Abstract
Objectives
The aim of this study was to determine the presence of protease-activated receptor 2 (PAR2) and matriptase proteins and quantify PAR2 and matriptase mRNA expression in the articular cartilage and synovial membrane of cats with and without osteoarthritis (OA).
Methods
A total of 28 articular cartilage samples from adult cats (14 OA and 14 normal), 10 synovial membranes from adult cats (five OA and five normal) and three cartilage samples from 9-week-old fetal cats were used. The presence of PAR2 and matriptase in the cartilage and synovial membrane of the adult samples was detected by immunohistochemical (IHC) staining, while real-time PCR was used for mRNA expression analyses in all samples.
Results
PAR2 was detected in all OA and normal articular cartilage and synovial membrane samples but confined to only a few superficial chondrocytes in the normal samples. Matriptase was only detected in OA articular cartilage and synovial membrane samples. PAR2 and matriptase mRNA expression were, however, detected in all cartilage and synovial membrane samples. PAR2 and matriptase mRNA expression levels in OA articular cartilage were five (P <0.001) and 3.3 (P <0.001) times higher than that of the healthy group, respectively. There was no significant difference (P = 0.05) in the OA synovial membrane PAR2 and matriptase mRNA expression compared with the normal samples.
Conclusions and relevance
Detection of PAR2 and matriptase proteins and gene expression in feline articular tissues is a novel and important finding, and supports the hypothesis that serine proteases are involved in the pathogenesis of feline OA. The consistent presence of PAR2 and matriptase protein in the cytoplasm of OA chondrocytes suggests a possible involvement of proteases in cartilage degradation. Further investigations into the PAR2 and matriptase pathobiology could enhance our understanding of the proteolytic cascades in feline OA, which might lead to the development of novel therapeutic strategies.
Introduction
Feline osteoarthritis (OA) is a common and complex progressive disease of feline appendicular joints. Clinically, it is defined as a slowly evolving articular disease characterised by the gradual development of joint pain, stiffness and the limitation of movement with obvious negative effects on the cat’s quality of life. Pathologically, it is defined as a degenerative disorder of synovial joints characterised by deterioration and loss of articular cartilage, and by the formation of new bone at the joint surfaces and margins. It is somewhat confusing and certainly misleading to regard OA purely as a degenerative disorder as inflammatory changes do occur to some extent, particularly within the synovium. Most cases of feline OA appear to be primary (or idiopathic), that is, there is no obvious underlying cause leading to the development of the disease, although secondary OA, where the disease is associated with some other existing joint pathology, is well documented.1 –6 Examples of underlying causes of secondary OA include trauma, hip dysplasia, patellar luxation, cranial cruciate ligament failure, acromegaly, osteochondrodysplasia and mucopolysaccharidosis. 6 Using the term primary (or idiopathic) OA almost definitely hides an ignorance of our understanding of the exact aetiology. Increasing age is the only significant risk factor for primary OA. 6 It often affects multiple joints in a bilaterally symmetrical fashion.
Overt lameness in cats with primary OA is often absent; it is alterations in their behaviour and lifestyle that are more relevant and the use of owner questionnaires is very helpful in identifying these. The most common alterations are reduced activity levels (eg, reduced interaction with owners and sleeping more), reduced mobility (eg, reduced jumping behaviour, reduced height of jumping, unwillingness to jump and stiff gait), reduced grooming, inappropriate toileting behaviour and changes in demeanour (eg, spending more time alone and aggressive behaviour towards other animals or owner).6 –10 A physical orthopaedic examination can be challenging in cats, but all the major limb joints should be carefully and gently palpated and manipulated. 6 Joint thickening, synovial effusion, reduced range of motion and crepitus may be detected but are less obvious than in other species and the common presence of bilateral disease means there is no contralateral normal joint for comparison making their detection even more difficult.
Protease-activated receptor 2 (PAR2) is the second member of the G protein-coupled group of receptors. Both are activated through proteolytic cleavage by serine proteases. Proteases cleave PAR2 at the extracellular N-terminal domain to expose the tethered peptide, which binds to the receptor’s second extracellular loop activating PAR2, thereby initiating an intracellular signal. 11 This activation is irreversible.12,13
Studies in human OA and murine models of OA have demonstrated a key role of PAR2 in mediating arthritic inflammation.14 –20 PAR2 has been found in OA chondrocytes, fibroblasts and macrophages, and reported to play a role in osteophyte formation, cartilage degradation and inflammation. 21 Increased expression of PAR2 mRNA and increased PAR2 protein levels have been observed in chondrocytes and synovial fibroblasts from OA vs normal joints.16,17,22,23 It is known that PAR2 activation can promote the release of proinflammatory cytokines such as interleukin (IL)-1, tumour necrosis factor (TNF)-alpha (α) and IL-6.24 –26 The secreted proinflammatory cytokines upregulate the expression of PAR2, stimulating more secretion of proinflammatory cytokines and metalloproteinases to enhance inflammatory responses and promote cartilage degradation.
Matriptase is a type II transmembrane serine protease and has been recognised as an activator of PAR2. In an OA model study, Milner et al 27 confirmed PAR2 as a substrate for matriptase. In a laser Doppler perfusion study in wild type (WT) and PAR2 deficient (PAR2–/–) mice, matriptase administered into the WT stifle joint resulted in a synovial vasodilator response, which was significantly reduced in PAR2–/– mice. 27 Immunohistochemical (IHC) analysis of the articular cartilage also confirmed the presence of PAR2 and matriptase expression in chondrocytes from the OA model but not in sham-operated mice.
With the demonstration that PAR2 and matriptase have a proinflammatory effect in human OA and murine arthritis, it is hypothesised that PAR2 could play an important role in the pathogenesis of feline OA, particularly in the initiation and progression of cartilage degeneration and loss. The purpose of this study was to determine the presence of PAR2 and matriptase protein and to quantify PAR2 and matriptase mRNA expression in the articular cartilage and synovial membrane of cats with and without OA.
Materials and methods
Sample collection
Ten adult (five OA and five normal) and three 9-week-old fetal cats submitted for post-mortem examination at the School of Veterinary Medicine, University of Glasgow, were used in this study. The cats were euthanased for reasons unrelated to the present study. The articular cartilage and synovium of the elbow joints were identified, based on gross assessment, as osteoarthritic or normal according to Bennett et al. 6 In all cases the appearance of the diseased joints was consistent with a primary OA. Articular cartilage and bone samples collected from osteoarthritic (n = 5) and normal (n = 5) elbow joints were placed into 10% formalin. Additionally, articular cartilage and synovium were harvested and placed into RNAlater (Ambion) and stored at 4°C until further analysis. Eighteen RNA samples (nine OA and nine normal) from the articular cartilage of feline elbow joints received from the Royal (Dick) School of Veterinary Studies, University of Edinburgh, were also included in the study. These samples were also obtained from the elbow joints of adult cats, euthanased for reasons unrelated to the study. The elbow joints were identified as normal or osteoarthritic in the same way as the Glasgow specimens. The RNA had been extracted from slices of articular cartilage in a similar way to that described below.
Tissue processing for IHC staining
The calcified tissues were decalcified in 14% EDTA solution, pH 8, at room temperature until the tissues were adequately soft to enable sectioning with the microtome knife. The decalcified and soft tissues were processed using the Shandon Excelsior (Thermo Scientific) tissue processor for 17 h. The processed tissues were embedded in paraffin wax, sectioned at 3.0 µm using the microtome (Shandon Finesse; Thermo Scientific) and mounted onto glass slides. The slides were dried at 60°C in an oven for 1 h and warmed at 37°C overnight.
IHC analysis
IHC staining was performed according to the protocol of Tindell et al. 28 One section each of cartilage and synovium from the elbow joint of each cat was stained. The optimal PAR2 (Santa Cruz Biotechnology) and matriptase (Sigma-Aldrich) antibody concentration was 2 µg/ml (1:100) and 4 µg/ml (1:50), respectively. Sections of the articular cartilage and synovium were deparaffinised, rehydrated and washed in phosphate buffered saline (PBS) for 5 mins. Antigen retrieval was undertaken in citrate buffer (pH 6.0) for 5 mins. Endogenous peroxidase was blocked by immersing the sections in 1% hydrogen peroxide in methanol for 30 mins. Sections were washed twice with PBS for 5 mins and blocked with supplemented blocking serum for 30 mins. Sections were incubated with primary antibody for 2 h, then secondary antibody for 30 mins and peroxidase-conjugated avidin for 30 mins. The sections were washed twice with PBS before each step. The antigen–antibody complex was visualised using 3,3-diaminobenzidine substrate. Sections were counterstained with haematoxylin, dehydrated, cleared in xylene and mounted with DPX. To determine non-specific staining, replicate sections were incubated with non-specific antibody as isotype control.
One cartilage section per joint was evaluated. Six randomly selected photomicrographs of each section stained for PAR2 and matriptase were taken at × 400 magnification. Included in the photomicrographs were the superficial, middle and deep layers of the articular cartilage and it was confirmed before evaluation that an intact cartilage surface was present. The total number of chondrocytes and the number of positively stained chondrocytes within the photomicrograph were recorded. Results were expressed as the percentage of the number of immunopositive chondrocytes per total number of chondrocytes present.
All synovial membrane sections with the lining and interstitial areas intact were evaluated. Only nucleated cells with a distinct cytoplasmic and interstitial area staining were considered as positive. PAR2 and matriptase synovial membrane immunostaining were scored as 0, 1, 2 and 4, according to Tindell et al. 28 Non-staining was considered as score 0; 1–25% staining as score 1; 26–50% staining as score 2; 51–75% staining as score 3; and 76–100% staining as score 4. The percentage was determined by the number of positively stained cells/nucleus per high-power field.
RNA isolation and cDNA synthesis
Total RNA was isolated using the RNeasy mini column (Qiagen) including an on-column DNase I (Qiagen) digest. Complementary DNA (cDNA) was synthesised from total RNA by using the Precision Reverse Transcription Premix (PrimerDesign) according to the manufacturer’s recommendations. cDNA synthesis was carried out on 1 ng RNA, at 55°C for 20 mins. Absence of contamination of RNA with genomic DNA was verified with minus-reverse transcriptase (minus-RT) controls.
Real-time PCR
Real-time PCR was performed using Precision MasterMix with SYBR green (PrimerDesign) following the manufacturer’s instructions. Each reaction was undertaken in a total volume of 20 μl consisting of 5 µl sample cDNA, 10 µl 2 × Precision MasterMix with SYBR green (PrimerDesign), 1 µl forward and reverse primers and 4 µl RNase free water. Each sample was measured in duplicate. The primer sequences for feline PAR2 were 5′-CTC TTT TCT CCC CAG ACC GC-3′ (forward) and 5′-GGG TAG CAA AGG GGT GTA TGT C-3′ (reverse), and for matriptase they were 5′-AGG ATT GTG ACT GTG GGC TG-3′ (forward) and 5′-GAG AGA TGA TCG AAG CCC CG-3′ (reverse). The reaction was conducted as follows: initial denaturation at 95°C for 10 mins followed by 40 cycles of denaturation at 95°C for 15 s and data collection at 60°C for 1 min. A negative control without a DNA sample was included for every assay. The melting curve was analysed and relative expression of each gene was normalised with the predetermined reference genes, ribosomal protein L18 (RPL18), beta-actin and hypoxanthine–guanine phosphoribosyltransferase and estimated as values of 2-∆∆Ct.
Statistical analysis
The normalised values that reflect the PAR2 and matriptase mRNA levels were analysed using a two-sample t-test to determine whether there was any difference in PAR2 and matriptase mRNA expression levels between the OA and healthy groups. P <0.05 was considered statistically significant. All analyses were performed using GraphPad Prism for Windows, version 8.
Results
PAR2 and matriptase immunoreactivity
Representative immunohistological images of PAR2 and matriptase in articular cartilage and synovial membrane are presented in Figures 1 and 2, respectively. PAR2 was detected in all OA (n = 5) and normal (n = 5) articular cartilage samples with a mean value of 33% (range 21–47%) in the OA group and 3.8% (range 5–14%) in the normal group. In normal articular cartilage, a positive cytoplasmic immunoreactivity of the chondrocytes was observed only in the superficial layer of the cartilage (Figure 1a). In OA articular cartilage positive PAR2 staining occurred in the superficial, middle and deep layers (Figure 1c). PAR2 immunoreactivity was also observed to be variable in different areas of the OA cartilage. In areas where PAR2 was present, the extracellular matrix was only faintly stained with haematoxylin (Figure 1c). Non-specific stained sections showed only background staining (Figure 1e,f).

Representative sections showing (a,c) protease-activated receptor 2 (PAR2) and (b,d) matriptase immunostaining in (a,b) normal and (c,d) osteoarthritic (OA) articular cartilage. (a–f) × 200 magnification. (a) In normal articular cartilage, PAR2 expression is seen in a few superficial chondrocytes (yellow arrows). (b) Matriptase immunoreactivity is not detected in normal articular cartilage. (c) PAR2 (yellow arrows) is diffuse from the superficial layer to the middle and deep layers of the OA articular cartilage. Only faint haematoxylin staining is seen in the extracellular matrix (asterisks). (d) Matriptase is present only in locations where PAR2 is also present. The extracellular matrix is only faintly stained with haematoxylin (asterisks). (e,f) There is no staining in articular cartilage when a primary antibody of irrelevant specificity was used
Positive immunostaining for PAR2 was observed in the synovial lining layer and interstitial area in all OA (n = 5) and normal (n = 5) synovial membrane samples (Figure 2a,c,d). Immunoreactivity was present in the cytoplasm of the synovial cells in three OA synovial membrane samples displaying score 3 staining and two displaying score 4 staining. All the normal synovial membrane samples showed score 2 staining.
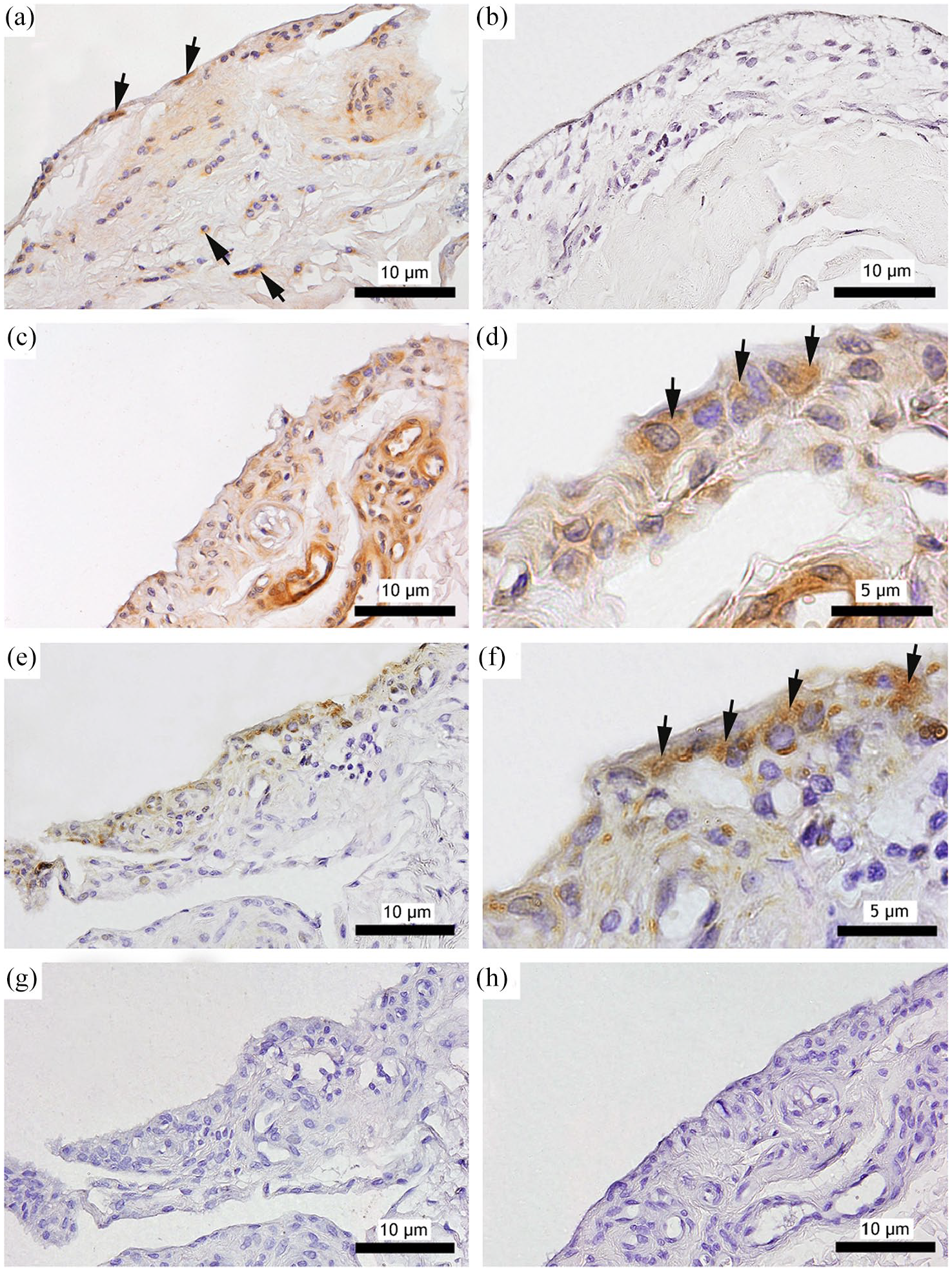
Presence of (a,c,d) proteinase-activated receptor 2 (PAR2) and (b,e,f) matriptase in (a,b) normal and (c–f) osteoarthritic (OA) synovial membrane. (a–c,e,g,h) × 200 magnification, (d,f) × 400 magnification. (a) PAR2 immunoreactivity (black arrows) is seen in the normal synovial membrane. (b) Matriptase immunoreactivity is not identified in normal synovial membrane. (c) PAR2 is expressed in the synovial lining layer from OA joints. (d) Higher magnification shows PAR2 is expressed in the cytoplasm of synovial lining cells (black arrows). (e) Matriptase protein is present in the synovial lining layer. (f) Higher magnification shows matriptase expression in the cytoplasm of the synovial lining cells (black arrows). (g,h) There is no staining in the synovium when a primary antibody of irrelevant specificity was used
Matriptase immunoreactivity was not detected in any of the normal cartilage (n = 5) and synovial membrane (n = 5) samples (Figures 1b and 2b). Matriptase staining was detected in all OA (n = 5) articular cartilage samples (Figure 1d). The mean percentage of matriptase-positive chondrocytes was 27% (range 12–42%). Chondrocyte distribution of matriptase immunoreactivity demonstrated considerable variation; in three cats the expression was intense particularly at the superficial and middle layers (Figure 1d), and in the other two cats virtually all the chondrocytes exhibited faint matriptase immunoreactivity in all layers. Despite the variability in the immunoreactivity, matriptase was seen only in locations where PAR2 was present. In the areas where matriptase was present, extracellular matrix was only faintly stained with haematoxylin (Figure 1d).
Matriptase immunoreactivity was also detected in all OA (n = 5) synovial membrane samples mainly in the cytoplasm of the lining cells (Figure 2e,f). All OA synovial membranes showed score 2 staining. Immunoreactivity was not detected in any of the normal synovial membrane samples (Figure 2b).
Comparative assessment of PAR2 and matriptase mRNA levels in cats with and without OA
PAR2 mRNA expression was detected in all articular tissues (14 OA, 14 normal and three fetuses). The PAR2 mRNA expression levels in OA and fetal articular cartilage were five and three times higher, respectively, than the normal adult group (Figure 3a). The PAR2 mRNA expression level in OA synovium was 1.5 times higher than in normal synovium (Figure 3b). A significantly higher expression of PAR2 mRNA (P = 0.0003) was seen in the OA articular cartilage group (Figure 3c). Synovial membrane samples did not show significantly different levels of expression between normal (n = 5) and OA (n = 5) samples (P = 0.3553) (Figure 3d). In view of the small sample size, statistical analysis for the fetal samples was not performed.

Protease-activated receptor 2 (PAR2) mRNA expression as determined by quantitative PCR in articular cartilage and synovium. (a) Bar graph shows the fold-change in PAR2 mRNA expression in osteoarthritic (OA) (n = 14) and fetal (n = 3) articular cartilage vs normal (n = 14). The PAR2 mRNA expression levels in OA and fetal articular cartilage were five and three times higher than the normal group, respectively. (b) Bar graph depicts the fold-change in PAR2 mRNA expression in OA synovium (n = 5) vs normal (n = 5). The PAR2 mRNA expression level in synovium was 1.5 times higher than the normal group. (c) The PAR2 mRNA expression of the OA articular cartilage was significantly higher than the normal group. (d) There is no significant difference in synovium PAR2 mRNA expressions between the OA and normal group. (c,d) Data are presented as mean ± SD (two-sample t-test, ***P = 0.0003).
Matriptase mRNA expression was identified in all articular tissues (14 OA, 14 normal and three fetuses). The matriptase mRNA expression levels in OA and fetal articular cartilage were 3.3 and 1.4 times higher, respectively, than the normal group (Figure 4a). Matriptase mRNA expression level in OA synovium was 1.1 times higher than the normal group (Figure 4b). The matriptase mRNA expression of the OA articular cartilage was significantly higher than the normal group (P = 0.0007) (Figure 4c). There was no significant difference in the OA synovium (n = 5) matriptase mRNA expression compared with the normal samples (n = 5) (P = 0.3801) (Figure 4d).
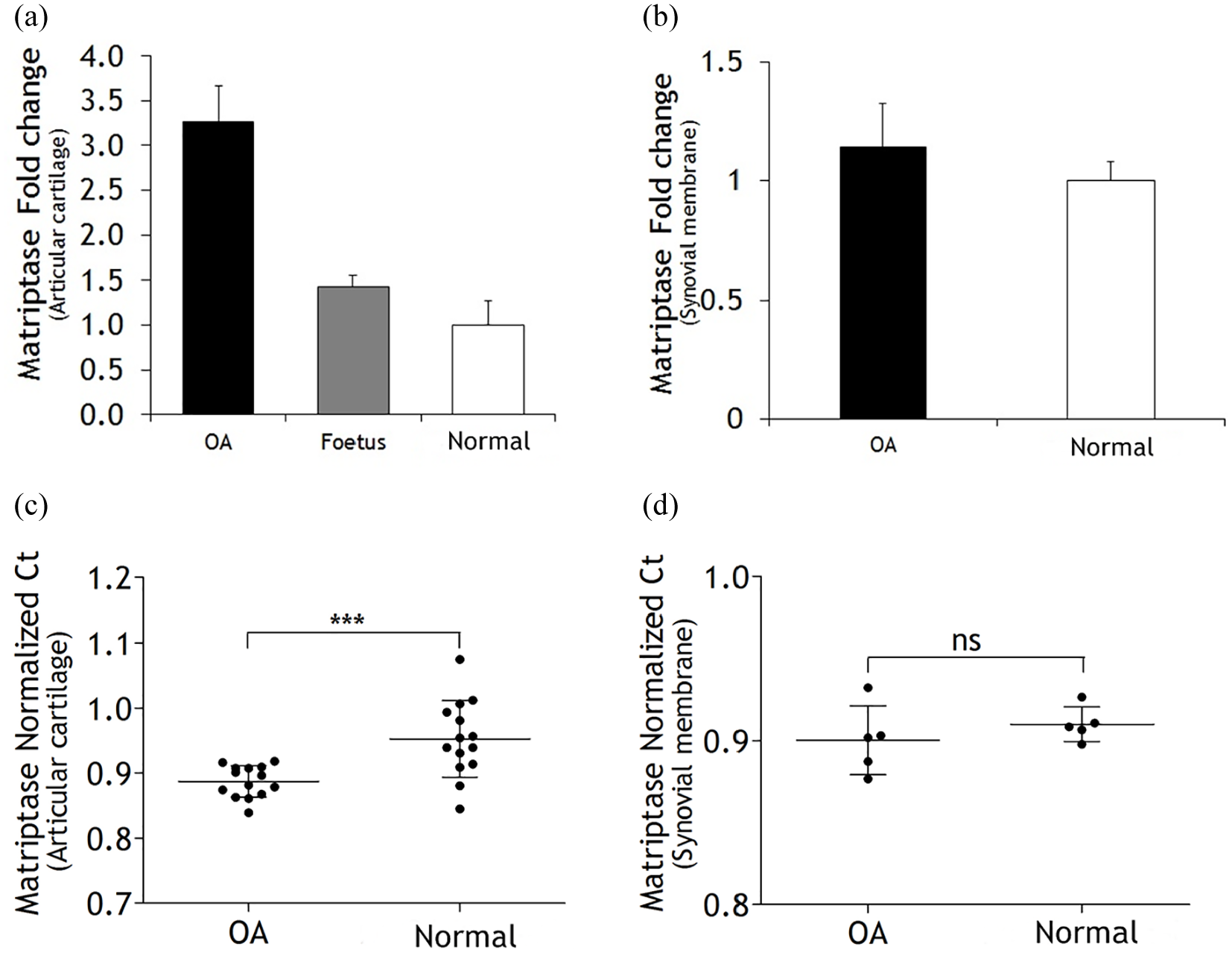
Matriptase mRNA expression as determined by quantitative PCR in articular cartilage and synovium. (a) Bar graph shows the fold-change in matriptase mRNA expression in osteoarthritic (OA; n = 14) and fetal (n = 3) articular cartilages vs normal (n = 14). The matriptase mRNA expression levels in OA and fetal articular cartilage were 3.3 and 1.4 times higher than the normal group, respectively. (b) Bar graph depicts the fold-change in matriptase mRNA expression in OA synovium (n = 5) vs normal (n = 5). The matriptase mRNA expression level in OA synovium was 1.1 times higher than the normal group. (c) The matriptase mRNA expression of the OA articular cartilage was significantly higher compared with the normal group. (d) There is no significant difference between the OA synovium matriptase mRNA expression and the normal group. (c,d) Data are presented as mean ± SD (two-sample t-test, ***P = 0.0007).
Discussion
The present study demonstrates for the first time that PAR2 is present in feline articular cartilage and synovial membrane of normal and OA cats. In normal articular cartilage PAR2 protein was found mainly in the superficial layers, while in OA articular cartilage PAR2 protein was present in the superficial, middle and deep layers of the cartilage. These results demonstrated that in OA, PAR2 protein is present throughout the layers of the cartilage matrix and upon activation it is very likely to be involved in cartilage degradation. The presence of PAR2 protein also varies in different areas of the OA cartilage, which might reflect local differences in severity of cartilage pathology.
Matriptase protein was consistently present in articular cartilage and synovial membrane of OA cats. Matriptase protein was seen in locations where PAR2 was also present. None of the normal cat samples showed immunoreactivity for matriptase. Matriptase was reported as absent in normal human articular cartilage. 29 Matriptase was also not detected in the chondrocytes of normal articular cartilage of laboratory mice but was consistently seen in OA articular cartilage. 27 Similarities in matriptase distribution in cat, human and mouse chondrocytes may reflect genuine physiological similarities in rodents, Felidae and primates.
PAR2 and matriptase are important mediators of collagenolytic activity in OA cartilage.17,27 Recently, matriptase was reported to induce the release of matrix metalloproteases (MMPs) from OA cartilage. 30 Collagenolysis initially occurs in the peri-cellular area of chondrocytes, suggesting proteases are associated with the chondrocyte cell surface. 31 Collagen breakdown allowing proteoglycans to ‘escape’ from the matrix is supported by the faint staining with haematoxylin in the areas where PAR2 and matriptase were present. Genetic disruption of PAR2 in murine OA models was associated with a significantly delayed progression of cartilage damage and inhibition of subchondral bone thickening, following destabilisation of the medial meniscus.19,32
Quantitative real-time PCR showed that PAR2 and matriptase mRNA were expressed in feline normal and OA articular tissues. PAR2 and matriptase expression levels were significantly increased in OA compared with normal feline articular cartilage. Specific mechanisms that trigger activation of matriptase are not completely understood. 33 However, PAR2 has been identified as one of the natural substrates for matriptase. 34 Matriptase activates PAR2 via the proteolytic release of a tethered ligand. 35 Matriptase has several functions as an inducer and activator of procollagenases, which lead to cartilage breakdown. 27 PAR2 activation has been associated with increased production of MMP-1 and MMP-13, as well as cyclooxygenase-2, which are key factors in OA pathophysiology. 17 In addition, proinflammatory cytokines such as IL-1beta (β) and TNF-α upregulate PAR2 expression in OA chondrocytes. 16 Thus, in OA cartilage, gene expression of high levels of matriptase could possibly indicate a role of matriptase and PAR2 in OA pathogenesis. However, further studies need to be carried out to characterise the pathobiological activities of matriptase and PAR2 in feline articular tissues with and without OA.
It has been reported that PAR2 expression is strongly correlated with synovitis in human OA and murine OA models.20,28 PAR2 expression has been shown in fibroblast-like synoviocytes and macrophages and lymphocytes derived from OA synovial membranes.16,28 PAR2 regulates the synovial release of proinflammatory cytokines, including IL-1β, which, in turn, upregulates PAR2 expression in OA chondrocytes.16,17,36 The present study showed that the fold-change expression levels of PAR2 and matriptase mRNA in OA synovium was higher than that in normal synovium. However, there was no significant difference between the expression levels of PAR2 and matriptase mRNA in the synovium samples from the normal and OA cats. This finding could be associated with the degree of inflammatory cell infiltration in the feline synovium, which, even in advanced OA, is generally mild. 37 Another explanation could be the limited sample quantity, in which case a larger number of synovial membrane samples needs to be studied. Although the PAR2 and matriptase gene expression were minimally different between normal and OA synovium samples, protein expression was not, which suggests that gene transcription is initiated/increased in the synovium of OA joints at some stage of the disease process.
PAR2 expression during embryonic development has been reported in murine tissues. 38 In developing chondrocytes, PAR2 expression was detectable at 17 days of gestation but, interestingly, was absent in adult cartilage. However, the study did not specifically state the age of the adult mice. The authors suggested that chondrocytic expression of PAR2 appears to be associated with metabolically active cartilage. Our study showed an increase fold-change in PAR2 expression in fetal cartilage (9 weeks of age), which supports the study by Jenkins et al. 38 Our study also showed an increased expression of matriptase mRNA vs normal adult tissue. However, no statistical data could be provided to support these observations owing to the small fetal sample size. The rapid growth of cartilage in the fetus with increased ‘turnover’ of cartilage matrix (degradation followed by replacement) could involve PAR2 and matriptase. Indeed, there is a continuous ‘turnover’ of normal cartilage throughout life and one might expect PAR2 and matriptase to be present even in normal tissue, albeit at lower levels.
Conclusions
The identification of PAR2 and matriptase proteins and gene expression in feline articular tissues is a novel and important finding supporting the hypothesis that serine proteases and protease ‘sensing’ receptors are involved in feline OA. The consistent presence of PAR2 and matriptase protein in the cytoplasm of OA chondrocytes suggests a possible involvement of proteases in cartilage degradation in OA. This suggests that cartilage degradation in feline OA could, in all probability, be stimulated by the activation of matriptase and PAR2 at the cellular level, which, in turn, facilitates pericellular collagenolysis of the matrix. Further investigations into PAR2 and matriptase pathobiology could enhance our understanding of the proteolytic cascades in feline OA, as well as the mechanism of cartilage ‘turnover’ in normal tissues, both of which might help our understanding of the aetiopathogenesis of feline OA and provide a basis for the future development of novel therapeutic strategies.
Footnotes
Acknowledgements
Special thanks are accorded to Professor William Ferrell, Institute of Immunity, Infection and Inflammation, University of Glasgow, for the use of his laboratory to undertake this study.
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors wish to thank the Ministry of Higher Education Malaysia for the award of a PhD scholarship under Skim Latihan Akademik IPTA (SLAI), which contributed towards the completion of this study. Financial support was also provided by the Programme Grant from Versus Arthritis (grant #20199).
Ethical approval
This work involved the use of non-experimental animals only (including owned or unowned animals and data from prospective or retrospective studies). Established internationally recognised high standards (‘best practice’) of individual veterinary clinical patient care were followed. Ethical approval from a committee was therefore not necessarily required.
Informed consent
Informed consent (either verbal or written) was obtained from the owner or legal custodian of all animal(s) described in this work (either experimental or non-experimental animals) for the procedure(s) undertaken (either prospective or retrospective studies). For any animals or humans individually identifiable within this publication, informed consent (either verbal or written) for their use in the publication was obtained from the people involved.
