Abstract
Background
The TP53 gene is estimated to be mutated in over 50% of tumors, with the majority of tumors exhibiting abnormal TP53 signaling pathways. However, the exploration of TP53 mutation-related LncRNAs in Hepatocellular carcinoma (HCC) remains incomplete. This study aims to identify such LncRNAs and enhance the prognostic accuracy for Hepatoma patients.
Material and Methods
Differential gene expression was identified using the “limma” package in R. Prognosis-related LncRNAs were identified via univariate Cox regression analysis, while a prognostic model was crafted using multivariate Cox regression analysis. Survival analysis was conducted using Kaplan–Meier curves. The precision of the prognostic model was assessed through ROC analysis. Subsequently, the Tumor Immune Dysfunction and Exclusion (TIDE) algorithm were executed on the TCGA dataset via the TIDE database. Fractions of 24 types of immune cell infiltration were obtained from NCI Cancer Research Data Commons using deconvolution techniques. The protein expression levels encoded by specific genes were obtained through the TPCA database.
Results
In this research, we have identified 85 LncRNAs associated with TP53 mutations and developed a corresponding signature referred to as TP53MLncSig. Kaplan–Meier analysis revealed a lower 3-year survival rate in high-risk patients (46.9%) compared to low-risk patients (74.2%). The accuracy of the prognostic TP53MLncSig was further evaluated by calculating the area under the ROC curve. The analysis yielded a 5-year ROC score of 0.793, confirming its effectiveness. Furthermore, a higher score for TP53MLncSig was found to be associated with an increased response rate to immune checkpoint blocker (ICB) therapy (
Conclusion
The present study aimed to identify and validate LncRNAs associated with TP53 mutations. We constructed a prognostic model that can predict chemosensitivity and response to ICB therapy in HCC patients. This novel approach sheds light on the role of LncRNAs in TP53 mutation and provides valuable resources for analyzing patient prognosis and treatment selection.
Introduction
Hepatocellular carcinoma (HCC) remains a significant global health concern, accounting for a substantial proportion of cancer-related deaths worldwide. 1 Despite advances in treatment, high rates of recurrence, metastasis and chemoresistance contribute to the poor prognosis of HCC patients. 2 Further investigation into the underlying mechanisms of HCC progression is essential to facilitate improved clinical diagnosis, therapy, and prognosis assessment. Long non-coding RNAs (lncRNAs) are RNA transcripts that exceed 200 nucleotides in length. They have been found to play a significant role in regulating gene expression. 3 Previous studies have identified stable and specifically expressed lncRNAs in HCC patients, suggesting their potential as non-invasive biomarkers for early diagnosis, risk stratification, and prognosis assessment. A comprehensive literature review on this topic is presented in Supplementary table 1.
As an essential tumor suppressor gene, TP53 (encoding p53 protein) is involved in the development and progression of most malignant tumors by regulating numerous signal transduction pathways. Therefore, it has been a major focus in the field of tumor molecular biology research. 4 More than half of all tumors are estimated to have TP53 mutations, resulting in a majority of tumors demonstrating aberrant TP53 signaling pathways. 5 The P53 protein encoded by TP53 is an important transcription factor. When cells stimulated by a variety of carcinogenic factors, P53 protein will be activated to block the cell cycle and induce the inhibition of cell proliferation, cell differentiation, aging and the cell apoptosis, involve in DNA damage repair, maintaining genomic stability, regulate energy metabolism, and inhibiting tumor angiogenesis, thus preventing the occurrence of tumor development. 6
Previous literature has reported that TP53 mutation is closely related to microvascular invasion, 7 tumor progression, early recurrence, and low survival time of HCC.8–10 Moreover, it has been reported that TP53 mutant status can affect tumor response to immunotherapy. 11 Therefore, it is important to probe into the role of TP53 in the etiopathogenesis of HCC. There are literatures points out that the unmutated TP53 binds to RAD51, an important role in DNA replication and the maintenance of genomic stability, in vivo and in vitro.12,13
Numerous prior studies have reported the crucial role of lncRNAs in maintaining genome stability. For instance, Mendell et al. demonstrated NORAD activation in response to DNA damage to assist with replication and repair. 14 Some other lncRNAs such as CUPID2 and DDSR1 will also produce similar functions.15,16 However, the association of lncRNAs in TP53 gene mutation and the clinical significance of these lncRNAs have not been thoroughly explored. In this study, we aimed to combine somatic mutation data and high-throughput sequencing data of HCC patients to investigate lncRNA signatures associated with TP53 mutations. Our goal was to explore the potential of these signatures in predicting prognosis and guiding treatment decisions.
Material and methods
Sources
The clinical information of the training group (

aChi square test.
Identification of TP53 mutation-associated lncRNAs
We utilized the “limma” R package to differentiate TP53 mutant and TP53 wild groups and isolate lncRNAs with differential expression between the two groups in the TCGA database. These lncRNAs satisfied our screening criteria of FDR <0.05 and |log2 FC| >1. 24 These lncRNAs were deemed TP53 mutation-related, and a heatmap depicting the top 30 lncRNAs was generated using the “Pheatmap” R package, sorted by |log2 FC| in descending order.
Statistical analysis
We applied the product-limit method and logrank test to analyze the survival data of patients, assessing the relationship between overall survival (OS) and TP53 mutation-related lncRNAs through Univariate and multivariate Cox proportional hazard regression analysis.25–30 By employing multivariate regression analysis, we established a prognosis signature (TP53MLncSig) based on TP53 mutation-derived lncRNAs, using the specific formula: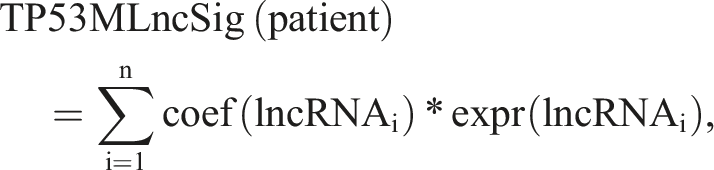
TP53MLncSig represents the risk score of HCC patients, which we determined using the median value as the cutoff. Patients whose risk score is higher than the cutoff are classified as the high-risk group, while those with a lower score are considered as the low-risk group. We evaluated the ability of TP53MLncSig to distinguish patients with poor outcomes by generating an ROC curve. For all analyses mentioned above, we considered
Predicting the biological processes of TP53 mutation-related LncRNAs
To evaluate the correlation strength between TP53 mutation-related lncRNAs and mRNAs, we utilized Pearson correlation coefficients. A Pearson correlation coefficient of greater than 0.5 and an FDR of less than 0.001 indicated co-expression between the mRNA and lncRNA. 35 To forecast potential biological functions of TP53 mutation-related lncRNAs, we employed the “clusterProfiler” R software package to identify the KEGG pathway and GO biological processes associated with co-expressed mRNAs.24,36 The resulting pathways were considered as the underlying biological processes of TP53 mutation-related lncRNAs. To determine the level of immune cell infiltration in each sample and compare infiltration differences between the low and high-risk score groups, we utilized the ssGSEA algorithm. 36
Unraveling the intimate correlation between TP53MLncSig and the activation of TP53-related pathways, alongside P53 protein expression levels
We employed The Cancer Proteome Atlas (TCPA) tool to investigate the expression level of P53 protein, encoded by TP53, in each sample of TCGA-LIHC cohort. 37 TCPA employs the reverse phase protein arrays (RPPA) technology to assess the selected protein levels in the samples efficiently, sensitively, and at high-throughput.
Identification of differentially expressed genes among distinct risk groups using the LIMMA R package, with a screening threshold of fold change >1.5 or <2/3 and FDR <0.05. These differential genes were input into the genetic perturbation similarity analysis (GPSA) tool using its enrichGPSA plugin. 38 The GPSA database comprises 6096 gene sets from 3048 RNA-seq datasets of gene perturbations, which include siRNA, shRNA, CRISPR/Cas9, and other methods. The enrichGPSA plugin explores which genes or proteins have a gene expression pattern similar to that of the input file after being knocked down.
We conducted functional enrichment analysis of gene sets with the Metascape database. For gene set enrichment analysis (GSEA), we used version 3.0 of the GSEA software from their Web site. The gene set limits were set to a minimum of 5 and a maximum of 5000, with 1000 resamplings, and any FDR threshold below 0.05 was considered significant. 39
The Molecular Signatures Database was used to search for P53 protein's function-related pathways, and their files were downloaded in the GMT format. 40 In addition, 22 cancer-related biological processes were obtained from Sanjeev Mariathasan et al.’s study, including but not limited to epithelial-mesenchymal transition (EMT) 1, 2, and 3, angiogenesis, antigen processing and presentation, antigen processing machinery, CD8 T effector, immune checkpoint, base excision repair, cell cycle, DNA damage repair, DNA replication, mismatch repair, extracellular matrix-receptor interaction, JAK-STAT signaling pathway, MAPK signaling pathway, NF-kappa B signaling pathway, nucleotide excision repair, Pan-F-TBRS, PI3K-Akt signaling pathway, TGF-beta signaling pathway, and Wnt signaling pathway. 41 The ssGSEA technique was applied to calculate the enrichment score of different pathways in order to assess their activation statuses.
Forecasting chemotherapy sensitivity
Maeser et al. have introduced the OncoPredict R package, designed to predict in vivo drug responses in cancer patients.
42
OncoPredict fits tissue gene expression profiles to half-maximal inhibitory concentration (IC50) of cancer cell line drugs sourced from the Genomics of Drug Sensitivity in Cancer (GDSC; https://www.cancerrxgene.org/) database and gene expression profiles from the Broad Institute Cancer Cell Line Encyclopedia (CCLE; https://portals.broadinstitute.org/ccle_legacy/home).43,44 We evaluated a total of 198 drugs and employed Wilcox tests to analyze drug sensitivities between high-risk and low-risk groups. Significance was based on a
Establishment of the nomogram
Nomogram Development with Tumor Staging and TP53MLncSig. To further improve clinical convenience, we constructed a nomogram using tumor staging and TP53MLncSig. We constructed the nomogram using the R software package rms. 45 Calibration curves were used to estimate the accuracy of the nomogram.
Real-time PCR (RT-PCR)
Total RNA was extracted from tissues and cells using TRIzol reagent (Invitrogen, CA, USA) and dissolved in RNase-free water (Sango Biotech, Shanghai, China). Next, the isolated total RNA was reverse-transcribed into complementary DNA with the HiFi Script cDNA Synthesis Kit (CWBIO, Beijing, China). Real-time PCR was performed using the SYBR Green I kit (Sango Biotech, Shanghai, China) according to the manufacturer’s protocols. Experiments were conducted in triplicate. The primers employed for RT-PCR were as follows: Forward Primer of PRRT3-AS1: 5’-TCAACAACGCCCCTCTGAAAG-3’; Reverse Primer of PRRT3-AS1: 5’- AGTGTGAGGCCGTATGAATGG-3’; Forward Primer of MIR210HG: 5’-GCTTGGTAGAGTGTCACGCC-3’; Reverse Primer of MIR210HG: 5’-CATCTGACCGAGCCAGTTTG-3’; Forward Primer of LINC00942: 5’-CCTGGATTGTGGGCCTTGAA-3’; Reverse Primer of LINC00942: 5’-TACTGTTTCCCGCAGACACC-3’; Forward Primer of AC099850.4: 5’-GGATTGCTTGAGCCTAGGAG-3’; Reverse Primer of AC099850.4: 5’-TTGCGGTAGTGGTTACATGA-3’; Forward Primer of MAFA-AS1: 5’-GGAGGGGAAGGAGAGCTACA-3’; Reverse Primer of MAFA-AS1: 5’-TGACTGCTTCACAGAACGCT-3’; Forward Primer of AC011297.1: 5’-GCCTGAAGTGGTTGTGTGAAG-3’; Reverse Primer of AC011297.1: 5’-GGCCTCCCAGACCTTGAAAT-3’; Forward Primer of AC105118.1: 5’-TTGGCAGAGTGGAGCAGATG-3’; Reverse Primer of AC105118.1: 5’-GCTGCCGTTGTTACCGTTTT-3’; Forward Primer of GAPDH: 5’-CCACATCGCTCAGACACCAT-3’; Reverse Primer of GAPDH: 5’-ACCAGGCGCCCAATACG-3’.
Four HCC patients were recruited from Anqing First People's Hospital affiliated with Anhui Medical University, and informed consent forms were obtained. HCC tumor tissue samples and adjacent normal liver tissue samples were collected from postoperative specimens of the HCC patients.
Results
TP53 mutation-related lncrnas in HCC patients
We conducted a comparative analysis of TP53 mutation-related lncRNAs in HCC patients using the TCGA dataset. We performed calculations using the “limma” R package by dropping the samples with TP53 mutation ( Heatmap of the dysregulated lncRNAs expression in tissues with TP53 mutation and those without TP53 mutation was analyzed using the TCGA datasets (a). Boxplots of somatic mutations in tissues with TP53 mutation (TP53-mut group) and tissues without TP53 mutation (TP53-wild group). Somatic cumulative mutations in the TP53-mut group are significantly higher than those in the TP53-wild group (b). Boxplots of RAD51 expression level in the TP53-mut group and TP53-wild group. The expression level of RAD51 in the TP53-mut group is significantly higher than that in the TP53-wild group (c). Horizontal lines means median values. Above the Boxplots is the 
Functional enrichment analysis of TP53 mutation-related lncrnas
A total of 2307 mrnas co-expressed with TP53 mutation-related lncrnas were identified (Supplementary table 3), and functional enrichment analysis was conducted on these mrnas (Supplementary table 4). Figure 2 displayed the first 20 terms with the smallest Functional enrichment analysis of GO and KEGG for the mRNAs co-expressed with the TP53 mutation-related lncRNAs.
Establishment of a TP53 mutation-derived lncRNA signature for prognostication of HCC Patients
To investigate the impact of TP53 mutation-derived lncRNAs on patient prognosis, we classified 369 HCC patients into two groups with no significant differences in clinical characteristics: the training group ( TP53MLncSig divided the patients in the training group into two groups: low or high risk, and Kaplan–Meier estimates of OS of the two groups (a). Univariate Cox analysis was used for statistical analysis. Time-dependent ROC curves analysis of the TP53MLncSig at 5 years (b). The expression level of the lncRNAs in TP53MLncSig and RAD51 gene changed with the increase of TP53MLncSig score in the training group (c). Forest plots based on univariate (d) and multivariate (e) survival analysis in patients with High TP53MLncSig risk score or low TP53MLncSig risk score. Distribution of RAD51 expression level in the high- and low-risk groups for training group patients. The red represents the high-risk group, and the blue represents the low-risk group for training group patients (f). Horizontal lines mean median values. Above the Boxplots is the 
Independent validation of TP53MLncSig
The current study aimed to verify the robustness of TP53MLncSig, a prognostic model for HCC, in an independent test set group ( TP53MLncSig divided the patients in the test group (a) and TCGA group (b) into two groups: low or high risk, and Kaplan–Meier estimates of OS of the two groups. Univariate Cox analysis was used for statistical analysis. The expression level of the lncRNAs in TP53MLncSig and RAD51 gene changed with the increase of TP53MLncSig score in the test group (c) and TCGA group (d). Distribution of RAD51 expression level in the high- and low-risk groups for the test group (e) and TCGA group (f) patients. The red represents the high-risk group, and the blue represents the low-risk group for training group patients. Horizontal lines mean median values. Above the Boxplots is the 
Combined prediction of prognosis using TP53MLncSig and TP53 mutation status in HCC patients
After excluding patients without somatic mutation data, we calculated the proportion of patients with TP53 gene mutation in the training group, validation group, and TCGA group, respectively, and the results are shown in Figure 5(a). In the training set, 43 patients (49%) in the high-risk group had mutations in TP53 and 45 patients (51%) did not, while 16 patients (18%) in the low-risk group had mutations in TP53 and 74 patients (72%) did not. The percentage of patients with TP53 mutations in the high-risk group was significantly higher than that in the low-risk group (chi-square test Relationship between the TP53MLncSig and TP53 somatic mutation. The proportion of TP53 mutation in high- and low-risk groups in the training set, testing set and the TCGA set (a). Kaplan–Meier curve estimates of OS is shown for TCGA group classified according to TP53 mutation status and the TP53MLncSig (b). The log-rank test was utilized for statistical analysis.
The TP53 mutant and TP53 wild groups were stratified into high-risk and low-risk groups based on TP53MLncSig scores. Our findings demonstrated worse patient prognosis in the high-risk group compared to the low-risk group, irrespective of TP53 mutation status (see Figure 5(b)). On the one hand, it showed that TP53MLncSig is robust in predicting prognosis, and it is not affected by the TP53 mutation status of patients. On the other hand, TP53MLncSig distinguished patients with worse prognosis in both TP53 mutation group and the TP53 wild group, suggesting that TP53MLncSig is more predictive of prognosis than the TP53 mutation status to some extent. In contrast, the TP53 mutation status of was able to distinguish the patients with poorer prognosis of HCC in the high-risk group, but it did not play a similar role in the low-risk group. This suggests that TP53 mutation status is not as robust as TP53MLncSig in prognostic prediction. Patients were categorized into four distinct groups based on TP53 mutation status and TP53MLncSig scores, as follows: TP53 mutated and high-risk group (
Validation of TP53MLncSig in independent HCC dataset
To assess the potential association between lncRNAs in TP53MLncSig and TP53 mutation status in HCC patients, we conducted a validation study using an independent dataset, LICA-FR, retrieved from the ICGC database. This dataset comprised 109 HCC patients with known TP53 mutation status, and only five lncRNAs from the TP53MLncSig panel (LINC00942, MAFIA-AS1, MIR210HG, PRRT3-AS1, and AC105118.1) were included in the validation set. Our analysis uncovered significantly elevated RAD51 expression levels in HCC patients with TP53 mutations, in comparison to those with wild-type TP53 ( Boxplots for RAD51 (a), MIR210HG (b), PRRT3-AS1 (c), MAFIA-AS1 (d), AC105118.1 (e) and LINC00942 (f) expression between patients with and without TP53 mutation in the LICA-FR of ICGC data sets. Above the Boxplots is the 
Evaluation of immune-checkpoint blockade (ICB) response
In this study, the TCGA dataset was stratified into high-risk and low-risk groups using TP53MLncSig and the median TP53MLncSig risk score in the training group. Figure 7(a) illustrates the variation in the fractions of 23 different immune cell types observed between the high-risk and low-risk groups, where Activated CD4 T cells and Type 2 T helper cells were predominantly more infiltrated in the high-risk group, whereas Activated B cells, Activated CD8 T cells, Type 1 T helper cells, Eosinophils, and Mast cells showed more infiltration in the low-risk group. To assess the enrichment scores of biological processes related to DNA damage repair in the high-risk and low-risk groups, the ssGSEA algorithm was utilized, and it was found that the scores were higher in the high-risk group. Additionally, the biological processes of angiogenesis scored higher in the low-risk group (Figure 7(b)). In order to examine whether patients in the high-risk and low-risk groups differed in their response to the immune checkpoint blockade (ICB) treatment, we used an online tool created by Peng Jiang et al. (http://tide.dfci.harvard.edu/) to investigate the response rate between the two groups (Supplementary table 7). We calculated the dysfunction score of tumor infiltration cytotoxic T lymphocyte (CTL) and exclusion of CTL using the tool and also determined the TIDE score of each patient. In the high-risk group, the tumor infiltration CTL dysfunction score was notably lower than that observed in the low-risk group ( Boxplots for the infiltration rate of 24 immune cells between high-risk (blue) and low-risk tumor tissues (red).*:
Intimate correlation between TP53MLncSig and the activation of TP53-related pathways, alongside P53 protein expression levels
The TCPA database indicated that protein expression levels of P53 were lower in the high-risk group (Supplementary Figure 2(a)). Furthermore, a significant negative correlation was observed between the risk scores obtained from TP53MLncSig and protein expression levels of P53 (Supplementary Figure 2(b)). These results are consistent with our earlier findings that the proportion of TP53 mutant patients was higher in the high-risk group compared to those with wild-type TP53 (Figure 5(a)). The occurrence of tumors involves mutually regulated mechanisms between RB and P53, both of which possess tumor suppressor characteristics and are functionally interdependent. Accordingly, we further explored the protein expression levels of RB in different risk score groups (Supplementary Figure 2(c)). Similar to P53, RB was observed to be downregulated in the high-risk score group, while a significant negative correlation was observed between the risk score and RB protein expression levels (Supplementary Figure 2(d)). These results further validate the close association of TP53MLncSig with TP53 mutations and P53 protein expression. Moreover, we found that more than half of the P53 protein-associated biological pathways exhibited differential activation in different risk groups (Supplementary Table 8).
The activation of TP53MLncSig-associated biological pathways
By performing the limma analysis, we identified 778 downregulated and 3247 upregulated genes in the high-risk score group (Supplementary Figure 3(a)). The top 50 upregulated and downregulated genes were depicted using a heatmap visualization (Supplementary Figure 3(b)). A total of 613 gene perturbation sets were enriched through the GPSA analysis, with the top 10 having the smallest adjusted
The ssGSEA algorithm was applied to appraise the activation levels of 22 tumor-related biological processes among high and low-risk groups. Remarkably, DNA damage repair and cell cycle-related pathways were significantly up-regulated in the high-risk group (Supplementary Figure 4(a); Supplementary table 11). This finding implies that the high-risk group experiences a higher degree of genomic instability, which may be associated with a higher level of TP53 mutations. To validate the aforementioned results, we conducted GSEA enrichment analysis on the DNA damage repair and cell cycle-related pathways in the KEGG database. Remarkably, all of these pathways were significantly up-regulated in the high-risk group (Supplementary Figure 4(b)).
The potential of TP53MLncSig as a guidance tool for treatment in HCC patients.
Assessing chemotherapy sensitivity for various drugs amongst patients in the TCGA-LIHC cohort using OncoPredict algorithm (Supplementary table 12). Notably, significant variations were observed in the sensitivity of commonly used chemotherapeutic and targeted drugs between the high-risk and low-risk groups, which include Gemcitabine, 5-Fluorouracil, Cisplatin, Docetaxel, Oxaliplatin, Erlotinib, and Sorafenib (Supplementary Figure 4(c)). In addition to its potential in predicting clinical outcomes, our TP53MLncSig also has the capacity to predict chemotherapy sensitivity in patients with HCC.
Establishment of the nomogram
We constructed a nomogram for clinical use by combining TP53MLncSig and tumor stage (Supplementary Figure 5(a)). Survival curves showed that HCC patients with higher scores in the nomogram had poorer prognosis and shorter median survival time (3.15 years vs 6.88 years; Supplementary Figure 5(b)). The calibration curve indicated a relatively high accuracy of the nomogram for predicting the 3-year survival rate of HCC patients (Supplementary Figure 5(b)).
Expression levels of TP53 mutation-associated LncRNAs between HCC tissues and normal hepatic tissues
We investigated the expression levels of TP53 Mutation-Associated LncRNAs in both HCC and normal liver tissues within the TCGA-LIHC cohort. With the exception of AC011297.1, the remaining six TP53 Mutation-Associated LncRNAs exhibited notably elevated expression levels in HCC tumor tissues (Supplementary Figure 6(a)). We further validated the expression levels of these LncRNAs using RT-PCR. The RT-PCR results revealed that among these LncRNAs, PRRT3-AS1, MIR210HG, LINC00942, AC099850.4, and MAFA-AS1 still exhibited significantly higher expression levels in HCC tumor samples compared to the adjacent normal tissues (Supplementary Figure 6(b)).
Discussion
The investigation of cancer occurrence, development, and prognosis has always been a research hotspo.1,46,47 In the diagnosis and treatment of HCC patients, traditional pathological features such as tumor size, regional lymph nodes with tumor metastasis, metastasis stage, and grade are still utilized as important prognostic indicators.48,49 However, due to the heterogeneity of HCC patients, these traditional pathological features may not effectively serve as prognostic indicators. TP53 gene mutation occurs in more than half of all tumors, and the majority of tumors have abnormalities in the TP53 signaling pathway.1,50 As a significant tumor suppressor gene, TP53 mutation plays a crucial role in tumor occurrence and developmen.51,52 In addition, patients with HCC and TP53 mutations have a shorter overall survival and a higher recurrence rate.9,10
Recent research suggests that lncRNAs play a crucial role in regulating gene expression levels in the form of RNA, and are essential regulators in tumor progression, with some serving as feasible prognostic markers for patients.53,54 In addition, lncRNAs are considered important for maintaining genome stability. For instance, NORAD is induced by DNA damage in a p53-dependent manner, and knocking out NORAD significantly reduces the expression of genes required to maintain genome stability. 14 Although numerous studies are focusing on the role of lncRNA in gene mutation, how lncRNAs are related to TP53 mutation in HCC remains in its infancy. 55 Some existing literature has begun to highlight the role of lncRNAs in regulating p53 protein function, thus impacting the proliferation and migration of HCC cells. The expression of lncRNA SNHG1 is elevated in HCC tumors compared to adjacent tissues. Furthermore, knockdown of SNHG1 suppresses the proliferation and invasive capabilities of liver cancer cell lines (SMMC-7721 and SK-HEP-10). 56 RIP and ChIP results indicate that SNHG1 can interact with DNMT1 and inhibit the expression of p53. 56 Juan et al. revealed that downregulation of lncRNA SAMD12-AS1 significantly enhances the expression of p21, p53, and E-cadherin, thereby promoting the occurrence, migration, and invasion of liver cancer. 57 Moreover, previous literature has reported that the overexpression of PLAC2 leads to the upregulation of p53, thereby enhancing apoptosis in cancer cells. 58 Additionally, it has been documented that the expression of Nrf2-lncRNA is downregulated during the initiation of apoptosis, functioning as a regulatory element in determining cell fate. 59 Hence, the interrelation between lncRNAs, p53 protein function, and the fate of HCC cells is closely intertwined. However, there remains a paucity of comprehensive analysis and synthesis of research pertaining to TP53-associated lncRNAs in the context of liver cancer.
In this study, we identified 85 TP53-mutation-related lncRNAs by combining the expression level of lncRNAs and TP53 mutation status of HCC patients in the TCGA database. We performed KEGG and GO enrichment analysis on mRNAs co-expressed with these 85 lncRNAs, and found that the enriched biological processes were highly relevant to the function of TP53 gene and associated with genomic instability.60–62
Furthermore, we constructed a TP53MLncSig containing seven lncRNAs to investigate the relationship between TP53-mutation-related lncRNAs and patient prognosis. Patients in the training set were divided into a high-risk group and a low-risk group by TP53MLncSig, and better outcomes were seen in the low-risk group. Such results were validated in an independent test set, where patients in the high-risk group had a higher proportion of TP53 mutations, and the expression of RAD51 gene also increased with the increase of TP53MLncSig score. The variation trend of RAD51 gene with TP53MLncSig indirectly indicated that TP53MLncSig was strongly correlated with TP53 mutation status.13,14 Moreover, we performed multidimensional validation, including GPSA and ssGSEA analyses, to confirm the close association between the identified TP53MLncSig and TP53 gene, which further supports the rationality and accuracy of TP53MLncSig. Moreover, it is worth emphasizing that TP53MLncSig is not only associated with TP53 mutations but also closely related to the expression level of P53 protein. This may be due to the higher level of TP53 mutations in patients in the high-risk group, resulting in reduced protein production. The deficiency of P53 not only affects cholesterol esterification, exacerbating hepatocarcinogenesis, but also leads to the progression of HCC. Therefore, the difference in P53 protein levels between the high- and low-risk groups may form the basis of TP53MLncSig's prognostic capabilities.
We further verified the relationship between lncRNAs in TP53MLncSig and TP53 mutation status in an ICGC dataset independent of the TCGA database, where only five of the seven lncRNAs in TP53MLncSig could be found. Notably, LINC00942, MIR210HG, and PRRT3-AS1 were significantly overexpressed in patients with TP53 mutations, whereas MAFIA-AS1 and AC105118.1 also exhibited the same trend of difference, although their
TP53MLncSig has been shown to serve as a valuable tool for detecting TP53 mutation status and predicting prognosis in patients with HCC, irrespective of whether they have TP53 mutation or not. Patients with higher TP53MLncSig scores were associated with poorer outcomes, suggesting that the predictive power of TP53MLncSig for prognosis is not compromised by the presence or absence of TP53 mutation. Conversely, TP53 mutant status was found to be an inadequate prognostic indicator in patients with low TP53MLncSig scores. These results indicate that TP53MLncSig outperforms TP53 mutant status as a prognosis indicator for HCC.
Furthermore, our study uncovered the significant impact of TP53MLncSig on immune cell infiltration in the tumor microenvironment. Specifically, we found that the high-risk group had significantly more infiltrated CD8 naive, NTREG, ITREG, DC, and B cells than the low-risk group, while the low-risk group had significantly more infiltrated CD4 naive, TC, TFH, MAIT, NK, and CD4T cells than the high-risk group. Prior studies have established that the tumor microenvironment plays a crucial role in determining the response to ICB therapy, and that TP53 mutation may serve as a guide for immunotherapy. 11 Our research suggests that TP53MLncSig, which is constructed using TP53mutation-related lncRNAs, may have a similar predictive effect on patients’ response to ICB treatment. Using the TIDE predictive framework, we confirmed that patients with high TP53MLncSig scores exhibited a greater degree of T cell dysfunction and higher response rates to ICB therapy. 23 Thus, TP53MLncSig not only identifies patients with a poor prognosis, but also those who may respond better to ICB treatment, providing valuable guidance for immunotherapy in patients with HCC. 11 In addition to ICB treatment, TP53MLncSig can also predict drug sensitivity to some common chemotherapeutic and targeted therapies in clinical settings. This expands the potential of TP53MLncSig in clinical applications. It should be noted that tumor staging remains the main prognostic indicator for HCC in current clinical practice. To further improve the predictive ability of TP53MLncSig for prognosis, we combined it with tumor staging to construct a nomogram for predicting the survival of HCC patients at 1, 3, and 5 years, which further facilitates clinical application.
While the present study on TP53 mutation status offers valuable insight into predicting patient prognosis and response to treatment, it is not without limitations. To ensure the robustness of the findings, further validation is needed using larger data sets. Furthermore, the study relied on high-throughput sequencing data, and additional functional verification is necessary to elucidate the regulatory mechanisms of the lncRNAs and TP53 mutation involved.
Conclusion
We employed high-throughput sequencing data to identify lncRNAs that are associated with TP53 mutation status in patients with HCC, and integrated this information with clinical data from the TCGA database to develop a signature comprised of TP53 mutation-derived lncRNAs (TP53MLncSig). The expression level of P53 protein is closely associated with TP53MLncSig, and patients in the high-risk group exhibit higher levels of TP53 mutations and genomic instability. The utility of TP53MLncSig as a prognostic indicator for patients with HCC was demonstrated in an independent patient cohort, while TP53MLncSig is also a valuable tool for guiding patient selection and combination of ICB therapy and chemotherapy. Furthermore, we also constructed a nomogram for predicting the prognosis of HCC patients at 1, 3, and 5 years.
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Supplemental Material
Supplemental Material - Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma
Supplemental Material for Identification and development of TP53 mutation-associated Long non-coding RNAs signature for optimized prognosis assessment and treatment selection in hepatocellular carcinoma by Chenghao Chu, Daoli Liu, Duofa Wang, Shuangjiu Hu, and Yongwei Zhang in International Journal of Immunopathology and Pharmacology
Footnotes
Author Contributions
Conception: Chenghao Chu and Yongwei Zhang; Interpretation or analysis of data: Daoli Liu and Shuangjiu Hu; Preparation of the manuscript: Chenghao Chua, Daoli Liua,and Duofa Wang; Supervision: Shuangjiu Hu and Yongwei Zhang. Conceptualization: Chenghao Chu and Daoli Liu; Formal analysis: Chenghao Chu and Yongwei Zhang; Methodology: Daoli Liu, Duofa Wang and Shuangjiu Hu; Writing – original draft: Chenghao Chu; Writing – review & editing: Yongwei Zhang, Duofa Wang and Shuangjiu Hu.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study were sponsored and funded by the Key Projects of Natural Science Research in Anhui Universities (Item no, KJ2021A0344) and Anqing Science and Technology Bureau science plan project fund (Item no, 2022Z3001).
Ethical statement
Data Availability Statement
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
