Abstract
Introduction
Endophytes are microorganisms such as bacteria and fungi, that colonize and inhabit plant tissues for all or part of their life cycle without causing harm or damage to the host plant. This community of microbes forms beneficial relationship with their plant host counterpart and exhibit plant growth and development properties through the production of unique and various biochemical compounds.1,2 Of interest in this study are bacterial endophytes, which like fungal endophytes are known to provide nitrogen-fixation benefits to the plant and may secrete secondary metabolites and siderophores, which allow for pathogenic resistance and absorption of iron in host plants. 3 Furthermore, the interactions between endophytic bacteria and plants promote plant’s ability to withstand extreme and unfavorable environmental conditions. 4
Bioremediation is a cleaning-up technique in polluted environments by transforming enzymes during the biosorption of different toxic metals.10,11 The environmental pollution due to various toxic metals is a serious concern in the food sector because of decreasing crop yields and food quality due to the overuse of fertilizer, mulching, and agricultural pesticides which cause heavy metal pollution of soils. 12 Bioremediation has been identified as a good technique for controlling pollution and treatment of toxins. Several studies have shown that endophytes can be used in bioremediation techniques because of their ability to improve plant response to a range of stressors. 13 Endophytes improve the ability of plants to survive in various environmental conditions such as drought, nitrogen deficiency, pathogens, and salinity, and such impact has a very important contribution to the bioremediation of heavy metals. 14
Whole genome sequence analysis of bacterial endophytes is a valuable tool that is necessary for the identification of genes that are responsible for key endophytic biochemical activities such as nitrogen fixation, nutrient acquisition, and phytohormone production. Individual genome sequences improve the study of plant-microbe interactions as well as data analysis in requisite endophytic metagenomics, proteomics, and transcriptomics studies.15,16 Furthermore, obtained whole genomic sequence data is more useful to advance taxonomical resolution of bacterial species in contrast to more traditional methods such as 16S rRNA gene sequencing analysis.
17
The objective of this study was to sequence, assemble, and annotate the genome of the endophytic bacterium
Materials and Methods
Bacterial sample preparation and genomic DNA extraction
The endophyte,
Genome sequencing, de novo assembly and annotation
The extracted DNA was sent for whole genome sequencing at the Agriculture Research Council (ARC) in Onderstepoort, South Africa, a commercial service provider. The Illumina HiSeq platform was used, and pair-ended libraries (2 × 150 bp) were created with NextEra DNA library preparation kit. A total of 781 234 paired-end reads at 25× coverage were obtained from this workflow. Pre-annotation analyses were done on the Galaxy web platform (https://usegalaxy.org) using default parameters.
20
The quality of the raw reads obtained was assessed with FastQC v 0.72. The
Phylogenomic analyses
The genomic data was uploaded on Type Strain Genome Server (TYGS) (https://tygs.dsmz.de) for whole genome based taxonomic comparison with other published type strains. 25 The average nucleotide identity (ANI) value was determined on the Orthologous Average Nucleotide Identity Tool (OAT) v 0.93.10. 26 Islandviewer 4 Web (https://www.pathogenomics.sfu.ca/islandviewer) was used to identify the genomic islands (GI) by screening the PGAP annotation file obtained from NCBI. 27 The shared and unique genes clusters of strain MHSD4 were determined by comparing it to other closely related species using the EDGAR 2.0 platform. 28
The whole genome shotgun project data has been deposited at DDBJ/ENA/GenBank with BioProject number PRJNA783158 and BioSample number SAMN23419518 under the accession number JAJNDH000000000. The version presented here is JAJNDH010000000.
Interprepation of Data Set
The assembly of the draft genome produced 41 contigs with N50 value of 550 575 base pairs (bp). The genome size obtained was 4 647 677 bp and the G+C content was 54.2%. The average genome sizes and G+C content of
Annotated genome statistics of

Abbreviations: bp, base pairs; ncRNA, non-coding RNA, tRNA, transfer RNA.
Phylogenomic classification of the draft genome
Based on 16S rDNA sequences, strain MHSD4 displayed a close relation to
Moreover, Figure 2 (Supplemental Data) shows that based on whole genome sequence analysis, strain MHSD4 was closely related to
Average nucleotide identity (ANI) and digital DNA-DNA hybridization (dDDH) values are both popular methods of discriminating between bacterial taxons and are imperative tools for genomic studies.31,32 The dDDH values calculated with 3 different formulas between
RAST identified a total of 1490 subsystem feature counts (Figure 1) with amino acids and derivatives subsystem category dominant with 23.8% followed by carbohydrate metabolism genes with 20%. The least number of genes identified on the genome were only 2 belonging to the phages, prophages, transposable elements, and plasmids subsystem category. Genes identified within the genome of strain MHSD4 that supported the endophytic lifestyle included siderophore production as well as other iron acquisition and metabolism genes, nitrogen, phosphorus, and potassium metabolism genes. 21 Furthermore, virulence, disease, and defense, phytohormone production, and stress tolerance genes play a significant role in the host plant growth promotion and development and the symbiotic association between bacterial endophytes and plants.10,35

The subsystem distribution of
The RAST sequence-based comparative tool was used to compare the genomes of strain MHSD4 to

(A)
Figure 3 shows the different OrthoANI values between various
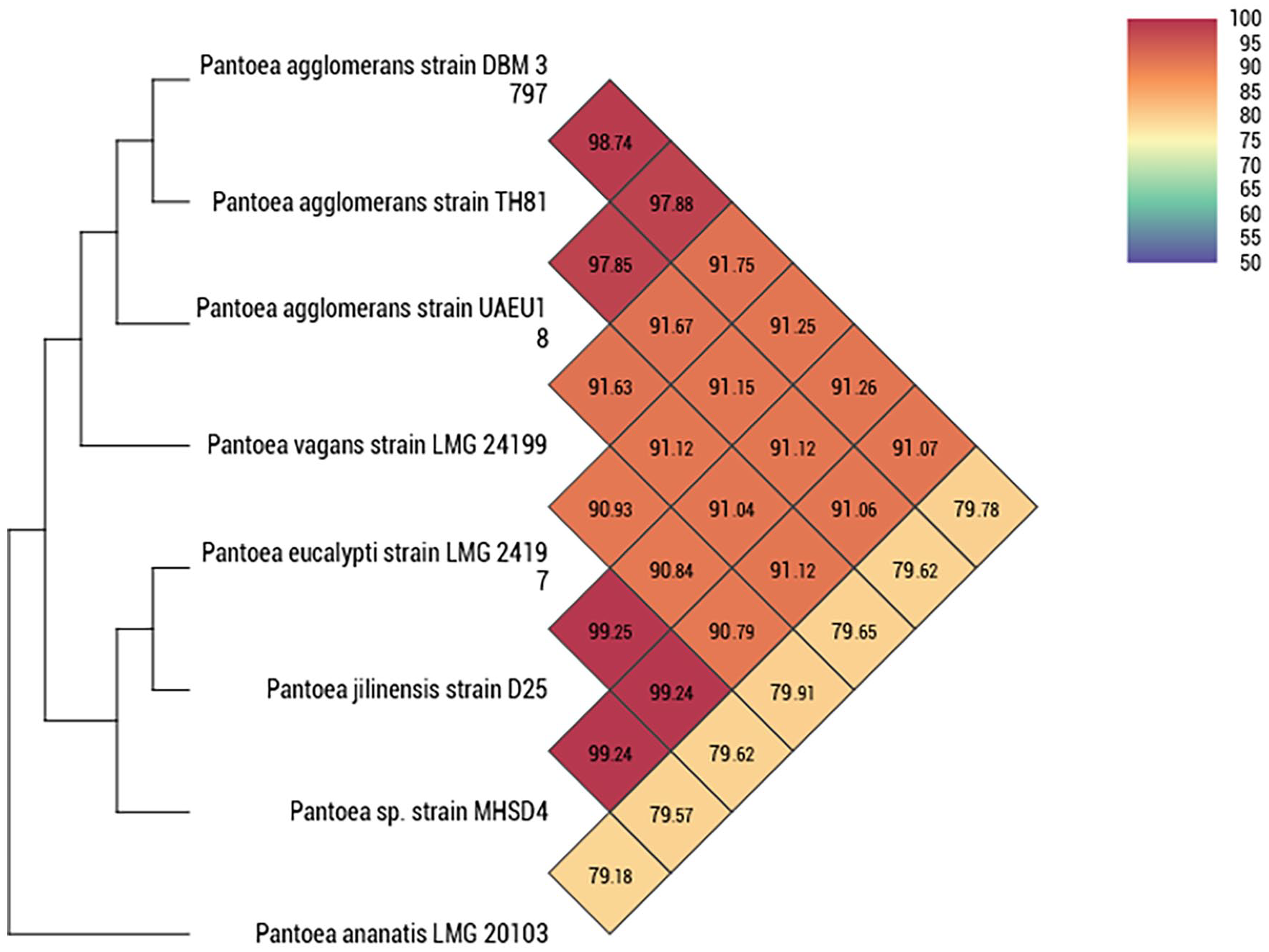
Heatmap generated with OAT software indicating OrthoANI values of

Venn diagram of the unique and shared genes of
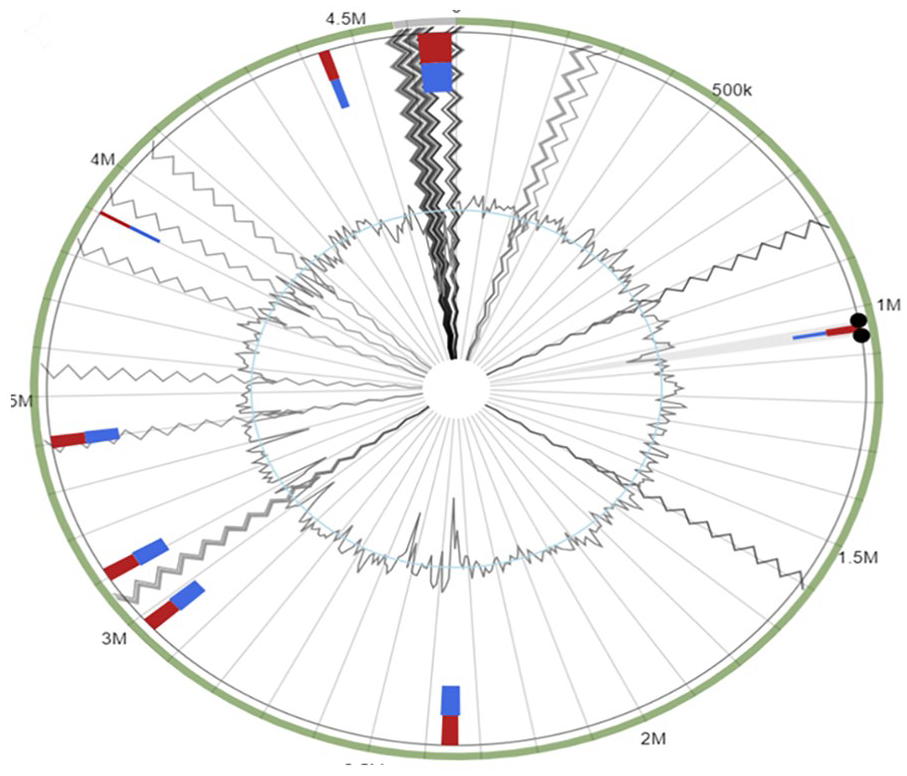
Genomic islands genes within
Biosynthesis pathways of phytohormones of
Genes responsible for Bioremediation that were identified from

In this study,
The presence of these genes may indicate biotechnological potential for this endophytic strain in terms of conferring resistance to host plants or as a biological key player for copper remediation in the environment. 14
High throughput sequencing of bacterial genomes allows for the identification and functional classification of genes within endophytic bacteria. This study reports on the draft genome of the bacterial endophyte,
Supplemental Material
sj-docx-1-evb-10.1177_11769343231217908 – Supplemental material for Draft Genome Sequence of Pantoea sp. Strain MHSD4, a Bacterial Endophyte With Bioremediation Potential
Supplemental material, sj-docx-1-evb-10.1177_11769343231217908 for Draft Genome Sequence of Pantoea sp. Strain MHSD4, a Bacterial Endophyte With Bioremediation Potential by Dimpho Michelle Morobane, Khuthadzo Tshishonga and Mahloro Hope Serepa-Dlamini in Evolutionary Bioinformatics
Footnotes
Author Contributions
Work was planned by S-DMH and executed by MDM, TK assisted with genome assembly, annotation and analysis.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Research Foundation of South Africa (Thuthuka Grant No. TTK210216586709).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
