Abstract
Rice (Oryza sativa) yield is correlated to various factors. Transcription regulators are important factors, such as the typical SHORT INTERNODES-related sequences (SRSs), which encode proteins with single zinc finger motifs. Nevertheless, knowledge regarding the evolutionary and functional characteristics of the SRS gene family members in rice is insufficient. Therefore, we performed a genome-wide screening and characterization of the OsSRS gene family in Oryza sativa japonica rice. We also examined the SRS proteins from 11 rice sub-species, consisting of 3 cultivars, 6 wild varieties, and 2 other genome types. SRS members from maize, sorghum, Brachypodium distachyon, and Arabidopsis were also investigated. All these SRS proteins exhibited species-specific characteristics, as well as monocot- and dicot-specific characteristics, as assessed by phylogenetic analysis, which was further validated by gene structure and motif analyses. Genome comparisons revealed that segmental duplications may have played significant roles in the recombination of the OsSRS gene family and their expression levels. The family was mainly subjected to purifying selective pressure. In addition, the expression data demonstrated the distinct responses of OsSRS genes to various abiotic stresses and hormonal treatments, indicating their functional divergence. Our study provides a good reference for elucidating the functions of SRS genes in rice.
Introduction
Rice (Oryza sativa) is considered as the major food of half the world’s population. Rice yield is not only multiplicatively determined by major agronomic traits, such as panicle number, grain per panicle and grain weight,1,2 but also impacted by environmental conditions. 3 Abiotic stresses, including drought, salinity, extreme temperatures, and wounding, as well as biotic stress, such as bacterial, fungal, and viral challenges, have great impacts on rice productivity and crop quality. 4 To adapt to extreme environments, genome-wide expression changes in plants have evolved in response to abiotic stresses. Transcription factors (TFs) critical to this process are also known as cis-acting elements, and they bind specifically to the promoters of target genes, playing significant roles in the regulation of the gene expression levels. 3 The structures and functions of TFs are important in regulating plant biotic and abiotic stress resistance. 5 TF gene families have expanded and evolved in plants to coordinate gene expression. SHORT INTERNODES (SHI)-related sequences (SRSs), which encode proteins with single zinc finger motifs, form a typical TF gene family. SRS proteins contain a putative zinc finger domain in a CX2CX7CX4CX2CX7 C arrangement, in which X represents variable amino acids. Rich glutamine residues adjacent to zinc finger domains have been found, as well as a short stretches of acid residues in the C-terminal third of the SHI proteins. 6 These features are important for their TF functions.
Multiple functions of SRS genes in plant organ growth, carpel development, photo-morphogenesis, metabolism-related plant architecture, photoperiodic regulation, and phytohormone responses have been reported. 7 For instance, the Arabidopsis SRS family contains 11 members, SHI, SRS3–8, STYLISH1 (STY1) and STYLISH2 (STY2), LATERAL ROOT PRIMORDIUM1, and another putative SHI protein. 7 The Arabidopsis SRS genes are highly sequence divergent, although they contain two conserved domains, one zinc finger and another IGGH domain. SHI and the closely related STY1 and STY2 genes act synergistically to promote gynoecium, stamen and leaf development, 8 and the overexpression of STY1, STY2, SHI or SRS7 inhibits stem elongation and the regulation of tapetum dehiscence. 9 In addition, SRS gene expression levels, such as that of the rice SHI gene, are correlated with plant stature through the gibberellin (GA) signal pathway. The function of OsSHI1 in modulating the transcriptional activity of IPA1 in rice was recently reported. 10
SRS genes have been identified and characterized in different plant species, such as Arabidopsis thaliana,6,8 O. sativa, 10 Populus euphratica, 11 and Hordeum vulgare. 12 However, our understanding of SRS gene family members and their evolutionary and functional characteristics in rice is limited. Here, based on the genomic and transcriptional data, we identified, characterized and functionally analyzed the SRS gene family in rice.
We identified only six SRS proteins in rice, which indicates that the SRS family is small. All of them contained another conserved domain-DUF702-that overlapped with the zinc finger domain. DUF stands for “domain of unknown,” which means their roles in a large number of genes remain unclear. 13 The functions of some DUF proteins have been elucidated in recent years, and some members are involved in plant development and stress responses. 14 The SRS genes were further divided into five major groups within the rice species having AA genomes, including three cultivars (indica, japonica and Oryza glaberrima) and six wild rice (Oryza nivara, Oryza rufipogon, Oryza barthii, Oryza meridionalis, Oryza glumaepatula, and Oryza logistaminata), rice species with BB genome (Oryza punctata) and rice species with FF genome (Oryza brachyantha). The gene structure, conservative motif, and phylogenetic analysis revealed that all the SRS genes in the same group have similar gene structures and motif arrangements. Japonica rice is the most studied and applied in agronomy; therefore, we thoroughly examined the gene features and structures of the OsSRS family. Segmental duplication appears to have played a crucial role in the expansion of the SRS gene family. The expression patterns in rice organs and tissues, as well as the expression responses to various hormones, demonstrated their divergent functions in accordance with their phylogenetic positions.
Materials and Methods
Data acquisition
The rice sub-species, maize, sorghum, Brachypodium genomic and proteomic sequences were downloaded from Phytozome (https://phytozome.jgi.doe.gov), Ensemble plants (http://plants.ensembl.org/index.html), and PlantTFDB (http://planttfdb.cbi.pku.edu.cn/) 15 ; the SRS protein sequences of Arabidopsis thaliana were obtained from the Arabidopsis genome database (TAIR 9.0 release, http://www.arabidopsis.org/) 16 ; and the OsSRS gene expression data were collected from the SRA database (https://www.ncbi.nlm.nih.gov/).
SRS gene identification and characterization
The hidden Markov model (HHM) file of the SRS domain (PF01542) was downloaded from Pfam protein family database (http://pfam.sanger.ac.uk/). 17 HMMER 3.0 was used to search the SRS genes from the maize, sorghum, Brachypodium, and Arabidopsis genome database. The default parameters were adopted and the cutoff value was set to 0.01. All candidate genes that may contain DUF247 domain or RING-like zinc finger domain based on HMMER 18 results were further examined by confirming the existence of SRS core sequences using the PFAM and SMART program (http://smart.embl-heidelberg.de/).19,20 Each potential gene was checked manually to ensure the conserved 28-peptide region of the predicted SRS domain. The incorrectly predicted genes were manually curated and the redundant sequences were discarded. Six transcripts were finally identified in the rice genome after comprehensive curation.
The other gene features such as protein length, molecular weight, isoelectric point, the possible sub-cellular localizations and trans-membrane area, were predicted using ExPASy (https://www.expasy.org/). 21
DNA-binding domain conservation analysis of OsSRS proteins
The complete protein sequences of OsSRS were retrieved by Quick Fasta Extractor or Filter software 22 (https://github.com/CJ-Chen/TBtools). Then, we identified all highly conserved domain for all OsSRS members by multiple sequence alignment analysis using ClustalW (https://www.genome.jp/tools-bin/clustalw) and visualized in JavieW software (http://circinus.ebi.ac.uk:6543/jalview/). 23
Phylogenetic Analysis
The SRS proteins from rice sub-species and other monocot plants, dicot plant A. thaliana were analyzed using ClustalW and phylogenetic trees were constructed using MEGAX by the neighbor-joining method, with the following parameters: Poisson model, partial deletion, and 1000-bootstrap replicates. 24
Gene structure of OsSRS proteins
The positional information of the OsSRS genes (intron-exon GTFF/GFF3 files and genome sequence files) was loaded into the Gene Structure Display Server (GSDS v2.0, available online: http://gsds.cbi.pku.edu.cn/) to obtain information on UTR/Intron/CDS structure. 25
Conserved motif detection
The conserved motifs of SRS proteins were detected using Multiple Expectation Maximization for Motif Elucidation (MEME v4.11.2, Available online: http://meme.nbcr.nrt/meme/) 26 with the parameters set as maximum width 5, minimum width 150, and maximum number 10.
Chromosomal localization, detection of gene duplication events, syntenic analysis and estimation of synonymous(Ks) and non-synonymous(Ka) substitution per site and their ratio
According to the gene positional information, we draw a map of the distribution of OsSRS genes throughout the rice genome using Amazing Gene Location software of TBtools. 22 The OsSRS gene pairs resulting from segmental duplication were linked by red lines.
Duplicated gene pairs of OsSRS genes within the japonica rice derived from segmental or tandem duplications were identified in the rice genome based on the method in the Plant Genome Duplication Database. An all-against-all BLASTP comparison with an E-value ⩽ 1 x 10-20 was performed for syntenic clustering determined by MCscanX (E-value ⩽ 1 x 10-20). 27 Pairs from segmental duplications were used to estimate Ka, Ks, and their ratio. The coding sequences from segmentally duplicated OsSRS genes were aligned by ClustalW and trimmed by Gblocks. The software of TBtools was then used to compute Ka, Ks values for each pair following the YN model. 28 The result is that if Ka/Ks > 1, it is positive selection pressure; if Ka/Ks = 1, it is neutral selection or natural selection pressure; if f Ka/Ks < 1, it is purification selection pressure. 29
Plant growth conditions, RNA sampling and qRT-PCR analysis of OsSRS genes
Indica rice IR26 seeds were grown at 28℃ and 12/12 hours light/dark period in the environmentally controlled greenhouse. Different organs of rice seedlings were collected after 20 days at the tillering stage. All the samples were kept in liquid nitrogen, and stored at -80℃ for further use. High-quality RNA was extracted using TRI REAGENT RT reagent (Molecular Research Center, Inc.) following the manufacturer’s instructions (http://www.mrcgen.com/tri-rt.ht).
The real-time PCR (qPCR) assay was completed to detect the relative expression of SRS genes in different rice organs as previously described. 30 Original expression data for OsSRS genes under different treatments including GA, paclobutrazol (PB), salt (PRJEB4672) and drought stress (PRJNA272723) were retrieved from NCBI SRA (Available online: https://www.ncbi.nlm.nih.gov/). All the data were analyzed using RNA-seq analysis process, including TopHat, cufflinks, cuffmerge, cuffcompare, cuffquant, and using cuffnorm to perform normalization and log transformation. Heat-map was produced by Amazing Heat Map software of serials TBtools 22 with average linkage hierarchical clustering.
Results
Identification and characterization of OsSRS genes in O. sativa subsp. japonica
To extensively understand the SRS gene family in japonica rice, we searched MSU rice databases with Pfam 17 Number PF05142 and verified by SMART software.19,20 Eventually, only six candidate DUF702 genes in japonica rice were identified; these genes coincidently were all later found to contain the SRS gene.
In addition, we systematically identified the basic protein properties of SRS candidates, including domain position, isoelectric point, molecular weight, cellular localization, and transmembrane region (Table 1). The length of the SRS proteins ranged from 303 aa (LOC_Os08g43410) to 360 aa (LOC_Os06g49830). Correspondingly, the molecular weights ranged from 30.69 kDa (LOC_Os08g43410) to 34.95 kDa (LOC_Os06g49830), and the predicted isoelectric points varied from 7.27 (LOC_Os01g71490) to 8.97 (LOC_Os09g36160). Four proteins encoded by SRS members were predicted to localize on the cell membrane and in the nucleus, while the other two were only predicted to localize in the nucleus. The cell membrane localization indicated that these proteins would have contact with external signals outside of the cells in the environment; however, there were no trans-membrane regions detected among these rice SRS proteins. The locations of the DUF702 and SRS domains were identified using SMART and positions of domains in similar SRS proteins, such as LOC_Os01g72490.1 and LOC_Os01g72490.2 (the model for alternative splicing of the SRS domain). LOC_Os08g43410 and LOC_Os09g36160 (phylogenetically closely related), encoded by the same or closely related genes, were similar, but quite different from the proteins encoded by phylogenetically alienated related genes, such as LOC_Os05g32070 and LOC_Os06g49830.
The information of SRS proteins in rice.
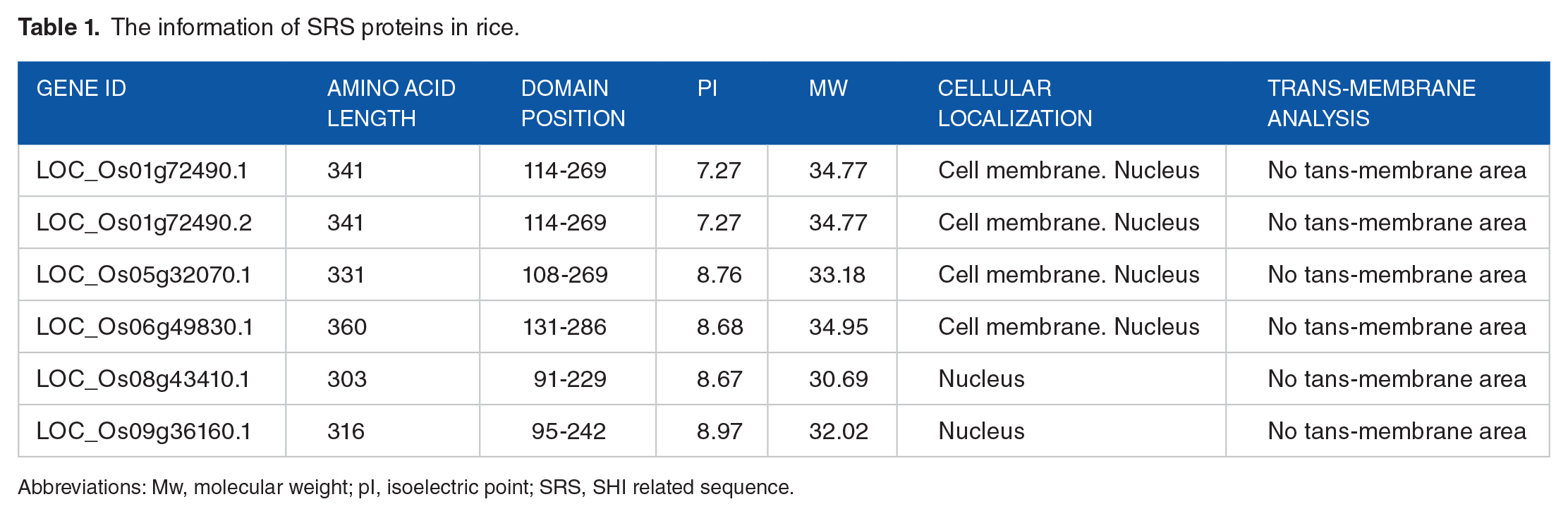
Abbreviations: Mw, molecular weight; pI, isoelectric point; SRS, SHI related sequence.
Multiple sequence alignment and phylogenetic analysis
SRS TFs usually have a highly conserved 138–161-aa region in the N-terminus, which contains a putative RING-like zinc finger unique to SRS proteins. The zinc finger protein has a C-X2 C-X7 C-X4 C-X2 C-X7 C arrangement (where X represents variable amino acids), and the conserved motif is essential for its regulatory function. In this study, the OsSRS family’s conserved domains were analyzed. Multiple alignments of the OsSRS protein sequences against SRS-binding domains revealed that they were highly conserved. In particular, a highly conserved domain structure C-QD-C-GNQAKKD-C-GHQR-C-RT-C-CKSRGFD-C was found in the OsSRS family (100% alignment) in a 28-aa region having the structure C-X2 C-X7 C-X4 C-X2 C-X7 C (Figure 1).

Multiple alignments of six OsSRS protein sequences against the putative zinc finger domain. The identical zinc finger domains of eight cysteine residues are shown on the top, and the dashed rectangle indicates the DUF702 domain area.
In addition, the domain positions in the OsSRS protein family were correlated with the members’ positions in the phylogenetic tree. The DUF702 domain contains two other conserved domains in the rice SRS proteins. Based on the phylogenetic tree, the domain structures in members of the same sub-group were nearly identical (Figure 2A).

The gene features of OsSRS genes. (A) The domain structure and phylogenetic tree of OsSRS protein family. (B) OsSRS gene structures and phylogenetic tree. UTR, CDS, and introns were depicted by filled blue boxes, yellow boxes, and single black lines. The phylogenetic tree was constructed by MEGAX with bootstrap 1000 times. (C) Motif 1, corresponding to C-X2 C-X7 C-X4 C-X2 C-X7 C zinc finger conserve structure, the detailed motif structure was shown in Supplementary Table S4. CDS indicates coding sequence; UTR, un-translated region.
To explore the phylogenetic relationships of OsSRS proteins, we carried out a phylogenetic analysis of SRS proteins from rice sub-species. A phylogenetic tree was constructed to include three cultivar rice species, six wild rice species, one BB-genome rice species, and one FF-genome rice species (Figure 3, Supplementary Table S1). For each gene, we chose all the proteins formed by alternative splicing. All the rice species contained five SRS genes, except O. barthii, O. nivara, O. longistaminata, and O. brachyantha. The phylogenetic tree revealed that all the SRS proteins in the various rice sup-species clustered into five major groups (A, B, C, D and E), and they were considered to be distinct based on bootstrap values greater than 80.

Phylogenetic relationships among AA genome rice with three cultivars as O. sativa japonica, O. sativa indica, O. glaberria, six wild rice as O. nivara, O. rufipogon, O. barthii, O. longistaminata, O. meridonalis, and O. glumaepatula. one BB genome rice O. punctate and one FF genome rice O. brachyantha. The neighbor-joining tree was created by MEGAX with bootstrap value setting at 1000. All the SRS proteins were divided into five major groups. Different color represents different groups divided based on the bootstrap values: the small red star represents values between 50% and 80% and the big red star represents bootstrap values between 80% and 100%. Detailed information was listed in Supplementary Table S1. SRS indicates SHI related sequence.
Group A contained 14 SRS proteins, groups B, C and E contained 10 protein members each, and group D had 7 members. Interestingly, the SRS proteins in the rice species were distributed evenly. In group A, SRS proteins from O. glaberrima and O. barthii clustered together with indica, derived from O. nivara, had a closer relationship with japonica, derived from O. rufipogon. The other three SRS proteins from the AA-genome rice species O. logistaminata, O. glumaepatula, and O. meridionalis were further apart. All the SRS proteins from AA-genome rice species clustered together, but SRS proteins of the BB- and FF-genome rice species O. punctata and O. brachyantha, respectively, clustered as out groups from AA-genome rice species. The results were similar for the B, C, D and E groups.
Furthermore, we constructed a phylogenetic tree of SRS proteins from one dicot plant A. thaliana and other four monocot plants rice, maize, sorghum, and Brachypodium (Figure 4A). The SRS proteins were divided into 11 major groups, from group A to K, supported by bootstrap values greater than 80%. Group A was the largest group, containing three, four, and two SRS proteins in rice, Brachypodium, sorghum, and maize, respectively. Groups B and C each contained seven SRS members with three maize SRS proteins in both clades. Group B also contained two, one and one members from rice, sorghum, and Brachypodium, respectively, while group C also contained one, two and one members from rice, sorghum, and Brachypodium, respectively. Group D contained three maize members and one member each from Brachypodium and sorghum. The groups from E to K were all A. thaliana members and contained no monocot protein members, except one Brachypodium member in group I. The SRS proteins in monocot and dicot plants diverged, which could result from species-/lineage-specific gene gain or loss events. We further checked the gene ontology (GO) annotations of these japonica OsSRS and Arabidopsis proteins. Some groups had specific annotations, such as group E, which was involved in “stigma development” and “xylem and phloem pattern formation.” In addition, group I contained only one A. thaliana (AT1G32730) member that annotated as “located in membrane.” Groups F and G contained members that were annotated as playing roles in “root development” and “gynoecium development” compared with the other groups, respectively. AT1G19790 was involved in “sequence-specific DNA binding” in group G (Supplementary Table S2). All these specific annotations in different groups from E to K imply possible divergent functions of the SRS genes in these groups. The phylogenetic tree showed that the SRS genes in monocot plants were duplicated several times before the divergence of these four species, and were highly conserved in rice, Brachypodium, maize, and sorghum. The SRS proteins from the monocot plants may be more closely related to each other evolutionarily to those from the dicot plant A. thaliana. In addition, the SRS gene numbers and proteins were also analyzed in Figure 4B. There were 11 gene members and 14 protein members in Arabidopsis; however, the genes members were only one less than protein numbers in monocot plants we had detected in this study. For example, in rice, there were only 5 SRS genes but 6 protein members, and in maize, 10 SRS proteins were found but 9 SRS genes were detected. Interestingly, the SRS members in dicot plants such as Arabidopsis were more than in monocot plants such as rice, maize, which may be because the annotation of the monocot genome was not too perfect.

Phylogenetic relationships among four monocot plants including rice, Brachypodium, maize and sorghum, and one dicot plant Arabidopsis thaliana. The neighbor-joining tree was created by MEGAX with bootstrap value setting at 1000. The SRS proteins from monocot plants were conserved and divided into four clades (clades A to D) and the rest of the SRS proteins were mainly from A.thaliana, except for one member from Brachypodium. The phylogenetic tree indicates the SRS protein function divergence between monocot and dicot plants. Different color represents different groups divided based on the bootstrap values: the small red star represents bootstrap values between 50% and 80% and the big red star represents bootstrap values between 80% and 100%. Detailed information was listed in the Supplementary Table S3. SRS indicates SHI-related sequence.
Gene structure and chromosome location analysis
Gene organization is important in the evolution of gene families, and the gene structure is the backbone. Therefore, we compared the introns and exons of OsSRS genes by aligning the genomic DNAs (Figure 2B). The results were consistent with the phylogenetic tree in that the intron/exon structure patterns were similar within the same group. For example, based on the gene structures, LOC_Os08g43410 and LOC_Os09 g 36160 contained 1 intron, 2 CDSs and 2 UTRs, which was different from the structures of the other groups. LOC_Os06g48830 and LOC_Os05g32070 had different structures, while LOC_Os01g72490.1 and LOC_Os01g72490.2 were two spliced forms of LOC_Os01g72490.
Our classification was also verified by the conserved motif analysis. All of the SRS proteins from various rice species were loaded into the MEME analysis tool to identify the conserved motifs. Information on all 10 motifs observed with statistically significant E-values below 5.10E-189 can be found in Supplementary Table S4 and Figure 5. The motifs identified by MEME were between 9 and 52 aa in length. One of the motifs (motif 1), corresponding to the C-X2 C-X7 C-X4 C-X2 C-X7 C zinc finger conserved structure, was found in all the SRS proteins in rice (Figures 2 C and 5). Motif 1 was also common in all the SRS proteins from various sub-species, except most of the SRS proteins from the BB- and FF-genome species O. punctata and O. brachyantha, respectively, making it a highly conserved motif of the SRS family. In the phylogenetic tree of SRS proteins from rice subspecies, groups C and D shared a close relationship with the same motif types, but their motif arrangements were different. For example, group C had a motif arrangement “10-3-8-1-5-2-8-4,” while group D had a motif arrangement “10-5-8-1-3-2-8-4.” However, some members did not comply with these arrangements, maybe because of the different genome types, such as OB09G24020.1 and OPUNC09G15790.1. The SRS proteins from group A contained the largest number of motifs and, specifically, contained both motifs 9 and 6. Nearly three of the SRS proteins that did not contain motif 1 were from group E. In group B, members had complicated motif patterns compared with the other groups, and the motif arrangement “3-2-8-4-7” was specific to this group.

Conserved motifs of SRS proteins in rice subspecies. The distribution of 10 conserved motifs in SRS proteins. Different colors represent the different groups (A to E) based on their protein sequences constructed a phylogenetic tree. SRS indicates SHI related sequence.
We have also investigated the chromosomal distribution of OsSRS genes which were plotted using gene location software (https://github.com/CJ-Chen/TBtools) (Figure 6). The results indicated that the OsSRS genes were distributed evenly in five chromosomes that were chromosome 1,5,6,8,9.

Chromosomal locations and gene duplication events of OsSRS. The scale on the left is basepair (bp). The SRS gene pairs of segmental duplications are linked by red lines. Detailed information of duplicated pairs is described in Supplementary Table S5. SRS indicates SHI-related sequence.
Gene duplications and synteny analysis of SRS genes
Gene duplication plays an important role during the process of plant evolution. Gene family expansion and genomic evolutionary mechanisms mainly depend on gene duplication events. In this study, gene duplication events were identified in the SRS gene family. The chromosomal distribution of OsSRS genes was plotted using gene location software (https://github.com/CJ-Chen/TBtools) (Figure 6). The duplication events were also examined, but no tandem duplications were detected in SRS genes from japonica. As shown in Figure 6, two segmental duplications occurred between two pairs of OsSRS genes. These two segmentally duplicated pairwise genes were evolutionary close together in the phylogenetic tree and had very high sequence similarities (Figure 2B), which indicated that the SRS gene expansion in rice mainly resulted from segmental duplications, while tandem duplications played no role. Gene duplicates are usually closely adjacent to each other in the same chromosome, a phenomenon that is speculated to result from unequal crossing over.
To further understand the evolutionary selection on this gene family, we calculated the non-synonymous substitution rate (Ka), synonymous substitution rate (Ks), and Ka/Ks for these two pairs of duplicated genes (Supplementary Table S5). These two pairwise genes’ Ka/Ks were less than one, which implied that the SRS genes in rice tended to be subjected to purifying selection. This may play an important role in the possible functional divergence among OsSRS genes.
To further infer the phylogenetic mechanisms of rice SRS family, we constructed three comparative syntenic maps of rice associated with three representative species including one dicot plant Arabidopsis.thaliana, two monocot plants Brachypodium and sorghum (Figure 7). Notably, all the rice SRS members showed syntenic relationship with those in Brachypodium and sorghum, but only one for dicot plant Arabidopsis and rice (Supplementary Table S6). The numbers of orthologous pairs between the other three species (Arabidopsis, Brachypodium, and sorghum) were 1, 6, and 6, which were LOC_Os01g72490.1-AT3G54430.1, LOC_Os01g72490.1-Bradi2g61120.1, LOC_Os01g72490.1-Bradi2g26823.1, LOC_Os05g32070.1-Bradi2g61120.1, LOC_Os06g49830.1-Bradi1g34990.1, LOC_Os09g36160.1-Bradi3g42197.1, LOC_Os09g36160.1-Bradi4g36030.1, LOC_Os01g72940.1-Sobic.003G429800.1, LOC_Os01g72940.1-Sobic.009G122600.1, LOC_Os05g32070.1- Sobic.009G122600.1, LOC_Os06g49830.1-Sobic.010G261700.1, LOC_Os09g36160.1-Sobic.002G274500.1, LOC_Os09g36160.1-Sobic.007G179000.1. All the rice SRS members were found to be associated with 6 gene pairs such as related to the phylogenetic relationship between rice and other 2 monocot species. Interestingly, the other 4 rice SRS members were not identified as collinear gene pairs between rice and Arabidopsis. Combined with the result of phylogenetic analysis, it may indicate that these orthologous pairs formed after the divergence of dicotyledonous and monocotyledonous plants. And the common identified between rice and 2 other monocot plants also indicated that these orthologous pairs may already exist before the ancestral divergence.

Syntenic analysis of SRS genes between rice and other three representative plant species. The gray lines indicate the collinearity blocks within japonica rice and other plant genomes. (A) Collinearity between rice and Arabidopsis. (B) Collinearity between rice and Brachypodium. (C) Collinearity between rice and sorghum. The detected gene pairs were also presented. at indicates Arabidopsis chromosome; Bd, Brachypodium chromosome; os, rice chromosome; sb, Sorghum chromosome; SRS, SHI-related sequence.
To better understand the evolutionary constraints acting on the SRS gene family, the Ka/Ks ratios of the SRS gene pairs were calculated. All the orthologous pairs had Ka/Ks < 1 except the Ka/Ks ratio between rice and Arabidopsis with unknown reasons, suggesting that the SRS family might have experienced strong purifying selective pressure during evolution.
Expression patterns of OsSRS genes under different abiotic stresses and hormonal treatments
To examine the possible divergent functions of OsSRS genes, we investigated their expression levels in different organs including root, shoot, leaf, panicle, seed, anther, and pistil with qRT-PCR. The results showed that LOC_Os01g72480 expressed higher than other members in the organisms of root, shoot, leaf, and panicle. All the SRS genes could be detected in the above-mentioned organisms except leaf and seed. In leaf, only three genes were detected, and LOC_Os01g72480 expressed highly, otherwise, LOC_Os05g32070 and LOC_Os08g43410 expressed extreme lowly. Moreover, LOC_Os08g43410 expressed extreme highly in seed, but the other three genes (LOC_Os05g32070, LOC_Os06g49830 and LOC_Os09g36160) were not detected in seed (Figure 8A to G).

Expression patterns of OsSRS genes in different organs and under different treatments. (A) to (G), Relative expression of SRS genes in different organs in rice including root, shoot, leaf, panicle, seed, anther and pistil; (H) The heatmap was generated by Amazing Simple HeatMap of TBtools software using the expression data of OsSRS genes and normalized log2 transformed values were used with hierarchical clustering represented by the color scale (1-6). Green indicates as low expression and the red indicates high expression. The samples were leaves of IR29 rice seedlings under GA, PB, salt (different times), and drought treatments. Detailed information of expression data is described in Supplementary Tables S6 and S7. GA indicates gibberellin; PB, paclobutrazol; SRS, SHI related sequence.
In addition, we also checked the SRS gene expression levels under various abiotic stress conditions and during hormonal treatments in the indica rice IR26 using the public transcriptome data from the NCBI SRA database (Supplementary Tables S7 and S8). We used a heat-map to visualize the global transcriptome profile of these OsSRS genes, and the results present the diverse responses to various treatments, including GA, PB, salt, and drought (Figure 8H).
In the GA- and PB-treatment experiments (gene expression data collected from leaves of rice seedlings), compared with the negative control mock, one gene, LOC_Os01g72490, was induced by GA but depressed by PB, while the other members were slightly depressed by GA, and the pattern was opposite for PB. We further checked the GO annotations of their Arabidopsis orthologs and found that they were annotated as “auxin responsive,” “root and pollen development,” “anther development” and “multicellular organism development,” which may imply possible functional roles for these genes.
We also further analyzed the expression patterns of OsSRS genes in 300 mM NaCl- and drought-stressed seedlings at the following different times: 1, 2, 5, 10 and 24 hours after salt stress within a day. The expression data were calculated between abiotic treatments and controls. Some of them showed significant changes under salt-stress conditions. LOC_Os01g72490 appeared to be down-regulated 2 and 24 hours after treatments but up-regulated 5 hours after the salt treatment, which indicates that its expression first increased, then decreased and increased again. LOC_Os06g49380’s expression was depressed by the salt treatment compared with the mock. The expression levels of other members did not change under salt-stress conditions. However, under drought-stress conditions, one gene (LOC_Os01g72490) was repressed, two genes (LOC_Os08g43410 and LOC_Os09g36160) were induced, one gene (LOC_Os05g32070) was insensitive and one gene (LOC_Os06g49830) was not detected. Notably, gene expression patterns consistent with the gene positions in the phylogenetic tree.
Discussion
The SRS gene family encodes proteins containing a conserved RING finger-like zinc finger domain at the N-terminus. In dicot plants, such as A. thaliana, the SRS gene family has been extensively studied in recent years. Members of this family are involved in apical gynoecium development and have important functions in stigma development. 7 The mechanism of SRS action has also been reported in some plants, and genes, such as AtSTY1, have transcriptional activity in auxin biosynthesis by directly binding to cis-elements in the promoter the YUCCA4 gene. In addition, the expression of SHI could increase the bioactive GA levels, 6 which are correlated with crop yields, such as “semi-dwarf 1” that led to the “green revolution.” Thus, the SRS gene family plays various roles in coordinating multiple hormonal pathways that modulate the growth and developmental networks. As a result, studies on the rice SRS gene family will provide insights into increasing rice yields.
A genome-wide gene family analysis is a basic key step in studying the gene functions in various developmental and hormonal pathways, along with the gene structure, evolution, and expression patterns. 31 The SRS genes play key roles in the regulation of growth and responses to diverse abiotic and biotic stresses. 10 ,32-35 Until now, the SRS genes have been identified in different plant species, but not systematically in rice. Therefore, we conducted a comprehensive analysis of the OsSRS family in rice, along with other rice sub-species, to determine their phylogenetic relationships and potential functions.
In total, we identified 1–5 SRS genes in 11 rice sub-species, consisting of 3 cultivars, 6 types of wild rice, and 2 other rice-genome types. Compared with the other TF families, the SRS gene family is very small. In the 11 rice sub-species, SRS proteins were divided into 5 groups (A–E), and the SRS members from these types of rice were distributed evenly, which indicated possible divergent functions. In addition, we constructed a phylogenetic tree of the SRS proteins from four monocot plants, japonica rice, maize, sorghum and Brachypodium, and one dicot, A. thaliana. Correspondingly, we discovered that members of groups E to K were syntenic with AtSRS proteins, except one Brachypodium protein. Almost all the monocot SRS proteins clustered together and were separated from those of the dicot plant, which implied that SRS proteins conserved within monocot and dicot plants. Notably, the Arabidopsis SRS proteins may be more diverse among themselves, which is consistent with the divergent functional research reported in recent decades. 36 In a monocot grass species (Poaceae), the number of SRS gene members does not correlate with genome size. However, the gene structures and conserved motif distribution patterns were consistent with the genes’ positions in the phylogenetic tree. Thus, SRS proteins may be conserved within the species but might have undergone species-/lineage-specific gain or loss events.
To determine whether the duplication events play roles in gene expansion or gene family variation, we have detected duplication events among the OsSRS gene family and the three other monocot species. Two segmental duplication events were found, and all the OsSRS members were associated with Brachypodium and sorghum SRS members. In addition, the gene orthologous pairs occurred in the same phylogenetic groups, such as LOC_Os01g72490-Bradi2g61120.1 and LOC_Os01g72490-Sobic.003G429800.1, which belonged to Group A (Figure 4A, Supplementary Table S6). Furthermore, the Ka/Ks ratios of all the involved genes pairs were less than one, implying that purifying selection played an important role in the evolution of OsSRS genes.
The high-throughput expression data analysis confirmed the conservation of OsSRS genes. OsSRS genes showed diverse responses to different abiotic stresses and hormonal treatments. One SRS gene (LOC_Os01g72490) could be induced by GA but depressed by PB, which indicated that GA and PB activated antagonistic mechanisms. Three SRS genes (LOC_Os08g43410, LOC_Os09g36160, LOC_Os01g72490) also responded to drought and salt stresses, suggesting their participation in the regulation of various stress- and hormone-related responses. Our work provides the first endeavor for elucidating the OsSRS genes’ detailed functions in plant growth, development and regulation, and also for better understanding the evolution of SRS genes in plants.
Conclusion
In conclusion, a total of only 5 OsSRS genes were identified from japonica rice, and SRS genes in other species are normally less than 10. This gene family is small but plays important roles in the growth, development, and biotic and abiotic stress of plants. And SRS proteins from 11 rice sub-species and other 3 monocot plants, A.thaliana were also analyzed. In different rice subspecies, their number of SRS genes is the same, but the number of proteins encoded by these genes is different, suggesting their potential functions in diversify of rice. Genome comparison revealed that the OsSRS gene expansion in rice mainly resulted from segmental duplications, suggesting their contribution to the recombination of the SRS genes in rice. And the expression profiles also suggested that two OsSRS genes (LOC_Os01g72490 and LOCOs05g32070) were involved in different hormonal and abiotic stress treatments. LOC_Os01g72490 has broadly participated in various treatments; LOC_Os06g489830 was the only response to salt stress. In summary, this study provides the first view for a better understanding of the SRS genes in rice.
Supplemental Material
Table_S1 – Supplemental material for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice
Supplemental material, Table_S1 for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice by Jun Yang, Peng Xu and Diqiu Yu in Evolutionary Bioinformatics
Supplemental Material
Table_S2 – Supplemental material for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice
Supplemental material, Table_S2 for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice by Jun Yang, Peng Xu and Diqiu Yu in Evolutionary Bioinformatics
Supplemental Material
Table_S3 – Supplemental material for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice
Supplemental material, Table_S3 for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice by Jun Yang, Peng Xu and Diqiu Yu in Evolutionary Bioinformatics
Supplemental Material
Table_S4 – Supplemental material for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice
Supplemental material, Table_S4 for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice by Jun Yang, Peng Xu and Diqiu Yu in Evolutionary Bioinformatics
Supplemental Material
Table_S5 – Supplemental material for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice
Supplemental material, Table_S5 for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice by Jun Yang, Peng Xu and Diqiu Yu in Evolutionary Bioinformatics
Supplemental Material
Table_S6 – Supplemental material for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice
Supplemental material, Table_S6 for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice by Jun Yang, Peng Xu and Diqiu Yu in Evolutionary Bioinformatics
Supplemental Material
Table_S7 – Supplemental material for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice
Supplemental material, Table_S7 for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice by Jun Yang, Peng Xu and Diqiu Yu in Evolutionary Bioinformatics
Supplemental Material
Table_S8 – Supplemental material for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice
Supplemental material, Table_S8 for Genome-Wide Identification and Characterization of the SHI-Related Sequence Gene Family in Rice by Jun Yang, Peng Xu and Diqiu Yu in Evolutionary Bioinformatics
Footnotes
Acknowledgements
We thank Catherine Perfect, MA (Cantab), from Liwen Bianji, Edanz Editing China (![]() ), for editing the English text of a draft of this manuscript. We also thank Phd student Jing Li from XTBG for providing advice and help for analysis data. We thank the central lab of the public technical service center for providing qPCR devices.
), for editing the English text of a draft of this manuscript. We also thank Phd student Jing Li from XTBG for providing advice and help for analysis data. We thank the central lab of the public technical service center for providing qPCR devices.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (31360330), “One-Three-Five” Strategic Planning of Chinese Academy of Sciences (2017XTBG-T02).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
P.X. and D.Y. conceived and supervised this study; J.Y. designed, carried out the experiments and analyzed the data, writing—original draft preparation. All authors reviewed, edited, and approved the final manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
