Abstract
Protein-protein interactions (PPIs) play a crucial role in the life cycles of living cells. Thus, it is important to understand the underlying mechanisms of PPIs. Although many high-throughput technologies have generated large amounts of PPI data in different organisms, the experiments for detecting PPIs are still costly and time-consuming. Therefore, novel computational methods are urgently needed for predicting PPIs. For this reason, developing a new computational method for predicting PPIs is drawing more and more attention. In this study, we proposed a novel computational method based on texture feature of protein sequence for predicting PPIs. Especially, the Gabor feature is used to extract texture feature and protein evolutionary information from Position-Specific Scoring Matrix, which is generated by Position-Specific Iterated Basic Local Alignment Search Tool. Then, random forest–based classifiers are used to infer the protein interactions. When performed on PPI data sets of
Introduction
Proteins play significant roles in the life activities of cells and organisms, such as neurotransmission, DNA replication, and cycle control. Most of the diversity of cellular functions is based on protein-protein interactions (PPIs). Detecting PPIs is highly critical for the exploration of biological cellular mechanisms. With the advent of high-throughput techniques, such as mass spectrometric protein complex identification, 1 protein chip, 2 and yeast 2-hybrid system,3,4 considerable PPI data have been generated. However, high-throughput experiments are usually accompanied by high false positive and false negative rates and high cost. Moreover, these methods can hardly predict the whole PPI networks. 5 Under this situation, developing a novel computational method to predict unknown PPIs is more urgent than adopting the traditional experimental method to identify PPI.6,7
It is important to make full use of available PPI experimental data to develop computational methods. Many PPI databases, such as Human Protein References Database (HPRD), 8 Database of Interacting Protein (DIP), 9 and Molecular INTeraction database (MINT), 10 have been built after a number of experiments depicting PPI network. However, there are differences in protein structure information,11,12 protein domains, and so on. With new protein amino acid sequence data explosively growing, computational methods are urgently needed to detect the information of protein sequence.
In recent years, a number of computational methods have been proposed to extract the feature vectors mainly from the amino acid sequence.13 -16 The discriminative feature can improve the performance of a classification model, and some computational methods were based on Chou’s pseudo amino acid composition (PseAAC)17 -19 that retains the information of protein sequence, although it only considers the influence of 3 kinds of characteristics. Furthermore, some new methods on feature vector extraction are based on kernels. The method proposed by Jaakkola et al 20 is the first to use Fisher kernel to detect homology. Shen et al 21 proposed the support vector machine (SVM)-based method to predict PPIs. Leslie et al 22 put forward the mismatch string kernel method, which detects protein amino acid sequence at a lower computational cost. The difference between a PseAAC-based method and a kernel-based method lies in the way of extracting the feature information, with the first extracting the feature directly from the protein sequence and the second retaining some prior information and extracting feature vectors more effectively.
In general, most of the computational methods use machine learning algorithms combining various descriptors of proteins. Concerning different kinds of protein data, the main existing computational approaches can be divided into 2 categories: one uses information from the structure of proteins and genomic context; the other uses information from protein sequences. Moreover, newly discovered protein sequences grow exponentially in many different types of databases, and to shorten the gap between known protein sequence data and their interaction statuses, it is important to develop computational methods that directly use the information in protein sequences.
In this work, a novel computational method for predicting PPIs from an amino acid sequence based on a random forest (RF)
23
classification and a Gabor feature descriptor was proposed. The major improvement of this method is that it extracts protein sequence features through Gabor texture representation. Specifically, we adopted the Gabor feature representation on a Position-Specific Scoring Matrix (PSSM)24,25 to extract the evolutionary information from protein sequence, and then a classification RF is applied to infer the PPIs. In this way, each protein sequence is represented as a PSSM. To obtain more feature descriptors, we use a Gabor descriptor to extract features in each protein PSSM, and then each protein sequence is represented by 100-dimensional feature vectors. Two corresponding feature vectors would be joint together and represent a protein pair as a 200-dimensional feature vector. Finally, we used RF as a machine learning classifier for classification. The method was adopted for 3 PPI data sets from
Materials and Methods
Golden standard data sets
From the public DIP, we collected
Two other PPI data sets were also collected. The first PPI data set was collected from the HPRD. The protein pairs with more than 25% sequence identity were removed. We constructed the golden standard positive data set with the remaining 3899 protein-protein pairs of experimentally verified PPIs from 2502 different human proteins. Following previous work,
28
we assumed that proteins in different subcellular compartments would not interact with each other. Therefore, 4262 protein pairs from 661 different human proteins were set as the golden negative data set. The complete human data set consists of 8161 protein pairs. The other PPI data set was constructed with 2916
Position-Specific Scoring Matrix
The PSSM is widely used in various biological research works, such as studies of subcellular localization, disordered regions, and protein secondary structure. The PSSM also has great potential in extracting evolutionary information from amino acid sequences. In this work, each protein sequence would be converted into PSSM by adopting a Position-Specific Iterated Basic Local Alignment Search Tool (PSI-BLAST). 24 The PSSM can be represented as follows:
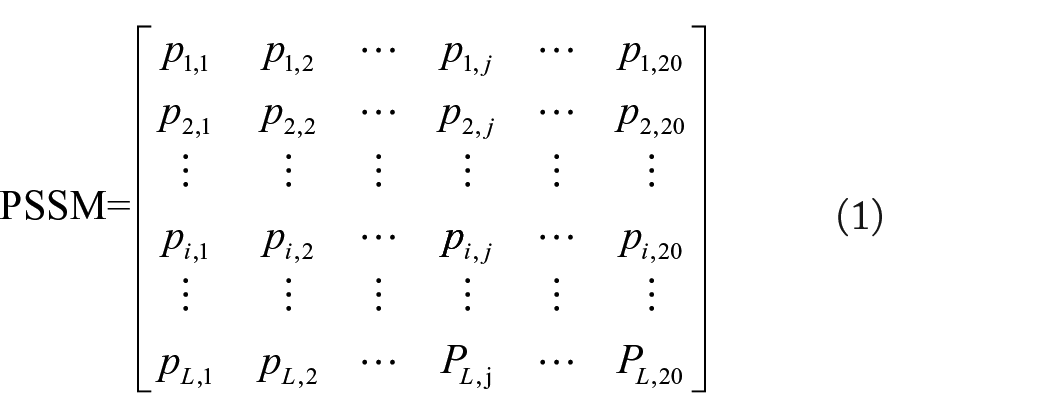
where
Gabor filter–based feature
First proposed by Gabor, 30 the Gabor filter is very similar to the visual stimulus response of cells in the human visual system. It has good characteristics in extracting local spatial and frequency domain information of targets. The Gabor feature is usually obtained by a convoluting image with a Gabor filter. Moreover, they have strong anti-interference ability in terms of image noise and illumination changes, and the most important advantages of Gabor filters are their translation, invariance to rotation, and scale. In image processing, the feature based on the Gabor filter is directly extracted from gray-level images. The 2-dimensional Gabor filter, in the spatial domain, is a Gaussian kernel function modulated by the complex sinusoidal plane wave. It can be defined as follows:
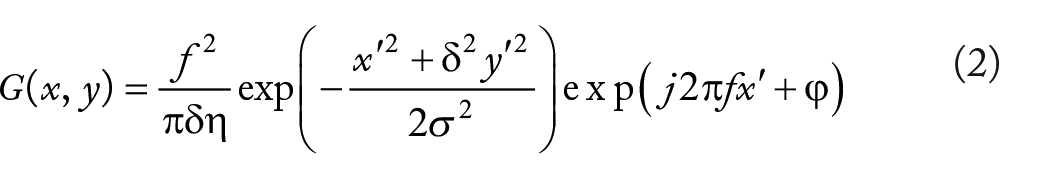


where
In our work, we use 40 Gabor filters in 5 scales and 8 orientations, which are shown in Figure 1. After using 40 Gabor filters, because of the high correlation of feature vectors, we can reduce the reduced feature data by way of downsampling for reducing information redundancy.31,32 Therefore, the protein sequence can be represented as Gabor feature vectors that are constructed by the first 100 coefficients.

Gabor filter in 5 scales and 8 orientations.
Random forrest classifier
At present, RF is one of the most popular prediction algorithms in data science. It was mainly developed by Breiman.
23
The RF model is one of the efficient ensemble classification algorithms, which uses multiple decision trees to reduce the output variance, thereby improving the accuracy of the classification. The RF classification makes full use of 2 powerful machine learning techniques. The first of RF classification is the selection of training examples, assuming that the original sample set has the total of examples
Results and Discussion
Evaluation measures
To better evaluate the proposed method, we calculated the following evaluation parameters: precision (PR), prediction accuracy (ACC), sensitivity (SN), specificity (SPC), and Matthew’s correlation coefficient (MCC). Their formation can be seen as follows:





where true negative (TN) is the number of true noninteracting pairs that are predicted correctly; true positive (TP) is the number of true samples that are predicted correctly; false positive (FP) is the number of true noninteracting pairs that are predicted to be interacting; and false negative (FN) is the correct number of samples that are predicted incorrectly. Moreover, the receiver operating characteristic (ROC) curves are one of the ways to evaluate the performance of the proposed method, and based on the prediction result, the area under an ROC curve (AUC) can also be computed to summarize ROC curve numerically.
Assessment of prediction ability
To ensure fairness of experiments, we conducted experiments in 3 different data sets of
Five-fold cross-validation prediction results obtained on
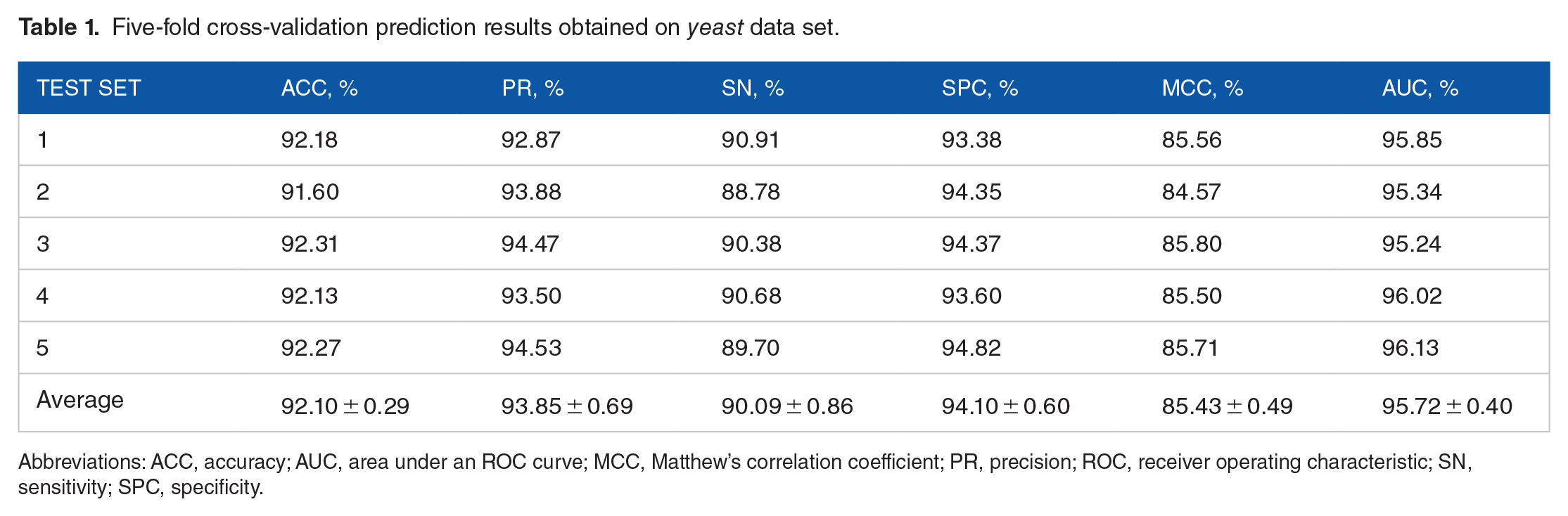
Abbreviations: ACC, accuracy; AUC, area under an ROC curve; MCC, Matthew’s correlation coefficient; PR, precision; ROC, receiver operating characteristic; SN, sensitivity; SPC, specificity.
Five-fold cross-validation prediction results obtained on

Abbreviations: ACC, accuracy; AUC, area under an ROC curve; MCC, Matthew’s correlation coefficient; PR, precision; ROC, receiver operating characteristic; SN, sensitivity; SPC, specificity.
Five-fold cross-validation prediction results obtained on

Abbreviations: ACC, accuracy; AUC, area under a ROC curve; MCC, Matthew’s correlation coefficient; PR, precision; ROC, receiver operating characteristic; SN, sensitivity; SPC, specificity.
As shown in the tables, when predicting PPIs of

ROC curves performed by the proposed method on

ROC curves performed by the proposed method on

ROC curves performed by the proposed method on
According to these results, the method is both practical and effective for predicting PPIs by combining the Gabor feature with RF classification. Furthermore, these criterion values in low deviations indicate that the method we proposed is stable and robust. The main advantage of the feature extraction method is that it can not only retain enough prior information of PSSM but also describe the sequence information of protein sequence efficiently. The ability of the Gabor feature in obtaining effective information in PSSM is outstanding. Besides, considering the influence of protein sequence order, the texture information extracted by the Gabor feature can retain the effective information of protein sequence well. The results show that the utilization of the Gabor texture feature to extract evolutionary information in predicting PPIs in the proposed method is effective.
Comparison with other feature extraction methods
To evaluate the effectiveness of the Gabor feature in extracting protein sequence information and identifying protein interactions, we further compared the results to DCT and LPQ with the same RF classification. The DCT algorithm is a popular linear separable transformation. It is mainly used for data or image compression, and DCT has a good performance of decorrelation due to its ability to convert signals from the spatial domain to the frequency domain. The LPQ is considered as an effective operator for texture feature descriptors, which remain the blur-invariant property, and the LPQ is also widely used in facial recognition and image processing. In our work, the DCT feature, the LPQ feature, and the Gabor feature were extracted from PSSM, and then we made a comparison in the same RF classification.
The results of

Performance comparison with 6 validation metrics using the Gabor feature (blue bar), the LPQ feature (green bar), and the DCT feature (yellow bar). (A) Accuracy rates, (B) sensitivity, (C) precision, (D) specificity, (E) MCC and (F) AUC. AUC indicates area under an ROC curve; DCT, discrete cosine transform; LPQ, local phase quantization; MCC, Matthew’s correlation coefficient; ROC, receiver operating characteristic.
Performance on the independent data sets
As we obtained good results on 3 PPI data sets of
Model prediction results of 4 species.

Abbreviation: ACC, accuracy.
According to the results in Table 4, when the PPI data set from
Comparison with other methods
In recent years, a large number of algorithms have emerged for predicting PPIs. In Table 5, we compared previous studies that proposed other methods to predict PPIs of
Performance comparison of different methods on the

Abbreviations: ACC, accuracy; ELM, extreme learning machine; HKNN,
Performance comparison of different methods on the

Abbreviations: ACC, accuracy; LD, local descriptor; MCC, Matthew’s correlation coefficient; PCA-EELM, principal component analysis-ensemble extreme learning machine; PR, precision; RF, random forest; SN, sensitivity; SVM, support vector machine.
Performance comparison of different methods on the

Abbreviations: ACC, accuracy; LDA, latent Dirichlet allocation; MCC, Matthew’s correlation coefficient; PR, precision; RF, random forest; RoF, rotation forest; SN, sensitivity; SVM, support vector machine.
From the table above, we can see that using an ensemble classifier such as the ensemble of HKNN (
Conclusions and Discussion
In recent years, the number of researchers requiring more knowledge to detect PPIs is increasing. Due to the complexity and high dimensionality of proteomic data, flexible and powerful statistical learning tools are needed for effective statistical analysis, which promotes the rapid development of computing methods for predicting PPIs. In this article, we proposed a novel computational method for predicting PPIs in which an RF classifier combined with the Gabor feature descriptor on the PSSM is used. The main improvements of the proposed method are that the Gabor feature can extract the discriminative information of protein sequence, especially enhancing the texture feature information of protein sequence that the interaction between proteins is more likely to occur in the region with higher energy. The experimental results demonstrated that the good performance of our proposed method in predicting PPIs. The results also showed that Gabor features perform better than LPQ and DCT in texture feature and protein sequence correlation extraction. In future studies, more effective feature extraction methods and machine learning techniques will be explored for PPI prediction.
Footnotes
Author Contributions
XKZ and ZHY conceived and designed the analysis. ZHY and LPL provided data. YL, ZW and XZK provided mathematical theory and performed simulations. XZK and JP wrote the manuscript. All authors reviewed the final manuscript.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research is funded by the National Natural Science Foundation of China under Grant (No. 61722212 and 61873212.)
