Abstract
Objectives
Polydactyly has been described in two breeds of domestic cats (Maine Coon and Pixie Bob) and in some outbred domestic cats (eg, Hemingway cats). In most cases, feline polydactyly is a non-syndromic preaxial polydactyly. Three variants located in a regulatory sequence involved in limb development, named ZRS (zone of polarising activity regulatory sequence), have been identified to be responsible for feline polydactyly. These variants have been found in outbred domestic cats in the UK (UK1 and UK2 variants) and in Hemingway cats in the USA (Hw variant). The aim of this study was to characterise the genetic features of polydactyly in Maine Coon cats.
Methods
Genotyping assay was used to identify the variant(s) segregating in a cohort of 75 polydactyl and non-polydactyl Maine Coon cats from different breeding lines from Europe, Canada and the USA. The authors performed a segregation analysis to identify the inheritance pattern of polydactyly in this cohort and analysed the population structure.
Results
The Hw allele was identified in a subset of polydactyl cats. Sequencing of two regulatory sequences involved in limb development did not reveal any other variant in polydactyl cats lacking the Hw allele. Additionally, genotype–phenotype and segregation analyses revealed the peculiar inheritance pattern of polydactyly in Maine Coon cats. The population structure analysis demonstrated a genetic distinction between Hw and Hw-free polydactyl cats.
Conclusions and relevance
Polydactyly in Maine Coon cats is inherited as an autosomal dominant trait with incomplete penetrance and variable expressivity, and this trait is characterised by genetic heterogeneity in the Maine Coon breed. Maine Coon breeders should be aware of this situation and adapt their breeding practices accordingly.
Introduction
Polydactyly refers to the occurrence of supernumerary digits and has been described in numerous species of vertebrates.1,2 Among mammals, a few breeds of dogs 3 and cats, and some outbred cats, may exhibit polydactyly. 4 In the Maine Coon breed, some breeding lines segregate for polydactyly. In the Pixie Bob breed, polydactyly is recognised as a morphological trait. Additionally, polydactyly segregates in Hemingway cats, an outbred breeding stock of domestic cats living at Hemingway’s home in Key West (Florida, www.hemingwayhome.com). Finally, this trait has been reported in outbred domestic cats in the USA, in Canada and in Europe. 4
Polydactyly may occur alone (non-syndromic polydactyly) or may be a sign of a developmental syndrome, associated with several congenital abnormalities (syndromic polydactyly). The extra digits may be located on the medial side of the limb (preaxial polydactyly), on the lateral side of the limb (postaxial polydactyly) or in the middle of the limb (central polydactyly). Polydactyly may involve a single limb, two, three or all four limbs.
The polydactyly observed in Maine Coon and Pixie Bob cats, and commonly observed in outbred cats, is a non-syndromic preaxial polydactyly. Cats usually have 18 digits, five on their front paws and four on their hindpaws.
The inheritance of feline non-syndromic preaxial polydactyly has been explored and pedigree data both from outbred domestic cats and Hemingway cats are consistent with an autosomal dominant inheritance pattern.4,5
Three variants have been identified as responsible for feline non-syndromic preaxial polydactyly. They are located in a regulatory sequence of the SHH gene, known to be involved in limb development. 6 This zone of polarising activity regulatory sequence, named ZRS, is located in the intronic sequence of the LMBR1 gene. 6 All three variants have been found in outbred European domestic cats living in the UK (variants named UK1 and UK2) and in Hemingway cats (Hw variant) living in the USA. 4
Here, we report the presence of one of the three ZRS feline variants in a cohort of Maine Coon cats bred in Europe (France, Denmark, Germany), Canada and the USA. We assessed the genotype–phenotype correlation and performed segregation and population structure analyses.
Materials and methods
Animals
Seventy-five Maine Coon cats, five Pixie Bob cats and three outbred domestic shorthair (DSH) cats aged from 3 months to 10 years were included in our study. They were examined in France, Canada and the USA between February 2009 and February 2014. Forty-eight Maine Coon cats, all five Pixie Bob cats and all three outbred DSH cats had extra digits. Twenty-seven Maine Coon cats had 18 digits, as is usually seen in the domestic cat. The polydactyl Maine Coon cats came from six breeding lines: a German line (n = 2), two Danish lines (n = 1, n = 1), a Canadian line (n = 29) and three lines originating from the USA, named US 1 (n = 5), US 2 (n = 3) and US 3 (n = 7).
All cats were included in the study at the owners’ request. Breeders and owners were made aware of the study through a breeder of polydactyl cats and at cat shows.
The DNA profiles from 339 cats were obtained from Antagene or determined using the custom 17 microsatellite marker panel described in the supplementary material. Profiles from Antagene were determined using the International Society for Animal Genetics 2006 panel (https://www.isag.us/committees.asp) for parentage testing purposes. All animals were purebred registered cats. Pedigrees were checked to identify related animals. All directly related cats (parents and littermates) were excluded. Breeds included Persian (n = 15), Exotic Shorthair (n = 17), Birman (n = 32), Maine Coon (43 cats belonging to the total cohort of 75 cats included in this study), Chartreux (n = 32), Norwegian Forest Cat (n = 30), British Shorthair (n = 24), Bengal (n = 18), Siamese (n = 17), Oriental (n = 6), Balinese (n = 1), Ragdoll (n = 24), Abyssinian (n = 21), Somali (n = 3) and Sphynx (n = 56).
Non-invasive buccal swabs were sent directly by owners or collected by a veterinarian (AH, FC, MF, BD, IV, MA or the regular veterinarian of the cat).
Ethics statement
As all the cats were client-owned and no harmful invasive procedures were performed, no animal experimentation according to the legal definition in Europe (Subject 5f of Article 1, Chapter I of the Directive 2010/63/EU of the European Parliament and of the Council) was involved.
Physical and radiographic examinations
Paw phenotype was assessed by physical examination and included visual inspection and palpation by the owner or by a veterinarian (AH, FC, MF, BD, IV or MA, n = 43) or was assessed by radiography as previously described (n = 40). 7
Genealogical data
Pedigree data provided by owners were mapped and drawn using Pedigree Draw 6.0 software (www.pedigree-draw.com).
DNA extraction
DNA was extracted from buccal swabs using a Maxwell 16 Instrument (Promega) and a Maxwell 16 Buccal Swab LEV DNA purification kit (Promega), according to the manufacturer’s protocol.
Sequencing
The pre-ZRS (pZRS) feline sequence was identified by comparing canine pZRS 3 with feline (ENSFCAG00000033659), human (ENSG00000105983) and mouse (ENSMUSG00000010721) LMBR1 intron 5 sequences (www.ensembl.org, feline genome assembly: Felis_catus_9.0) using BLAST 8 and VISTA. 9
PCR and sequencing primers for ZRS and pZRS were designed using the feline LMBR1 intron 5 sequence (ENSFCAG00000033659) and Primer3: 10 ZRSpart1F 5′-ATCGGGGGCAGATGAAATCAC-3′, ZRSpart1R 5′-CACAGGATAGAAACACATGGAACGA-3′, ZRSpart2F 5′-ACTGACCAGGTGGCAGCAAAG-3′, ZRSpart2R 5′-TCCAACAATTTATGGATGATCAGTGG-3′, ZRSpart3F 5′-TTTGTCTCAGGCCTCCGTCTT-3′, ZRSpart3R 5′-GGGGAGGGCAGAGGAATTCTA-3′, pZRSpart1F 5′-GGAGTTTCCTAGTGCACGTTCT-3′, pZRSpart1R 5′-CAGGGGAATCAATGGGAGTA-3′, pZRSpart2F 5′-TCGAGTCATGACAAACTTGAAA-3′, pZRSpart2R 5′-TTACGTTCTCCTACCGTCGAG-3′, pZRSpart3F 5′-GGGTTGACCGTGTCCTTTTA-3′ and pZRSpart3R 5′-CGGAGAGACATGTCCACAAA-3′.
PCR amplicons were Sanger sequenced in both forward and reverse directions by GATC Biotech (Konstanz, Germany). Electropherograms were manually inspected with Chromas Lite (Technelysium). Multiple alignments were performed using Multalin. 11
Genotyping
Genotypes of single nucleotide polymorphisms (SNPs) were determined using Sanger sequencing or a pyrosequencing method adapted from Ahmadian et al 12 on a Biotage PSQ96 pyrosequencer.
PCR amplicons were Sanger sequenced in both forward and reverse directions using ZRSpart1 (UK1 variant) and ZRSpart2 (UK2 and Hw variants) primers by GATC (GATC Biotech). Electropherograms were manually inspected with Chromas Lite.
For the pyrosequencing method, forward and reverse PCR primers for Hw were 5′-BIOTIN-GTCTCAGGCCTCCGTCTTAAA-3′ and 5′-ACCACGCTGGACTTCCTACTC. Forward and reverse PCR primers for UK1 and UK2 were 5′-BIOTIN-ACCAGGTGGCAGCAAAGAGC-3′ and 5′-CGCTCAGCTTTATAGGCCTTCC-3′, and 5′-BIOTIN-GGCCTCCGTCTTAAAGAGACAC-3′ and 5′- TCTGAGACCACGCTGGACTTC-3′, respectively. The sequencing primer for Hw was 5′-TGGACTTCCTACTCATTTC-3′. The sequencing primers for UK1 and UK2 were 5′-TTATAGGCCTTCCCAG-3′ and 5′-GCTGGACTTCCTACTCAT-3′, respectively. In each run, positive and negative control DNA was used, except for UK2 (no UK2 heterozygous DNA was available).
Accession numbers
ZRS and pZRS genomic sequences of Maine Coon cats from the Canadian line (Felis catus) were submitted to GenBank. The accession numbers are [GenBank: KX010805] for ZRS and [GenBank: KX010806] for pZRS.
Statistical analyses
Results were compared by applying a χ2 test. Statistical significance was set at P < 0.05.
Population structure
Cat population structure and the number of clusters (K) in the data set were determined using STRUCTURE 2.3.3 software (https://web.stanford.edu/group/pritchardlab/structure.html). The analysis involved an admixture model with correlated allelic frequencies and included detection of distinct genetic population (K), assignment of individuals to populations and identification of migrant individuals. The appropriate number of inferred clusters to model the data set was selected by using 2–17 inferred clusters. In each Bayesian clustering analysis, 10,000 iterations were carried out after a 10,000 burn-in period, under the admixture model. The best number of clusters fitting the data set was determined by checking the mean LnP(D) over two independent runs.
Results
Variable expressivity of polydactyly
Eighty-three cats were examined, including a cohort of 75 Maine Coon cats (23 polydactyl females, 25 polydactyl males, 17 non-polydactyl females and 10 non-polydactyl males) among which 30 polydactyl cats (13 females, 17 males) and eight non-polydactyl cats (five females, three males) had previously been described. 7 The remaining eight cats included five polydactyl Pixie Bob cats (four females, one male) and three polydactyl outbred DSH cats (one female, two males). For the 56 polydactyl cats, the additional digits were observed during the physical examination. No additional digit was apparent in the 27 non-polydactyl Maine Coon cats on physical (n = 17) or radiographic (n = 10) examination (Figure 1a). The polydactyl phenotype was characterised by broad phenotypic diversity (summarised in Figure 1b–d; see Hamelin et al 7 for more details). The five Pixie Bob cats and the three outbred DSH cats were mitten paw (Figure 1b). Four of them had additional digits only on their forepaws. One outbred DSH cat had extra digits on all four paws (Figure 1b,d).
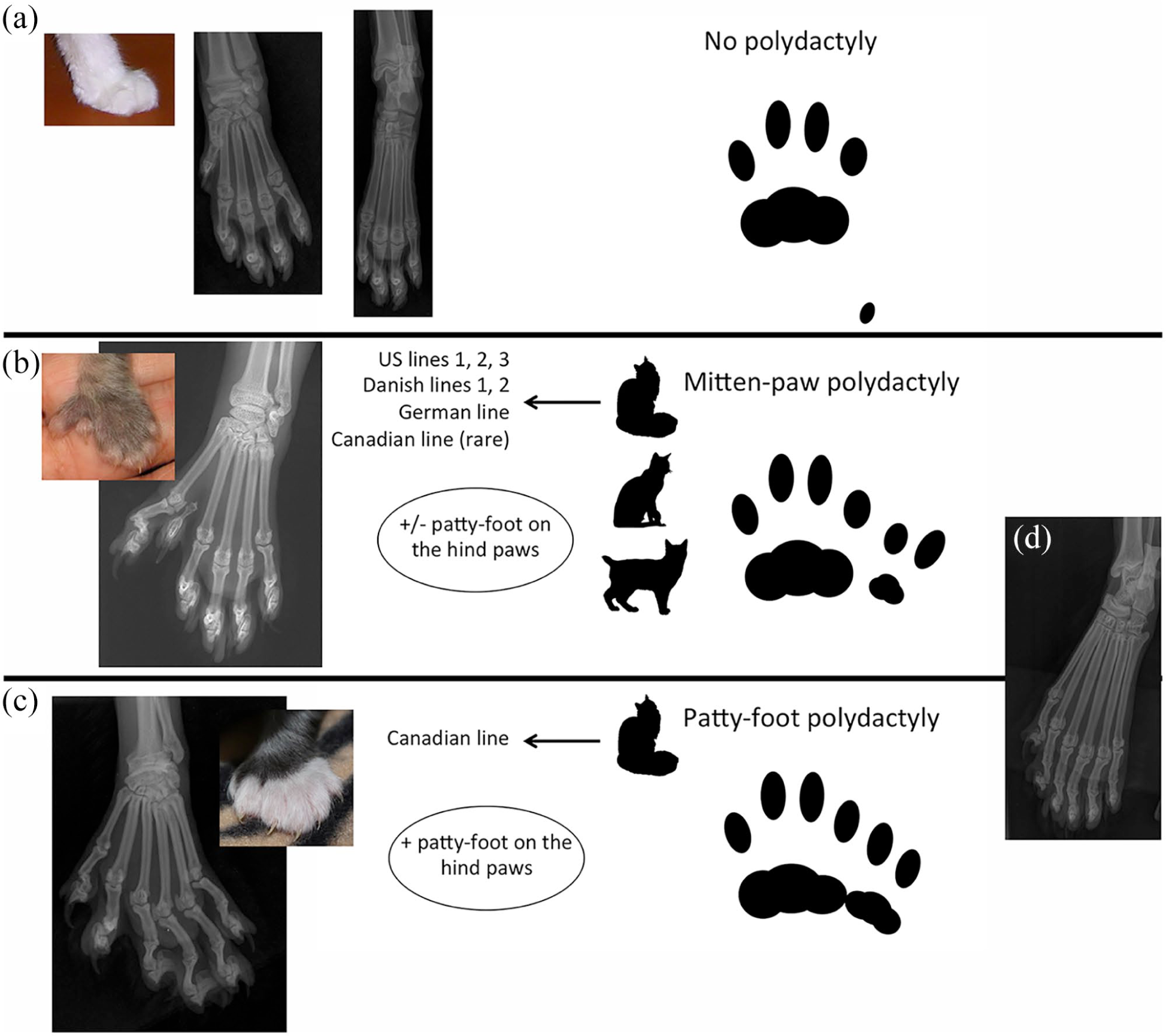
Phenotypic diversity of feline polydactyly. (a) Control cats had 18 digits: five on each front paw and four on each hindpaw. From left to right: left forepaw of a 3-year-old cat; radiographs of left front and hindpaws of a 4-month-old cat; schematic pattern of friction pads on the left forepaw. (b) Mitten-paw polydactyly on forepaws is characterised by the presence of extra digit(s) shorter than the other digits and separate from them (the distance between the extra digit and the first normal digit is greater than the distance between two normal digits). This phenotype was observed in Pixie Bob cats, outbred domestic cats, Maine Coon cats from the US lines, the Danish lines, the German line and, rarely, in Maine Coon cats from the Canadian line (n = 1/29). From left to right: left forepaw of a 10-day-old kitten; radiograph of the left forepaw of a 2-year-old cat; example of schematic pattern of friction pads on a polydactyl left forepaw. (c) Patty-foot polydactyly on forepaws is characterised by the presence of extra digits of the same length and continuous with the normal digits (same distance between extra digits and normal digits and between two extra digits). This phenotype was present in Maine Coon cats belonging to the Canadian line. From left to right: radiograph of the left forepaw of a 4-year-old cat; left forepaw of a 10-day-old kitten; example of schematic pattern of friction pads on a polydactyl left forepaw. (d) Radiograph of a patty-foot polydactyl left hindpaw of a 4-year-old cat
Hw variant in Maine Coon cats
Three variants have previously been described in polydactyl outbred domestic cats 4 bred in the UK (UK1 and UK2 variants) and in the USA (Hw variant). We explored the presence of these variants in our Maine Coon cohort, in five polydactyl Pixie Bob cats and in three polydactyl outbred DSH cats. A total of 83 cats were genotyped for the three variants. One outbred cat (DSH1) was heterozygous for the UK1 variant. The other two outbred cats (DSH2 and DSH3) were heterozygous for the Hw variant. All five polydactyl Pixie Bob cats were heterozygous for the Hw variant. Eighteen of the 48 polydactyl Maine Coon cats were Hw/+ heterozygous, one cat was Hw/Hw homozygous and 29 cats were +/+ wild-type homozygous for the three variants. Among the non-polydactyl Maine Coon cats (n = 27), two cats were Hw/+ heterozygous and 25 cats were +/+ wild-type homozygous for the three variants (Table 1). None of the Maine Coon cats carried the UK1 or UK2 allele. Thus, the Hw allele was associated with polydactyly in a subset of our polydactyl Maine Coon cohort.
Genotypes for the Hw, UK1 and UK2 variants

Genotype–phenotype correlation in Hw cats
Because the Hw variant had previously been reported in outbred domestic cats with a forepaw-restricted polydactyly, 4 we analysed the genotype–phenotype correlation for this variant in our cohort. The Hw/Hw homozygous Maine Coon cat belonged to the US line 2 (see ‘Materials and methods’ section) and was polydactyl only on its hindpaws with two additional digits present on each paw. Among the 18 polydactyl Hw/+ heterozygous Maine Coon cats, polydactyly in 13 cats was restricted to the forepaws. The remaining five Maine Coon cats were polydactyl on all four paws. The five Pixie Bob cats and the two outbred polydactyl Hw/+ cats carried extra digits only on their forepaws. Two non-polydactyl Maine Coon cats were Hw/+ heterozygous. Their phenotype was confirmed by radiography.
In conclusion, the genotype–phenotype correlation for the Hw variant in Maine Coon cats was not strict and the phenotypic expression of polydactyly in cats carrying the Hw allele was broad.
Incomplete penetrance of the Hw allele
We identified two Hw/+ heterozygous non-polydactyl Maine Coon cats. Both cats were isolated breeding stock males living with females in indoor catteries. No other male was present in either cattery. The first cat (male A; Figure 2a) sired two litters. The second cat (male B, Figure 2b) sired three litters. Both cats, mated with non-polydactyl females belonging to non-polydactyl breeding lines (females 1 to 5; Figure 2a,b), produced polydactyl kittens (Figure 2a,b). Seven relatives of the two males were available and we genotyped them for Hw, UK1 and UK2 variants. The six polydactyl relatives were Hw/+ heterozygous, while the non-polydactyl relative was wild-type homozygous for all three variants (Figure 2a,b). Thus, the Hw allele was characterised by an incomplete penetrance in some Hw/+ heterozygous Maine Coon cats.

Partial pedigrees of two Hw/+ non-polydactyl Maine Coon males. Circles represent females, squares represent males. Polydactyl cats are shown in black, non-polydactyl cats are shown in white. When available, genotypes for the Hw variant are shown. (a) Partial pedigree of the first male (A) from US line 3. This male was mated with two females (4 and 5). (b) Partial pedigree of the second male (B) from US line 3. This male was mated with three females (1, 2 and 3). Crossed symbols represent non-Maine Coon cats (Chartreux cats)
Genetic heterogeneity of polydactyly
Of the 48 polydactyl Maine Coon cats genotyped for Hw, UK1 and UK2, 29 cats were wild-type homozygous for the three variants. All of them belonged to the Canadian line. To explore the molecular aetiology of their polydactyly, we sequenced the ZRS locus 4 in four polydactyl Maine Coon cats from the Canadian line, two non-polydactyl wild-type control Maine Coon cats, the three polydactyl outbred cats and a Hw/+ Maine Coon cat from US line 1 (Table 2). Three cats (two polydactyl cats and one control cat) were heterozygous for a single nucleotide polymorphism (SNP) at position 149 in ZRS according to Lettice et al 4 (GenBank: KX010805) and at position 169533175 on chromosome A2 according to Felis_catus_9.0 reference genome sequence (ensembl.org, g.169533175A>G, Table 2). No other variant was found in ZRS.
Sequence variations in ZRS, pZRS or located in the vicinity of pZRS

US1.3, DSH2 and DSH3 cats were Hw/+ heterozygous. DSH1 cat was UK1/+ heterozygous. Positions are shown according to Felis_catus_9.0 genome assembly. Dashes represent non-analysed sequences. For ZRS, nine cats were analysed. For pZRS, two Canadian polydactyl Maine Coon cats, one control Maine Coon cat and one domestic shorthair (DSH1) were analysed
ZRS = zone of polarising activity regulatory sequence; pZRS = pre-ZRS; DSH = domestic shorthair
The feline pZRS sequence, located in LMBR1 intron 5, was obtained by aligning the dog pZRS, known to be involved in canine polydactyly, 3 on feline, human and mouse sequences (GenBank: KX010806; Figure 3a). Sequencing of pZRS (chromosome A2: 169533698 bp to 169534678 bp, reverse strand) in two polydactyl cats from the Canadian line, one control Maine Coon cat and one outbred polydactyl cat did not reveal any variation among these cats (Figure 3b). Three SNPs were identified in the vicinity of pZRS: g.169533780G>C, g.169533798C>T and g.169533800C>T (Figure 3b, Table 2). None of them was specific to polydactyl Maine Coon cats. We concluded that polydactyly in Maine Coon cats was aetiologically heterogeneous. A second locus, distinct from ZRS and pZRS, is likely involved in Maine Coon polydactyly.

Feline pre-zone of polarising activity regulatory sequence (pZRS). (a) Presentation of highly conserved regions between the cat and the dog, human and mouse LMBR1 intron 5, analysed using VISTA (http://genome.lbl.gov/vista). Regions with >50% identity analysed with windows of 100 base pairs (bp) appear on the plot, while those >70% are shaded in red. ZRS and pZRS are conserved between cat, dog, human and mouse. (b) Partial alignment of canine (cLMBR1int5), feline (fLMBR1int5) and human (hLMBR1int5) LMBR1 intron 5 performed with MultAlin, showing the high conservation of pZRS in the three species. Conserved residues are shown with dots. Dashes represent deletions. Canine pZRS (from Park et al3) is surrounded. Primers flanking the amplified and sequenced feline locus (feline chromosome A2: 169533698 bp to 169534678 bp, reverse strand, genome assembly: Felis_catus_9.0) are underlined (pZRSpart1F and pZRSpart3R, see ‘Materials and methods’). Single nucleotide polymorphisms identified in the low-conserved vicinity of pZRS (g.169533800C>T, g.169533798C>T and g.169533780G>C) are shown with stars (Table 2)
Inheritance pattern of polydactyly in Maine Coon cats
Polydactyly in outbred domestic cats was initially described as an autosomal dominant trait with variable expressivity.4,5 To explore the inheritance pattern of the trait in our Maine Coon cohort, pedigree data from polydactyl Maine Coon cats were mapped and drawn (Figures 2 and 4) and segregation analyses were performed (Table 3). Three types of matings were distinguished. The first category of litters that we analysed included at least one non-polydactyl kitten and resulted from mating a polydactyl cat with a non-polydactyl cat (Figures 2a,b and 4a–c). The parental genotypes were inferred to be ‘polydactyl heterozygous’ and ‘wild-type homozygous’, respectively. No statistical difference between the observed and expected classes was detected under the hypothesis of an autosomal dominant inheritance pattern with complete penetrance (Hw-associated polydactyly P = 0.16; Canadian line polydactyly P = 0.78; Table 3). The second category of litters included only polydactyl kittens and resulted from mating a polydactyl cat with a non-polydactyl cat (Figure 4a,c). The parental genotypes were inferred to be ‘polydactyl homozygous’ and ‘wild-type homozygous’, respectively. Two litters with an Hw-associated polydactyly born to the same mother (Figure 4a) were observed (n = 17) in this category. The mother was born to two polydactyl parents. Four litters from the Canadian line were also observed (Figure 4c) in this category. The first litter was born to a polydactyl female and the three other litters were born to a polydactyl male. This female and male were littermates and were born to two polydactyl parents (Figure 4c). A third category of litters born to two non-polydactyl parents and including polydactyl kittens was also identified (Figure 2a,b). Genotyping analysis revealed that in each mating one of the non-polydactyl parents carried the Hw allele.

Pedigrees from polydactyl Maine Coon cats. (a) US line 2. (b) US line 1. (c) Canadian line. Circles represent females, squares represent males. Polydactyl cats are shown in black, non-polydactyl cats are shown in white
Segregation analysis of polydactyly in Maine Coon cats

Data were collected from three families (Figure 4). Two litters with only one kitten were excluded from the analysis
Thus, the inheritance pattern that best fitted our data was autosomal dominant with incomplete penetrance for cats segregating the Hw allele and autosomal dominant with complete penetrance for cats from the Canadian line.
Two subpopulations of Maine Coon cats
We assessed the genetic relationship between cat breeds using STRUCTURE. A range of cluster numbers was examined (K = 2–17). The likelihood LnP(D) increased from K = 2 to K = 11 and stabilised between K = 12 and K = 13. Assuming K = 12, we found a remarkable degree of population substructures (Figure 5). Siamese, Oriental and Balinese cats were clustered in the same group. Persian and Exotic Shorthair cats and Abyssinian and Somali cats were also clustered. The Maine Coon cohort was split into two distinct subpopulations (Figure 5).
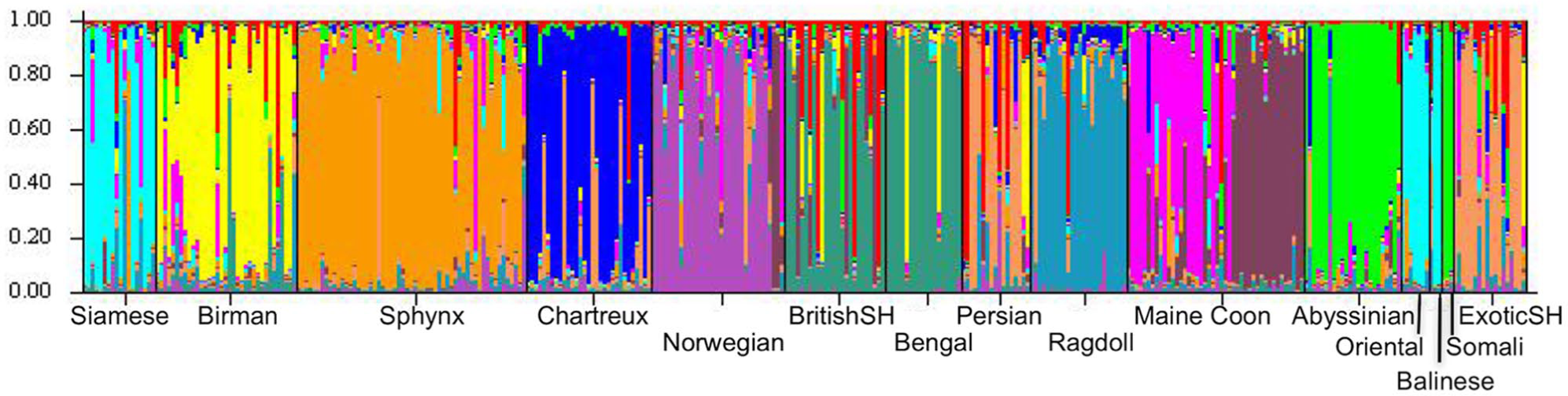
Graphical presentation of genetic population structure for the 339 cats. K = 12. Each cat is represented by a vertical line divided into K colours. The colours correspond to predicted genetic clusters. The y-axis represents the individual’s estimated proportion of membership to a particular cluster. Black lines separate breeds. Cats showed a remarkable population substructure. Siamese, Oriental and Balinese cats were clustered in the same group (in light blue). Persian and Exotic Shorthair cats (in light orange) were clustered; Abyssinian and Somali cats (in light green) were also clustered. Maine Coon cats were split into two distinct subpopulations (in pink and purple). Norwegian = Norwegian Forest Cat; BritishSH = British Shorthair; ExoticSH = Exotic Shorthair.
Discussion
Polydactyly in Maine coon cats is genetically heterogeneous
None of the polydactyl Maine Coon cats from our cohort carried the UK1 or UK2 allele. Only a subset of the cohort carried the Hw allele (n = 19/48). All these 19 polydactyl cats belonged to breeding lines that were founded using American polydactyl cats (the three US lines, the two Danish lines and the German line). The remaining 29/48 polydactyl Maine Coon cats belonged to the Canadian line. Lettice et al genotyped a small number of American polydactyl outbred cats and showed that they all carried the Hw allele, suggesting that this allele was common in polydactyl cats in North America. 4 We confirm this hypothesis here. Indeed, all the polydactyl Maine Coon and Pixie Bob purebred cats from our cohort that were bred in the USA carried the Hw allele. The Hw allele may have spread in these populations thanks to a founder effect. However, we found that polydactyl Maine Coon cats bred in Canada lacked a known polydactyly allele. Finally, we detected the Hw allele in two polydactyl outbred cats sampled in France. One of these cats was born in the USA, the other was born in France. It can be hypothesised that the Hw allele was introgressed from US cats into the French outbred feline population, but, currently, the small number of French polydactyl cats sampled prevents more in-depth analyses.
Polydactyly is an autosomal dominant trait with incomplete penetrance in Hw cats
Studies performed on outbred cats concluded that polydactyly was inherited as an autosomal dominant trait.5,13 Segregation analyses performed in Maine Coon cats are in agreement with an autosomal dominant inheritance pattern, but we also described two litters including polydactyl kittens that were born to two non-polydactyl parents. In both cases the sire was heterozygous for the Hw variant and the non-polydactyl phenotype was confirmed radiographically. These two males were indoor cats living only with females. Thus, the hypothesis of another male being the sire of the polydactyl kittens was ruled out.
The Hw variant was discovered in outbred domestic cats known as Hemingway’s cats. 4 The writer Ernest Hemingway became a famous lover of polydactyl cats, after being offered a six-toed cat by a ship’s captain in the 1930s (www.hemingwayhome.com). The restricted number of cats genotyped by Lettice et al prevented evaluation of the penetrance of the Hw allele. 4 However, large studies aimed at deciphering the inheritance pattern of polydactyly were conducted during the 1940s and 1960s in outbred cats bred in the USA. These studies failed to reveal an incomplete penetrance of the trait.5,13 Here we ascertained that polydactyly is characterised by an incomplete penetrance in cats carrying the Hw allele. Breeders owning polydactyl Maine Coon cats derived from American breeding lines will need to be aware of this finding. By genotyping their breeding stocks they will be able to better manage polydactyly in their catteries.
We found that in Maine Coon cats belonging to the Canadian line, the inheritance pattern that best fitted our pedigree data was autosomal dominant with complete penetrance; however, as long as the molecular aetiology of polydactyly in this line remains undeciphered, it will not be possible to confirm this complete penetrance.
At least two loci underlie polydactyly in Maine Coon cats
Genotyping polydactyl Maine Coon cats for the three Hw, UK1 and UK2 variants revealed that all animals in the Canadian line were wild-type homozygous for all three variants. Sequencing ZRS and pZRS failed to identify the variant underlying polydactyly in these cats. Numerous loci are involved in syndromic polydactyly, but some loci have been identified for non-syndromic polydactyly in mammals.1,14 In humans, they include SHH regulatory sequences and four genes: GLI3, ZNF141, MIPOL1 and PITX1. 15 In dogs, variants in pZRS 3 and in the ALX4 gene 16 have been described. Additionally, engineered mice and chicken embryo models have revealed numerous genes involved in limb patterning.17–19 The strongest candidates for polydactyly in Canadian Maine Coon cats are the SHH regulatory sequences. SHH coordinates limb growth and digit patterning simultaneously. SHH maintains growth from the posterior limb, resulting in a digit pattern that is regulated by the concentration and length of exposure to SHH. At later stages of limb development, SHH regulates expansion of the limb field. The localisation, timing of SHH expression and strength of SHH signalling is tightly controlled to create a localised morphogen source, which is a mandatory step in creating a signalling gradient in order to specify digit identity. When mutated, SHH limb-specific regulatory sequences involved in this control are known to produce polydactyly.3,4,6,20 The whole SHH regulatory domain covers ~1000 kilobases, but limb-specific regulatory sequences have been found in the LMBR1 intronic sequence (ZRS, pZRS) and recently in an intergenic region located ~240 kilobases upstream from the SHH promoter. 20
Two founder effects hypothesised for polydactyl Maine Coon cats
Analysis of the population structure revealed three clusters of breeds (Persian and Exotic Shorthair; Abyssinian and Somali; Siamese, Oriental and Balinese), as previously highlighted and consistent with the history of these breeds.21,22 The analysis also showed that our cohort of Maine Coon cats split into two distinct clusters. On consulting the pedigrees, we found that the first cluster was composed of animals originating from the USA and German lines (cats from the Danish lines were not included in the analysis) and from classical non-polydactyl Maine Coon lines. The second cluster was made up of cats belonging to the Canadian line.
These two distinct clusters suggest that polydactyl cats from the same breed are genetically divergent. US polydactyl cats are genetically similar to classical Maine Coon cats, which are known to have originated from farm feral cats living in several US states, including Maine (www.pawpeds.com). Polydactyl feral cats may have been included in the initial foundation stock. They may also have participated in the creation of the Hemingway outbred stock. However, a recent introduction of the Hw allele in classical Maine Coon lines cannot be excluded owing to the permitted use of foundation cats in some US stud books (ie, US outbred cats appearing as Maine Coon cats). Hemingway cats and US bred Maine Coon cats with recent foundation ancestors need to be sampled to confirm one of these hypotheses. Finally, the Canadian line seems to be a separate Maine Coon line in which polydactyly may have arisen through to a second founder effect.
Conclusions
Polydactyly in Maine Coon cats is inherited as an autosomal dominant trait with incomplete penetrance and variable expressivity. Some polydactyl Maine Coon cats carry an allele previously described in Hemingway cats, but Maine Coon cats from a Canadian line carry an allele that remains unidentified to date. Thus, polydactyly in the Maine Coon breed is characterised by genetic heterogeneity.
Maine Coon breeders should be aware of this peculiar genetic situation when dealing with polydactyly in their catteries.
Supplemental Material
Supplementary Material
Custom microsatellite marker panel
Footnotes
Acknowledgements
The authors wish to thank the practitioners who supplied the radiographs; the radiology technicians from the Alfort, Nantes and Toulouse veterinary schools; Maine Coon breeders and owners who contributed, especially Maryline Compagnon from the Eole Coon cattery; and Diana Warwick, who edited the manuscript.
Supplementary material
The following file is available online: Custom microsatellite marker panel
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. Antagene is a for-profit company selling DNA tests for animals.
Funding
This work was supported by the French Felinotechnic Society (Société Française de Félinotechnie). Feline DNA samples are part of the Feli-DNA biobank, which forms part of the CRB-Anim infrastructure (ANR-11-INBS-0003, within the framework of the ‘Investing for the Future’ programme [PIA]).
Ethical approval
This work involved the use of non-experimental animals only (including owned or unowned animals and data from prospective or retrospective studies). Established internationally recognised high standards (‘best practice’) of individual veterinary clinical patient care were followed. Ethical approval from a committee was therefore not necessarily required.
Informed consent
Informed consent (either verbal or written) was obtained from the owner or legal custodian of all animal(s) described in this work (either experimental or non-experimental animals) for the procedure(s) undertaken (either prospective or retrospective studies). No animals or humans are identifiable within this publication, and therefore additional informed consent for publication was not required.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
