Abstract
Inbred mouse strains have long proved useful as tools for biomedical research. They remove the effects of genetic background as an experimental variable. Within all mouse colonies, genetic drift is a recognised phenomenon and monitoring and documenting changes is important for experimental design and consistency. This communication documents the initial characterisation through SNP analysis of the inbred mouse strains bred and used at the time at the Medical Research Council National Institute for Medical Research (MRC-NIMR), Mill Hill, now The Crick Institute. These inbred strains were part of the foundation colonies for the many genetically modified mouse strains made at Mill Hill. We found small genetic changes in four of the nine inbred strains. Although phenotypic differences have not yet been found between the NIMR and the correspondent parental strains, I cannot discard that these may arise or have already arisen. This work has also authenticated the 129/SvJEvNimr-Gpi1c strain that was widely used at MRC-NIMR for gene targeting. All these inbred strains have been renamed according to The International Committee on Standardized Genetic Nomenclature for Mice.
The Medical Research Council National Institute for Medical Research (MRC-NIMR), Mill Hill was one of the world’s leading medical research institutes. Research at MRC-NIMR covered a broad spectrum of basic biomedical science and was underpinned by small animal facilities. Since the late 1950s until its closure in 2015, MRC-NIMR had maintained a rodent breeding unit managed as a closed colony, which provided inbred and outbred animals for all scientists working at the institute and collaborators around the globe. Inbred mouse strains bred at MRC-NIMR included A/J, C57BL/6J, C57BL/10ScSnJOla, 129/SvJEv, 129P2/OlaHsd, BALB/cJ, CBA/CaH, C3H/HeJ and FVB/NHsd. Work with these inbred strains had been and is still being published in leading scientific journals.1 –9
While microbiological monitoring is a well-established practice, the same level of familiarity often does not extend to genetic monitoring. As a result, the production of a genetic monitoring report for animals being shipped is a less common occurrence. However, many of us find assurance in the correct genetic identification and naming of animals, adhering to the Guidelines for Nomenclature of Mouse and Rat Strains established by the International Committee on Standardized Genetic Nomenclature for Mice and the Rat Genome and Nomenclature Committee.10 –12 Some of the methods employed for genetic identification are purely phenotypic and can still be effectively used, provided proper records are maintained at the colony of origin.
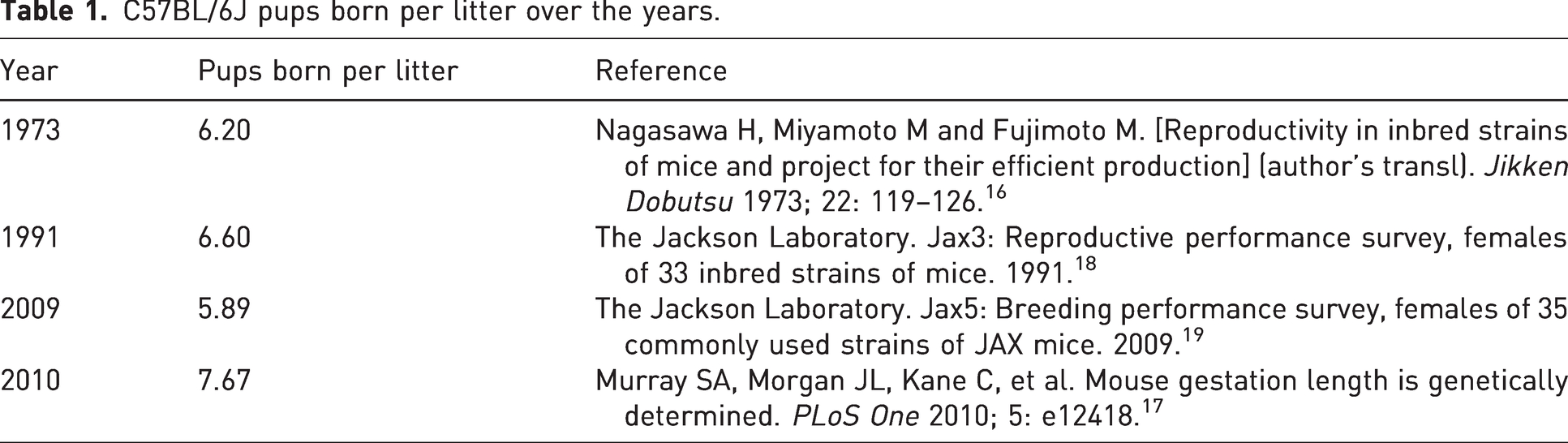
A rodent inbred strain is defined as one that has been mated brother to sister for 20 or more consecutive generations, and individuals of the strain can be traced to a single ancestral pair at the 20th or subsequent generation.13,14 Inbreeding can also be achieved by a sequence of mating offspring to the younger parent. 15 As more inbred strains were developed, a greater appreciation for the variability between individual strains became recognised. A fully inbred strain represents a unique combination of largely homozygous alleles that are responsible for the characteristics of that strain. Thus, the genetic identity of these inbred animals needed to be defined and authenticated. Different methods have been employed in the past to verify the authenticity of various isogenic strains of laboratory mice and, to some extent, laboratory rats. Early on, careful breeding practices and visible characteristics were the only available tools to define and maintain genetic purity. Examples include physical characteristics such as coat colour, the frequency of bent tails, the occurrence of microphthalmia or hydrocephalus, and strain-specific behavioural or neurological traits. Another easily implemented check for the integrity of these isogenic animals involves strain performance monitoring. This approach examines various parameters typically assessed for the strain at the facility and can detect deviations from the norm. One illustrative example is monitoring reproductive parameters, which can be easily recorded and analysed for deviations from the standard for the isogenic strain. However, caution is warranted when drawing conclusions based on these parameters, as they may fluctuate over time. To illustrate this issue, I have pulled out data on C57BL/6J number of pups born per litter over the years, and present them in Table 1.16 –19
C57BL/6J pups born per litter over the years.

In 1974, a first attempt to genetically authenticate inbred strains was done by the Medical Research Council by launching an accreditation scheme for genetic monitoring. 20 This genetic monitoring scheme consisted of a binding series of rules plus a genetic analysis consisting of different mandible measures and its statistical analysis. 21 Michael Festing established this method in 1972.22 –28 Other osteometric traits had also been employed to ascertain the purity of these animals. 29
Biochemical marker genes were also used for the authentication and introduced in 1971.21,30 –32 At the Jackson Laboratory, their quality control programme used a set of different biochemical markers (isoenzymes) and proteins. They employed four to five biochemical markers and the coat colour to distinguish most common mouse inbred strains, and 10 markers for a unique profile for each of the inbred strains. Nevertheless, biochemical markers were not able to discriminate between close strains or substrains.33,34
Immunology has also played a pivotal role in ensuring the genetic quality control of mice, particularly due to immunologists’ vested interest in maintaining the purity of mouse lines. In the 1960s and early 1970s, the technique of skin grafting emerged as a means of genetic quality control in mice.26,35 Initially established in mammals during the 1950s by Billingham and Medawar, this technique encountered initial technical challenges in its application and interpretation in mice.36,37 Subsequently, Festing introduced the utilisation of strain-specific polyvalent alloantisera in combination with a dye-exclusion microcytotoxic test, offering a method for routine genetic quality control. 38
Later advancements included the development of Strain Restricted Typing Sera (SRTS), an immunological technique exploiting antigenic disparities in white blood cells among different strains of mice and rats. By generating antisera specific to the major histocompatibility complex of these strains (H-2, RLA), SRTS evaluates the compatibility of animals presumed to belong to the same strain. 39 Widely employed in the 1980s, this technique proved instrumental in verifying the integrity of congenic lines and found application in a lineage of minipigs homozygous for the major histocompatibility complex (MHC genotypes, Swine Lymphocyte Antigen, SLA).39 –43
None of these methods were completely satisfactory for genetic authentication. DNA based methods for genetic monitoring such as DNA finger printing or Simple Sequence Length Polymorphisms (SSLPs) were introduced in the early 1990s as these molecular techniques became more user friendly.44,45 However, it was the microsatellite, also known as the simple sequence repeat (SSR), that revolutionised the way we characterise these animals. Since 1990, Todd’s group in Oxford led the pioneering use of microsatellites for genetic analysis in mice.46 –50 Microsatellites consist of one, two, three or four nucleotide repeats that are specific in length for each inbred strain. These repeats are distributed within the mouse genome at a density of up to one every centiMorgan (cM). The advent of the polymerase chain reaction allowed for the efficient amplification of these microsatellites. 51 By 1994, a map of more than 4000 SSRs had been constructed, with 90% of the genome lying within 1.1 cM of a marker. There was an average polymorphism rate of 50% between laboratory strains. 52
More recently, single nucleotide polymorphisms (SNPs) have emerged as a valuable tool for genetic monitoring, offering a comprehensive scan of the entire mouse genome.53 –57 The feasibility of using SNPs for genetic analysis of mouse strains was realised in the early 21st century, following the publication of a collection of 2848 SNPs by the Eric Lander group. 58 By 2005, a panel of 673 SNPs was developed for genetic analysis of mice, with applications ranging from the generation of congenic mice to positional cloning and mapping quantitative or qualitative trait loci. 59 Within just a year, a panel of 10,000 SNPs evenly distributed across the mouse genome facilitated the construction of genetic maps for both outbred and inbred mice, and separately for males and females. 60
Expanding beyond mice, advancements in SNP genotyping also extended to rats. By 2008, knowledge of approximately three million SNPs in the rat genome enabled the genotyping of 167 rat inbred strains using a 20,238 SNP panel developed by the STAR Consortium. 61 Subsequent endeavours included the development of the Mouse Diversity Array in 2009 by The Jackson Laboratory in collaboration with Pardo-Manuel de Villena’s group at the University of North Carolina. This array boasted over 600,000 SNPs, capturing all known genetic variations in the laboratory mouse. 62
Continuing the trajectory of SNP array development, the Mouse Universal Genotyping Array (MUGA) was introduced with 7851 SNPs initially, designed as a cost-effective genotyping platform on the Illumina platform. 63 Its successor, the MegaMUGA, featured 77,808 SNPs, representing a 10-fold increase in marker density and optimized for discriminating haplotypes. 64 The evolution culminated in the third generation of the MUGA family, the GigaMUGA, with an even higher marker density of 143,259 SNPs. 57
Here, I report the genetic analysis of the inbred mouse strains bred at the MRC-NIMR using a 768 SNP panel. I had found that four out of nine inbred strains have accumulated changes in their genome during their time at MRC-NIMR.
Materials and methods
Ethical considerations
Legal and ethical requirements have been met with regard to the humane treatment of animals.
Animals
A, C57BL/6, C57BL/10ScSn, 129/SvEv-Gpi1c, 129P2/Ola, BALB/c, CBA/CaH, C3H/He and FVB/N mouse inbred strains were bred in-house and kept according to the UK Animals (Scientific Procedures) Act 1986 and following the Home Office Code of Practice.65
–67 The origins of these strains are as follows:
A/J – initially acquired from The Jackson Laboratory and subsequently transferred to Allington Farm, Porton Down, before being transferred to MRC-NIMR in 1963; C57BL/6J – obtained directly from The Jackson Laboratory in 1986; C57BL/10ScSnJ – originally sourced from The Jackson Laboratory (as C57BL/10ScSnJ) and subsequently transferred to the Clinical Research Centre (CRC) in 1970, and from CRC transferred to the Oxford Laboratory Animal Centre (OLAC) in 1976, and from OLAC transferred to the Laboratory Animals Centre (LAC) at Carshalton in 1976, and from there ultimately to MRC-NIMR in 1984; 129/SvJEv-Gpi1c – initially obtained from The Jackson Laboratory (as 129/SvJ) and later modified by Martin Evans at Cambridge, where the Gpi1c allele from a 101 strain background was introduced. The strain was then transferred to the MRC Mammalian Development Unit (MDU) and subsequently to MRC-NIMR in 1989; 129P2/OlaHsd – acquired from Harlan in 1998; BALB/cJ – originating from the National Institutes of Health, Bethesda, in 1959;
68
CBA/CaH – initially sourced from Harwell and subsequently transferred to the University of Edinburgh under the laboratory of NA Mitchison in 1954. It was then relocated to MRC-NIMR in 1961 (F? + 25); C3H/HeJ – obtained directly from The Jackson Laboratory in 1982; FVB/NHsd – acquired from Harlan Olac in 1994.
All of these strains were bred and maintained at the highest-level barrier facility dedicated to breeding at MRC-NIMR after their arrival. No additional mice from the same strains were introduced afterward.
Our breeding programme followed the guidelines outlined in the UFAW handbook. 69 Briefly, continuous brother–sister breeding pairs were established at 6–8 weeks of age. Breeding pairs were typically replaced every six months. Selection of future breeders was primarily done from animals born in the second to fifth litter, although occasionally new breeders were chosen from the first or sixth litter, albeit infrequently.
DNA extraction
Animals were euthanized according to the Schedule 1 of the Animals (Scientific Procedures) Act 1986. 65 DNA was extracted from tails using PureGene Tissue kit (Gentra systems, Minnesota, USA) according to the manufacturer’s instructions. The DNA was dissolved in 50 µl of hydration buffer (DNA hydration solution, Gentra systems, Minnesota, USA) and stored at –70°C. DNA was quantified using Nanodrop (Labtech International Ltd, East Sussex, UK).
SNP analysis
Samples from five randomized animals (using Excel Rand function) from the nucleus colony of each of nine inbred strains of mice bred at MRC-NIMR were sent for SNP analysis. Genomic DNA from referenced inbred strains C57BL/10ScSnJ (stock 476, preparation P32983, The Jackson Laboratory, Bar Harbor, Maine, USA) and CBA/CaJ (stock 654, preparation P37635, The Jackson Laboratory, Bar Harbor, Maine, USA) was also sent for SNP analysis. For each animal, 4 µg of DNA at 100 ng/µl were sent to the Genetics Division at Brigham and Women’s Hospital (Harvard Medical School, Boston, Massachusetts, USA) for a 768 SNP panel analysis using the Illumina multiplexed allele extension and ligation method (Golden Gate) (Illumina, San Diego, California, USA). 53
Results
Inbreeding generations at MRC-NIMR
We re-started our generations’ records from October 1996 as previous maintained records were archived and lost during the refurbishment of our breeding unit. Table 2 shows the number of generations for each of the inbred strains until May 2008. The 129P2/OlaHsd strain was frozen down in May 2007.
MRC-NIMR strains brother x sister inbreeding generations.
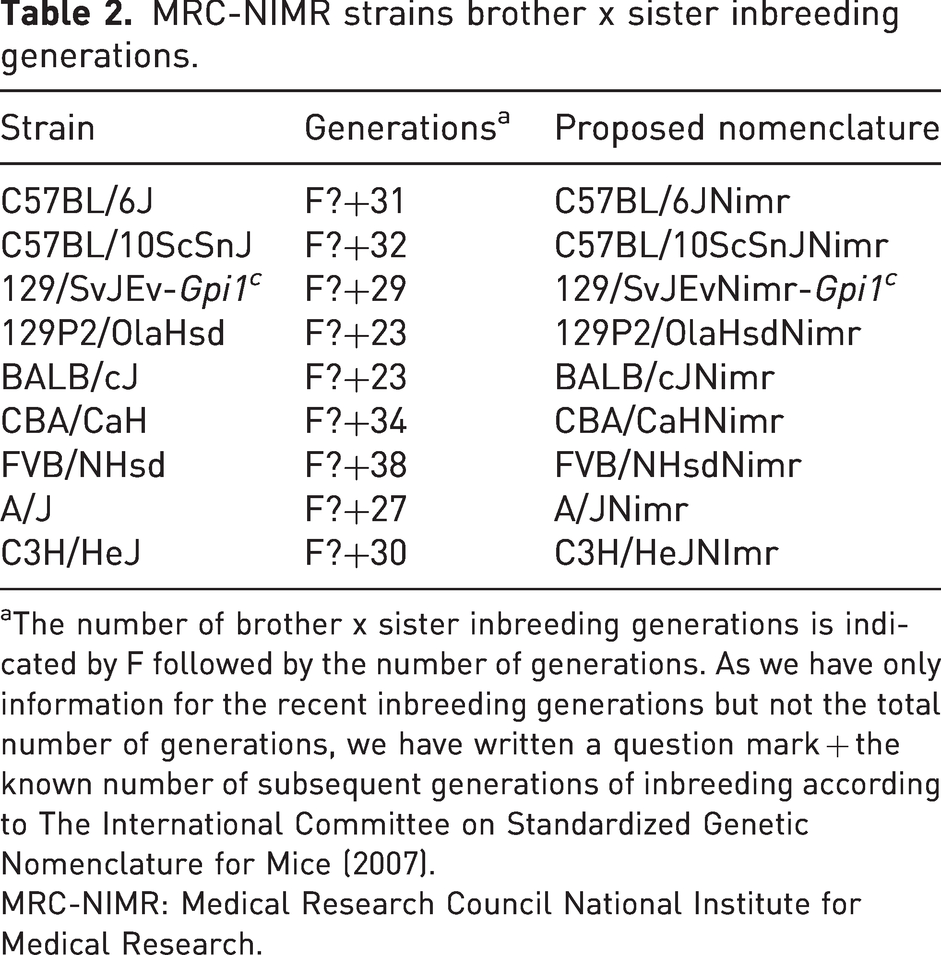
The number of brother x sister inbreeding generations is indicated by F followed by the number of generations. As we have only information for the recent inbreeding generations but not the total number of generations, we have written a question mark + the known number of subsequent generations of inbreeding according to The International Committee on Standardized Genetic Nomenclature for Mice (2007).
MRC-NIMR: Medical Research Council National Institute for Medical Research.
SNP analysis
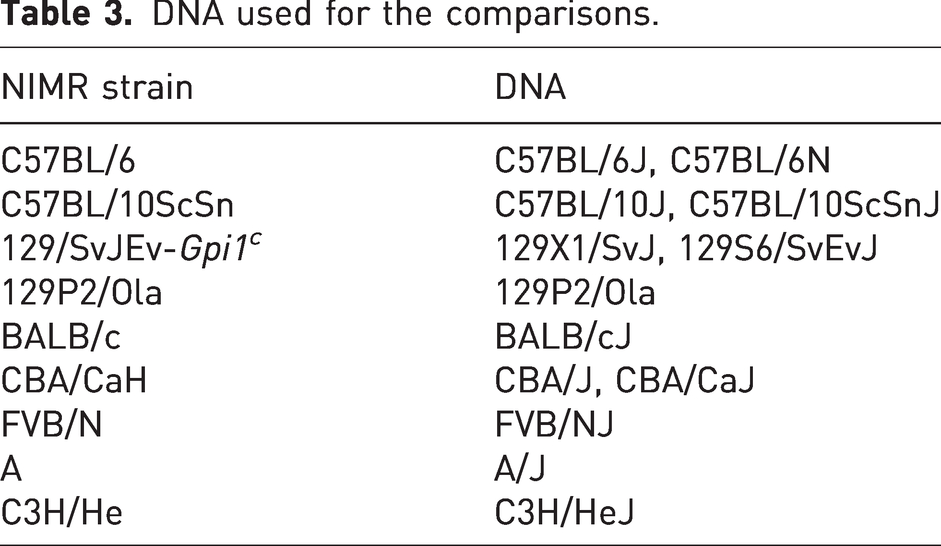
Table 3 shows the strains with which NIMR strains were compared. Table 4 shows the SNP differences found.
DNA used for the comparisons.
MRC-NIMR SNP differences with strain of provenance.
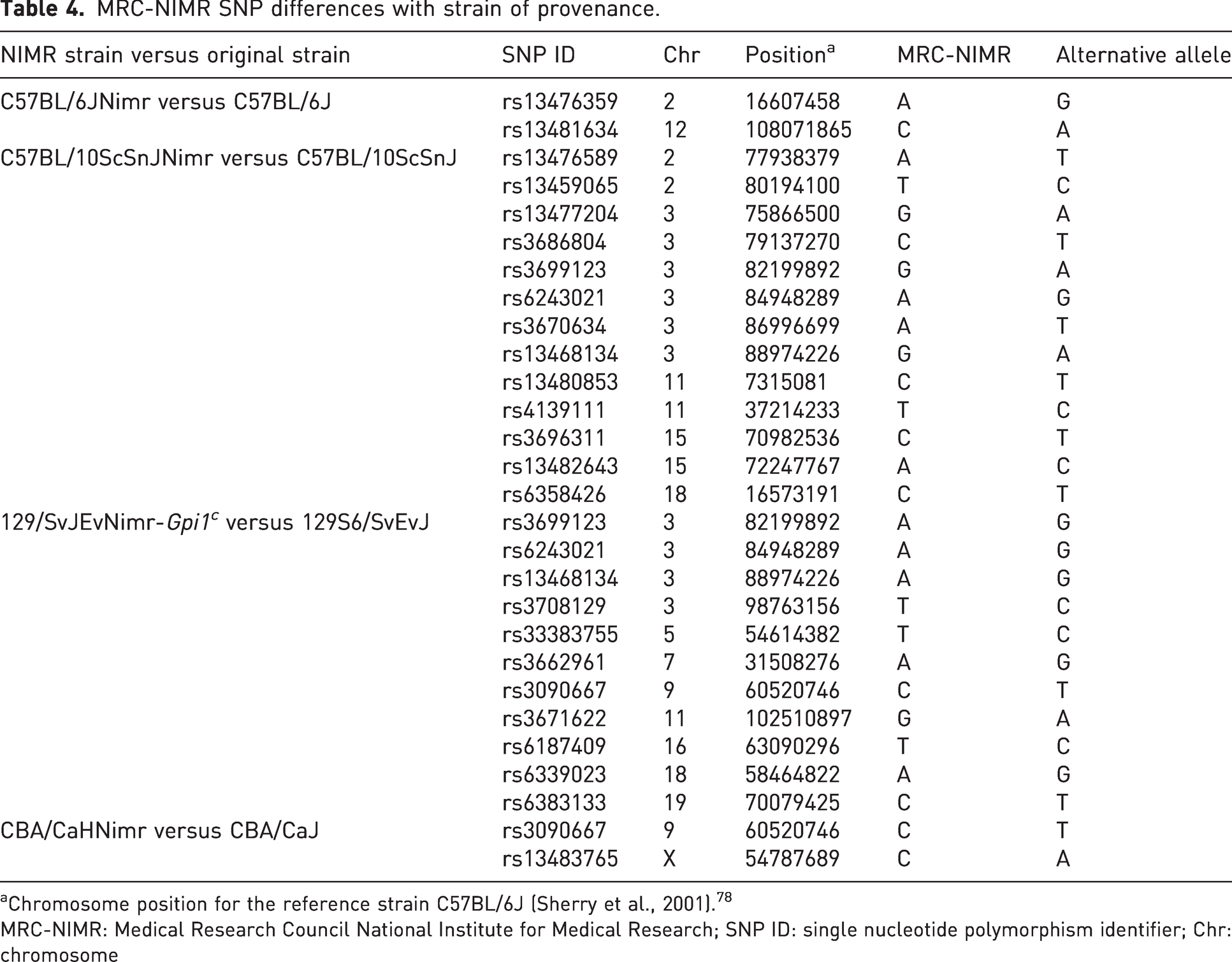
Chromosome position for the reference strain C57BL/6J (Sherry et al., 2001). 78
MRC-NIMR: Medical Research Council National Institute for Medical Research; SNP ID: single nucleotide polymorphism identifier; Chr: chromosome
None of the768 SNPs analysed for the MRC-NIMR stocks of A, 129P2/Ola, BALB/c, C3H/He and FVB/N differed from the original strain maintained at The Jackson Laboratory. Even though no difference was found I have named the inbred strains according to the Rules and Guidelines for Nomenclature of Mouse and Rat Strains as A/JNimr, 129P2/OlaHsdNimr, BALB/cJNimr, C3H/HeJNimr and FVB/NHsdNimr. 14
Our C57BL/6 differs in just two SNPs of the 768 from the original strain at The Jackson Laboratory even though the strain has been bred at MRC-NIMR as a closed colony for more than 20 years. Accordingly, it has been named as C57BL/6JNimr. In contrast, the C57BL/10ScSn MRC-NIMR strain differs in 13 SNPs from the parental strain C57BL/10ScSnJ. We found 11 SNP differences between the 129/SvJEv-Gpi1c maintained at MRC-NIMR and the 129S6/SvEvJ strain maintained at The Jackson Laboratory. I also checked this strain against the 129X1/SvJ and found 48 SNP differences. The CBA/CaH strain bred at MRC-NIMR had two SNP differences when compared with CBA/CaJ, and 78 SNP differences compared with CBA/J.
Discussion
The origin of inbred mouse strains can be traced back to Miss Abbie EC Lathrop. She was a retired schoolteacher living near to the Bussey Institute directed by William E Castle. Miss Lathrop began, around 1900, to breed mice for sale as pets from her home in Granby, Massachusetts.13,70 It was in 1909 when Clarence Cook Little, studying the inheritance of coat colour in mice under William E Castle’s supervision, started the inbreeding of a pair of mice carrying the recessive genes for dilution, brown and non-agouti, which produced the first inbred strain of mice, DBA. In 1921, the A strain was started by Leonell C Strong from a cross between a mouse from Halsey J Bagg albino stock and Little albino stock at Cold Spring Harbor. The A strain arrived at The Jackson Laboratory in 1947 and in 1963 at MRC-NIMR. Also in 1921, Little obtained mice from Lathrop and mated litter mates female 57 with male 52, progeny from which segregated as brown and black. The black progeny were the foundation for the C57BL strain, with substrains 6 and 10 being separated prior to 1937. C57BL/6J has now more than 150 generations of inbreeding whereas C57BL/10J has more than 158 and C57BL/10ScSnJ has more than 136. The C57BL/6 and C57BL/10ScSn strains came to MRC-NIMR from The Jackson Laboratory in 1986 and from LAC at Carshalton in 1984, respectively. The 129 strain originated from crosses established by LC Dunn in 1928 between coat colour stocks obtained from English fanciers and a chinchilla stock obtained from William E Castle.71,72 Strain 129/SvJ was genetically contaminated in about 1978 by an unknown strain, and differs from other 129 substrains at about 25% of SSLP genetic markers. 72 Martin Evans backcrossed the Gpi1c marker (from the 101 strain) into 129/SvJ to create the 129/SvEv-Gpi1c strain; 73 MRC-NIMR obtained this strain in 1989 from MRC MDU. At Cold Spring Harbor, E Carleton MacDowell inbred the Bagg albinos, from which George D Snell had some around 1932. Snell used the letter ‘c’ in his laboratory records to indicate that animals were white. The letter became attached to ‘Bagg alb’ and the designation evolved to BALB/c.70,71 BALB/cJ has been inbred for more than 150 generations. The BALB/cJ strain came to MRC-NIMR in 1959 from the National Institues of Health (NIH). 68 In 1920, Leonell C Strong started the CBA and C3H strains from a cross of a Halsey J Bagg albino stock female and a DBA male. Strain CBA was selected for low mammary tumour incidence and C3H for high incidence.70,71 Our CBA/CaH strain came in 1961 from Harwell. The C3H/He strain came in 1982 from The Jackson Laboratory and it has more than 130 inbreeding generations. The FVB strain originated from a colony of Swiss mice at NIH. Animals from this colony were selected for their sensitivity or resistance to histamine diphosphate in a vaccine challenge and some of the sensitive animals were discovered later to carry a mutation (Fv-1b), which makes them susceptible to infection by the B strain of Friend leukaemia virus. These animals were designated FVB for Friend virus B-type susceptibility. 74 This strain was brought to the MRC-NIMR colony in 1994 from Harlan.
I would have expected to find at least some differences with the original strains in A/JNimr, BALB/cJNimr and C3H/HeJNimr as they have been bred separated from the colonies of origin for a very long time at MRC-NIMR. The fact that I have not seen differences in the 768 SNPs analysed does not mean they do not differ from the original strain. More work must be done on different SNPs to document any potential differences. In the case of FVB/NHsdNimr, even though the strain has been bred for more than 38 generations we again could not find differences in the SNPs analysed.
Another strain without differences from the parental strains is 129P2/OlaHsd, which has been archived and is no longer used at MRC-NIMR.
The C57BL/6JNimr and the CBA/CaHNimr have been documented as differing in two SNPs each from the parental strain.
The 129/SvJEvNimr-Gpi1c strain is a congenic strain that must have residual sequences from the 101 strain, which served as donor strain for the glucose phosphate isomerise 1 gene variant c (Gpi1c). I would have liked to check these differences against the 101 strain but the Jackson does not keep DNA or animals from this strain. The 129/SvJEvNimr-Gpi1c strain was not characterised in the original article describing the 129 substrain genetic variation. Thus, this work authenticates this strain that was widely used at MRC-NIMR for gene targeting.
The SNP panel used in this work does not allow distinguishing between C57BL/10ScSnJ and C57BL/10J substrains. I have named the strain as C57BL/10ScSnJNimr, but more works must be done on different SNPs to document any differences.
It is also worth noting that the number of breeding pairs varied between the strains and over the years. This variation could be one reason why SNP differences were observed in some but not all of the inbred strains. However, I cannot rule out that the absence of differences in some analysed strains might be due to the relatively low number of SNPs analysed at the time.
Although some groups have claimed that there could be a direct correlation between strain phenotype and SNP variation, so far SNP differences between substrains have not been linked to phenotype differences. 75 Nevertheless, there are examples of spontaneous single gene mutations that affect the phenotype.76,77 This fact alone justifies the authentication of substrains used for research. With the increase in number of SNPs analysed it will be easier to point out to specific regions in the genome for analysis in the case of uncovering of phenotypic differences.
Phenotypic differences have not been found between the NIMR and the correspondent parental strains. Nevertheless, scientists using these MRC-NIMR strains should be aware that they have already started to diverge.
Footnotes
Acknowledgements
We would like to thank Dr Paul S Burgoyne and Dr Kathleen Mathers for critical reading of the manuscript.
Data availability
All data presented in this manuscript is available as a supplementary Excel file or upon request.
Declaration of competing interests
The authors have no conflicts of interest to declare.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the MRC-National Institute for Medical Research, the Mutation Mapping and Developmental Analysis Project (MMDAP) NICHD (grant no. U01 HD43430) and the National Center for Research Resources (grant U54 RR020278). MRC-National Institute for Medical Research provided the departamental budget to carry out the SNP analysis but this finance was not part of a grant.
