for the content and writing of this article.
