Abstract
Obligate intracellular chlamydiae diverged into pathogenic and environmental chlamydiae 0.7-1.4 billion years ago. While pathogenic chlamydiae have adapted to a wide range of vertebrates, environmental chlamydiae inhabit unicellular amoebae, the free-living
Introduction
Obligate intracellular chlamydiae separated into the environmental chlamydiae (eg,
A number of recent studies have revealed that the genomes of environmental chlamydiae (2.0-3.0 Mb) are more than double the size of those of pathogenic chlamydiae (1.0-1.2 Mb).1,9–12 It is thus clear that environmental chlamydiae still possess certain genes that pathogenic chlamydiae have lost. Meanwhile, similar to pathogenic chlamydiae, environmental chlamydiae undergo a unique developmental process, consisting of two distinct forms: the elementary body, its infectious form, and the reticulate body, its replicative form.12,13 We also found that some environmental chlamydiae could grow in immortalized human cells.12,13 It is thus clear that these two types of chlamydiae share similar backgrounds. However, the environmental factors that are responsible for promoting the divergence that occurred during chlamydial evolution and resulted in these two groups remain unknown.
During the last 10 years, giant viruses, which can be visualized under a light microscope, have been discovered and shown to have similar genes to those in other organisms, particularly those in several types of bacteria and in eukaryotes.14–16 The giant viruses consist of two distinct groups, protozoan-related and protozoan-unrelated types. The protozoan-related giant viruses include the families
In the present study, we thus attempted to visualize genes of giant viruses in chlamydial genomes by bioinformatic analysis mainly with comparative genome and phylogenic analysis, seeking genes present in chlamydiae that are specifically shared with protozoa-related giant viruses. For the first time, we show a linkage between chlamydiae and protozoan-related giant viruses.
Materials and Methods
Data sets
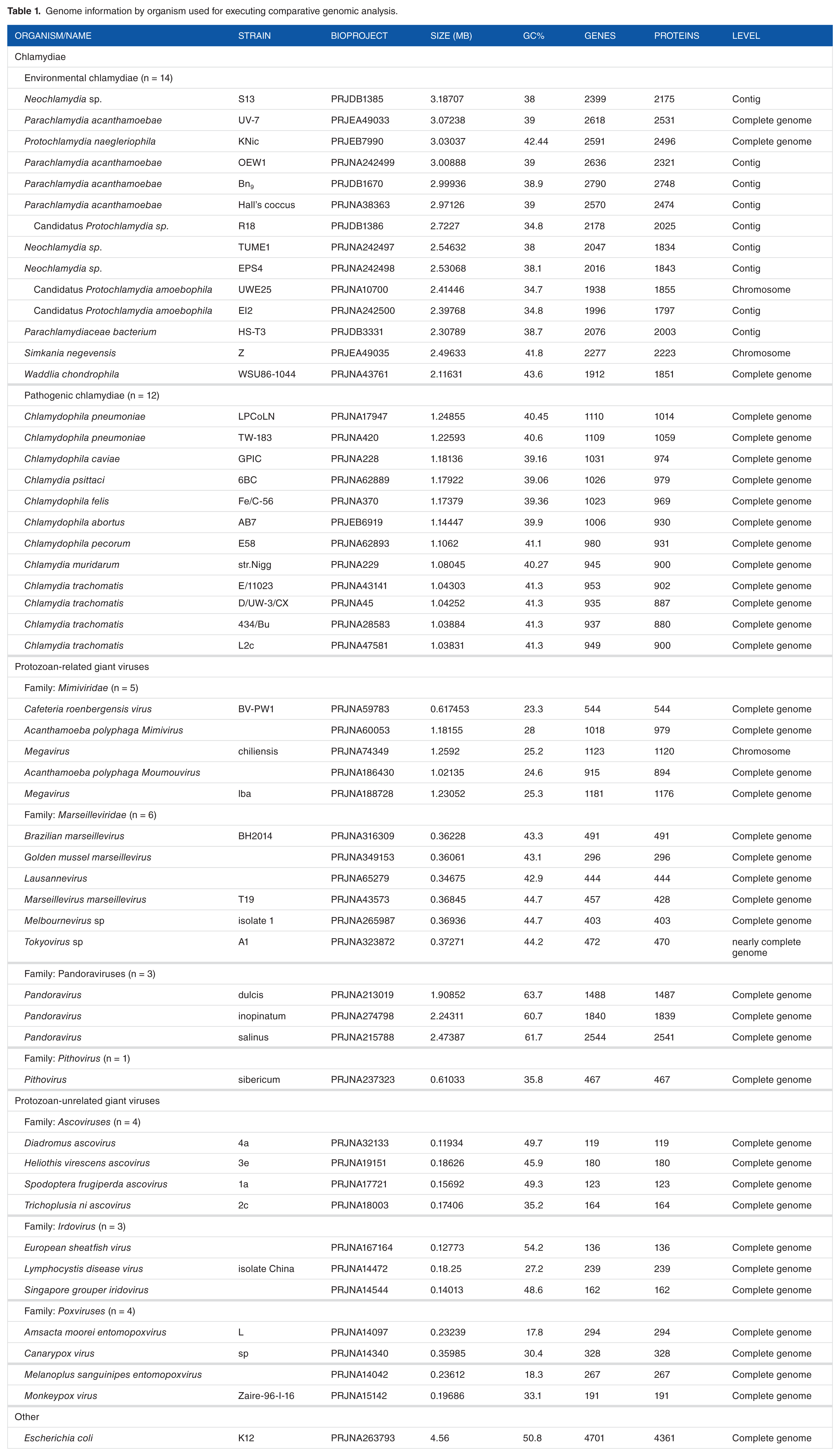
Chlamydiae and others (environmental chlamydiae n = 14, pathogenic chlamydiae n = 12, protozoan-related giant viruses n = 15, protozoan-unrelated giant viruses n = 11, others n = 1 (
Genome information by organism used for executing comparative genomic analysis.
Analysis flow
The genome (or contig) information was obtained from the National Center for Biotechnology Information (NCBI) database (http://www.ncbi.nlm.nih.gov/genome/browse/), and the obtained sequence information was reconstructed as data sets with functional annotations into Rapid Annotation using Subsystem Technology (RAST) (http://rast.nmpdr.org/), which is an open-access genomic analysis tool that acts as a fully automated service for genomic annotation with Basic Local Alignment Search Tool (BLAST) analysis.
19
These reconstructed RAST data sets with annotated amino acid sequences are shown into Tables S1 to S4 (Table S1, protozoan-related giant viruses; Table S2, protozoan-unrelated giant viruses; Table S3, pathogenic chlamydiae; Table S4a and b, environmental chlamydiae). Then, BLAST analysis was performed using the RAST data sets with the default settings (cut-off 10−10 identity >10%), and these sequences were furthermore selected with bidirectional hits and length cut-off (>30 amino acid residues). Numbers of orthologs were normalized with genome sizes of both chlamydiae and viruses. Specifically, the normalized numbers were obtained from raw numbers divided with each of the chlamydia and virus genome sizes; it is shown as ortholog numbers of giant virus assumed with 1 Mbp of genome size per 1 Mbp of chlamydial genome. Also, the cut-off value (>1.48%) as a background was defined by the prevalence of genes from
Statistical analysis
Comparison of the prevalence of giant virus genes between pathogenic and environmental chlamydiae was performed by Mann–Whitney’s U test. The presence of a correlation between the prevalence of genes from giant viruses within chlamydial genomes and annotated chlamydial ORF numbers was determined by Pearson’s single regression analysis. A correlation coefficient value of >0.5 or <−0.05 with a
Results and Discussion
Several genes of protozoa-related giant viruses of the family Mimiviridae are significantly conserved in the genomes of both chlamydiae
To explore the traces of an encounter with giant viruses in chlamydiae, the prevalence of genes from giant viruses in chlamydiae was assessed by BLAST analysis using RAST with genomic information from multiple species from each group (environmental chlamydiae n = 14, pathogenic chlamydiae n = 12, protozoan-related giant viruses n = 15, protozoan-unrelated giant viruses n = 11)(see Table 1). To ensure a uniform annotation of all the genes, pre-existing annotations in the database were re-annotated by RAST (http://rast.nmpdr.org/), which is an open-access genomic analysis tool that acts as a fully automated service for genomic annotation with BLAST analysis. Also, the cut-off value (>1.48%) as a background was defined by the prevalence of genes from

Comparisons of the prevalence rates of giant virus genes in chlamydial genomes and of the trend of dispersion on the prevalence of giant virus genes between pathogenic and environmental chlamydiae. Panels (A) and (B) show protozoa-related giant viruses and protozoa-unrelated giant viruses, respectively. Blue and red bars show the prevalence of giant virus genes in environmental and pathogenic chlamydial genomes, respectively. Comparisons of the prevalence rate were conducted using Mann–Whitney’s U test. Stars show a significant difference (

Phylogenic analysis with most prevalent sets (
As expected, we found that, in contrast to the protozoa-unrelated viruses, several genes of protozoa-related giant viruses, the family
The prevalence of genes from protozoa-related giant viruses in chlamydiae is negatively and specifically correlated with chlamydial ORF numbers
If the prevalence of genes from giant viruses in chlamydial genomes specifically revealed that chlamydiae had encountered protozoa-related giant viruses presumably in ancestral amoebae, this would also suggest that this encounter resulted in specific modifications of the chlamydial genome, such as changes of the ORF numbers. To assess this hypothesis, the correlation between the prevalence of giant virus genes in chlamydial genomes and the chlamydial ORF numbers was assessed by Pearson’s single regression analysis. The results showed significant correlation coefficients of <−0.5 with a
The prevalence of the genes from almost giant viruses in each of the chlamydiae was negatively and specifically correlated with the number of chlamydial ORFs. These results suggest that these giant viruses changed the chlamydial genome and influenced chlamydial evolution. Interestingly, studies have shown that
In contrast to the pathogenic chlamydiae, the environmental chlamydiae specifically possess genes conserved among the Mimiviridae (Megavirus chiliensis)
Since information on the specific genetic material that was shared would be critical for understanding the forces driving the evolution and divergence of chlamydiae, we explored the specific genes of chlamydiae commonly shared with protozoa-related giant viruses, the

Total number of functional genes in each of the chlamydial genomes shared among protozoa-related giant viruses in the
Thus, these findings indicated that, in contrast to the pathogenic chlamydiae, the environmental chlamydiae specifically possessed functional genes conserved among the
Conclusions
Altogether, our study showed a putative linkage between chlamydiae and protozoa-related giant viruses, in particular
Supplemental Material
Supplemental_Material – Supplemental material for Lateral Gene Transfer Between Protozoa-Related Giant Viruses of Family Mimiviridae and Chlamydiae
Supplemental material, Supplemental_Material for Lateral Gene Transfer Between Protozoa-Related Giant Viruses of Family Mimiviridae and Chlamydiae by Takanori Watanabe, Sumire Yamazaki, Chinatsu Maita, Mizue Matushita, Junji Matsuo, Torahiko Okubo and Hiroyuki Yamaguchi in Evolutionary Bioinformatics
Footnotes
Acknowledgements
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants-in-aid for Japan Society for scientific research KAKENHI (grant number: 16K15270). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
HY conceived and designed the project; TW, SY, CM, MM, JM, TO, and HY contributed toward the analysis and confirmation; YH, SY, JM, TO, and HY contributed toward critical editing; HY wrote the manuscript. All authors read and approved the final manuscript.
Ethical approval
The study reported in this manuscript did not involve any human participants, human data, human tissue, data on specific individuals, or animal experiments.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
