Abstract
In recent times, diverse agriculturally important endophytic bacteria colonizing plant endosphere have been identified. Harnessing the potential of
Keywords
Introduction
Insights into plant endosphere communities have provided several opportunities in ensuring sustainable agriculture.
1
Endosphere represents the entire internal tissue of plants colonized by microorganisms called ‘microbial endophytes’, which is beneficial to the host plants without eliciting any pathological effect.
2
The role of microbial endophytes has been postulated with significant interest in agricultural biotechnology for plant growth and protection against environmental stress adaptors.
3
Remarkably, identifiable bacterial species in the genus
Genetic modification of
The genomic analysis of
The bacteria isolated from sunflower roots were sourced from farmlands in Lichtenburg, South Africa (26°4′31.266″S, 25°58′44.442″E) in February 2020. The healthy sunflower plants were carefully uprooted, placed inside sterile zip-lock bags, transported into the laboratory and stored at 4°C. The root samples were cut into small sizes with a sterile scalpel and washed in sterile distilled water. To ensure complete removal of the epiphytic bacteria, surface sterilization was achieved by soaking the samples in 70% ethanol for 3 minutes, followed by 3% sodium hypochlorite for 3 minutes, 70% ethanol for 30 seconds and rinsed with sterile distilled water. The level of sterility of the samples was assessed by pour plating on Luria-Bertani (LB) agar using the last water used to rinse plant samples. One gram (1 g) of plant material was weighed, suspended in 1 M phosphate-buffered solution (PBS) and manually macerated in a mortar and pestle until a smooth suspension was obtained. Sample suspensions were serially diluted up to 10−9 dilutions and 0.1 mL of an aliquot from dilutions 10−5 and 10−6 were pipetted into Petri dishes and pour plated with sterilized LB agar. Inoculated Petri plates were incubated at 28°C for 24 hours. Distinct bacterial colonies formed on the plates were counted and selected based on morphological appearance. Pure culture of the bacterial isolate was obtained by repeated streaking onto sterile LB agar and incubated at 28°C for 24 hours. The pure bacterial strains were kept on an agar slant and stored at 4°C for further use.
The stored purified
Genomic sequences were analysed on the KBase platform.
8
The quality assessment of the sequence reads was evaluated by FastQC (version 0.11.5),
9
while the removal of sequence adaptor and low-quality bases was processed with trimmomatic (version 0.36).
10
Furthermore, sequence reads were assembled with SPAdes (version 3.13.0).
11
Gene annotation and prediction were performed on RASTtk (Rapid Annotations using Subsystems Technology toolkit – version 1.073) and the publicly available NCBI (https://www.ncbi.nlm.nih.gov/) Prokaryotic Genome Annotation Pipeline (PGAP).
12
All analyses were performed using default parameters unless otherwise specified. Secondary metabolites were determined by antiSMASH (version 6.0.0).
13
The circular genome visualization was obtained from KBase (https://kbase.us/), while the phylogeny analysis was performed using MrBayes (http://www.phylogeny.fr/one_task.cgi?task_type=mrbayes) (version 3.2.6).8,14 The genomic comparison of strain T4S with other
The strain T4S responded positively to almost all the plant growth-promoting tests (Table 1).
Plant growth-promoting features of

Abbreviations: IAA, indole acetic acid; ZOC, zone of clearance measurement.
Mean ± standard deviation values with different superscripts (small letters) within the same column represent a significant difference.
− = negative reaction.
+ = positive reaction.
The comparison of genomic features of
Comparison of genomic features of
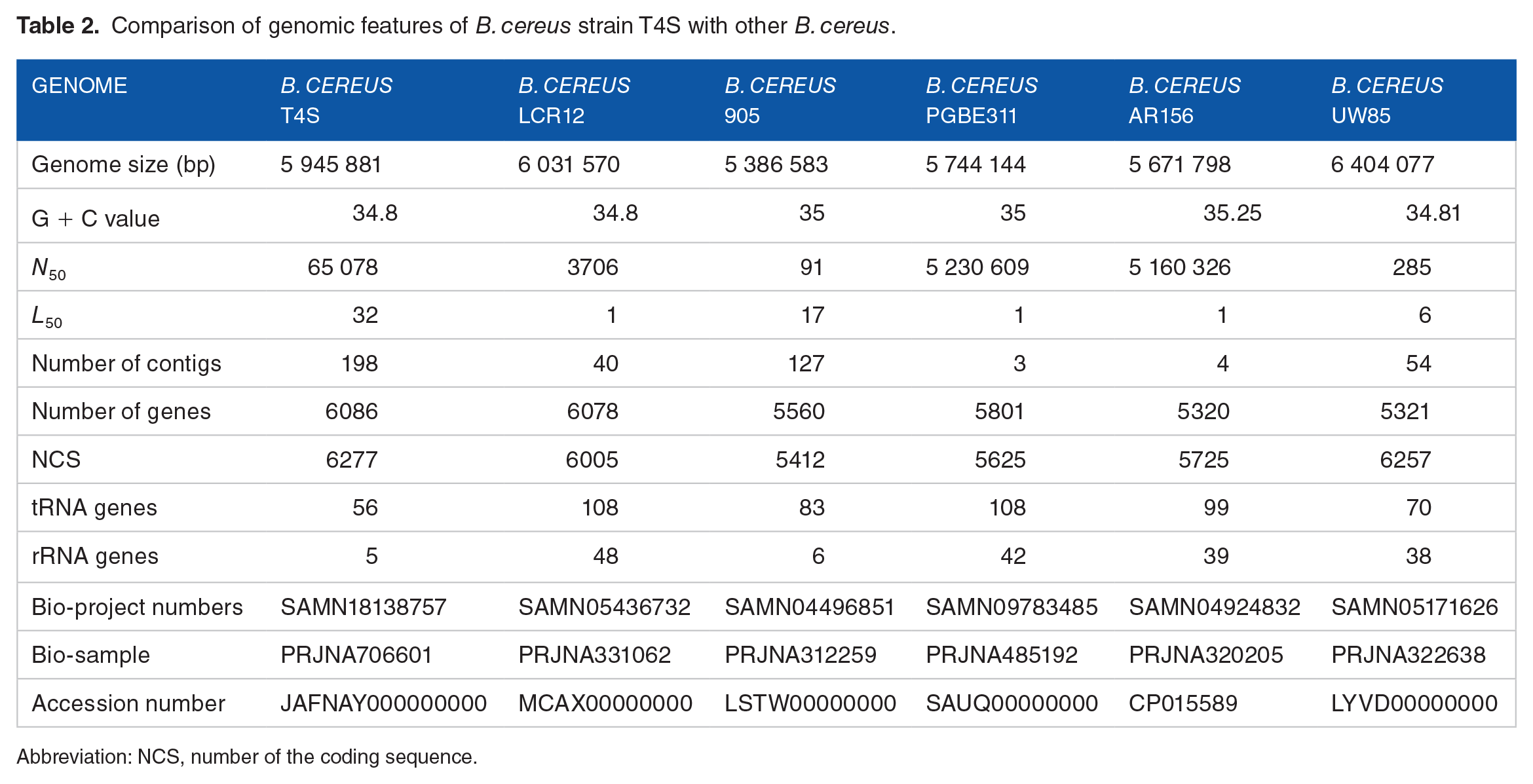
Abbreviation: NCS, number of the coding sequence.

Phylogeny sequence data of
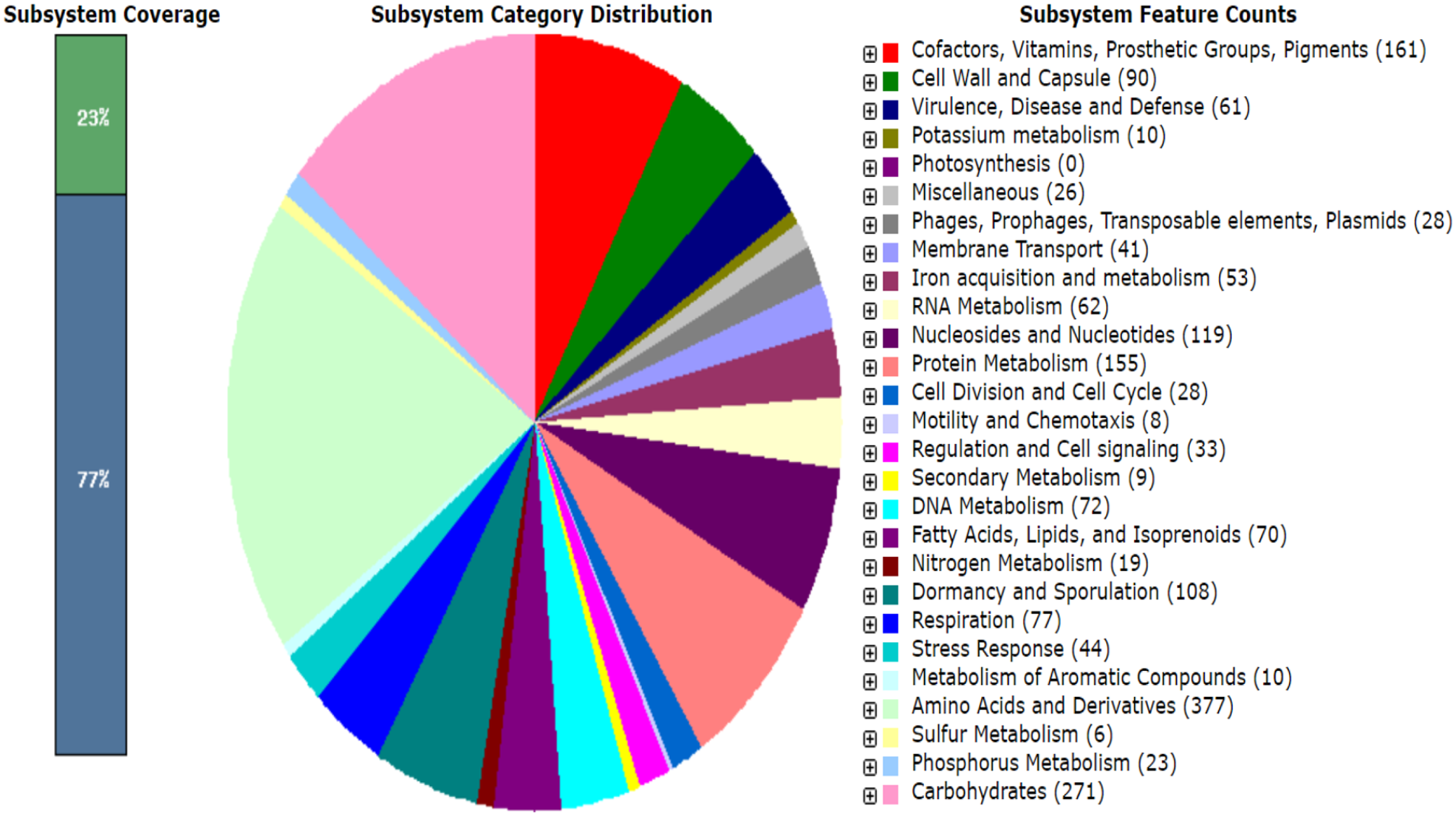
Subsystem category distribution of key PCG of

Circular representation of the draft genome of
The secondary metabolite gene clusters, namely, non-ribosomal peptides (NRPS) and siderophore of strain T4S with 100% similarity selected and information on the functions of notable secondary metabolite (siderophore) genes (AsbABCDEF) that code for petrobactin synthesis/petrobactin-synthetic aryl carrier protein and 3,4 dihydroxybenzoic acid-AMP ligase, specific for additional biosynthetic, core synthetic and other genes were presented (Table 3).
Information on the siderophore similar known gene cluster.

Also, in Figure 4, the core structure of NRPS with a functional group, thiol (SH), carboxyl (COOH), amine (NH2), carbonyl (C=O) is represented. Genomic analysis revealed various genes involved in nitrogen fixation and nitrogen metabolism, iron transport and siderophore production, growth hormone synthesis, phosphate solubilization and transport, motility, biofilm formation and chemotaxis, biological control, and oxidative and nitrosative stress (Table 4).

Predicted core structure of NRPS synthesis by
Genes involved in plant growth promotion (PGP).

The results obtained from this study revealed the genetic capacity of strain T4S, which suggests its bioprospecting in agriculture in enhancing plant growth. The predicted genes on the other hand may contribute to plant-bacterial interactions and ensure endosphere competence. Therefore, genome analysis of strain T4S provides information on the genes involved in sustainable plant growth and health.
Sequence Data Information
From the NCBI database output, the Bioproject is https://www.ncbi.nlm.nih.gov/bioproject/PRJNA706601; BioSample number is https://www.ncbi.nlm.nih.gov/biosample/SAMN18138757 while the whole genome accession number is JAFNAY000000000.
Footnotes
Acknowledgements
OOB acknowledges the National Research Foundation of South Africa for the grants (Grant numbers: 123634; 132595), supporting research in her laboratory. BSA thanked the National Research Foundation of South Africa and The World Academy of Science (TWAS) for the NRF-TWAS African Renaissance Doctoral scholarship (UID:116100). ASA is grateful to North-West University for a postdoctoral fellowship award.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the National Research Foundation of South Africa grants (UID: 123634; 132595 to OOB).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
All the mentioned authors contributed substantially and intellectually to this work. OOB designed the research, revised the work critically for important intellectual content, performed quality assurance, provided funding, project administration and resources. BSA was involved in data curation, formal analysis, investigation, visualization of data and writing of the original draft of the manuscript. ASA was involved in data curation, visualization of data, reviewing and thoroughly editing of the initial draft, validation and formal analysis.
