Abstract
Pearl millet (Pennisetum glaucum (L.)) is a remarkable cereal crop known for its ability to thrive in challenging environmental conditions. Despite its resilience, the intricate molecular mechanisms behind its toughness remain a mystery. To address this knowledge gap, we conducted advanced next-generation RNA sequencing. This approach allowed us to compare the gene expression profiles of pearl millet seedlings exposed to heat stress with those grown under standard conditions. Our main focus was on the shoots of 13-day-old pearl millet plants, which we subjected to a brief heat stress episode at 50°C for 60 seconds. Within the vast genomic landscape comprising 36 041 genes, we successfully identified a set of 10 genes that exhibited significant fold changes, ranging from 11 to 14-fold compared to the control conditions. These 10 genes were previously unknown to have such substantial changes in expression compared to the control. To uncover the functional significance hidden within these transcriptomic findings, we utilized computational tools such as MEME, String, and phylogenetic tree analysis. These efforts collectively revealed conserved domains within the transcriptomic landscape, hinting at potential functions associated with these genetic sequences. Of particular note, the distinct transcriptomic patterns specific to pearl millet leaves under thermal stress shed light on intricate connections to fundamental biological processes. These processes included the Ethylene-activated signaling pathway, Regulation of intracellular signal transduction, Negative regulation of signal transduction, Protein autophosphorylation, and Intracellular signal transduction. Together, these processes provide insight into the molecular strategies employed by pearl millet to overcome thermal stress challenges. By integrating cutting-edge RNA sequencing techniques and computational analyses, we have embarked on unraveling the genetic components and pathways that empower pearl millet’s resilience in the face of adversity. This newfound understanding has the potential to not only advance our knowledge of plant stress responses but also contribute to enhancing crop resilience in challenging environmental conditions.
Keywords
Introduction
Agriculture has had a rocky past because of the wide range of climates around the world. Particularly vulnerable to abiotic factors such as salinity, drought, heat, and cold are crop yields in arid and semi-arid environments. Because of its exceptional resistance, pearl millet (Pennisetum glaucum) stands out as an important grain crop, especially in these severe environments. Its importance in halting global warming and guaranteeing food supply cannot be overestimated.1 -4
In their natural environments, plants face a number of threats that can have devastating effects on their growth, development, and crop yields, ultimately leading to food shortages and economic losses. However, plants have been endowed by nature with complex mechanisms to endure and adapt to harsh conditions. This article focuses on heat tolerance as examples of abiotic stresses that this study investigates. Growing crops that can withstand climate change is becoming increasingly important around the world. For this purpose, pearl millet has much promise because of its potential to provide sustainable agriculture and food security despite changes in weather patterns.
About 90 million people living in poverty rely on pearl millet as their primary source of nutrition, making it the world’s sixth most essential cereal. It extends over 30 000 000 hectares of farmland in the dry and semi-dry tropics of Asia and Africa. The Committee on Agriculture (COAG) of the Food and Agriculture Organization of the United Nations (FAO) has declared 2023 the “International Year of Millets.” 4 Sustainable agriculture in arid and semi-arid countries could benefit greatly from more study in this field, which could lead to the development of improved pearl millet varieties with increased stress tolerance. A previous study on pearl millet examined the trnL-F region of chloroplast DNA to explore the genetic diversity among 6 native pearl millet varieties in Saudi Arabia. The findings revealed variations among these varieties, shedding light on the adaptation of these plants to the harsh climate conditions of the Arabian Peninsula.5,6 Increased crop resilience and food security can be achieved through better understanding the processes of stress tolerance in plants. The resilience of plants in the face of abiotic stress is a testament to nature’s flexibility. Adapting the plant’s hardiness will help us meet the problems of feeding the world’s growing population in the future. 7 Numerous studies have delved into the molecular mechanisms governing stress tolerance in pearl millet. For instance, Yan et al 8 research delved into the roles of RWP-RK genes under thermal stress at 45°C in pearl millet. Their examination of pearl millet leaves and roots’ transcriptomes revealed that key parameters such as relative water content, relative electrical conductivity (REC), and malondialdehyde (MDA) content remained largely stable (P > .05) until the 21st day of heat treatment. In a similar vein, a separate investigation led by Jin et al in their study 9 highlighted how the RWP-RK gene family could bolster the heat adaptability of elephant grass by improving nitrogen use efficiency and enhancing peroxidase gene expression. Meanwhile, Sun et al 10 study subjected pearl millet variety Tifleaf 3 to a rigorous regimen of 14 hours of light at 40°C and 10 hours of darkness, resulting in the identification of 1881 differentially expressed genes that were common to both heat and drought conditions.
Lastly, another study, conducted by Awan et al, 11 demonstrated that the application of exogenous melatonin, through both soil application and foliar treatment, ameliorated the toxicity of cadmium (Cd). This treatment not only enhanced plant growth but also bolstered the antioxidant defense system. The exogenous melatonin exhibited distinct regulatory effects on the expression of antioxidant-responsive genes, including superoxide dismutase SOD-[Fe] 2, Fe-superoxide dismutase, Peroxiredoxin 2C, and L-ascorbate peroxidase-6, leading to significant increases in plant height, chlorophyll content, and carotenoid levels when compared to plants exposed to Cd treatment alone.
In our current study, our objective was to unravel the intricate molecular responses of pearl millet to environmental stresses, particularly elevated temperatures, which have significant implications for its survival and adaptability. Specifically, we focused on pearl millet (Pennisetum glaucum) and examined the top 10 thermally expressed RNA sequences that exhibited fold changes ranging from 11 to 14 compared to the control. We subjected these sequences to in silico analysis.
In pursuit of this goal, we meticulously analyzed a comprehensive transcriptomic dataset of pearl millet under thermal stress conditions. The key outcome of this endeavor was the discovery of conserved domains within the proteins that were highly expressed under thermal stress in pearl millet. These conserved domains, identified through the use of bioinformatics tools, were found to collectively belong to the amino acid transporter family. The involvement of this family suggests a crucial role in the cellular response to thermal stress, potentially related to the maintenance of metabolic balance or the facilitation of cellular adaptation.
Material and Methods
Plant materials
Seeds of Pearl millet [Pennisetum] glaucum subsp. monodii, cultivar (Pennisetum glaucum (L.) R. Br. ssp. monodii (Maire) Brunken), were collected from Naif Bakheet almutairi, the farm owner located in Almazre, Alhamanah, Mahd Adh Dhahab Governorate, Madinah Province, Saudi Arabia (23.026438 N 39.896101 E) [link to Google Maps: https://goo.gl/maps/uSXj5YqJmf3pcMg69].
Pearl millet (Pennisetum) glaucum subsp. monodii seeds were used in this study. They were given the registration number 1319 at the Centre of Genetic Resource in the Ministry of Environment, Water, and Agriculture in Saudi Arabia. Under the terms of the International Treaty on Plant Genetic Resources for Food and Agriculture, both Saudi Arabian researchers and agricultural producers have access to these seeds.
Before the seeds were placed on the moist sheets to germinate, they were first washed with 5% sodium hypochlorite and then with distilled water. After 48 hours, the seedlings were transplanted into a medium that contained equal parts vermiculite and soil in a ratio of 1 to 1. In a conservatory with a temperature of 25°C, each pot measuring 6 in by 17 in. Twenty milliliters of distilled water was used to irrigate each pot (6 × 17 in) in a conservatory at 25°C. Pearl millet [Pennisetum] glaucum subsp. monodii, cultivar (Pennisetum glaucum (L.) R. Br. ssp. monodii (Maire) Brunken), after 10 days of growth samples were divided into 2 groups:
Control group (no treatment).
Heat stress group: treated with 50°C for 60 seconds at the age of 13 days.
All samples were collected from 13-day-old seedlings following treatment, immersed in RNAlater Stabilization Solution derived from the Lader 12 ; Lader 13 method, and stored at 4°C. The RNA of stems was isolated using the Trizol method developed by Chomczynski and Mackey. 14 The experimental design included a control group of 14 seedlings grown under normal conditions and abiotic stress group of 14 seedlings subjected to heat stress.
Transcriptome sequencing and data analysis
Library protocol
The quality of genomic DNA and double-strand cDNA was assessed using Qubit dsDNA estimation and agarose gel electrophoresis. For each sample, 500 ng of genomic DNA and 200 ng of cDNA were combined and used for library preparation. The libraries for all samples (3 per group) were created using the NEBNext UltraII DNA library preparation kit for Illumina (Cat no: E7770, New England Biolabs) following the manufacturer’s recommended protocol. Subsequently, sequencing was performed on the S4 flowcell of NOVASEQ 6000, utilizing 150 bp paired-end chemistry. Both the library preparation and sequencing processes were outsourced to commercial service providers, specifically NGB Diagnostics located in Noida, India.
The DNA samples were shared with an insert size of approximately 300-400 bp for each sample. The fragmented genomic DNA was then subjected to end-repair and A-tailing, followed by the ligation of indexed adapters. After purification and PCR enrichment, the final genomic DNA library was obtained. To quantify the libraries, a Qubit Fluorometer (Invitrogen, Life Technologies, Grand Island, NY, USA) was used. Additionally, the distribution of library fragments was examined using the HSDNA kit on Tapestation (Agilent Technologies, USA). The tagged genomic DNA libraries were pooled in equal ratios and loaded onto the c-bot automated system for cluster generation. Post-cluster generation, the libraries were loaded onto the Illumina S4 Flow Cell of the Illumina Novaseq 6000 Sequencing system, and sequencing was conducted using 2 × 150 bp paired-end chemistry. After sequencing, the samples were demultiplexed, and the indexed adapter sequences were trimmed using the CASAVA v1.8.2 software from Illumina Inc.
Read quality check and de novo assembly
To assess the quality of the reads, FastQC version 0.11.1 (www.bioinformatics.babraham.ac.uk/projects/fastqc/) was employed. To preprocess the data, the Universal Illumina Adapters (AGATCGGAAGAGC) were removed using trim galore version 0.6.2 (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/). This tool serves as a wrapper script for automated quality and adapter trimming, as well as quality control. Additionally, any sequences shorter than 20 bp were discarded. Apart from adapter removal, low-quality ends of the reads were trimmed, keeping a Phred score of 20.
The preprocessed reads were subjected to De Novo assembly using Trinity version 2.11.0 (https://github.com/trinityrnaseq/trinityrnaseq/wiki) with default settings and a kmer value of 25. Trinity incorporates 3 independent software modules—Inchworm, Chrysalis, and Butterfly—sequentially, enabling the processing of large volumes of RNA-seq reads. To minimize redundancy, the assembly was clustered at a sequence similarity of 90% using CD-HIT software version 4.7 (https://www.bioinformatics.org/cd-hit/). The longest isoform from each cluster was selected and considered as unigenes.
Differential analysis
The number of reads mapped to genes was quantified using samtools version 0.1.19. For the differential analysis between the Control and thermal stress condition, DESeq2 version 1.38.3, an R package tailored for analyzing count data from high-throughput sequencing assays like RNA-Seq, was utilized. Genes with an absolute log2 fold change (log2FC) greater than or equal to 2 and an adjusted P-value (Padjusted) less than or equal to .05 were considered highly differentially significant genes (DEGs).
Data submission in National Center for Biotechnology Information (NCBI)
Pearl millet gene assembly v.1.1 reference genome was aligned using HISAT2 (http://daehwankimlab.github.io/hisat2/). 15 The resultant SAM files were then converted to BAM format using Samtools (http://www.htslib.org). 16 For effective downstream analysis, the BAM files were indexed using the corresponding GTF (Gene Transfer Format) file.
The combined count matrix was normalized using the DESeq2 package (https://bioconductor.org/packages/release/bioc/html/DESeq2.html) 17 in RStudio. This normalization procedure took library size differences into account and utilized appropriate normalization techniques. The RNA sequences of pearl millet seedlings from this study were deposited in the database of the National Center for Biotechnology Information (NCBI) as Sequence Read Archive (SRA) data under the accession number PRJNA991076.
Pearl millet transcriptomic processing and phylogenetic analysis
The RNA sequences from the top 10 uncharacterized pearl millet leaves (refer to Table 1) were employed to conduct a search for homologous proteins via PSI Blast (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins). This process yielded 334 protein sequences. These sequences underwent filtration using the Cluster Database at High Identity with Tolerance (CD-HIT) Suite: Biological Sequence Clustering and Comparison (http://weizhongli-lab.org/cdhit_suite/cgi-bin/index.cgi) with a 90% identity cut-off. Employing the 90% cut-off led to a final selection of 110 protein sequences for incorporation into the multiple sequence alignment (MSA). The MSA was generated using MEGA7 (Molecular Evolutionary Genetics Analysis) software version 6. 18 Subsequently, a phylogenetic tree was constructed within MEGA 6, and alignment was conducted utilizing the Muscle program across the set of 334 proteins, where 10 of them were previously uncharacterized upregulated proteins in pearl millet. The alignment of protein sequences was achieved through both neighbor-joining 19 and UPGMA (Unweighted Pair Group Method), while employing a bootstrap method involving 500 replications for phylogeny testing. The interactive tree of life (iTol) online web tool (http://itol.embl.deg) was then employed to visualize and make adjustments to the tree’s color scheme. 20
De Novo assembly of the top 10 genes exhibiting the greatest fold change in pearl millet leaves, as compared to the control, during thermal stress.

Identification of conserved motifs and functional enrichment analysis
In order to delve deeper into the structural variations of the thermally expressed proteins in pearl millet, we subjected the protein sequences to the online tool MEME 21 (Multiple Expectation Maximization for Motif Elicitation) v. 4.12.0 (http://meme-suite.org/tools/meme) to characterize conserved motifs/domains. With the exception of setting the maximum number of motifs to 10, all default parameters were retained. Functional protein association and Enrichment Analysis were conducted using STRING 11.0 (string-db.org, accessed on August 9, 2023) 22 for the top 10 RNA sequences, with sorghum bicolor as the reference organism. The analysis was performed with a configuration of 10 nodes, 13 edges, an average node degree of 2.6, a local clustering coefficient of 2.6, and an average local clustering coefficient of 5.
Results and Discussion
De novo assembly metrics
The refined reads were subjected to De Novo assembly via Trinity version 2.11.0 (https://github.com/trinityrnaseq/trinityrnaseq/wiki), using default settings and a kmer value of 25. Trinity leverages 3 independent software modules—Inchworm, Chrysalis, and Butterfly—consecutively to process substantial quantities of RNA-seq reads. The resulting assembly was subsequently clustered to eliminate redundancy at a sequence similarity threshold of 90%, employing CD-HIT software version 4.7 (https://www.bioinformatics.org/cd-hit/). The longest isoform from each cluster was extracted utilizing Trinity and designated as unigenes. The top 10 genes exhibiting the greatest fold change in pearl millet leaves, as compared to the control, during thermal stress were listed in Table 1.
Pearl millet transcriptomic phylogenetic analysis
In the phylogenetic tree analysis, the results revealed distinct clustering patterns among the genes. Specifically, the following observations were made: The genes TRINITY_DN27468_c4_g1_i8, TRINITY_DN34944_c1_g1_i1, and TRINITY_DN39286_c0_g1_i1 were found to form individual clusters, indicating their distinct evolutionary relationships. Conversely, a group of 5 genes, namely TRINITY_DN33701_c2_g1_i2, TRINITY_DN38907_c1_g1_i1, TRINITY_DN60262_c0_g1_i2, TRINITY_DN38667_c0_g1_i1, and TRINITY_DN27468_c4_g1_i8, exhibited a close evolutionary relationship as they clustered together. This suggests a shared lineage or functional similarity among these genes (Figure 1).
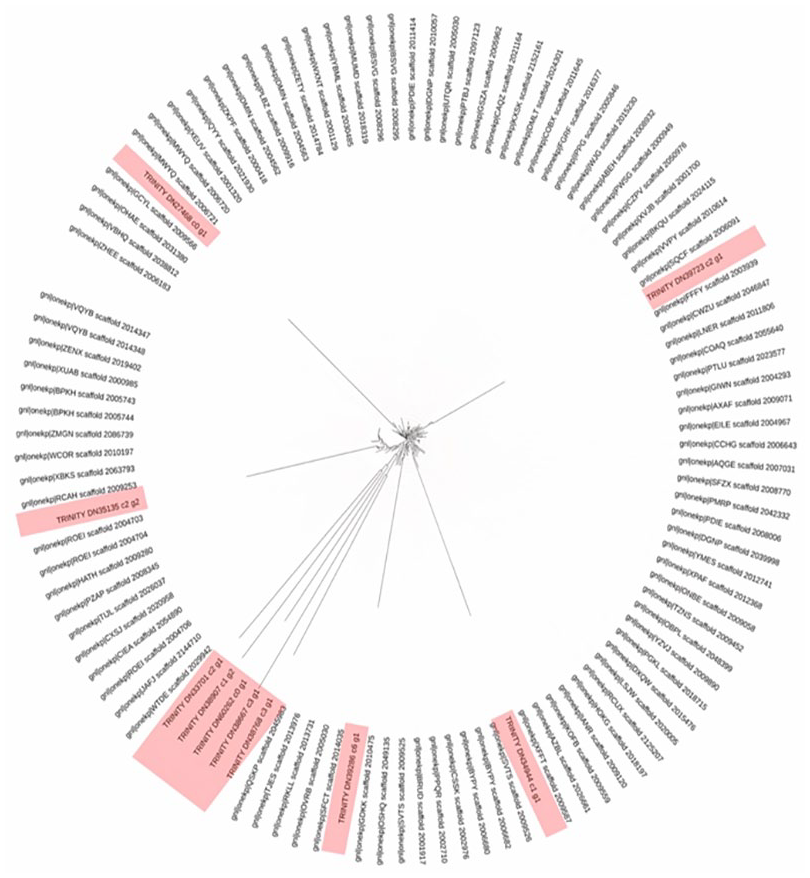
Phylogenetic tree was generated using 110 RNA sequences, which included the top 10 upregulated genes in pearl millet under thermal stress. The red color highlights the positioning of pearl millet RNA sequences within the distribution of clades.
Identification of conserved motifs
Amino acid (AA) residues play a crucial role in protein function and structure. Their significance is often determined by their evolutionary conservation. When AA residues are conserved throughout the evolutionary history of a protein, it suggests that they hold important functional or structural roles. The findings from the MEME analysis shed light on the conservation of domains within a set of 10 thermal-expressed pearl millet proteins (Figure 2). This analysis revealed that these 10 proteins share common and conserved domains, pointing toward a potential functional relevance. Notably, these conserved domains were identified as belonging to the amino acid transporter family. Among the set of 10 conserved domains that were characterized, a particularly intriguing discovery was made concerning Motif 7. This motif corresponds to the AA sequence of the gene TRINITY_DN38907_c1_g1_i1. Impressively, Motif 7 exhibited the highest degree of identity among these conserved domains (as depicted in Figure 2). This observation implies that the AA sequence of Motif 7 is remarkably well-preserved across evolutionary time, highlighting its critical role within the protein’s function or structure.
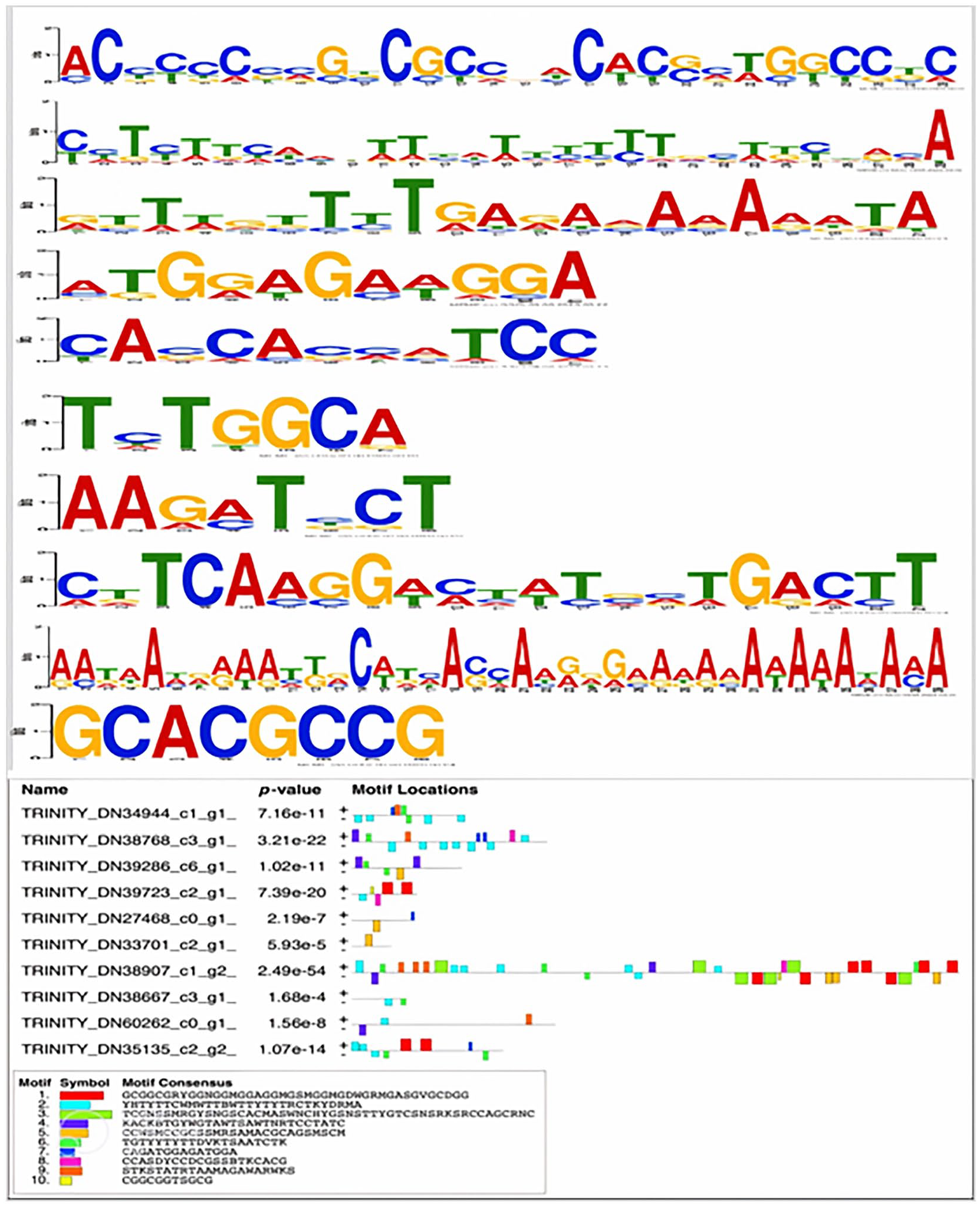
The discovery and analysis of conserved motifs within the top 10 upregulated proteins in pearl millet leaves under thermal stress were conducted. This involved molecular identification and sequence characterization of these conserved motifs. Diverse colors in the boxes represent distinct conserved motifs (motif 1-15), where larger font sizes denote higher conservation levels of the motifs. The original submitted sequence names are listed below the motifs.
Based on this remarkable conservation, the AA sequence of Motif 7 holds significant potential as an indicator for identifying uncharacterized thermal-expressed pearl millet proteins. This sequence could serve as a valuable tool for distinguishing these proteins from others, potentially aiding researchers in understanding their functions and roles in the context of thermal stress response. In essence, the investigation of these conserved AA residues not only provides insights into the evolutionary history of these proteins but also offers a promising avenue for studying their functional and structural characteristics.
Enriched functional categories of thermally induced proteins in pearl millet
Thermal stress biological processes and key domains
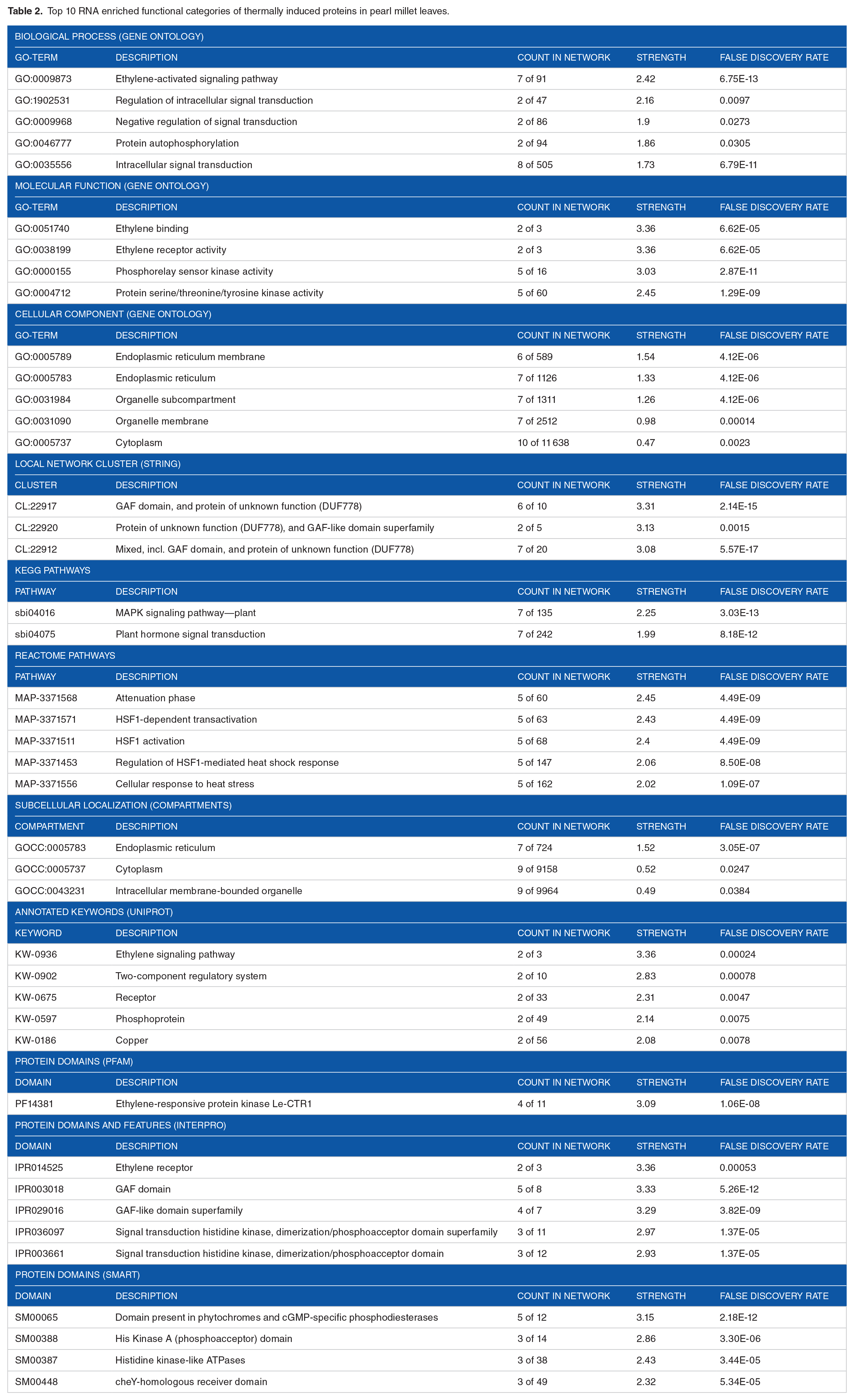
The enrichment analysis encompasses a diverse array of factors, including biological processes, molecular functions, cellular components, local network structure, KEGG pathways, Reactome pathways, subcellular localization (COMPARTMENTS), annotated keywords (UniProt), protein domains (Pfam), and protein domains (SMART). These findings are presented in Table 2 and Figure 3, showcasing the top 10 thermally induced proteins in pearl millet. The biological processes underlying the thermal response in pearl millet leaves involve intricate cellular mechanisms orchestrating the plant’s reaction to heat stress. Several pivotal processes contribute to the plant’s capacity to withstand and adapt to higher temperatures:
Top 10 RNA enriched functional categories of thermally induced proteins in pearl millet leaves.
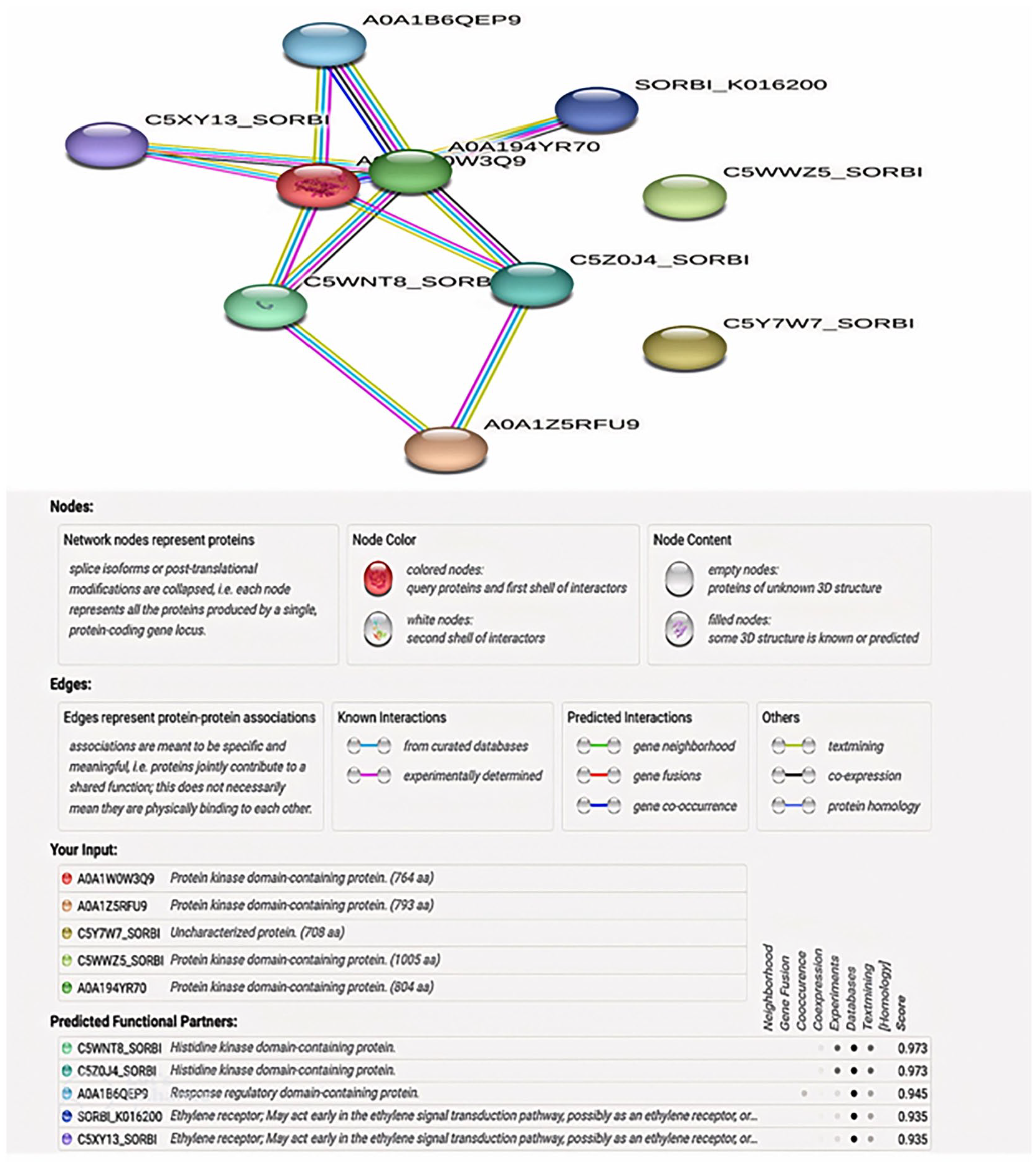
Protein-protein interaction networks of the top 10 RNA sequences from pearl millet leaves under thermal stress. String website used to perform the analysis and produce the image. Network nodes represent proteins. The network is clustered into 3 clusters, which are represented with red, green, and blue nodes, respectively. Colored nodes: query proteins and first shell of protein interactors; white nodes: the second shell of protein interactors. Edges represent protein-protein associations.
Ethylene-Activated Signaling Pathways and Ethylene Receptors: Ethylene, a plant hormone, regulates diverse growth and developmental processes, including stress responses. 23 Ethylene receptors are integral membrane proteins that perceive ethylene signals. These receptors share a common domain architecture among different isoforms, characterized by 3 transmembrane α-helices at the N-terminus forming the ethylene-binding domain, followed by a GAF (cGMP-specific phosphodiesterases, adenylyl cyclases, and FhlA) domain and a kinase domain. 24 These receptors play a pivotal role in modulating the plant’s response to thermal stress. In our study, exposing pearl millet to elevated temperatures induced changes in ethylene receptors. Conformational shifts might alter the receptors’ sensitivity to ethylene binding, thereby influencing downstream signaling events governing heat stress responses. This encompasses gene expression, physiological adjustments, and stress tolerance. The ethylene-activated signaling pathway serves as a conduit for transmitting the ethylene signal from the cell membrane to the nucleus, initiating a cascade of responses. During thermal stress, this pathway is likely activated as part of the plant’s adaptive strategy. Ethylene receptors perceive the stress signal, prompting downstream components to regulate gene expression, enzyme activities, and physiological alterations that bolster heat stress tolerance. The initial step involves ethylene binding proteins, enabling the plant to detect the hormone’s presence and trigger appropriate stress-related reactions. Ethylene receptors, situated in the cell membrane, facilitate ethylene signal perception. Their activity involves binding to ethylene molecules and undergoing conformational changes that drive downstream signaling events. In the context of thermal stress, the activity of ethylene receptors becomes crucial in conveying temperature-related cues to intracellular pathways, ultimately culminating in stress-responsive gene expression and physiological adjustments (as depicted in Table 2).
Environment sensing and transmitting signals domains
Plant leaves serve as intricate organs crucial for photosynthesis and various physiological processes vital to a plant’s survival. However, their well-being becomes vulnerable when exposed to thermal stress, such as high temperatures. In response, plants initiate a series of molecular and cellular reactions to counteract the harm inflicted by excessive heat. Central to these responses is the perception and transduction of signals, facilitated by diverse protein domains. In the ongoing study, proteins expressed under thermal conditions in pearl millet exhibited associations with specific domains. Table 2 and Figure 3 depict the Protein Domains (Pfam) linked to thermally expressed RNA in pearl millet leaves.
Plant environmental sensing and signal transduction are fundamental processes allowing plants to perceive and react to various external cues and stressors. Specialized domains play pivotal roles in these processes, receiving sensory input and transmitting signals within the plant.
Signal transduction histidine kinase, dimerization/phosphoacceptor domain superfamily
Histidine kinases are integral to 2-component signal transduction systems in plants and other organisms, enabling cells to respond to changing environments. The dimerization/phosphoacceptor domain superfamily encompasses the section of histidine kinases responsible for dimerization and subsequent autophosphorylation upon signal recognition. In thermally stressed plant leaves, these domains are likely initiators of signaling cascades that trigger adaptive responses. Phosphorylation events influenced by these domains can lead to changes in gene expression, protein activity, and cellular processes crucial for stress tolerance. Notably, PgB12D-Like, a protein implicated in signal transduction, exhibited heightened expression during cold, drought, and heat stress, suggesting its involvement in abiotic stress responses in Pennisetum glaucum. 25
In our study, GAF Domain and GAF-like Domain
Transcriptome analysis of pearl millet leaves exposed to thermal stress revealed the presence of GAF Domain and GAF-like Domain. The GAF domain is a conserved structural motif found in diverse proteins across various organisms, including plants. It stands for “cGMP-specific phosphodiesterases, adenylyl cyclases, and FhlA,” representing its presence in functionally diverse proteins. In thermally stressed plant leaves, the GAF domain could be part of proteins engaged in sensing environmental cues and transmitting signals. Conformational shifts induced by temperature variations in the GAF domain might trigger downstream responses, such as altered gene expression or enzyme activities, aiding the plant’s adaptation to thermal stress. The GAF-like domain superfamily encompasses a range of domains sharing structural similarities with the GAF domain. Similarly, the N-terminal GAF domain of AtNLP7 has a significant role in responding to nitrate deficiency and initiating ROS signaling. 26 These domains often engage in small molecule binding or protein-protein interactions. In thermally stressed plant leaves, members of the GAF-like domain superfamily likely contribute to temperature-responsive signaling pathways. They serve as intermediaries, transmitting temperature-related information and coordinating downstream reactions that facilitate appropriate stress responses.
Domain present in cGMP-specific phosphodiesterases
In response to stress, thermally treated pearl millet leaves revealed the presence of cGMP-specific phosphodiesterases. These enzymes regulate cellular levels of cyclic guanosine monophosphate (cGMP), a crucial secondary messenger involved in various signaling pathways, including stress responses. Within these enzymes, a “cheY-homologous receiver domain” is present. This domain operates within a 2-component signaling system akin to histidine kinases. During thermal stress, this domain could potentially perceive or transmit temperature-related signals, leading to changes in cGMP levels and consequent stress-related reactions. Although the fundamental biochemical processes governing cyclic nucleotides lack direct identification, extensive pharmacological and physiological investigations underscore cAMP and cGMP’s roles in regulating vital plant functions. Studies indicate their participation in stomatal opening mechanisms. 27 Evidence from both plant protoplast experiments28 -31 and intact tissue studies supports direct effects of cyclic nucleotides on cation fluxes, hinting at their potential as regulators for K+, Ca2+, and Na+ homeostasis. Notably, recent research has shown that exposure of intact plants to NaCl results in a rapid surge in cellular cGMP levels within seconds. 32 Additionally, cGMP has been identified as a contributor to phytochrome-mediated activation of the gene responsible for chalcone synthase, a pivotal enzyme in anthocyanin biosynthesis during chloroplast development. 33 Exposure of barley aleurone layers to gibberellic acid temporarily elevates cGMP levels, subsequently stimulating gene transcription and α-amylase production for starch-to-sugar conversion. 34 Furthermore, defense-related genes like PAL and PR-1 exhibit transcription linked to nitric oxide (NO) response, with cGMP as an intermediary. 35 Treating plants with membrane-permeable cGMP has shown to impact the transcription of various genes, especially those encoding transporter proteins. 31
Domain present in phytochromes
Phytochromes serve as photoreceptors enabling plants to perceive and react to light, encompassing both red and far-red wavelengths. Within this framework, Phytochrome-interacting factor (PIF) is classified as a basic helix-loop-helix (bHLH) type transcription factor. PIF functions as a central hub for intracellular signal regulation, participating not only in plant growth and development but also in abiotic stress resistance, such as coping with low and high temperatures. 36 While their primary role is in light sensing and signaling, they also engage in responses to thermal stress. A pivotal domain found in phytochromes is the “Histidine kinase A (phosphoacceptor) domain.” This domain is instrumental in phosphorylation events where a histidine residue is autophosphorylated in response to specific cues, including light and potentially thermal stimuli. Activation of this domain initiates downstream signaling cascades governing gene expression, physiological adjustments, and stress adaptation in pearl millet leaves. The “histidine kinase-like ATPase” domains are recognized for their involvement in 2-component signal transduction systems, sharing structural parallels with histidine kinases. This domain participates in ATP binding and hydrolysis, vital for transmitting signals in response to thermal stress. It potentially modifies downstream effectors’ activity or mediates responses to temperature shifts, thereby triggering diverse physiological responses. This process, involving phosphorylation, facilitates oligomerization, dimerization, interactions with other proteins, interactions with DNA, or combinations of these mechanisms. 37
In the context of thermal stress in pearl millet leaves, these domains collectively contribute to the plant’s capacity to perceive, convey, and react to temperature-related cues. They serve as intermediaries initiating cellular responses, modulating gene expression, and orchestrating adaptive mechanisms that bolster the plant’s resilience and survival in challenging thermal conditions. To summarize, the aforementioned domains—ethylene receptor, GAF domain, GAF-like domain superfamily, and the signal transduction histidine kinase dimerization/phosphoacceptor domains—play indispensable roles in enabling plant leaves to detect, transmit, and counteract thermal stress. These molecular components are integral to the intricate signaling network that empowers plants to adapt and thrive amid demanding environmental circumstances.
Thermally Expressed Biological Process
Heat stress poses a significant threat to plants and their growth. The systems plants have developed to respond to heat stress are intricate and finely tuned, encompassing both heat signaling transduction and photosynthesis. To safeguard crop yield, comprehending the molecular mechanisms behind heat stress signaling transduction and photosynthesis is essential. Modern proteomics studies have unveiled pathways related to plant heat-responsive signaling and photosynthesis modulation. Notably, components like GTP-binding protein, nucleoside diphosphate kinase, annexin, and brassinosteroid-insensitive I-kinase domain interacting protein 114 have been implicated in heat signaling transduction. Diverse protein patterns within photosynthetic proteins underscore that plant heat tolerance hinges on the adjustment of factors such as stomatal CO2 exchange, photosystem II, Calvin cycle, ATP synthesis, and chlorophyll biosynthesis. 38
Pearl millet’s elaborate signaling networks enable it to sense and adapt to elevated temperatures. These routes rely on fundamental mechanisms:
Regulation of Intracellular Signal Transduction: This mechanism is crucial for the plant’s response to temperature stress, controlling signal propagation. Intracellular signal transduction involves intricate protein complexes and pathways that finely adjust signal transmission, ensuring the appropriate cellular response to stress.
Negative Regulation of Signal Transduction: An essential aspect of minimizing prolonged stress effects, negative control modulates signal intensity and duration. Feedback loops and inhibitory mechanisms are vital for maintaining cellular balance and energy efficiency.
Autophosphorylation of Protein Kinases under Heat Stress: Heat stress triggers autophosphorylation in protein kinases. This alteration in activity and interaction with downstream effectors transmits temperature information, activating stress-responsive genes and pathways.
Metabolic Processes and Intracellular Signal Transduction: Thermal stress initiates metabolic processes that transmit signals from the cell membrane to the nucleus, instigating intracellular signal transduction. Consequently, gene expression and physiological processes undergo changes. These pathways likely leverage heat to activate transcription factors that govern gene expression related to heat shock proteins, antioxidants, and stress responses. In unison, these processes equip pearl millet with the ability to sense, comprehend, and respond effectively to elevated temperatures. This capacity enhances the plant’s resilience and survival by adjusting molecular, cellular, and physiological functions to confront challenging thermal conditions.
Molecular Function Associated With Thermal Stress
A study demonstrated that the Arabidopsis multistep-phosphorelay (MSP) signaling system employs histidine kinases, phosphotransfer proteins, and response regulators. Although the 2-component system (TCS) is a well-studied prokaryotic signal transduction mechanism, the plant MSP was only characterized in 1993. Despite some progress, MSP remains insufficiently understood. Recent findings in Arabidopsis thaliana revealed significant similarities between bacterial TCSs and plant MSPs in terms of protein phosphorylation, dephosphorylation, degradation, and dimerization, which govern TCS/MSP signaling. 39
In pearl millet leaves experiencing thermal stress, various molecular functions are pivotal for the plant to sense, process, and respond to higher temperatures. These functions are integral to the signaling and regulatory processes enabling the plant to adapt to challenging thermal conditions. The molecular function analysis of thermally treated pearl millet leaves indicated a link with Phosphorelay Sensor Kinase Activity and Protein Serine/Threonine/Tyrosine Kinase Activity.
Phosphorelay sensor kinases are central components of plant 2-component signal transduction systems that detect external stimuli and initiate appropriate responses. These kinases play a critical role in sensing temperature changes during thermal stress. They autophosphorylate in response to such cues, triggering a cascade of phosphorylation events that transmit the thermal signal along the signaling pathway, ultimately leading to various stress adaptations. Meanwhile, protein kinases are enzymes regulating cellular processes by phosphorylating specific amino acid residues in target proteins. Under thermal stress, protein kinases with specificity for serine, threonine, and tyrosine residues become particularly active. They facilitate the phosphorylation of downstream targets involved in stress responses. These kinases modify protein activity, stability, and interaction partners, influencing the plant’s capacity to withstand and manage elevated temperatures.
Conclusion
The primary objective of this investigation was to gain insights into the molecular mechanisms underlying the response of pearl millet leaves to thermal stress. The study focused on 2 primary areas of investigation: the preservation of protein domains and the enhanced functional classifications of proteins generated by heat conditions. Utilizing various in silico approaches, such as phylogenetic analysis, distinct clustering patterns were identified among the genes, leading to significant observations. Specifically, TRINITY_DN27468_c4_g1_i8, TRINITY_DN34944_c1_g1_i1, and TRINITY_DN39286_c0_g1_i1 formed separate clusters, indicating their distinct evolutionary relationships. In contrast, another cluster comprised 5 genes: TRINITY_DN33701_c2_g1_i2, TRINITY_DN38907_c1_g1_i1, TRINITY_DN60262_c0_g1_i2, TRINITY_DN38667_c0_g1_i1, and TRINITY_DN27468_c4_g1_i8, suggesting a close evolutionary connection and potential functional similarity among them. This analysis unveiled a shared and conserved domain among these 10 proteins, strongly implying potential functional significance. Notably, these conserved domains belong to the amino acid transporter family. Among these 10 characterized conserved domains, a particularly intriguing finding pertains to Motif 7. This motif corresponds to the amino acid sequence of the gene TRINITY_DN38907_c1_g1_i1. Impressively, Motif 7 exhibited the highest degree of identity among these conserved domains, as depicted in Figure 2. This observation underscores the exceptional preservation of Motif 7’s amino acid sequence across evolutionary time, highlighting its pivotal role in the protein’s function or structure.
The enrichment analysis encompasses a diverse array of factors, encompassing biological processes, molecular functions, cellular components, local network structure, KEGG pathways, Reactome pathways, subcellular localization (COMPARTMENTS), annotated keywords (UniProt), protein domains (Pfam), and protein domains (SMART).
The biological processes orchestrating the thermal response in pearl millet leaves are intricate, underpinning the plant’s ability to effectively respond to heat stress. Several pivotal processes contribute to the plant’s capacity to withstand and adapt to higher temperatures:
Ethylene-Activated Signaling Pathways and Ethylene Receptors: During thermal stress, this pathway likely becomes activated as part of the plant’s adaptive strategy. Ethylene receptors detect the stress signal, prompting downstream components to regulate gene expression, enzyme activities, and physiological adjustments that enhance heat stress tolerance.
Signal Transduction Histidine Kinase Dimerization/Phosphoacceptor Domain Superfamily: The dimerization/phosphoacceptor domain superfamily, responsible for dimerization and subsequent autophosphorylation upon signal recognition, likely initiates signaling cascades in thermally stressed plant leaves. Phosphorylation events influenced by these domains can lead to changes in gene expression, protein activity, and cellular processes crucial for stress tolerance.
GAF Domain and GAF-like Domain: Temperature-induced conformational shifts in these domains potentially trigger downstream responses, such as altered gene expression or enzyme activities, aiding the plant’s adaptation to thermal stress. These domains serve as intermediaries that transmit temperature-related information and coordinate downstream reactions.
Domain Present in cGMP-Specific Phosphodiesterases: Domains within these enzymes, such as the “cheY-homologous receiver domain,” potentially perceive or transmit temperature-related signals, leading to changes in cGMP levels, and consequent stress-related reactions. Cyclic nucleotides like cGMP impact various aspects of plant function, from stomatal opening mechanisms to ion flux regulation and gene transcription.
Domain Present in Phytochromes: Phytochromes serve as photoreceptors enabling plants to perceive and respond to light across different wavelengths. The “Histidine kinase A (phosphoacceptor) domain” within phytochromes plays a pivotal role in phosphorylation events upon signal recognition, potentially initiating downstream signaling cascades governing gene expression, physiological adjustments, and stress adaptation in pearl millet leaves.
Furthermore, the molecular function analysis reveals that thermally treated pearl millet leaves possess Phosphorelay Sensor Kinase Activity and Protein Serine/Threonine/Tyrosine Kinase Activity. These molecular functions are pivotal for the plant to sense, process, and respond to higher temperatures, as they play crucial roles in initiating signaling cascades and modifying downstream effectors’ activity, ultimately contributing to the plant’s resilience and survival in challenging thermal conditions.
Future research may involve breeding programs or genetic engineering approaches to develop heat-resistant varieties. Additionally, comparing the molecular responses to thermal stress in pearl millet with those in other crops can reveal unique adaptive mechanisms. Research in this field has the potential not only to enhance our understanding of pearl millet’s response to thermal stress but also to contribute to the development of resilient crop varieties capable of thriving in the face of climate change-induced temperature fluctuations. This, in turn, can significantly bolster food security in vulnerable regions.
Footnotes
Acknowledgements
I would like to express my sincere gratitude to Dr. Zainab M. Almutairi from Prince Sattam bin Abdulaziz University m for generously providing me with the seeds necessary for the current study. Their support and contribution have been invaluable in enabling the progress and success of this research project.
Author Contribution
FD: Conceptualization; investigation; methodology; formal analysis; visualization; writing original draft; writing review and editing preparation.
Funding:
The author received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests:
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability
The RNA sequences of the current study pearl millet shoots for all samples were submitted to the National Center for Biotechnology Information (NCBI) database under the Sequence Read Archive (SRA) data, with the accession number PRJNA991076.
