Abstract
Objectives
Autosomal dominant polycystic kidney disease (ADPKD) is a common inherited disease in cats. In most cases, the responsible abnormality is a nonsense single nucleotide polymorphism in exon 29 of the PKD1 gene (chrE3:g.42858112C>A, the conventional PKD1 variant). The aim of this study was to conduct a large-scale epidemiological study of ADPKD caused by the conventional PKD1 variant in Japan and to search for novel polymorphisms by targeted resequencing of the PKD1 using a next-generation sequencer.
Methods
A total of 1281 cats visiting the Veterinary Medical Center of the University of Tokyo were included in this study. DNA was extracted from the blood of each cat. We established a novel TaqMan real-time PCR genotyping assay for the conventional PKD1 variant, and all cases were examined for the presence of this variant. Targeted resequencing of all exons of the PKD1 was performed on the DNA of 23 cats with the conventional PKD1 variant, six cats diagnosed with cystic kidneys but without this variant, and 61 wild-type normal cats.
Results
Among the 1281 cats examined in this study, 23 (1.8%) harboured the conventional PKD1 variant. The odds of having the conventional PKD1 variant were significantly higher in Persian cats, Scottish Folds and Exotic Shorthairs than in the other breeds, although the number of cases in each breed was small. Furthermore, we identified four variants unique to cats with cystic kidneys that were not found in wild-type normal cats, all of which were in exon 15. In particular, two (chrE:g.42848725delC, pGly1641fs and chrE:g.42850283C>T, pArg2162Trp) were candidate variants.
Conclusions and relevance
This study revealed that the conventional PKD1 variant was prevalent in Scottish Fold, Persian and Exotic Shorthair breeds in Japan, and variants in exon 15 of PKD1, in addition to the conventional variant in exon 29, would be key factors in the pathogenesis of ADPKD in cats.
Introduction
Autosomal dominant polycystic kidney disease (ADPKD) is a common inherited disease in cats and is characterised by the development of multiple fluid-filled cysts dispersed in the kidney and occasionally in the liver or pancreas. 1 The cysts gradually increase in number and size, causing deterioration of the renal function through compression of the renal parenchyma. Most patients remain clinically normal for years until chronic kidney disease (CKD) develops later in life (>7 years). 2 Ultrasonographic examination is routinely performed for the diagnosis and evaluation of feline ADPKD. Owing to its hereditary nature, no fundamental treatment options preventing the onset of the disease have been found to date. Hence, feline ADPKD is generally treated by following the standard CKD treatment protocols, such as subcutaneous fluid administration. 3
Feline ADPKD is caused by a heterozygous variant that was identified by Lyons et al 4 in 2004 and is located in the PKD1 gene with a nonsense single nucleotide polymorphism (SNP) in exon 29 (chrE3:g.42858112C>A, the conventional PKD1 variant). This variant causes a stop codon, truncating the protein up to 25% at the C-terminus. 4 This variant in a homozygous state is lethal in the embryonic stage. PKD1 encodes polycystin 1 (PC1), which is a membrane-bound large multidomain glycoprotein with a G-protein-coupled receptor proteolytic site. Its physiological function is unclear; however, it is known to be located on the primary cilia, a flagellar organelle protruding from the cell surface used for mechanotransduction. PC1 is thought to perform an important role in transmitting signals from outside the cell.5,6 The loss of functional PC1 caused by the variant appears to contribute to the cystogenesis in ADPKD, and much research has been conducted to identify the underlying mechanisms. Some have proposed the influence of various upregulated or downregulated intracellular signals, while others have argued that macrophage recruitment by the primary cilia plays a huge role in cystogenesis. 7 Despite these efforts, the exact mechanism remains unclear. However, two models of the relationship between PC1 dosage and cystogenesis have been established: the two-hit model and the threshold model. The two-hit model suggests that after inactivation of one allele by germline mutation, a somatic mutation occurs, inactivating the other allele and thereby causing cystogenesis. 8 On the other hand, the threshold theory is the more recent belief, which suggests that the disease occurs when the PC1 levels drop below a certain threshold and do not necessarily have to be at zero.9,10 Since the variant responsible for feline ADPKD truncates PC1, it can be assumed that the truncated abnormal protein is unable to function properly, leading to a lack of functional PC1. Subsequent to the discovery of the responsible variant, PKD1 genetic tests have also been developed for the diagnosis of feline ADPKD, which allows for easier diagnosis, even in the asymptomatic early stages of ADPKD. 11 However, these tests cannot predict the severity of the disease or evaluate its progression, suggesting that genetic tests should be used together with ultrasonographic examination.
Previous studies in the USA, Europe, Australia and Taiwan have suggested that ADPKD in cats is most common in Persian cats and related breeds1,12–15; the same trend has been reported in Japan along with reports of ADPKD in other breeds, such as Scottish Folds and American Shorthairs. 16 The prevalence of the conventional PKD1 variant has been investigated in very few studies, but is reported as 13.5% in China (n = 111), 33.3% in Slovenia (n = 24, limited to Persian cats) and 40% in Japan (n = 377, limited to cats suspected of ADPKD).15–17 These studies have mostly focused on Persian cats or cats already suspected of ADPKD. There are no data on the prevalence of the disease or the conventional PKD1 variant among the cat population as a whole, even though comprehending the frequency of the conventional PKD1 variant and confirming predisposed breeds in every region is crucial for the prevention of ADPKD. In addition, some studies have revealed that cats without the conventional PKD1 variant can develop ADPKD. Three cases of an ADPKD diagnosis based on ultrasonographic examination in the study in China did not have the conventional PKD1 variant. 15 Another study on Maine Coons in Switzerland reported that none of the six cases with ADPKD had the PKD1 conventional variant. 18 These observations suggest that an unknown variant may serve as another cause of ADPKD.
Given this background of feline ADPKD, we conducted a large-scale epidemiological study to understand the nature and current situation of ADPKD in Japan accurately. In addition, we performed target resequencing of all exons of PKD1 using next-generation sequencing to identify new variants.
Materials and methods
Feline cases and samples
There were 7536 visits and 5716 admissions of feline cases at the Veterinary Medical Center of the University of Tokyo between April 2015 and March 2018 (cumulative total 13,252). After excluding repetitive visits and patients that did not undergo blood tests, 1281 cats were included in this study.
Clinical information on the age, breed, sex and diagnosis of all cases was obtained from the electronic medical records system. The cats were aged 0–20 years, comprising 717 males (101 intact and 616 neutered), 560 females (94 intact and 466 neutered) and four of unknown sex. There was a wide variety of breeds, as follows: random breed (n = 791); American Shorthair (n = 101); Scottish Fold (n = 87); Persian cats (n = 37); Russian Blue (n = 34); Maine Coon (n = 31); Norwegian Forest Cat (n = 29); Munchkin (n = 28); Abyssinian (n = 24); Somali (n = 20); Ragdoll (n = 16); British Shorthair (n = 14); American Curl (n = 13); Himalayan (n = 13); Bengal (n = 13); Chartreux (n = 6); Exotic Shorthair (n = 6); Singapura (n = 6); Ocicat (n = 2); Ragamuffin (n = 2); Siamese (n = 2); Tonkinese (n = 2); Birman (n = 1); European Shorthair (n = 1); La Perm (n = 1); and Selkirk Rex (n = 1). The body weight, ultrasonographic findings and values of blood urea nitrogen (BUN), creatinine and urine-specific gravity were assessed in the cats with the conventional PKD1 variant according to genotyping from the electronic medical records system. The electronic medical records system was scanned for the mention of feline ADPKD in the same period (April 2015 to March 2018) to identify samples diagnosed with cystic kidneys that did not have the conventional PKD1 variant according to genotyping. Ultrasonographic images were retrieved and reviewed for the applicable samples.
DNA was extracted from EDTA-anticoagulated blood using a NucleoSpin Blood Quick Pure kit (Macherey-Nagel) and was stored in the gene bank run by Azabu University (Kanagawa, Japan).
Real-time PCR PKD1 genotyping
We conducted a real-time PCR genotyping assay using TaqMan probes, which are probes with fluorescein amidite (FAM) and 2’-chloro-7’-phenyl-1,4-dichloro-6-carboxy-fluorescein (VIC) on one end and a non-fluorescent quencher (NFQ) on the other. The assay was performed with 5.0 μl TaqPath ProAmp Master Mix (Thermo Fisher Scientific), 0.5 μl Custom TaqMan SNP Genotyping Assay (Thermo Fisher Scientific), >20 ng DNA and ultrapure water up to 10 μl. The Custom TaqMan SNP Genotyping Assay included forward (CCTCGGAGCCGCTTCAC) and reverse (TTGGCGCCCAGGAAGAG) primers and two probes, VIC–ACGAGGAGGACGCAACA–NFQ and FAM–CGAGGAGGACTCAACA–NFQ, for wild-type and mutant alleles, respectively. Real-time PCR was performed on the fast cycle on a StepOnePlus Real-time PCR system (Thermo Fisher Scientific) for one cycle at 60°C for 30 s and 95°C for 5 mins. This was followed by 40 cycles of 95°C for 5 s and 60°C for 30 s and finally one cycle of 60°C for 30 s.
To determine the specificity of this assay, two samples each of wild-type and heterozygous mutant samples selected randomly and a water control were run on the genotyping assay. The genotypes of these samples were confirmed using Sanger sequencing (Azenta Life Sciences) after PCR amplification and extrication from the agarose gel using Wizard SV Gel and a PCR Clean-Up System (Promega). The threshold was set to 0.6 ΔRn, and threshold cycle values were calculated automatically on the StepOnePlus Real-time PCR system. A value >40 was assigned to samples whose threshold cycle values were undetermined after 40 PCR cycles.
The real-time PCR assay was performed as described above for all DNA feline samples to determine their genotype. The genotypes of each sample were determined by examining the amplification of the reporter dye (FAM or VIC) on the amplification plot.
Target resequencing of PKD1 using next-generation sequencing
This study included 23 cases of cats with the conventional PKD1 variant (chrE3:g.42858112C>A), six cases diagnosed with cystic kidneys but without the conventional PKD1 variant, and 62 control cases without renal cysts and without the conventional PKD1 variant. DNA was extracted from EDTA-anticoagulated blood using a NucleoSpin Blood Quick Pure kit and was stored with help from the gene bank run by Azabu University. Data regarding age, breed, sex, diagnosis and ultrasonographic images of all patients were obtained from the electronic medical records system.
We designed two primer pools of 106 primer pairs with DesignStudio (Illumina) to amplify PKD1. The sequencing library was constructed using the AmpliSeq Library PLUS for Illumina. The PCR product size distribution was checked using Bioanalyzer (Agilent Technologies) after adapter ligation. Paired-end sequencing (300 bp) with a dual 8-bp barcode was performed using MiSeq (Illumina). The sequence reads were sorted individually using each barcode.
Raw sequence data were processed according to the Broad Institute’s Genome Analysis Tool Kit (GATK) best practices workflow for small germline variants. 19 Raw sequence reads were trimmed to remove adaptors using Trimmomatic (version 0.38) 20 and mapped to the feline reference sequence (Ensembl Genome assembly felCat9 GCA_000181335.4) using bwa mem (version 0.7.17). 21 Samtools (version 1.9) was used to sort and index each mapped file. 22 These data were subjected to VariantFiltration using GATK (version 4.2.2). Moreover, the data were applied to VariantFiltration using GATK (version 4.2.2) with the filter-expression parameter of ‘QD <5.0 || FS >50.0 || SOR >3.0 || MQ <50.0 || MQRankSum <–2.5 || ReadPosRankSum <–1.0 || ReadPosRankSum >3.5’. In individual samples, the coverage rate was defined as the proportion of the target region covered by ⩾20 sequencing reads. Variants were called using HaplotypeCaller by GATK. Annotation was performed using SnpEff (version 4.3) 23 and Ensembl Variant Effect Predictor, 24 with all transcripts registered in Ensembl release 104.
Statistical analyses
Statistical analyses were performed using GraphPad Prism software (version 5.0.1; GraphPad Software). The odds ratio was calculated for each breed to determine the breeds predisposed to the conventional PKD1 variant. Significance was confirmed using a two-tailed Fisher exact test. The relationship between the frequency of the conventional PKD1 variant and sex was evaluated using the χ2 test. The age, plasma creatinine and BUN levels were compared between two groups with and without renal cysts in the cats with the conventional PKD1 variant using a two-tailed Mann–Whitney U test. The relationship between the age and plasma creatinine or BUN levels in the conventional PKD1 variant was determined using the Spearman rank correlation coefficient. P values <0.05 were considered statistically significant.
Results
Establishment of a novel TaqMan real-time PCR genotyping assay
To determine the specificity of the real-time PCR PKD1 genotyping assay, two samples each of heterozygous PKD1 mutants and wild-type cases, whose genotypes had previously been confirmed by Sanger sequencing, were randomly selected. The two wild-type samples produced threshold cycle values in the VIC channel of 29.165 and 28.124, respectively. However, these samples did not produce threshold cycle values within 40 cycles in the FAM channel (>40 cycles were needed). On the other hand, the two heterozygous mutant samples had similar threshold values in both the VIC (29.564 and 29.957, respectively) and FAM channels (28.041 and 28.684, respectively). Figure 1 shows the production of threshold cycle values for water, PKD1 wild-type and PKD1 variant samples. Wild-type samples showed a slight increase in the ΔRn in FAM, which could be considered non-specific (Figure 1b). This increase was within the threshold range (ΔRn = 0.6), even within 40 cycles, and was clearly different from the increase in FAM in PKD1 mutants (Figure 1c). These results suggest that the assay was able to discriminate between wild-type and the conventional PKD1 variant with high specificity.
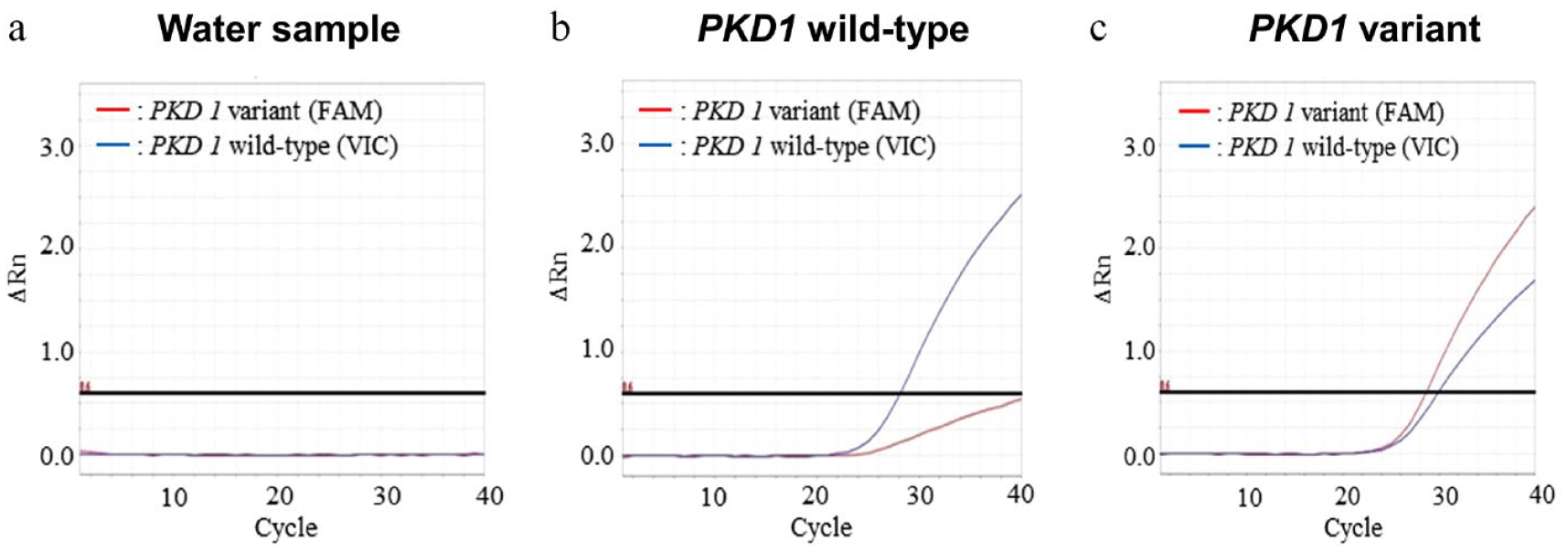
Representative amplification plots from real-time PCR genotyping. The red graph shows amplification of the reporter dye on the conventional PKD1 variant probe (FAM) and the blue graph the wild probe (VIC). (a) In the water sample, no changes were noted in either graph. (b) In the wild-type samples, only the blue graph showed a significant increase. The slight increase in the red graph is presumably non-specific. (c) In the DNA samples with the conventional PKD1 variant (chrE3:g.42858112C>A), both graphs showed increased values, reflecting the heterozygous variant. FAM = fluorescein amidite; VIC = 2′-chloro-7′-phenyl-1,4-dichloro-6-carboxy-fluorescein
Prevalence of the conventional PKD1 variant and clinical evaluation
Out of 1281 samples, 23 (1.8%) had the conventional PKD1 variant (chrE3:g.42858112C>A), while six diagnosed with cystic kidneys did not have the conventional PKD1 variant (Table 1). The 23 samples comprised Scottish Folds (n = 6), random breed (n = 6), Persian cats (n = 4), American Shorthairs (n = 3), Exotic Shorthairs (n = 2) and Munchkins (n = 2). The breed with the highest positive rate for the variant was Exotic Shorthair at 33.3%. Persian cats also had a high positive rate of 10.8%, followed by Munchkin (7.14%), Scottish Fold (6.90%) and American Shorthair (2.97%) breeds (Table 2). The odds of having the conventional PKD1 variant were significantly higher in Persian cats, Scottish Folds and Exotic Shorthairs than in the other breeds (Table 2). Cases with the conventional PKD1 variant were dispersed across different age and sex groups, with an age range of 0–13 years and comprising 14 females (three intact and 11 neutered) and nine males (all neutered). There was no significant difference in the frequency of the conventional PKD1 variant according to the sex by the χ2 test (P = 0.0988). All six cases diagnosed with cystic kidneys but without the conventional PKD1 variant had multiple cysts in one kidney or both kidneys confirmed by ultrasonographic images.
Cases with the conventional PKD1 variant and cases with cystic kidneys without the PKD1 variant in this study

CM = castrated male; SF = spayed female
NE = not evaluated
Prevalence of the conventional PKD1 variant and odds in different breeds

Bold text indicates significant differences
Renal cysts were confirmed on ultrasonography in 16/23 cats with the conventional PKD1 variant, four cats did not have renal cysts, and the remaining three cats did not undergo ultrasonography (Table 1). There was no statistically significant relationship between age and the presence of renal cysts; however, there was a higher tendency for renal cysts to occur in older cats than in younger cats (P = 0.0598) (Figure 2). Two out of the four cats without renal cysts were aged under 1 year, while the other two were relatively older (5 and 8 years, respectively); one of the older cats had multiple cysts in the liver and the other in the pancreas.

Association between the age and presence of renal cysts in cats with the conventional PKD1 variants (n = 20). Of the 23 cases with the conventional PKD1 variants, three did not undergo ultrasound examination. Results are expressed as means ± standard errors. Data were evaluated using two-tailed Mann–Whitney U test (P = 0.0598)
Plasma creatinine and BUN levels were compared between the patients with and without renal cysts (Figure 3). BUN levels were significantly higher in cats with renal cysts than in those without (P = 0.0178), and even plasma creatinine levels tended to be higher in the former (P = 0.0992). The age did not correlate with plasma creatinine (P = 0.7837, r = 0.06374) and BUN levels (P = 0.2143, r = 0.2827) in cats with the conventional PKD1 variant.

Association between presence of renal cysts and the (a) plasma creatinine (n = 17) and (b) blood urea nitrogen (BUN) (n = 18) levels in cats with the conventional PKD1 variants. Of the 23 cases with the conventional PKD1 variants, three did not undergo ultrasound examination. In addition, the plasma creatinine values were not recorded in three cases, while BUN values were not recorded in two; hence, they were excluded from this analysis. Results are expressed as means ± standard errors. Data were evaluated using two-tailed Mann–Whitney U test (plasma creatinine: P = 0.0992; BUN: P = 0.0178). BUN = blood urea nitrogen
Target resequencing of PKD1 using next-generation sequencing
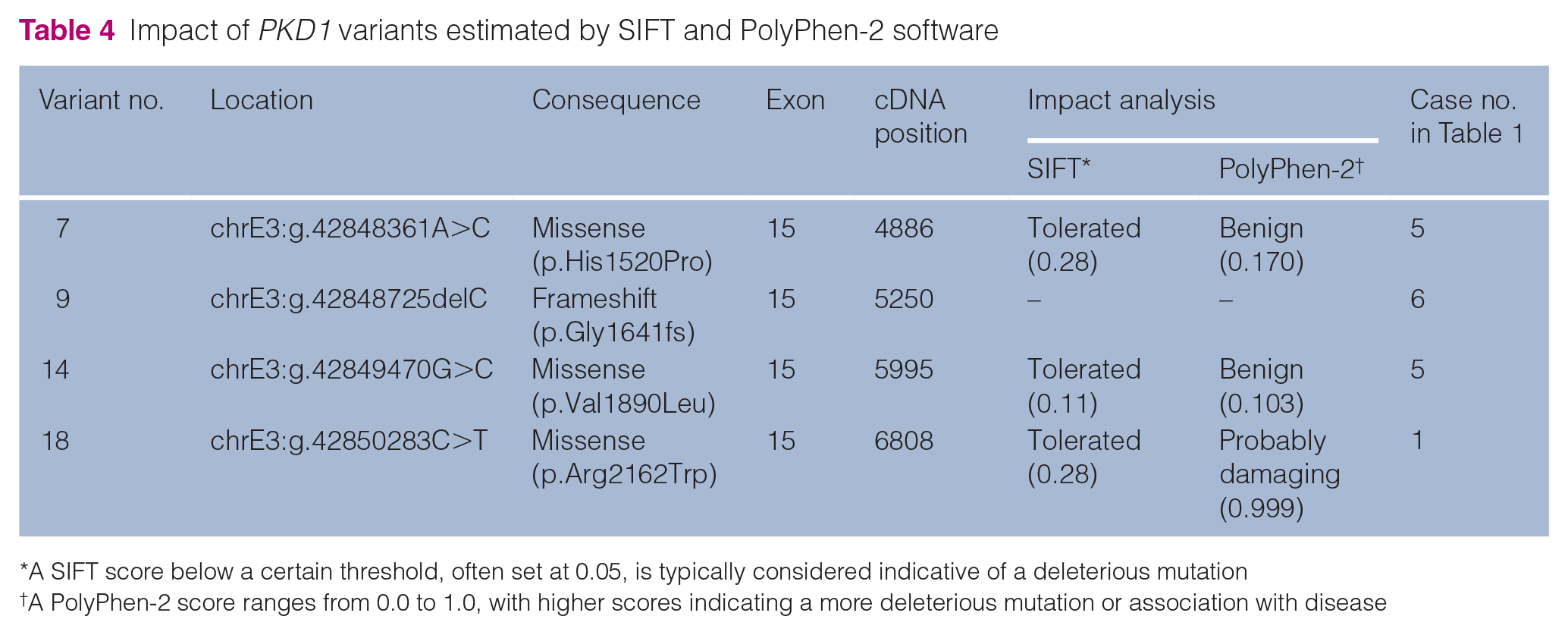
One of 62 controls without the conventional PKD1 nonsense variant (chrE3:g.42858112C>A) and one of 23 ADPKD cats with the conventional PKD1 variant were excluded because their DNA concentrations were low. We identified 86 non-synonymous variants of PKD1, including 18 protein-truncating variants (high impact estimated by Ensembl) and 68 missense variants, among the included samples (see Table S1 in the supplementary material). Of these variants, 27 non-synonymous variants (two protein-truncating variants and 25 missense variants) were identified in six cases of cystic kidneys without the conventional PKD1 variant (Table 3; see also Table S2 in the supplementary material). We focused on four singleton variants that were found only in six cases of cystic kidneys without the conventional PKD1 variant but not in any of the control (wild-type) cases (variants 7, 9, 14 and 18). All four of these variants were located in exon 15, of which one was a single-base deletion (chrE3:g.42848725delC), which causes immense damage to the resulting protein, and the other three were missense variants (Table 4). The three missense variants were analysed using SIFT 25 and PolyPhen-2 26 tools to estimate their impact on amino acid substitutions. We found that chrE3:g.42848361A>C (p.His1520Pro) and chrE3:g.42849470G>C (p.Val1890Leu) were predicted as benign in both software packages. However, chrE3:g.42850283C>T (p.Arg2162Trp) was predicted as damaging by PolyPhen-2 (Table 4). Variants chrE3:g.42848361A>C and chrE3:g.42849470G>C were present in the same cat. We used the UCSC LiftOver tool (https://genome.ucsc.edu/cgi-bin/hgLiftOver) to compare the variant locations with the human gene variants reported in human ADPKD. We found that chrE3:g.42850283C>T was identical to the human variant rs373256534 (p.Arg2162Trp).
Novel PKD1 gene variants found in six cases of cystic kidneys without the chrE3:g.42858112C>A nonsense variant
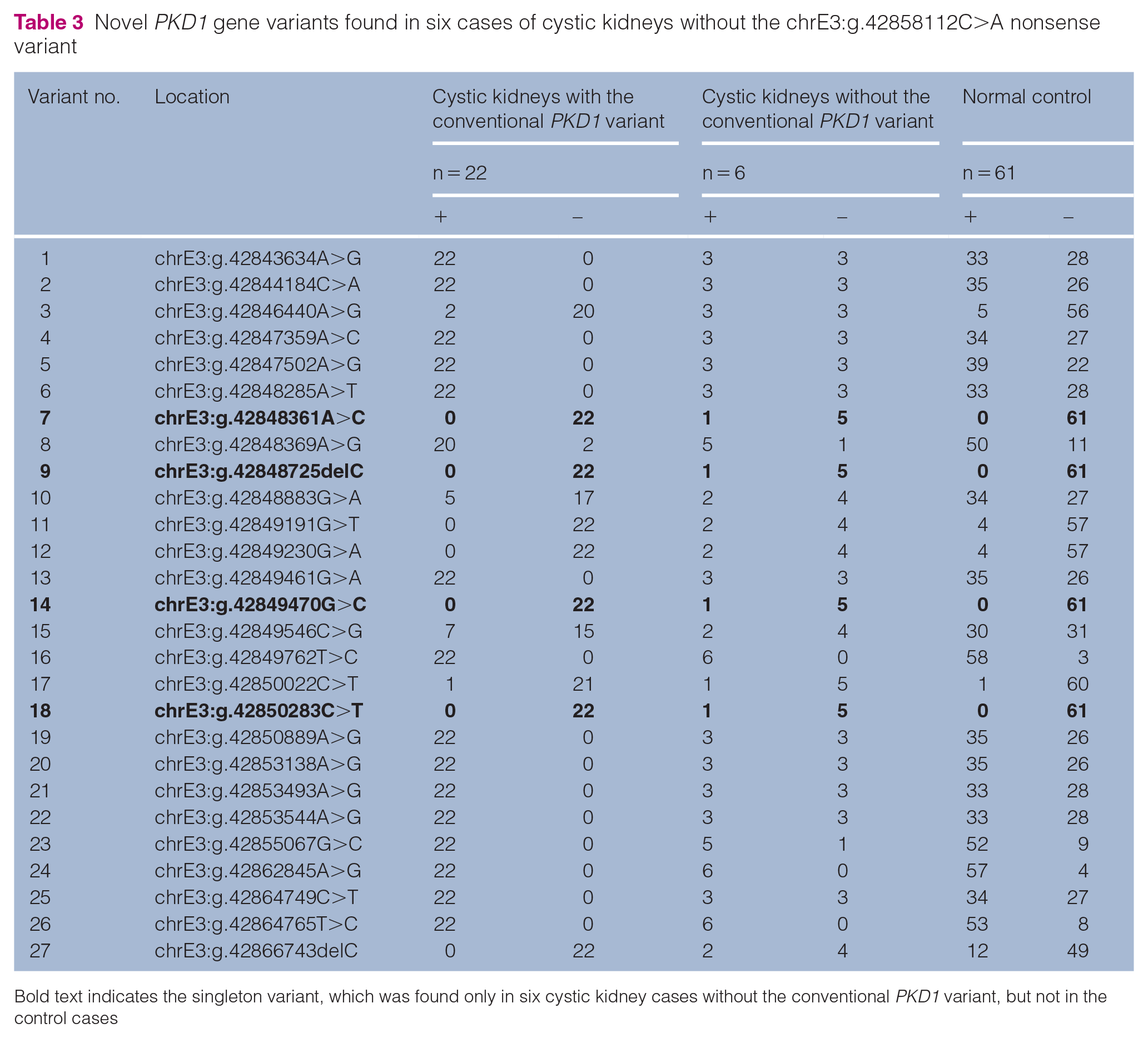
Bold text indicates the singleton variant, which was found only in six cystic kidney cases without the conventional PKD1 variant, but not in the control cases
Impact of PKD1 variants estimated by SIFT and PolyPhen-2 software
Discussion
Feline ADPKD first featured in case reports in the 1960s as a disease observed in Persian and Persian-cross cats. Subsequent research on the disease has been based on this presupposition, mainly focusing on Persian cat-related breeds.27,28 Moreover, the conventional PKD1 variant (chrE3:g.42858112C>A) was initially discovered in Persian cats. 4 Occurrence of the disease in other breeds has also been reported more recently, 15 shedding light on the importance of investigating the disease on a wider scale to capture the whole picture. Although a recent study explored the prevalence of the conventional PKD1 variant in Japan (n = 377), it had limitations in that it focused only on cases suspected of ADPKD based on family history or ultrasonographic observations. 16 In contrast, this study investigated 1281 cats, regardless of the case history, chief complaint, diagnosis or breed, making it the first large-scale epidemiological study assessing the conventional PKD1 variant prevalence in Japan using a novel real-time PCR genotyping assay.
The TaqMan real-time PCR genotyping assay established in this study can be considered a quicker method with higher throughput for performing genotyping for PKD1 when compared with pre-existing methods. Sanger sequencing is the most traditional and concrete method for decoding sequences and is still used for confirmation because of its high accuracy. While it is important to refer to Sanger sequencing when establishing a new genotyping method, there is no need to read the entire sequence when detecting SNPs, and easier methods can be used for genotyping. Recently, the PCR-restriction fragment length polymorphism method has been used widely for SNP detection because of its reproducibility. However, both these methods involve many steps, which could take several hours. On the other hand, our genotyping method can be run on a fast setting on real-time PCR instruments and takes only up to 40 mins. In addition, this method allows the analysis of 93 samples at once (on a 96-well plate, with one negative control and two positive controls), making genotyping of multiple samples remarkably faster and more convenient. The TaqMan real-time PCR genotyping method has sufficiently high specificity and is compatible with other methods, making it a very useful potential substitute for pre-existing methods.
According to the real-time PCR genotyping assay, the conventional PKD1 variant was observed in 1.8% of all included cases. The cases included in this study were those who visited the Veterinary Medical Center of the University of Tokyo, which specialises in secondary care, is located in the city centre and receives cases from many different primary care clinics in the surrounding area. Therefore, this 1.8% is considered to largely reflect the situation in Japan. This result was relatively lower than that in previous reports. The prevalence of the variant was 13.5% in China (n = 111), 33.3% in Slovenia (n = 24) and, most surprisingly, 40% in the study conducted in 2019 in Japan (n = 377).15–17 The huge difference between the results of our study and the 2019 Japanese study is most likely due to the difference in the research methods; the previous study focused on cats already suspected of ADPKD based on imaging or family history, as opposed to our study, which included cats regardless of their medical history. The previous study can be considered more inclined toward relationship analysis between ultrasonographic and genetic test results rather than an epidemiological study. Thus, the overall prevalence of the variant in Japanese cats is presumably closer to the 1.8% obtained in this study. Similarly, the study in Slovenia was limited to Persian cats, which could explain the high prevalence reported. The method used in the study from China was closest to ours; however, the number of cases included was much less, which may have increased the reported prevalence of the variant. However, this could also be a reflection of regional differences.
The odds of having the conventional PKD1 variant were significantly higher in Persian cats and Exotic Shorthairs, a cross between Persians, and short-haired breeds such as American or British Shorthairs, corroborating the findings of previous studies.1,12–15 Scottish Folds also had a significantly high odds ratio, similar to that reported in the previous research conducted in Japan. 16 Although some American Shorthairs had the conventional PKD1 variant, their odds ratios were not significant due to their large population. Furthermore, 6/718 random breed cats presented the variant, suggesting that random breeds are also potential carriers of the conventional PKD1 variant in Japan. However, the number of cats in each breed evaluated in this study was small, and these results may need to be considered in relation to the effects of regional differences in Japan.
In cats with the conventional PKD1 variant, the BUN levels were significantly higher in those with renal cysts than in those without, suggesting that the presence of renal cysts is associated with renal function deterioration. This finding supports the notion that renal function deteriorates in ADPKD. However, there was no significant difference in the plasma creatinine levels of patients with and without renal cysts, although they tended to be higher in the former than in the latter (P = 0.0989). This could be due to the small number of ADPKD cases in this study, and further research regarding renal function in ADPKD cases in a larger population is warranted. We could not evaluate the relationship between the cyst size or number and the deterioration of renal function, and determining whether the increase in cysts actually correlates with the decline in renal function is important for developing effective treatment methods. Future research is needed to clarify this, ideally over the course of a few years, in the same individuals.
The results of the real-time PCR PKD1 genotyping assay mostly reflected the results of the ultrasonographic examination, with a few exceptions. Four cats with the conventional PKD1 variant did not have renal cysts, which could be attributed to the stronger tendency for cysts to be present in older cats than in younger cats. Notably, these four cats were aged 9 months, 11 months, 5 years and 8 years, respectively; the two older cats had cysts in the liver and pancreas, respectively, which are common complications of ADPKD. Interestingly, there were nine negative cases with cysts in the kidneys, liver or pancreas suggestive of ADPKD. This suggests the possibility of a different variant causing the disease, as indicated in previous studies. 18 Since human ADPKD, a disease very similar in pathology and mechanism to feline ADPKD, is caused by many different variants of PKD1 and even some other genes involved in transportation of PC1 protein, 29 it is only natural that feline ADPKD may have other causative variants too.
To explore unknown PKD1 variants that could be the additional cause of ADPKD, we performed target resequencing using next-generation sequencing. As a result, 86 novel variants were detected in total, and when we focused on cases with cystic kidneys without the conventional PKD1 variant (chrE3:g.42858112C>A), 27 variants were extracted. Of these variants, four singleton variants were found only in cases with cystic kidneys and in none of the unaffected cats, indicating their association with the disease. All four variants were in exon 15, suggesting that, along with exon 29, exon 15 also could play an important role in the cystogenesis of ADPKD. ChrE3:g.42848725delC was the only variant that caused a frameshift, and, although its impact could not be calculated using prediction tools such as PolyPhen-2, it would definitely damage the PC1 structure and function substantially. Of the remaining three missense variants, chrE3:g.42850283C>T was the only one to be probably damaging, as predicted by PolyPhen-2. Interestingly, when we compared the location of human genes using the UCSC LiftOver tool, this variant was the only variant that had a complete match with a known variant of the human PKD1. This suggests that the same variant can lead to ADPKD in both humans and cats, thus taking the resemblance between feline and human ADPKD and the role of feline ADPKD as a pathological model to another level.
Our study was based on patients referred to the University of Tokyo Veterinary Medical Center, which specialises in secondary care. Thus, it may have had a higher number of sick animals than would a primary care clinic or the Japanese animal population as a whole. We focused on variants that were present only in cases with cystic kidneys and did not further analyse variants that were found in both ADPKD cases and a few control cases. As shown in Table 3, several variants from ADPKD cases were also seen in fewer than five control cases. We cannot deny the possibility of those cases being misdiagnosed as ‘healthy’ or simply being too young to develop detectable cysts, as most of the samples were scanned by ultrasonographic examination, meaning that there may be overlooked variants that actually were the cause of ADPKD. To rule out this possibility, we need to monitor these ‘control cases’ carefully to determine if they will develop ADPKD in the future. The four new variants proposed in this study were only found once each in three out of six samples with cystic kidneys lacking the nonsense transversion on chrE3:g.42858112C>A. In future research, the collection of more ADPKD samples lacking the nonsense variant is required to study the frequency of the four variants. In addition, although two of these variants were predicted to be benign, considering that they were found in cats with ADPKD, there is a possibility that these variants have a destructive effect on the resulting protein. Hence, protein analysis is necessary to evaluate the impact of these variants correctly.
Conclusions
The conventional PKD1 variant (chrE3:g.42858112C>A) was found to be most common in Scottish Folds, Persian cats and Exotic Shorthairs in Japan, and four new variants were suggested as the cause of feline ADPKD. These new variants were all found in exon 15 of PKD1, indicating that exon 15 along with exon 29 are key factors in ADPKD pathogenesis. By promoting further PKD1 analysis, fundamental prevention of the disease can be achieved by controlling the breeding of PKD1 variant carriers.
Supplemental Material
Table 1
A total of 86 non-synonymous variants of PKD1 identified by targeted resequencing in this study.
Supplemental Material
Table 2
Correspondence of each case in a total of 27 variants identified from 6 cases with cystic kidneys.
Footnotes
Acknowledgements
We would like to thank Drs M Yoshida and M Sakaguchi (Azabu University) for providing the feline DNA samples and Dr M Arahori and Ms K Koinuma for helping with target sequencing.
Supplementary material
The following files are available as supplementary material:
Table 1: A total of 86 non-synonymous variants of PKD1 identified by targeted resequencing in this study.
Table 2: Correspondence of each case in a total of 27 variants identified from 6 cases with cystic kidneys.
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by JSPS KAKENHI, a Grant-in-Aid for Science Research (grant numbers 19H00968 and 23H00357), and the Anicom Capital Research Grant (EVOLVE).
Ethical approval
The work described in this manuscript involved the use of non-experimental (owned or unowned) animals. Established internationally recognised high standards (‘best practice’) of veterinary clinical care for the individual patient were always followed and/or this work involved the use of cadavers. Ethical approval from a committee was therefore not specifically required for publication in JFMS. Although not required, where ethical approval was still obtained, it is stated in the manuscript.
Informed consent
Informed consent (verbal or written) was obtained from the owner or legal custodian of all animal(s) described in this work (experimental or non-experimental animals, including cadavers) for all procedure(s) undertaken (prospective or retrospective studies). No animals or people are identifiable within this publication, and therefore additional informed consent for publication was not required.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
