Abstract
Objectives
The aim of this study was to evaluate changes in the conjunctival microbiota of shelter-housed cats with time, upper respiratory disease (URD) and famciclovir administration.
Methods
Cats were assigned to treatment groups on shelter entry. Healthy cats or cats with URD received ~30 mg/kg or ~90 mg/kg of famciclovir or placebo PO q12h for 7 days, or were untreated. Swabs were collected from ventral conjunctival fornices prior to (day 1) and immediately after (day 8) the treatment period. Microbiota analysis was conducted on 124 randomly selected swabs from healthy (56 swabs) or URD-affected (68 swabs) cats. Following DNA extraction and amplification of the V4 region of the 16S rRNA gene, sequences were assembled into operational taxonomic units (OTUs). Over-represented OTUs (as determined by linear discriminate analysis effect size), alpha and beta diversity, and median relative abundance of known feline ocular surface pathogens were assessed for the entire population and in 10 clinically relevant subpopulations of cats.
Results
Bacteria from 33 phyla and 70 genera were identified. Considering all cats, median relative abundance of Mycoplasma increased from day 1 to day 8, while Proteobacteria decreased. Community membership and structure (beta diversity) differed between days 1 and 8 for all famciclovir-treated cats (regardless of health status or dose) and healthy or URD-affected cats (regardless of famciclovir dose). Differences in taxonomic diversity within a sample (alpha diversity) between day 1 and day 8 were not detected in any subpopulations.
Conclusions and relevance
Within 1 week of shelter entry, there were significant changes in community structure and membership of the feline conjunctival microbiota, with a shift towards over-representation of feline ocular surface pathogens. Although famciclovir may impact beta diversity of the feline conjunctival microbiota, absence of change in alpha diversity suggests minimal shift in individual cats.
Introduction
The community of living microorganisms in a particular environment, including its inhabitant bacteria, viruses, fungi and protozoa, is termed the microbiota.1–4 By contrast, the microbiome describes all genetic material within the microbiota.4,5 Initial investigation of the microbiota and microbiome was limited to culture-based and similar techniques. The advent of metagenomic techniques, such as 16S ribosomal RNA (16S rRNA) gene sequencing, as well as bioinformatics systems, have allowed broader and more detailed investigation of the microbiome and microbiota, and their associations with various host factors and pharmacologic interventions.2,3,5–9
Evaluation of the bacterial microbiota can involve various approaches, including comparison of specific members (eg, different phyla and genera), alpha diversity indices (ecological indicators of the bacterial community in a sample) and assessment of beta diversity (evaluation of differences in overall bacterial communities between groups). Additional approaches can be used to provide different insights, such as the use of linear discriminate analysis effect size (LEfSe) to identify operational taxonomic units (OTUs) that are differentially abundant in species groups. Microbiota analysis frequently begins following assembly of sequences with greater than 97% nucleotide identity into groups called OTUs.3,10 Alpha diversity metrics indicate the taxonomic diversity (inverse Simpson’s index), evenness (Shannon’s evenness index) and richness (number of species observed [observed species richness: SOBS] and Chao1 index).3,10–12 Beta diversity metrics include measurements of microbial community membership (number of shared genera; Jaccard’s index) and structure (number of shared genera and their relative abundances; Yue and Clayton index) between samples. 10 A wide range of methods can be used to compare statistically the composition of microbiota communities, such as analysis of molecular variance (AMOVA) and unweighted UniFrac. Briefly, UniFrac measures differences in branch lengths between taxa in a phylogenetic tree, 13 and AMOVA measures differences between the genetic diversity of multiple communities and their pooled genetic diversity. 14 While there are no standard approaches to microbiota assessment, a combination of assessment of taxonomic differences, alpha diversity and beta diversity allow for detailed assessment of specific and broad aspects of the microbiota.
Host–microbiome interactions are complex, and whether changes in the host precede changes in its microbiome, or vice versa, is largely unknown; however, the relationship is undoubtedly important. For example, studies in humans have proposed a connection between the microbiome and cancer, obesity, asthma, atherosclerosis and diabetes.2,15 The ocular microbiome has also received attention, with blepharitis, meibomian gland disorders, chronic inflammation, ocular rosacea, 16 sleep duration 17 and contact lens use16,18 each associated with alterations in composition of the human ocular microbiome.
The feline microbiota has received some research attention. In clinically normal cats, the conjunctival microbiome (lower conjunctival sac/fornix, eyelid margins, and palpebral conjunctiva of third eyelid) has been examined using culture-based19–21 and metagenomic/bioinformatics 22 techniques. Changes to the nasal microbiome have been identified in cats of different ages and those living in different environments, 5 and as a function of season. 19 Likewise, the microbiome of cats affected by feline immunodeficiency virus (FIV), 22 acute and chronic gastrointestinal disease, 23 and conjunctivitis 20 have all been demonstrated as quantitatively or qualitatively different than those of their healthy counterparts. However, to our knowledge there are no published genomic studies assessing the microbiota of cats housed within a shelter environment, or the effect of upper respiratory disease (URD) or the administration of antiviral medications. Considering the ubiquitous nature of URD infection in shelter-housed cats,24,25 the impact of host and environmental factors on the host microbiome and development of feline URD, alterations in the microbiome of cats with conjunctivitis, 20 and the effects of time in a shelter on shedding of viral and bacterial pathogens associated with feline URD,26,27 we were interested in assessing the conjunctival microbiota of cats in a shelter environment. The objectives of the present study were to evaluate the conjunctival microbiota of clinically healthy and URD-affected, shelter-housed cats for changes associated with time, URD status and treatment effect of two doses of the antiviral agent famciclovir.
Materials and methods
Cats and treatment groups
This study used conjunctival swabs collected as part of a project assessing the prophylactic and therapeutic effects of famciclovir on infectious URD in shelter-housed cats. 28 All samples were collected between March and November 2016 from cats housed in a county animal shelter in Northern California. All cats were held for a legally mandated 72 h period after arriving at the shelter, during which time they underwent behavioral testing, and intact cats were neutered. Cats were considered for study enrollment within 48 h of being listed for adoption. The study was approved by the Institutional Animal Care and Use Committee of the University of California–Davis (IACUC #18929) and designed in accordance with the Association for Research in Vision and Ophthalmology statement for the ethical use of animals in ophthalmic research.
Inclusion criteria were body weight ⩾1 kg, body condition score ⩾3/9, negative serum feline leukemia virus antigen and FIV antibody test results (SNAP Combo FeLV Ag-FIV Antibody test; IDEXX) and aged ⩾3 months old (as estimated based on dental examination). Any cat that that lost >15% of its entry body weight was excluded from the study. 28 Cats meeting all the inclusion criteria were graded by masked observers for clinical signs of URD according to an established semi-quantitative scoring system for clinical evidence of FHV-1-associated disease. 29 All ocular (conjunctivitis, blepharospasm and ocular discharge) and non-ocular (sneezing and nasal discharge) scores were summed to determine the total clinical disease score, which ranged from 0 to 24. Cats were then randomized to one of seven treatment groups by a single researcher uninvolved in subsequent clinical assessments. Randomization was performed such that all groups in both arms had approximately equivalent sex and age distribution. Cats without evidence of URD (ie, clinical disease score = 0) received either no treatment or ~30 mg/kg or ~90 mg/kg PO q12h for 7 days famciclovir. Cats with evidence of URD (ie, clinical disease score ⩾1) received either no treatment, or lactose (placebo), or ~30 or ~90 mg/kg of famciclovir PO q12h for 7 days. 28
Swab collection and microbiota analysis
On day 1 (study entry) and day 8 (cessation of famciclovir/placebo administration), cats in all seven treatment groups received one drop of 0.5% proparacaine (Akorn) in each eye, and a dry polyester swab (Fisher Scientific) was rolled vigorously in the ventral conjunctival fornix of both eyes. All swabs were immediately placed into cryovials, stored on ice for 1–3 h, and then at −80°C until genomic analysis. Samples were collected for microbiota assessment from 158 cats from which a convenience subset of 151 swabs from 99 cats was selected by laboratory personnel masked to all information about sample source.
The DNA was extracted from swabs using the Qiagen DNeasy Power Soil Kit with an extra step of heating at 70°C before continuing with step 5. The DNA was eluted into a 30 ml volume of elution buffer. Quantification of DNA was performed using a NanoDrop ND-1000 UV-Vis Spectrophotometer (NanoDrop Technologies). The V4 region of the 16S rRNA gene was amplified using the modified primers 515-F/806-R.30,31 Each 25 µl mix consisted of 12.5 µl of KAPA2G Fast HotStart ReadyMix (KAPA Biosystems), 0.5 µl (10 pM/µl) of each of the primers, 2 µl of DNA template and 9.5 µl of PCR-grade water. The following PCR conditions were used: 3 mins at 94°C for denaturing, followed by 30 cycles of 45 s at 94°C, 60 s at 52°C and 60 s at 72°C, with a final period of 10 mins at 72°C. The PCR products were purified with Mag Bind RXNPure Plus beads (Omega Bio-tech). A second PCR reaction was used for attachment of Illumina adapter primers (N716-N729 and S513-S522). A final mix containing 5.0 µl of DNA template, 4.0 µl of primers (2.5 pM/µl) and 12.0 µl of PCR-grade water underwent cycling for 3 mins at 94°C followed by eight cycles of 30 s at 94°C, 30 s at 55°C and 30 s at 72°C, plus a final period of 10 mins at 72°C. After purification (Mag Bind RXNPure Plus; Omega Bio-tech), sequencing was performed using Illumina MiSeq and 2 × 250 chemistry.
Forward and reverse sequence reads were assembled and subjected to a series of quality control steps using Mothur 1.42.3 to remove primers, sequences that were of abnormal size or contained ambiguous base calls, sequences not consistent with the V4 regions and chimeras. 32 Sequences were assembled into OTUs using a de novo (open OTU) picking approach with a 3% dissimilarity cut-off. Individual sequences and OTUs were classified using the Ribosomal Database Project Classifier. 33
Data analysis
Prior to analysis, data were assessed using the Shapiro–Wilk test of normality. Because a large proportion of data sets were non-normally distributed, all data were converted to ranks using commercially available software (SigmaPlot 14.0; Systat Software). Sequence data were unmasked and linked to the individual cat and swab from which they were collected. Data were excluded from any cat that developed illness requiring medical intervention. This included development of URD and/or ocular disease treated with antibiotics administered orally and/or topically (15 cats), diarrhea treated with sulfadimethoxine (one cat), and/or gastrointestinal parasitism treated with fenbendazole (one cat). Additionally, one cat developed liver failure and was humanely euthanized. According to these criteria, 27 samples from 18 cats were excluded from subsequent analyses.
To evaluate for microbial changes within the entire study population, data from swabs collected on day 1 were compared with those collected on day 8 in all cats (n = 81) collectively. To assess for other clinically relevant changes, sequence data were re-grouped to allow more granular comparisons among 10 subpopulations: healthy untreated cats; URD-affected untreated cats; URD-affected placebo-treated cats; URD-affected and healthy famciclovir-treated cats; URD-affected famciclovir-treated cats; URD-affected cats receiving 30 mg/kg famciclovir; URD-affected cats receiving 90 mg/kg famciclovir; healthy famciclovir-treated cats; healthy cats receiving 30 mg/kg famciclovir; and healthy cats receiving 90 mg/kg famciclovir.
Alpha diversity was analyzed by comparing diversity (inverse Simpson index), evenness (Shannon’s evenness index) and richness (number of species observed and Chao1 index) among subpopulations. Differences in alpha diversity and median relative abundance between days 1 and 8 were assessed using the Kruskal–Wallis one-way ANOVA on ranks with Dunn’s all multiple comparison procedures for difference in ranks. Differences in alpha diversity and median relative abundance between doses of famciclovir were each assessed using the Mann–Whitney rank sum test with a Yates continuity correction. Beta diversity was analyzed by UniFrac analysis of Jaccard and Yue and Clayton indices so as to compare microbial community membership (unweighted) and structure (weighted), respectively, and via AMOVA.
To ensure that specific pathogens were individually assessed, median relative abundance of selected genera or phyla containing known feline ocular surface pathogens (Mycoplasma, Streptococcus, Pseudomonas, Chlamydiae and Proteobacteria) were compared. The LEfSe method was used to identify differentially abundant OTUs for each time point in each population. An OTU with a linear discriminate analysis >4 was considered to be over-represented. For assessment of alpha and beta diversity, sequence numbers were normalized through random selection of 6480 sequences per sample. Beta diversity was also analyzed using non-subsampled data; values obtained were very similar to those for the subsampled data; therefore, no further analysis of non-subsampled data was performed.
All analyses of alpha diversity and median relative abundance were performed using commercial software (SigmaPlot 14.0; Systat Software), and P values were adjusted for false discovery rate using the Benjamini–Hochberg technique. All beta diversity analyses were performed using open access software (Mothur v1.42.3). 32 For all statistical analyses, a P value of ⩽0.05 was considered significant.
Results
Following exclusion of 27 samples from 18 cats that received medications other than famciclovir during the study period, 124 conjunctival swab samples that passed all quality control steps were assessed. These comprised 56 swabs from healthy cats and 68 swabs from URD-affected cats (Figure 1) from which bacteria belonging to 33 phyla and 70 genera were identified.
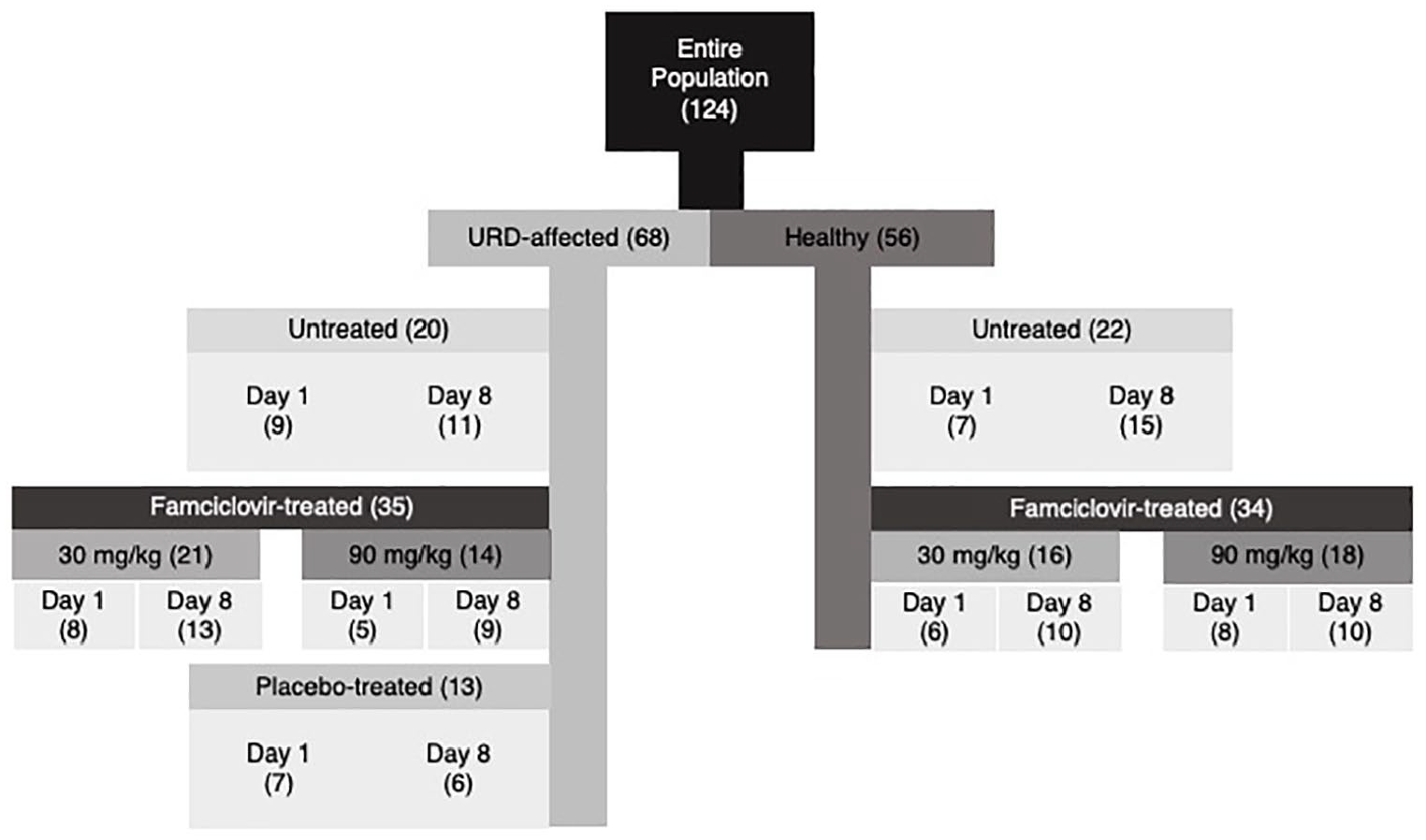
Number of conjunctival swabs (in parentheses) undergoing microbiota analysis from 81 shelter-housed cats with or without clinical evidence of upper respiratory disease (URD) that were untreated or received orally administered famciclovir at one of two doses or placebo (lactose) q12h for 7 days. Swabs were collected prior to (day 1) and immediately after (day 8) the 7-day treatment period
Effect of time in the shelter on feline ocular surface pathogens and the conjunctival microbiota of the entire study population
Considering the entire population (irrespective of cat health status and treatment group), Mycoplasma, Streptococcus and Pseudomonas were isolated from all 124 samples, and their median relative abundance was 0.0117, 0.0268 and 0.0021, respectively (Figure 2). Proteobacteria were detected in all 124 samples analyzed and Chlamydiae in 48/124 (39%). Median relative abundance for Proteobacteria and Chlamydiae was 0.1596 and 0, respectively (Figure 3). Of the five ocular surface pathogens analyzed, significant differences in relative abundance between days 1 and 8 were detected only for Mycoplasma and Proteobacteria. For all comparisons of clinical interest (Table 1), median relative abundance of Mycoplasma tended to increase from day 1 to day 8, and did so significantly in three populations assessed – all 81 cats when considered as a whole, those receiving famciclovir regardless of health status or famciclovir dose, and URD-affected untreated cats. By contrast, median relative abundance for Proteobacteria decreased significantly over the study week only when the entire study population was considered as a whole (Table 1).

Distribution of 11/70 bacterial genera isolated from 124 conjunctival swabs collected from 81 shelter-housed cats with (68 swabs) or without (56 swabs) clinical evidence of upper respiratory disease (URD). Cats were untreated or received orally administered famciclovir at 30 mg/kg or 90 mg/kg or placebo (lactose) q12h for 7 days. Genera selected for graphic representation were those over-represented at any time for any population and those that contained ocular surface pathogens. Swabs were collected prior to (day 1; left bar of each pair on chart) and immediately after (day 8; right bar of each pair on chart) the 7-day study period. Data are presented for all swabs combined (ALL), ie, day 1 and day 8 samples from the entire study population, and for the 11 clinically relevant populations described in Figure 1: (A) entire population; (B) healthy untreated cats; (C) URD-affected untreated cats; (D) URD-affected placebo-treated cats; (E) healthy and URD-affected famciclovir-treated (30 mg/kg or 90 mg/kg) cats; (F) healthy famciclovir-treated (30 mg/kg or 90 mg/kg) cats; (G) URD-affected famciclovir-treated (30 mg/kg or 90 mg/kg) cats; (H) URD-affected famciclovir-treated (30 mg/kg) cats; (I) healthy famciclovir-treated (30 mg/kg) cats; (J) URD-affected famciclovir-treated (90 mg/kg) cats; and (K) healthy famciclovir-treated (90 mg/kg) cats

Distribution of 6/33 bacterial phyla isolated from 124 conjunctival swabs collected from 81 shelter-housed cats with (68 swabs) or without (56 swabs) clinical evidence of upper respiratory disease (URD). Cats were untreated or received orally administered famciclovir at 30 mg/kg or 90 mg/kg or placebo (lactose) q12h for 7 days. Phyla selected for graphic representation were those with a median relative abundance >0.046 and those that contained ocular surface pathogens. Swabs were collected prior to (day 1; left bar of each pair on chart) and immediately after (day 8; right bar of each pair on chart) the 7-day study period. Data are presented for all swabs combined (ALL), ie, day 1 and day 8 samples from the entire study population, and for the 11 clinically relevant populations described in Figure 1: (A) entire population; (B) healthy untreated cats; (C) URD-affected untreated cats; (D) URD-affected placebo-treated cats; (E) healthy and URD-affected famciclovir-treated (30 mg/kg or 90 mg/kg) cats; (F) healthy famciclovir-treated (30 mg/kg or 90 mg/kg) cats; (G) URD-affected famciclovir-treated (30 mg/kg or 90 mg/kg) cats; (H) URD-affected famciclovir-treated (30 mg/kg) cats; (I) healthy famciclovir-treated (30 mg/kg) cats; (J) URD-affected famciclovir-treated (90 mg/kg) cats; and (K) healthy famciclovir-treated (90 mg/kg) cats
Median relative abundance of Mycoplasma and Proteobacteria from 124 conjunctival swabs collected from 81 shelter-housed cats with (68 swabs) or without (56 swabs) clinical evidence of upper respiratory disease (URD) that were untreated or received orally administered famciclovir at one of two doses or placebo (lactose) q12h for 7 days

Swabs were collected prior to (day 1) and immediately after (day 8) the 7-day study period. Data are compared across the entire population and for all clinically relevant subpopulations. The values in bold are significant at P ⩽0.05. No significant differences were detected for the remaining three ocular surface pathogens (Chlamydiae, Streptococcus and Pseudomonas)
BH = Benjamini–Hochberg
LEfSe analysis revealed that Escherichia/Shigella was over-represented on day 1 and Clostridium cluster XI, Mycoplasma, Staphylococcus and Streptococcus were over-represented on day 8 (Table 2). Both community membership and community structure differed significantly between days 1 and 8, whether assessed by AMOVA or UniFrac (Figures 4 and 5; Table 3). However, no significant differences in observed or estimated richness, diversity or evenness were identified among all cats between days 1 and 8 (Figures 6 and 7).
Over-represented operational taxonomic units (OTUs) of bacterial genera from 124 conjunctival swabs collected from 81 shelter-housed cats with (68 swabs) or without (56 swabs) clinical evidence of upper respiratory disease (URD)

Swabs were collected prior to (day 1) and immediately after (day 8) a 7-day study period during which cats were untreated or received orally administered famciclovir at 30 mg/kg or 90 mg/kg or placebo (lactose) q12h. For analysis, cats were considered within the following 11 subgroups: (A) entire population; (B) healthy untreated cats; (C) URD-affected untreated cats; (D) URD-affected placebo-treated cats; (E) healthy and URD-affected famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (F) healthy famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (G) URD-affected famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (H) URD-affected famciclovir-treated (30 mg/kg) cats; (I) healthy famciclovir-treated (30 mg/kg) cats; (J) URD-affected famciclovir-treated (90 mg/kg) cats; and (K) healthy famciclovir-treated (90 mg/kg) cats. OTUs were defined as those with linear discriminate analysis (LDA) values >4.00 based on LDA effect size (LEfSe) analysis

Beta diversity of the conjunctival microbiota analyzed using Jaccard’s index (community membership) performed on 124 conjunctival swabs collected from 81 shelter-housed cats with (68 swabs) or without (56 swabs) clinical evidence of upper respiratory disease. Each point represents the microbial community of a conjunctival swab collected prior to (day 1; blue dots and ellipse) or immediately after (day 8; red dots and ellipse) a 7-day study period, during which the cats were untreated or received orally administered famciclovir at 30 mg/kg or 90 mg/kg, or placebo (lactose) q12h. Data for each day represent the entire study population irrespective of treatment group or disease status. Non-metric multidimensional scaling (NMDS) Axis 1 and Axis 2 represent the axes of the two-dimensional ordination space. Ellipses represent the 95% confidence intervals at each time point
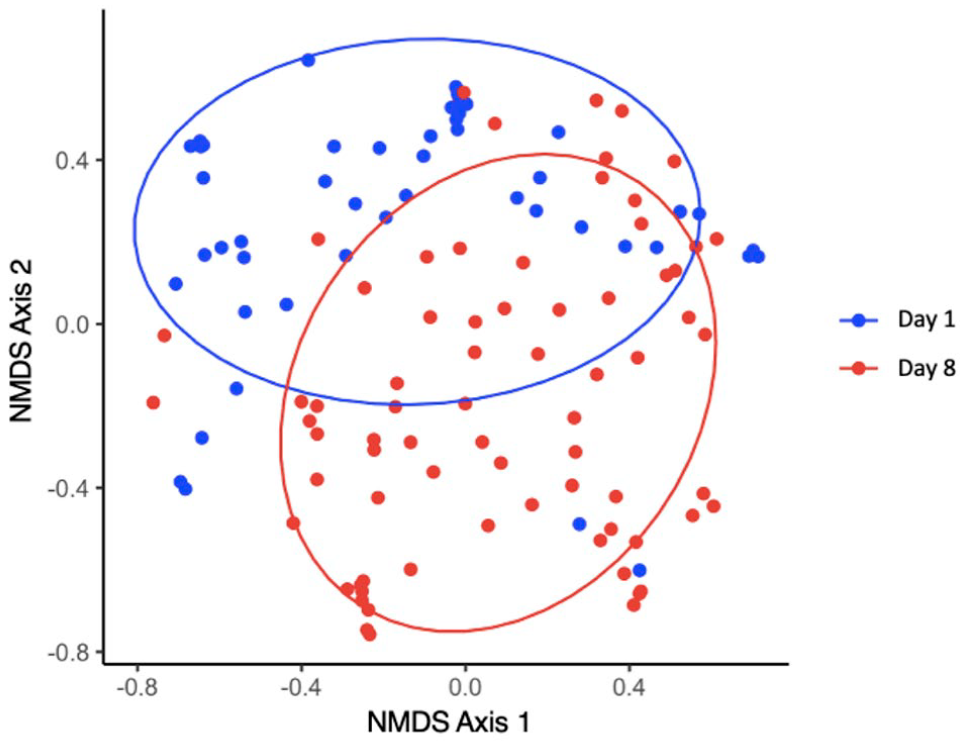
Beta diversity of the conjunctival microbiota analyzed using Yue and Clayton’s index (community structure) performed on 124 conjunctival swabs collected from 81 shelter-housed cats with (68 swabs) or without (56 swabs) clinical evidence of upper respiratory disease. Each point represents the microbial community of a conjunctival swab collected prior to (day 1; blue dots and ellipse) or immediately after (day 8; red dots and ellipse) a 7-day study period, during which the cats were untreated or received orally administered famciclovir at 30 mg/kg or 90 mg/kg, or placebo (lactose) q12h. Data for each day represent the entire study population irrespective of treatment group or disease status. Non-metric multidimensional scaling (NMDS) Axis 1 and Axis 2 represent the axes of the two-dimensional ordination space. Ellipses represent the 95% confidence intervals at each time point
Community membership (Jaccard index) and structure (Yue and Clayton index) by analysis of molecular variance (AMOVA) and UniFrac from 124 conjunctival swabs collected from 81 shelter-housed cats with (68 swabs) or without (56 swabs) clinical evidence of upper respiratory disease (URD) that were untreated or received orally administered famciclovir at one of two doses or placebo (lactose) q12h for 7 days

Swabs were collected prior to (day 1) and immediately after (day 8) the 7-day study period. Data are compared across the entire population and for all clinically relevant subpopulations. The values in bold are significant at P ⩽0.05
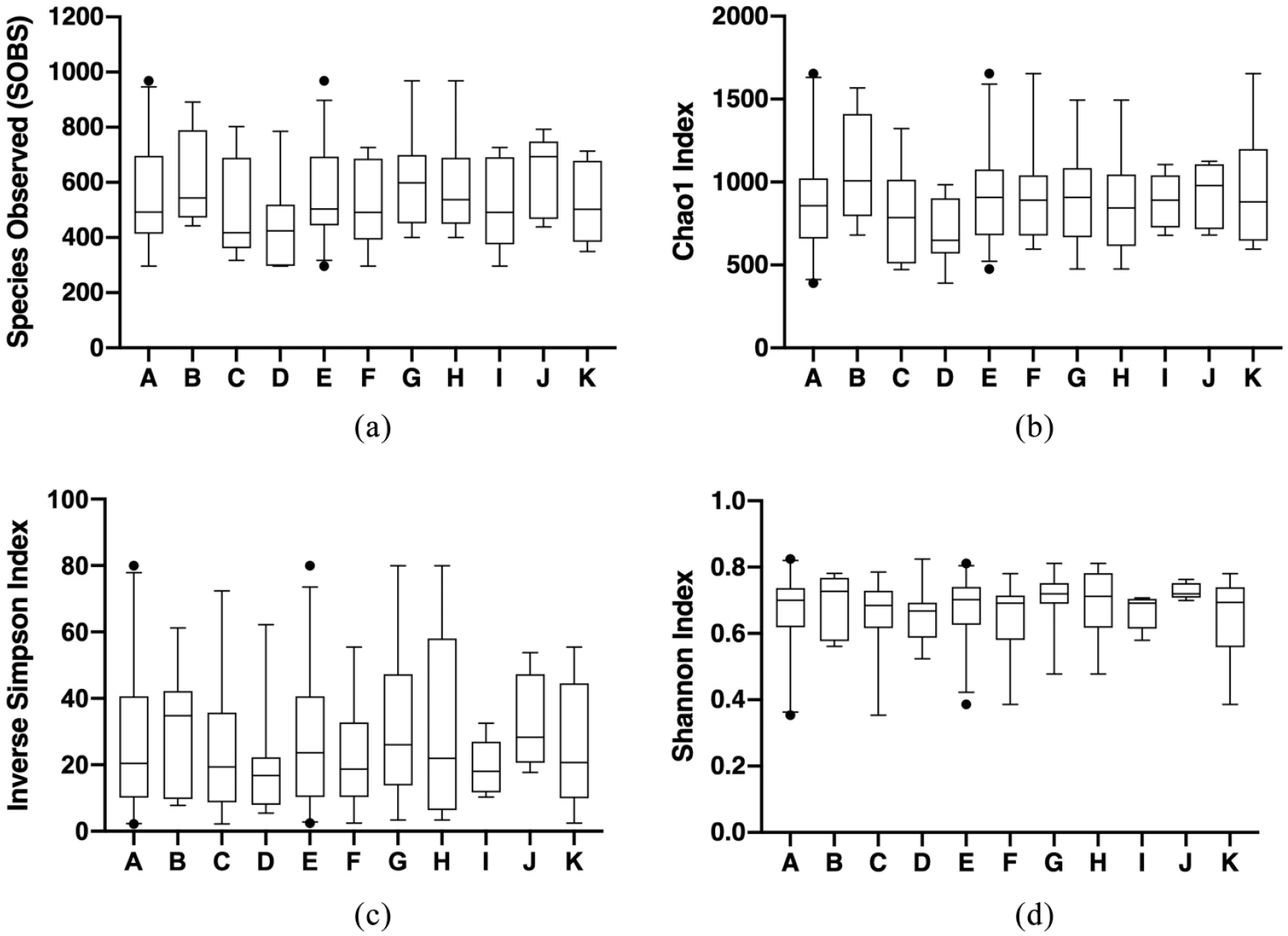
Alpha diversity of the conjunctival microbiota assessed using 50 swabs collected at shelter entry (day 1) from 50 cats with (n = 29) or without (n = 21) clinical evidence of upper respiratory disease (URD). Cats subsequently underwent a 7-day study period during which they were untreated or received orally administered famciclovir at 30 mg/kg or 90 mg/kg or placebo (lactose) q12h, and alpha diversity was assessed again on day 8 (see Figure 7). For data analysis, cats were considered within 11 populations as described in Figure 1: (A) entire population; (B) healthy untreated cats; (C) URD-affected untreated cats; (D) URD-affected placebo-treated cats; (E) healthy and URD-affected famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (F) healthy famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (G) URD-affected famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (H) URD-affected famciclovir-treated (30 mg/kg) cats; (I) healthy famciclovir-treated (30 mg/kg) cats; (J) URD-affected famciclovir-treated (90 mg/kg) cats; and (K) healthy famciclovir-treated (90 mg/kg) cats. (a) Species observed represents observed species richness, (b) Chao1 index represents estimated species richness, (c) inverse Simpson index represents diversity and (d) Shannon index represents species evenness. For each figure, boxes represent the range between the 25th and 75th percentiles, whiskers represent the 2.5th and 97.5th percentiles, the horizontal line inside the boxes defines the median, and closed circles represent outliers

Alpha diversity of the conjunctival microbiota assessed using 75 swabs collected immediately after a 7-day study period initiated at shelter entry, during which cats were untreated or received orally administered famciclovir at 30 mg/kg or 90 mg/kg or placebo (lactose) q12h. Swabs were collected from 75 cats with (n = 35) or without (n = 40) clinical evidence of upper respiratory disease (URD). For data analysis, cats were considered within 11 populations as described in Figure 1: (A) entire population; (B) healthy untreated cats; (C) URD-affected untreated cats; (D) URD-affected placebo-treated cats; (E) healthy and URD-affected famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (F) healthy famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (G) URD-affected famciclovir-treated (30 mg/kg and 90 mg/kg) cats; (H) URD-affected famciclovir-treated (30 mg/kg) cats; (I) healthy famciclovir-treated (30 mg/kg) cats; (J) URD-affected famciclovir-treated (90 mg/kg) cats; and (K) healthy famciclovir-treated (90 mg/kg) cats. (a) Species observed represents observed species richness, (b) Chao1 index represents estimated species richness, (c) inverse Simpson index represents diversity and (d) Shannon index represents species evenness. For each figure, boxes represent the range between the 25th and 75th percentiles, whiskers represent the 2.5th and 97.5th percentiles, the horizontal line inside the boxes defines the median, and closed circles represent outliers
Effect of time in the shelter on the conjunctival microbiota of untreated cats
In URD-affected untreated cats, LEfSe analysis indicated that Escherichia/Shigella was the only OTU over-represented on day 1, whereas Mycoplasma and Streptococcus were over-represented on day 8 (Table 2). There was a significant increase in the median relative abundance of Mycoplasma between day 1 and day 8 (Table 1). The only significant change in beta diversity was in microbial community structure between day 1 and day 8 as measured by AMOVA (Table 3). There were no significant differences between days 1 and 8 in alpha diversity in this population (Figures 6 and 7).
By comparison, in healthy untreated cats, over-represented OTUs at day 1 were Escherichia/Shigella and Bacteroides, and at day 8 were Clostridium cluster XI and Staphylococcus (Table 2). No significant differences in median relative abundance were apparent between days 1 and 8 for any of the common ocular surface pathogens analyzed in this population (Table 1). There were significant differences between days 1 and 8 in microbial community structure (as measured by unweighted UniFrac and AMOVA) and community membership (as compared by unweighted UniFrac; Table 3). There were no significant differences in the alpha diversity measures of observed or estimated richness, diversity or evenness (Figures 6 and 7). Additionally, there were no significant differences in community membership or structure between URD-affected untreated and healthy untreated cats at either day 1 or day 8 (Table 4).
Community membership (Jaccard index) and structure (Yue and Clayton index) by analysis of molecular variance (AMOVA) and UniFrac from 124 conjunctival swabs collected from shelter-housed cats with (68 swabs) or without (56 swabs) clinical evidence of upper respiratory disease (URD) that were untreated or received orally administered famciclovir at one of two doses or placebo (lactose) q12h for 7 days

Swabs were collected prior to (day 1) and immediately after (day 8) the 7-day treatment period. Data are compared between healthy and URD-affected cats for clinically relevant subpopulations at each timepoint
Effect of placebo treatment on the conjunctival microbiota of URD-affected cats
The predominant OTUs in URD-affected placebo-treated cats, based on LEfSe analysis, were Bacteroides, Fusobacterium, Escherichia/Shigella and Gallibacterium at day 1, and Clostridium cluster XI at day 8 (Table 2). No significant differences in median relative abundance of ocular surface pathogens were detected between days 1 and 8 in this population (Table 1). Considering just URD-affected placebo-treated cats, significant differences between days 1 and 8 were detected only in community structure as assessed by AMOVA (Table 3). No significant differences were detected over time in community structure by UniFrac, community membership or alpha diversity. Additionally, on neither day 1 nor day 8 was a significant difference detected in community membership or structure between URD-affected cats that were untreated and those that were placebo-treated (Table 4).
Effect of famciclovir administration on the conjunctival microbiota of healthy and URD-affected cats
The over-represented OTUs for all famciclovir-treated cats considered collectively (regardless of health status or famciclovir dose) were Meiothermus at day 1, and Mycoplasma, Staphylococcus and Clostridium cluster XI at day 8 (Table 2). There was also a significant increase in median relative abundance of Mycoplasma from day 1 to day 8 (Table 1). Median relative abundance did not differ significantly between these two time points for any other ocular surface pathogens. No significant differences were detected between day 1 and day 8 in the beta diversity parameters of community membership and structure (as analyzed by UniFrac and AMOVA; Table 3) or for any measures of alpha diversity in this population (Figures 6 and 7).
LEfSe analysis indicated that the over-represented OTUs for cats that were healthy at shelter entry and treated with either dose of famciclovir were Bacteroides at day 1 and Clostridium cluster XI, Mycoplasma and Staphylococcus at day 8 (Table 2). There was a significant increase in median relative abundance of Mycoplasma between days 1 and 8 (Table 1). There were also significant differences in community structure and membership by AMOVA and UniFrac between day 1 and day 8 for this population (Table 3). No significant changes in measures of alpha diversity, evenness, or richness between days 1 and 8 were apparent (Figures 6 and 7).
The over-represented OTUs for URD-affected cats treated with either dose of famciclovir were Escherichia/Shigella at day 1 and Clostridium cluster XI and Staphylococcus at day 8 (Table 2). There were no significant differences between day 1 and day 8 in the median relative abundance (Table 1) of the ocular surface pathogens assessed for this population. There were significant differences in community structure (as assessed by AMOVA and UniFrac), and in community membership (as assessed by UniFrac) between day 1 and day 8 in this population (Table 3). No significant difference was detected between day 1 and 8 in the alpha diversity of any of the ocular surface pathogens assessed (Figures 6 and 7).
Considering only URD-affected cats receiving 30 mg/kg famciclovir, the over-represented OTUs were Bacteroides at day 1 and Mycoplasma, Clostridium cluster XI and Staphylococcus at day 8 (Table 2). There were no significant differences in the median relative abundance of ocular surface pathogens between day 1 and 8 for this population (Table 1). Regardless of whether the data were analyzed by AMOVA or by UniFrac, no significant differences in community structure or membership were detected between days 1 and 8 (Table 3), or in any measures of alpha diversity (Figures 6 and 7).
In healthy cats treated with 30 mg/kg famciclovir, the over-represented OTUs were Gallibacterium and Pasteurella at day 1 and Staphylococcus and Clostridium cluster XI at day 8 (Table 2). No significant differences were present in the median relative abundance of ocular surface pathogens between days 1 and 8 in this population (Table 1). Community membership only differed significantly between days 1 and 8 by AMOVA in this population. Community structure was not significantly different between days 1 and 8 (Table 3). There were also no significant differences in alpha diversity between day 1 and day 8 (Figures 6 and 7).
For URD-affected cats treated with 90 mg/kg famciclovir, the only over-represented OTU was Fusobacterium at day 1 (Table 2). There were no significant differences detected between day 1 and day 8 in the median relative abundance of any ocular surface pathogens (Table 1). No significant differences in beta diversity, community structure or community membership were observed between days 1 and 8 in this population (Table 3).
Considering only healthy cats receiving 90 mg/kg famciclovir, there were no over-represented OTUs at day 1; however, the over-represented OTUs in this population at day 8 were Bacteroides, Staphylococcus and Clostridium cluster XI (Table 2). There were no significant differences between days 1 and 8 in median relative abundance of ocular surface pathogens (Table 1). Significant differences were apparent in community membership between days 1 and 8 as measured by AMOVA and UniFrac (Table 3). There were no significant changes in community structure between these time points.
Alpha diversity did not significantly differ between day 1 and day 8 for either the healthy or URD-affected population (Figures 6 and 7). Additionally, there were no significant changes in community structure or membership when comparing healthy and URD-affected cats treated with 90 mg/kg famciclovir at either day 1 or day 8 (Table 4).
To evaluate the effect of famciclovir dose, conjunctival microbiota from swabs collected on day 1 were compared between healthy cats treated with 30 mg/kg famciclovir and those treated with 90 mg/kg famciclovir, and between URD-affected cats treated with 30 mg/kg famciclovir and those treated with 90 mg/kg famciclovir. No significant differences in the alpha diversity or beta diversity were detected (Table 5; Figures 6 and 7), and no difference in median relative abundance of the five ocular surface pathogens was detected for any population (Table 6).
Community membership (Jaccard index) and structure (Yue and Clayton index) by analysis of molecular variance (AMOVA) and UniFrac from 69 conjunctival swabs collected from shelter-housed cats with (35 swabs) or without (34 swabs) clinical evidence of upper respiratory disease (URD) that were untreated or received orally administered famciclovir at one of two doses q12h for 7 days

Swabs were collected prior to (day 1) and immediately after (day 8) the 7-day treatment period. Data are compared between famciclovir doses at each timepoint
Median relative abundance of Mycoplasma and Proteobacteria from 69 conjunctival swabs collected from healthy (34 swabs) or upper respiratory disease (URD)-affected (35 swabs) shelter-housed cats receiving 30 mg/kg or 90 mg/kg of orally administered famciclovir q12h for 7 days

Swabs were collected prior to (day 1) and immediately after (day 8) the 7-day treatment period. No significant differences were detected for the remaining three ocular surface pathogens (Chlamydiae, Streptococcus and Pseudomonas)
Discussion
In the present study, we were consistently able to detect bacteria from conjunctival swabs collected from various populations of shelter-housed cats at study entry and 1 week later. This resulted in a large and complex data set, which permitted assessment of alpha diversity, beta diversity and median relative abundance of the conjunctival microbiota as a whole and of phyla or genera containing ⩾1/5 selected feline ocular surface pathogens. The methodology for best interpreting these results remains under development; however, using currently accepted techniques we were able to assess potential effects of time in a shelter environment, famciclovir administration and dose, and URD status on the feline conjunctival microbiota generally, as well as Mycoplasma, Streptococcus and Pseudomonas, Chlamydiae, and Proteobacteria.
Our data suggest that within 1 week in a shelter environment there were changes in community structure and membership of the conjunctival microbiota, with a shift towards over-representation of some ocular surface pathogens. At shelter entry, only Escherichia/Shigella was over-represented; these two phyla are difficult to differentiate by 16S rRNA gene analysis and are therefore reported combined. By day 8, Mycoplasma, Clostridium cluster XI, Staphylococcus and Streptococcus were all over-represented. Simultaneously, changes were also identified in beta diversity measures which were, at least in part, related to alterations in the median relative abundance of Proteobacteria (which was decreased at day 8 vs day 1) and Mycoplasma (which increased from day 1 to day 8). By contrast, differences in the alpha diversity of the conjunctival microbiota during the same period were not detected. This suggests that changes within the conjunctival microbiota of individual cats were relatively minor, with assessment at the population or subpopulation level providing greater insight.
Differences in community membership and structure (beta diversity) between days 1 and 8 were also observed for multiple subpopulations of famciclovir-treated cats, including all cats treated with famciclovir (regardless of URD status or dose), and healthy or URD-affected cats receiving either dose of famciclovir. By contrast, the only subpopulation of untreated cats that experienced changes in both community structure and membership between days 1 and 8 was the healthy untreated group. The similar changes noted in healthy untreated and all famciclovir-treated cats may suggest a direct or indirect beneficial effect of famciclovir on the conjunctival microbiota as has been noted in germ-free mice, where antiviral effects, independent of the microbiota, were observed following topical aminoglycoside administration. 34 Meanwhile, median relative abundance of Mycoplasma also increased in famciclovir-treated cats in the present study when considered as one group regardless of dose or URD status. However, this same change was also seen in URD-affected untreated, but not URD-affected placebo-treated, cats. There is no apparent clinical rationale for the difference between placebo-treated and untreated cats; sampling error cannot be definitively ruled out in this instance. Overall, it is unclear whether famciclovir administration affected the median relative abundance of Mycoplasma, or if this was a reflection of the increase seen in the entire population, especially given that famciclovir-treated cats formed the second-largest subpopulation. However, concurrent changes in community structure and membership of the three largest famciclovir-treated subpopulations may provide some evidence that famciclovir was associated with this change. While our data suggest that a 1-week course of famciclovir affects the feline conjunctival microbiome, no difference due to famciclovir dose was detected. Previous work by our group in the same population of cats revealed that treatment with famciclovir had no significant impact on URD clinical disease scores. 28 It is possible that famciclovir has a minor impact on the beta diversity of the conjunctival microbiota but that this difference is too small to cause clinically distinguishable alterations in disease status.
It is reasonable to assume that cat handling or stress during famciclovir administration may also have affected the microbiome, as shelter-housed cats with higher stress scores were 5.6 times more likely to develop URD than those with lower stress scores. 35 However, our data do not support such a correlation. At neither day 1 nor 8 were there any differences in community membership or structure of URD-affected placebo-treated cats vs URD-affected untreated cats, suggesting that twice-daily handling for oral medication administration did not significantly alter the conjunctival microbiota.
The only subpopulations in which only community structure differed between days 1 and 8 were the URD-affected untreated and the URD-affected placebo-treated cats, while changes in only community membership were detected in healthy cats treated with 30 mg/kg or 90 mg/kg famciclovir. This raises the possibility that famciclovir administration or health status are primarily responsible for these changes. Given that the two doses of famciclovir assessed in the present study did not differentially impact the microbiota, and because URD status was the common factor for cats with altered community membership, it seems more likely that URD status may be responsible for this change. This is similar to data from a study in which the conjunctival microbiota of heathy cats receiving erythromycin for 7 days was stable overall. 36 In humans, altered relative abundance of microbial species, lower bacterial diversity and different ocular surface microbiota have been demonstrated in affected and fellow eyes of patients with fungal keratitis vs healthy eyes. 37 Similarly, human eyes with bacterial conjunctivitis had a higher overall bacterial count but fewer mean number of species cultured than did healthy eyes. 38 By contrast, in the present study a difference was not detected at day 1 or day 8 between the conjunctival microbiotas of healthy vs URD-affected untreated cats, or healthy compared with URD-affected cats treated with 30 mg/kg or 90 mg/kg famciclovir. Therefore, while differences in community structure of URD-affected cats and in community membership of healthy cats are detectable after 1 week in a shelter, these changes may not be large enough to reveal a significant difference between healthy and URD-affected cats. This suggests that the feline conjunctival microbiota was relatively conserved over the entire study population, with small and similar changes occurring in healthy and URD-affected cats. This is consistent with previous data from cats with or without conjunctivitis. 20
Limitations of this study include those inherent to all microbiota analyses, including uncertainty in determining cause-and-effect relationships due to the multivariate nature of host–microbiome interactions. 16S rRNA sequencing has limited ability to characterize sequences to lower taxons, 11 which restricted sequence characterization in the present study to the phylum or genus level. Investigation of feline ocular surface pathogens at the species level would have allowed more in-depth comparison of the conjunctival microbiota among subpopulations. Finally, the cats evaluated in the present study were from a shelter population, which may limit application of these data to client-owned cats maintained in smaller groups; however, shelter populations represent a clinically important group with respect to infectious disease and were selected for this reason. Though environment, treatment and sample collection were standardized following shelter entry, prior history (medical and otherwise) is unknown, as is the effect these factors may have had on the conjunctival microbiota.
Conclusions
Our data reveal that beta diversity of the conjunctival microbiota changes after cats have been housed within a shelter for 1 week. The complexity and number of potential interactions among factors studied here do not permit determination of definitive causal relationships for this change. Although treatment with famciclovir appears to exert an impact on the feline conjunctival microbiota, the dose administered did not alter the beta diversity of the conjunctival microbiota of URD-affected or healthy cats. The absence of change in alpha diversity suggests that changes within the conjunctival microbiota of individual cats are minimal.
Footnotes
Acknowledgements
We thank the Solano County Sheriff’s Office Animal Care Division and Sheriff Thomas A Ferrara for use of their animal adoption facility; IDEXX Laboratories for performance of the qPCR; Monica Motta BS, RVT, RLAT, for technical support; Kate Hurley DVM, and Michael Bannasch BS, RVT, for assistance with project design; and the following undergraduate students for preparation of study medications and data collection: Christina Ball, Connor Chang, Marisa Ciccozzi, Megan Coyne, Nadya Dooley, Alyssa Hoehn, Anny Huang, Winter Keith, Jessica Kwok, Jayden Li, Kira Lin, Kimberly Ly, Ariana Marangakis, Hannah Morris, Huong Nguyen, Josefina Nordenstahl, Allison O’Donnell, Sanskruti Potnis, Mengyue Wang and Isabel Warner.
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by grants from the Morris Animal Foundation and the UC Davis Centre for Companion Animal Health and RM Cello Endowment.
Ethical approval
This work involved the use of non-experimental animals (owned or unowned) and procedures that differed from established internationally recognised high standards (‘best practice’) of veterinary clinical care for the individual patient. The study therefore had ethical approval from an established committee as stated in the manuscript.
Informed consent
Informed consent (either verbal or written) was obtained from the owner or legal custodian of all animal(s) described in this work (either experimental or non-experimental animals) for the procedure(s) undertaken (either prospective or retrospective studies). No animals or humans are identifiable within this publication, and therefore additional informed consent for publication was not required.
