Abstract
Keywords
Introduction
Clinically, MIRI is still thought to be one of the worst threats to cardiac patients worldwide due to the unclear mechanisms and serious complications involved if reperfusion injury is too severe. Until now, the method to cope with MIRI has been to restore blood flow in a timely manner after myocardial ischaemia, such as coronary artery bypass grafting and percutaneous coronary intervention, to avoid a second hit called reperfusion injury.1,2 Nevertheless, these salvage strategies tend to further endanger the heart, and the clarified mechanisms include inflammatory reactions, oxidative stress and calcium overload. 3 Because IRI is unavoidable under certain circumstances, protocols should be implemented to protect the injured myocardium from further injury by those downstream molecules, or more attention should be placed on studying the mechanisms of downstream biological processes.
With the development of mechanistic research, researchers tend to realize that MIRI often precedes various pathological conditions, 4 such as calcium overload, oxygen paradox, proinflammatory response and apoptosis, which directly determine the course, prognosis and outcome of the disease. Moreover, these pathologic events are partially correlated with ERS, 5 leading to the opening of mitochondrial permeability transition pores, which is marked as the common end of cell lysis and death. Therefore, during MIRI, many different deleterious events could be potential targets to effectively resolve pathologic processes. Among them, ERS occurs at the later stage of MIRI almost ubiquitously. Additionally, cardiac ERS usually follows the overactivation of the unfolded protein response (UPR), which is activated during reperfusion of the ischaemic heart and is accompanied by ROS generation and the release of numerous inflammatory cytokines.6,7 During this procedure, many efforts have been made to keep the UPR at an appropriate level by using specific inhibitors, 8 herbal medicine, 9 etc., or even to try to protect against MIRI by regulating ERS in the heart.8,10–12 Unfortunately, few players in this field have been validated as useful in regulating ERS. In this regard, the key target molecules and the underlying mechanisms between ERS and MIRI should be clarified. First, the general genetic expression profile should be organized to help us screen the potential genes during ER stress in myocardial I/R injury.
Microarray chip analysis is an effective method to screen and even identify promising DEGs for clinical problem solving under specific conditions, including MIRI and ERS, but few studies have shed light on this subject in rats. The present study aimed to identify DEGs in the ERS-related signalling pathway in an in vivo MIRI model.
Methods
Gene data profile
The key words “(myocardial ischaemia reperfusion injury) AND (rat)” were selected to search for a matched GEO database website (http://www.ncbi.nlm.nih.gov/geo), and three microarray array datasets were identified. The criteria used for selection were as follows: (i) raw data were completely derived from rats, (ii) samples were collected from heart tissues, and (iii) MIRI was induced with 45 min of ischaemia and 24 h of reperfusion by the LAD method. Finally, the GSE122020 database was preferable for data extraction, and subsequently, the gene expression matrix was downloaded from the NCBI website. The dataset profile was obtained as follows: GSE122020 was produced on the GPL22388 platform (Affymetrix Rat Transcriptome Array 1.0 [transcript (gene) CSV version]), which is an integrated high throughput screening gene chips including information about mRNA and lncRNA. After downloading the fully attached datasets, the stratified mRNA data were extracted from the control and IRI groups, and each group was analysed in triplicate.
Analyses and enrichment of DEGs
With the help of the website attached GEO2R tool and R studio, the gene expression data were pooled according to the group allocation. After normalization, principal component analysis was conducted to depict the primary source of variability in the data. Then, the gene expression matrix was organized with corresponding DEGs on the condition of p value <0.05 and | fold change (FC)| ≥ 1.2. Available tools, such as Kyoto Encyclopedia of Genomes and Genes (KEGG) and GO (http://geneontology.org/), were utilized to define each gene’s cellular function, cellular components, and biological processes. Moreover, as an important analysis method to predict possible signalling pathways, the KEGG website was explored, and the outcomes were exhibited with a plot depicted by R studio for visualization of details between the MIRI and control groups.
Potential genes in the ERS-related signalling pathway
After conducting GO and KEGG enrichment analyses, the 20 most valuable signalling pathways were identified and analysed. Moreover, GO enrichment analyses, such as cellular function, cellular components, and biological processes, were also included, and the upregulated and downregulated genes were analysed as well. For target genes involved in the ERS pathway, molecules belonging to the category of biological processes such as “negative regulation of response to endoplasmic reticulum stress”, “response to endoplasmic reticulum stress”, “intrinsic apoptotic signalling pathway in response to endoplasmic reticulum stress” and “negative regulation of endoplasmic reticulum stress-induced intrinsic apoptotic signalling pathway” were chosen.
The definition of hub genes by PPI network analysis
The Search Tool for the Retrieval of Interacting (STRING, https://cn.string-db.org/, version 11.5) database is commonly adopted for analysing PPIs. 13 In brief, the list of DEGs was uploaded onto the STRING website to predict the correlation of those proteins on the condition of a confidence score of ≥0.4. Subsequently, an SVC format file containing the concluded genes was imported into Cytoscape. 14 By calculation, the network module containing specific biological functions was constructed, and then the Cytoscape Molecular Complex Detection (MCODE) plug-in was applied to identify divergent modules in the PPI network. The parameters used were set as node score cut-off = 0.2, degree cut-off = 2, K-core = 2, and max depth = 100. 15 Finally, the cyto-Hubba plugin was applied to identify hub genes in the present PPI network according to rank. Altogether, the top six nodes representing the potential mediators ranked by the maximal clique centrality (MCC) method were selected for validation.
MIRI model establishment
To verify the predicted results derived from the above bioinformatics analysis, the experimental study was conducted on rats with an in vivo LAD model induced by MIRI (n = 5, each group/each experiment), and each experiment was performed in triplicate. Thirty SPF grade male adult Sprague‒Dawley rats weighing 200–300 g were raised in the experimental animal centre of Jia Xing Medical College (SYXK-2017-0006) under standard conditions and randomized into two groups. All experimental procedures were strictly in accordance with the “Consensus of author guidelines on animal ethics and welfare” and approved by the Laboratory Animal Ethics Committee of Jia Xing Medical College (JUMC2022-107). According to the experimental protocol, the in vivo I/R injury model was established according to YANG’s reports. 10 Briefly, rats were anaesthetized with intraperitoneal 1% pentobarbital sodium 40 mg/kg, 2 min after which tracheal intubation was completed to begin mechanical ventilation with a composition of oxygen and air by connecting an animal ventilator. With minimally invasive thoracotomy, the left anterior descending artery was isolated, and a thread was placed beneath the vessel. For the MIRI group, the artery of rats was occluded for 45 min to induce myocardial ischaemia. Then, the thread was removed, and the incision was closed for 24 h of reperfusion. For the control group, nothing was different from the MIRI group except for the lack of ligation of the vessel. A heating carpet was used to keep the body temperature as normal as possible during the entire procedure. To verify the success of the in vivo model of MIRI, the rats were anaesthetized at the end of 24 h of ligation, echocardiography was conducted, and myocardial samples were collected immediately for RNA extraction. Blood was taken for serum cardiac troponin I (cTnI) detection. All animals were euthanized by carbon dioxide asphyxiation at a flow rate of 5 L/min with 50% volume displacement/min after the necessary experiment in a chamber for approximately 20 min. If the animal was not found dead, cervical dislocation was applied.
RT‒PCR examination
To verify the mRNA expression levels of six major genes in animals, all tissue samples from the two groups were collected with the same procedure. The RT‒PCR procedure was performed as follows: the left ventricle of the rat heart was removed and washed three times with saline. Then, total RNA was extracted with TRIzol reagent, and the total RNA concentration and 260/280 ratio were determined using a nanodrop machine (Thermo Fisher Scientific, USA). After, using a commercial reverse transcription RNA kit (PrimeScript RT Master Mix (for Real Time, RR036A) and qPCR kit (TB Green Premix DimerEraser [Perfect Real Time], RR091A) purchased from Takara Corporation (Takara Biomedical Technology [Beijing] Co., Ltd), the relative mRNA expression level was calculated according to the manufacturers’ protocols. Briefly, approximately 1 μg of total RNA was reverse transcribed to cDNA. The real-time assays were conducted with a 20 μl reaction system containing the Master Mix and 1 μl of cDNA product. The reaction parameters were as follows: (1) stage 1 reaction at 95°C for 0.5 min, (2) stage 2 reaction included 95°C for 10 s and 60°C for 0.5 min, reacted for 40 cycles, (3) stage 3 reaction is for the melting curve, consisting of 95°C for 15 s, 60°C for 1 min and 95°C for 15 s. Subsequently, the relative gene expression levels of each gene were calculated by using the 2−ΔΔCT method and compared to the housekeeping gene glyceraldehyde-phosphate dehydrogenase accordingly. The primers used in this procedure were as follows: Atf3 Forward Primer 5′-TTTGCTAACCTGACACCCTTTG-3′, Reverse Primer 5′-AGAGGACATCCGATGGCAGA-3′. Ppp1r15a Forward Primer 5′-GCCTGCAAGGGGCTGATAAG-3′, Reverse Primer 5′-TTTGTATCCCGGAGCTATGGA-3′. Casp12 Forward Primer 5′-TAGGGGAAAGTGCGAGTTTCA-3′, Reverse Primer 5′-GGGCCAATCCAGCATTTACCT-3′. Eif2ak2 Forward Primer 5′-ATGCACGGAGTAGCCATTACG-3′, Reverse Primer 5′- TGACAATCCACCTTGTTTTCGT-3′. Ifng Forward Primer 5′-GCCACGGCACAGTCATTGA-3′, Reverse Primer 5′-TGCTGATGGCCTGATTGTCTT-3′. Hspa1a Forward Primer 5′-TGGTGCAGTCCGACATGAAG-3′, Reverse Primer 5′- GCTGAGAGTCGTTGAAGTAGGC-3′.
ELISA and Echocardiography
To verify the success of the in vivo model of MIRI, the rats were anaesthetized at the end of 24 h of ligation, and echocardiography was conducted for both groups of rats. Briefly, an ultrasonic workstation (MYLAB™ X5 VET, Esaote, Italy) was used to detect the volume of the systolic and diastole periods of the heart in M-mode. The formula EF (%) = (EDV − ESV)/EDV (%) was adopted to calculate the left ventricle ejection fraction (LVEF, %), where EDV stands for end diastolic volume and ESV stands for end systolic volume. Moreover, the serum cTnI level was measured after peripheral blood was taken according to the protocol of ELISA kits (Elabscience, China). Briefly, 0.5 mL blood was collected from the retro-orbital sinus following anaesthesia and immediately before euthanasia. After the buildup of the standard curve, the original cTnI concentration was calculated based on the absorbance values at 450 nm.
Statistical analysis
The results are expressed as the mean ± standard error (SE) for all variables if the dataset is normally distributed. Statistical analysis was performed using GraphPad Prism (version 7, GraphPad Software, San Diego, USA). Student’s t test was adopted to detect the statistical significance between the MIRI and control groups, and p < 0.05 was considered statistically significant.
Results
Identification of DEGs in the GEO database
From GSE122020, the profiles of six samples (three control samples: GSM3453064, GSM3453065, and GSM3453066 and three treatment samples: GSM3453058, GSM3453059, and GSM3453060) were reviewed and selected thereafter for comparison. All data expression results are described in Figure 1. Volcano plots showed that 1647 mRNAs, including 790 upregulated and 857 downregulated genes, were identified as DEGs in the GSE122020 dataset under the mRNA category when compared between groups (Figure 1(a)). The clustered data profile, which showed obvious biological variability between the control and MIRI treatment samples, can be seen in Figure 1(b) with 2D PCA plots. After normalization, the gene expression in both control samples and IRI treatment samples was compared under t test examination. The range of gene expression values is illustrated in the box plot in Figure 1(c), of which the middle black lines indicate the median gene expression values (Figure 1(c)). For convenience, the top 300 DEGs in each set between different groups are displayed in the heatmap (Figure 1(d)). Representative plot of outcomes from GSE122020. (a) Volcano plot of overall DEGs; the red and blue dots represent upregulated and downregulated genes, respectively. (b) 2D PCA plot between six samples. The purple and green dots represent the treatment and control samples, respectively. (c) Description of the general matrix expression of six GSM examples after normalization. (d) Heatmap of the first 300 DEGs cited; the red and blue colours represent upregulated and downregulated genes compared to the control group, respectively. GSE, the Gene Expression Omnibus databases; DEGs, differentially expressed genes.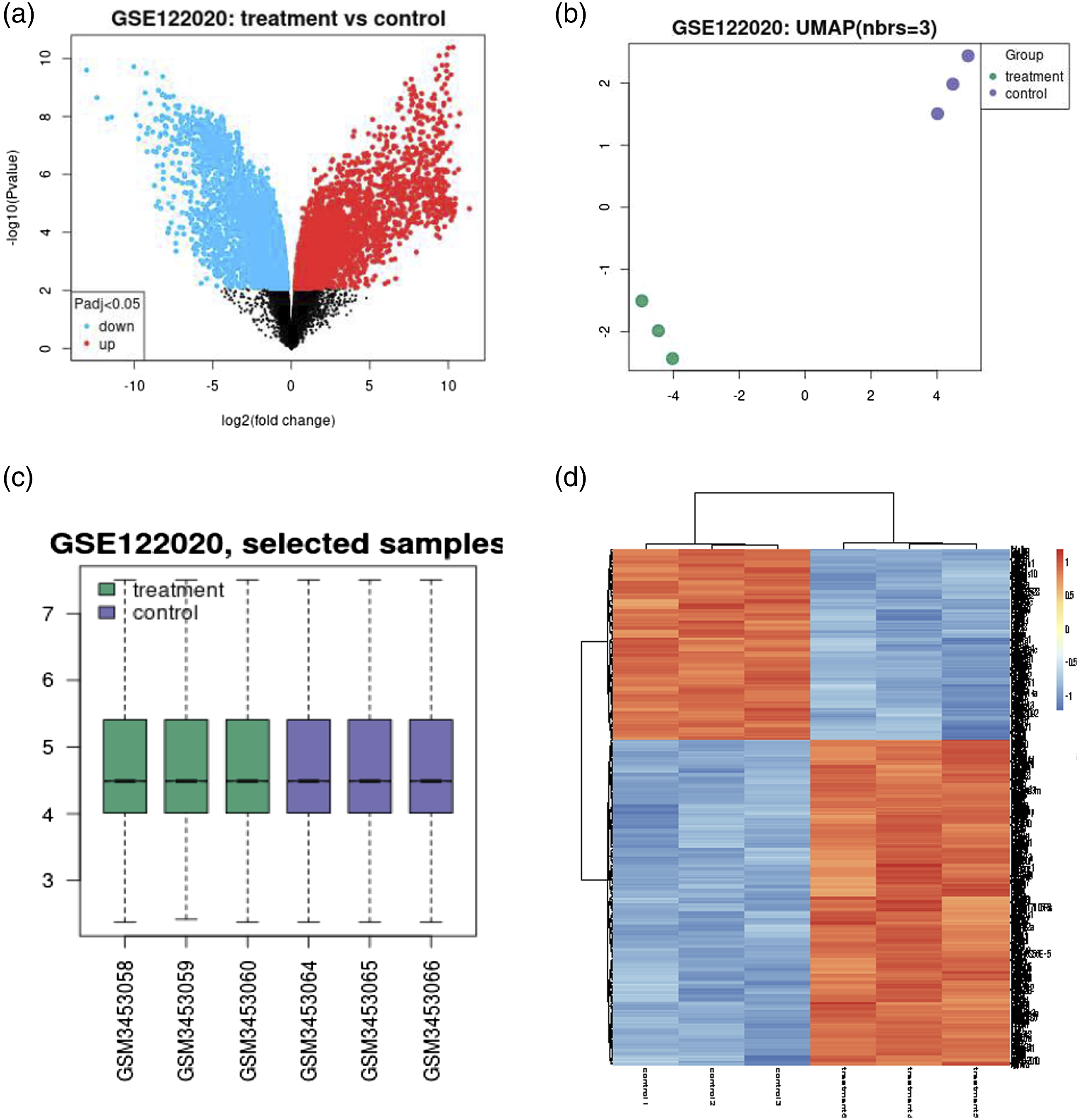
GO and KEGG pathway enrichment analysis
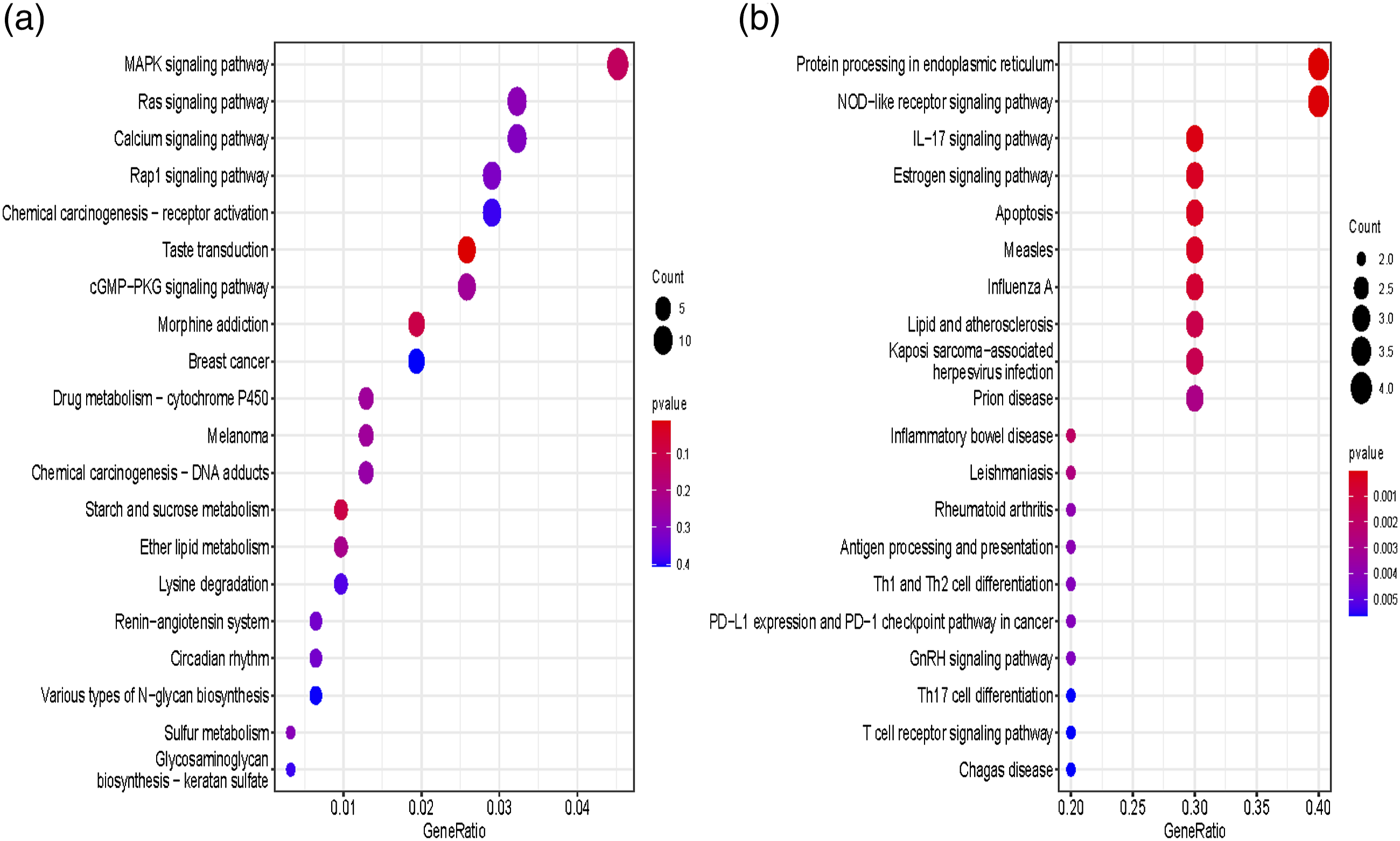
To further elucidate gene function, GO and KEGG signalling pathway enrichment analyses were conducted with the upregulated and downregulated DEGs, respectively. The analysed terms included three classical categories: biological process (BP), cellular component (CC), and molecular function (MF). Through enrichment of GO terms, the top 10 significantly meaningful terms are listed in Figure 2 according to the gene count and p value. Figure 2 shows that most of the upregulated genes were anchored at the extracellular matrix or the external side of the membrane, exerted the molecular function of actin binding and were involved in biological processes, including positive regulation of cytokine production and regulation of the inflammatory response (Figure 2(a)). The downregulated genes were not closely associated with specific functions or biological processes (Figure 2(b)). A smaller number of genes were observed to exert odourant binding functions and participate in the biological processes of “response to pheromone”. For the KEGG analysis, 20 out of all functional signalling pathways were pooled and depicted by R-studio in Figure 3(a). The most important signalling pathway shared by these DEGs was the MAPK signalling pathway, which included more than 10 genes with statistically significant p values. In addition, the Ras signalling pathway and calcium signalling pathway were also candidates. The precise details can be seen in. Supplementary file 1. Gene Ontology enrichment of significantly expressed DEGs in terms of molecular function, cellular component and biological process. (a) The upregulated DEGs in terms of molecular function, cellular component, and biological process according to the changes in p values and gene count. (b) The downregulated DEGs in terms of molecular function, cellular component, and biological process according to the changes in p values and gene count. DEGs, differentially expressed genes. Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment of related genes. (a) The 20 most significant signalling pathways from the 1647 mRNAs are illustrated in a bubble plot; the rank was determined by the p value, genotype and gene count. (b) The 20 most significant signalling pathways, from datasets of ERS-related biological processes, are illustrated with bubble plots; the rank was determined by the p value, genotype and gene count. ERS, endoplasmic reticulum stress.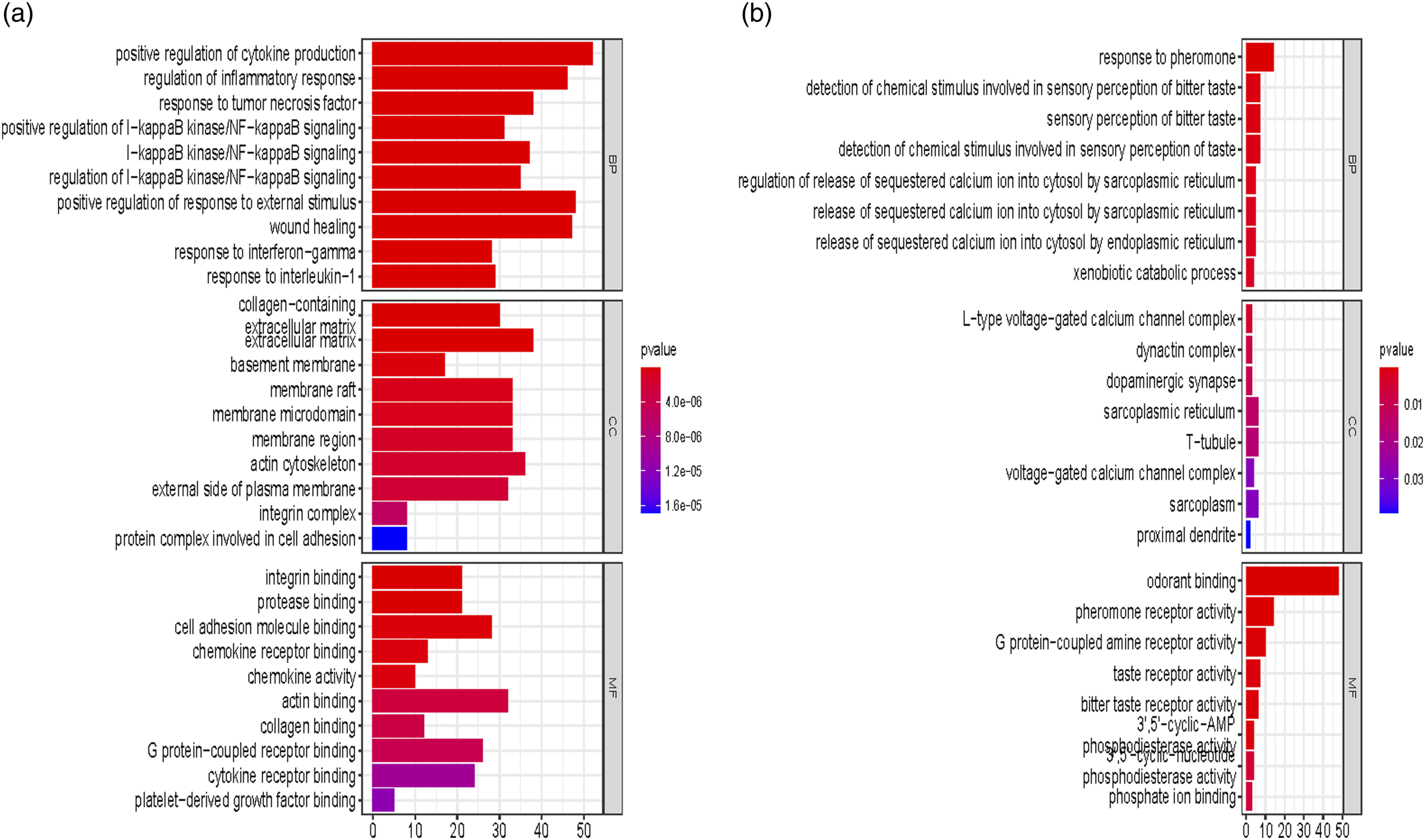
PPI network and significant modules
The biological processes correlated to “endoplasmic reticulum stress” were found in the upregulated GO enrichment dataset, the genes list involved could be seen in below table.

After the selection of the 15 DEGs, they were entered into the STRING website to complete PPI analysis, and the result was visualized in Cytoscape (Figure 4). In total, 15 DEGs constructed an interaction network containing 15 nodes and 16 edges under relatively high confidence (0.4), making the average node degree reach 2.3 and the p value <6.91 × 10−11 (Figure 4(a)). Eventually, 10 genes formed the first modules with tight connections by using the Cytoscape MCODE plugin under the condition of k-core equals two and cut off is 2. Therefore, the module contains the following genes: Itpr1, Jun, Casp12, Eif2ak2, Trim25, Ifng, Atf3, Hspa1a, Ppp1r15a and Creb3 (Figure 4(b)). The representation of the PPI network and hub genes of 15 DEGs. (a) The connected 15 genes in the ERS pathway were entered into the PPI network on the STRING website, and the interaction was illustrated. (b) The first cluster was presented with Cytoscape. (c) The six hub genes were calculated with the MCC method and ranked by score. (d) GO enrichment analysis of the six hub genes. PPI, protein‒protein interaction; STRING, Search Tool for the Retrieval of Interacting Genes/Proteins; DEGs, differentially expressed genes.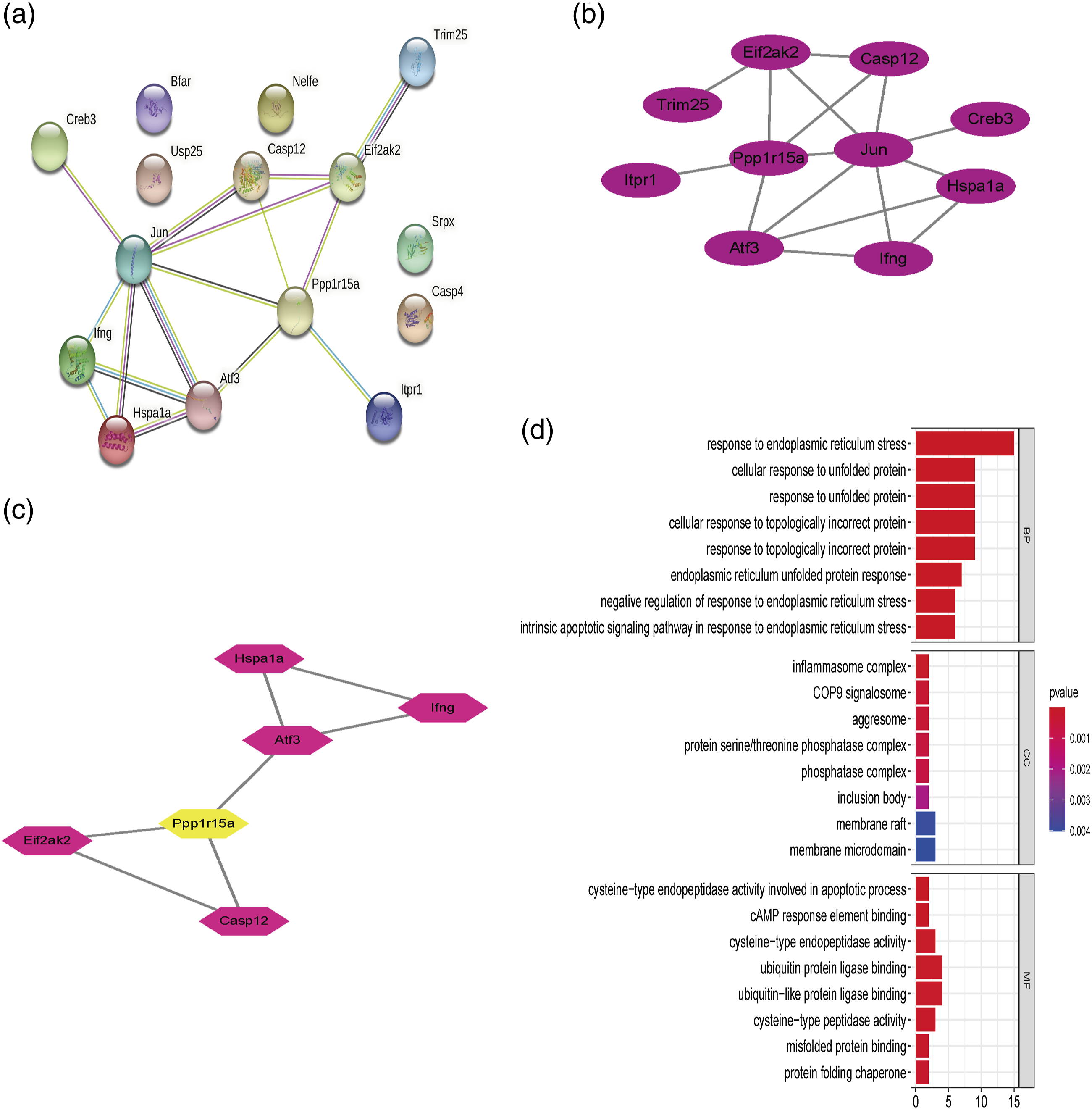
Confirmation of the hub genes
After establishing the first module in the PPI network, the cytoHubba plugin of Cytoscape was used to select the top six hub genes from these 10 genes, which were ranked by the MCC method. These six genes were Atf3, Ppp1r15a, Casp12, Eif2ak2, Ifng and Hspa1a according to the scores (Figure 4(c)). Among them, Atf3 and Ppp1r15a were the most important genes. Subsequently, the enrichment analysis was completed. The GO enrichment revealed that the genes were mainly associated with the biological processes of “response to endoplasmic reticulum stress”, “cellular response to unfolded protein”, or “negative regulation of response to endoplasmic reticulum stress” (counted genes >10, and p value <0.001) (Figure 4(d)). Additionally, these biological processes were fulfilled primarily by executing the molecular function of “ubiquitin protein ligase binding” or “ubiquitin−like protein ligase binding”. From these results, we concluded that these genes may interfere with the degradation of protein by ubiquitin protein ligase binding to influence ERS in cells. In addition, the online analysis of KEGG enrichment revealed that four of the genes, Ppp1r15a, Casp12, Eif2ak2 and Hspa1a, were closely involved in the “protein process in endoplasmic reticulum” pathway, and Eif2ak2 may be the upstream regulator in this respect. The specific details of the related pathway can be seen in Supplementary File 2.
The conformation of the MIRI model
None of the included animals were found dead. The LVEF of rat heart was evaluated by echocardiography at the end of 24 h of reperfusion (Figure 5(a)). Compared to the animals in the control group, rats in the MIRI model group exhibited lower LVEF (75.8 ± 2.85 vs 47.6 ± 4.51, p = 0.0007) (Figure 5(b)). Similarly, the concentration of cTnI in rat serum increased markedly in the MIRI model group compared to that in the control group (n = 5, 9.01 ± 0.46 vs 2.72 ± 0.27, p < 0.0001) (Figure 5(c)). In addition, the colour of myocardial ischaemia and S-T segment changes were also confirmed by staff during the experiment. The parameters for the in vivo MIRI model. (a) Representative echocardiography images of mice in both the control and MIRI groups (n = 5). (b) The statistical results of LVEF of mouse heart from picture a. (c) The average serum concentration of cTnI detected at the end of 24 h after the experiment. Data represent means ± SEMs. MIRI, myocardial ischaemia reperfusion injury; LVEF, left ventricle ejection fraction, cTnI, serum cardiac troponin I; *p < 0.05 compared to the control group.
Experimental verification of six hub genes
From the analysis of myocardial tissue, the mRNA expression of these six targeted genes examined by RT‒PCR markedly increased in the MIRI model group compared to that in the control group. The expression ranged from several-to dozens of-fold (p < 0.05). Moreover, Figure 6 shows that Ppp1r15a and Ifng had the greatest increase under IRI stimulation. The remaining four increased relatively less but were statistically significant (p < 0.05, Figure 6). These results were consistent with those of predictions from bioinformatics analysis, but further verification is needed. Relative mRNA expression of six potential genes. (a–f) Bar chart illustration of the specific real-time quantitative gene expression after normalization to GAPDH between the control and LPS groups (n = 5). Data represent means ± SEMs. MIRI, myocardial ischaemia reperfusion injury; *p < 0.05 compared to the control group.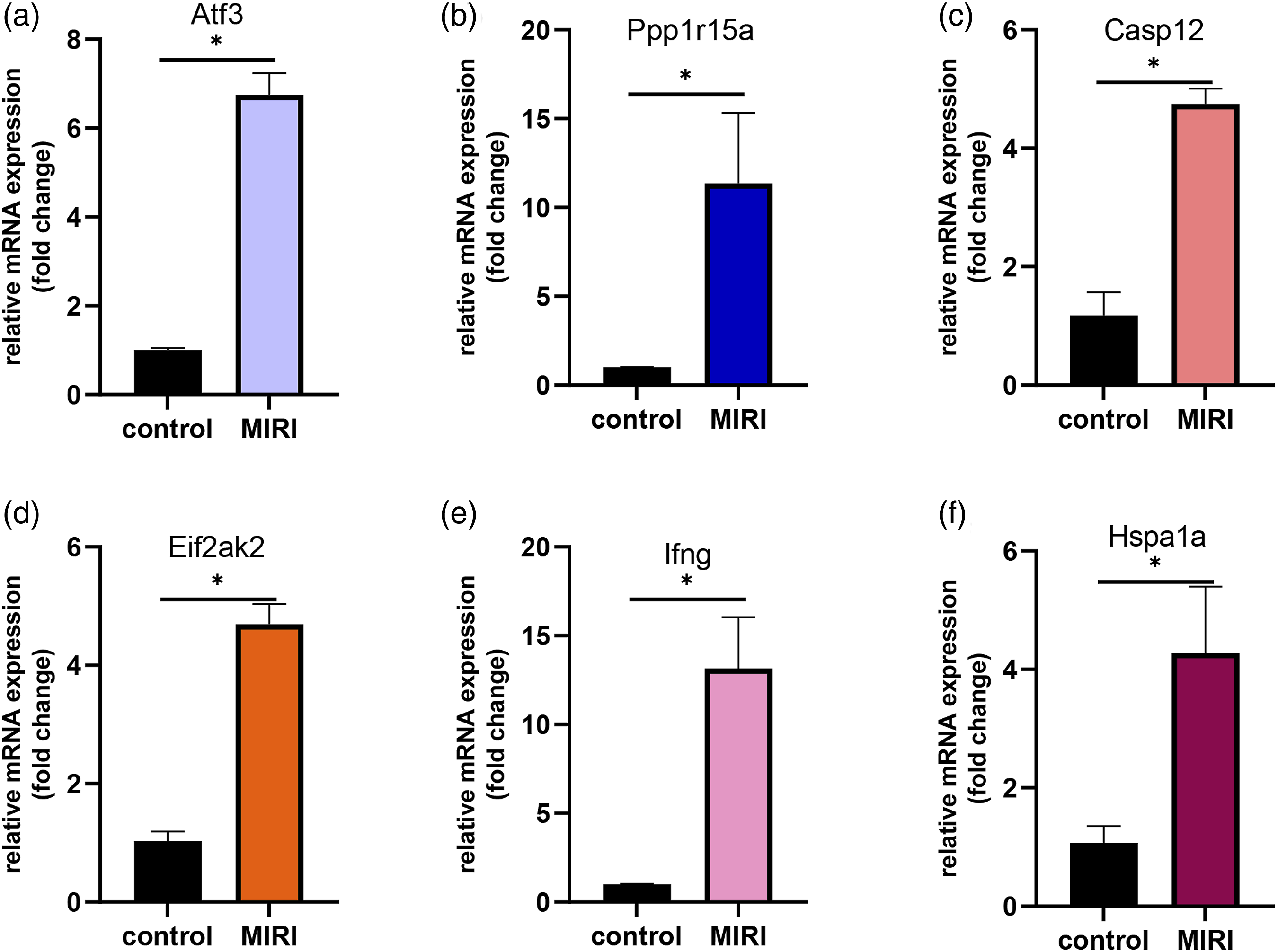
Discussion
There are multiple parallel processes, such as ferroptosis, 16 oxygen-sensitive transcription, 17 redox balance, 8 and ERS, 10 connected to the pathogenesis of MIRI. For prevention or therapy, heart injury can be prevented or attenuated by regulating these mediators. Among these methods, targeting the ERS pathway is a potential way to cope with MIRI.8,9,11,12,18 In this study, a transcriptomic study based on the rat MIRI model was conducted, and 790 upregulated genes were identified as meaningful genes by bioinformatics analysis. Through further enrichment analysis and fundamental experimental validation, the final six hub genes, Atf3, Ppp1r15a, Casp12, Eif2ak2, Ifng, and Hspa1a, were identified as potential genes that are closely involved in ERS biological processes and exhibit significant mRNA overexpression in the left anterior descending coronary artery ligation (LAD) model.
MIRI can lead to multiple biochemical mechanisms based on previous studies; ERS is correlated with cardiomyocyte apoptosis, oxidative stress, the inflammatory response, and calcium imbalance induced by reperfusion.19,20 These underlying reasons disrupt the homeostasis of the endoplasmic reticulum if the reaction is long lasting. GSE122020 is a microarray analysis database, aiming to find the DEGs for treating IRI damage in rats, presented by WANG et al. 21 Based on their supplementary files, the mRNA expression matrix was split from the whole set and reanalyzed by researchers. Compared to rats in the control group, rats in the MIRI group showed distinct gene expression in the MAPK signalling pathway, Ras signalling pathway and calcium signalling pathway. The molecular functions of those DEGs were mainly interpreted as “positive regulation of cytokine production” and “regulation of inflammatory response”. These enrichment conclusions were consistent with Xu’s experimental results in the field of MIRI, 22 illustrating an inflammatory response following MIRI. To further determine the overlap between MIRI and ERS, GO:1903573, GO:0034976, GO:0070059 and GO:1902236 were found to be closely related to ERS under the subunit of upregulated genes, and finally, 15 highly correlated genes were found, including Bfar, Itpr1, Srpx, Hspa1b, Usp25, Jun, Casp12, Eif2ak2, Casp4, Trim25, Ifng, Atf3, Hspa1a, Ppp1r15a, and Creb3. Using PPI network analysis and Cytoscape software, 10 out of 15 DEGs formed the first modules with tight protein connections, including Itpr1, Jun, Casp12, Eif2ak2, Trim25, Ifng, Atf3, Hspa1a, Ppp1r15a and Creb3. Finally, the six hub genes were retrieved for further verification.
Unlike other studies either based on existing targets to predict downstream signalling pathways 23 or analyse diverse disease models, 24 the present study identified promising genes in ERS following MIRI. To our knowledge, this is the first microarray analysis on the classic LAD model to identify ERS targets in rat. To test the predictions in an animal model, we adopted Xu’s method to create the in vivo model 25 with a 45 min ischaemia and 24 h reperfusion. The results showed that all six genes were overexpressed more or less in IRI myocardial tissue. Atf3, a membrane-located transcription factor, is predicted to be part of the CHOP-ATF3 complex. Hayner and Lu’s report26,27 indicated that the appearance of ERS could induce the activation of ATF3, further regulating the transcription of stress molecules. Ppp1r15a,28–30 a cytoplasmic protein phosphatase, enables protein kinase binding and is involved in the endoplasmic reticulum unfolded protein response. Ppp1r15a also corelated with the activation of ER, is a product of ERS and could influence tissue apoptosis. Casp12 is a positive regulator of apoptotic processes in striated muscle cells involved in the intrinsic apoptotic signalling pathway in response to ERS.31,32 Eif2ak2, which is coded as protein PERK, can mediate upregulation of ERS-induced autophagy and may govern signalling pathways through UPR-independent functions. Eif2ak2 is reportedly involved with autophagy and apoptosis in ER.33–35 From the above explanations, these hub genes may be involved in the development of ERS, but extensive mechanisms are still unknown.
This study has some limitation. First, we did not examine the myocardial infarction size on the rats due to a reagent shortage, so the alternative echocardiography was used instead. Second, we did not examine the time points for gene expression in the animal model and chose the 24 h reperfusion to perform the experiment based on literature reports. Fulfilling the ideal model parameters of interest would be best.
Conclusion
From the bioinformatics and experimental aspects, the final 6 DEGs were found to be involved in the regulation of ERS following the MIRI event and were overexpressed in the in vivo MIRI model. Based on the results of GO and KEGG enrichment analyses, these hub genes are potential targets for treating MIRI.
Supplemental Material
Supplemental Material - The key mediators involved in myocardial endoplasmic reticulum stress induced by ischaemia reperfusion injury in rats
Supplemental Material for The key mediators involved in myocardial endoplasmic reticulum stress induced by ischaemia reperfusion injury in rats by Jiayue Qing, Hongmei Zhou, Li Hu, Zhipeng Zhu in European Journal of Inflammation.
Supplemental Material
Supplemental Material - The key mediators involved in myocardial endoplasmic reticulum stress induced by ischaemia reperfusion injury in rats
Supplemental Material for The key mediators involved in myocardial endoplasmic reticulum stress induced by ischaemia reperfusion injury in rats by Jiayue Qing, Hongmei Zhou, Li Hu, Zhipeng Zhu in European Journal of Inflammation.
Supplemental Material
Supplemental Material - The key mediators involved in myocardial endoplasmic reticulum stress induced by ischaemia reperfusion injury in rats
Supplemental Material for The key mediators involved in myocardial endoplasmic reticulum stress induced by ischaemia reperfusion injury in rats by Jiayue Qing, Hongmei Zhou, Li Hu, Zhipeng Zhu in European Journal of Inflammation.
Footnotes
Author’s contributions
Conceptualization, Zhipeng Zhu; methodology, Jiayue Qin; software, Zhipeng Zhu; resources, Hongmei Zhou; writing—original draft preparation, Zhipeng Zhu; supervision, Li Hu. All authors have read and agreed to the published version of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
