Abstract
Neonicotinoid insecticides, known for their selectivity and low mammalian toxicity, have been widely used in recent years as alternatives to organophosphate insecticides. Although neonicotinoids are generally considered to be safe, data show that they can cause harmful effects on human and environmental health. Due to the lack of information on their mechanism of toxicity, the effects of imidacloprid and thiamethoxam on DNA methylation as the most used marker for epigenetic effects were investigated in human neuroblastoma (SH-SY5Y) cells. The cells were exposed to imidacloprid and thiamethoxam in concentrations of 100, 200, and 500 μM for 24 hours, then global DNA methylation and expression of genes involved in global DNA methylation (DNMT1, DNMT3a and DNMT3b) were investigated. Global DNA methylation significantly increased after imidacloprid exposure at 100 μM, and thiamethoxam exposures at 200 µM and 500 μM (>1.5-fold). Imidacloprid significantly decreased the expression of DNMT1 and DNMT3a, whereas thiamethoxam did not cause any significant changes in the expression of DNMT genes. Our findings suggested that alteration in global DNA methylation may be involved in the toxic mechanisms of imidacloprid and thiametoxam.
Introduction
Neonicotinoid insecticides bind to nicotinic acetylcholine receptors on the post-synaptic membrane in both mammals and insects nervous systems. 1 The affinity of neonicotinoids to nicotinic acetylcholine receptors in mammals is lower than in insects, which makes their predicted toxicity in mammals and aquatic organisms lower than in the insects.1,2 Since neonicotinoids reportedly have less toxicity in insect populations than organophosphates and other insecticides, they have been used extensively. Due to the long-term intensive use of organophosphates and carbamates in the insect control, resistance to these insecticides has developed. This consequence stimulated the production and use of a new class of insecticides, the neonicotinoids, with high activity at the low concentrations, selectivity to insects and low toxicity to humans and ecosystems. 3 Imidacloprid, belonging to the chloronicotinyl insecticides, became the world’s best-selling neonicotinoid. Thiamethoxam, a thianicotinyl sub-group of insecticide, was second in terms of sales of neonicotinoids. 4 The widespread use and misuse of these two neonicotinoid insecticides have caused problems that significantly affect the health of bees and mammals. The US Environmental Protection Agency (EPA) classified imidacloprid as a Group E chemical, with “evidence of non-carcinogenicity for humans” and thiamethoxam is classified as “not likely to be carcinogenic to humans”.5,6
Previous data indicated that imidacloprid has cytogenetic and DNA damaging effects on rat bone marrow;7,8 induced oxidative stress and lipid peroxidation in rat and mouse liver, brain, and kidney9-13 and in rat ovary; 14 impaired learning and memory 15 and induced other neurotoxic effects.16,17 Studies have shown that thiamethoxam caused oxidative damage, antioxidant response, DNA damage, histological changes, cell necrosis, apoptosis, inflammatory cell infiltration.18-24 Thiamethoxam leads to hepatocellular hypertrophy, fatty changes in liver, and suppression of humoral and cellular immune responses such as leucopenia and lymphocytopenia in mouse.18,19,25 Thiamethoxam also had adverse effects on the development in the embryos of mouse and rabbit 26 and caused neurobehavioral and biochemical changes in rats.2,27 In addition, thiamethoxam caused antioxidant system changes, increase in biochemical markers, apoptosis and necrosis in rat myocardial. 28
Since the effects of imidacloprid and thiamethoxam on DNA damage are not clearly demonstrated, it was hypothesized that non-genotoxic (epigenetic) alterations may play a role in their neurotoxic effects. Therefore, in present study the effects of imidacloprid and thiamethoxam on global DNA methylation and the expression of genes involved in global DNA methylation were evaluated in human neuroblastoma SH-SY5Y cells.
Materials and Methods
Chemicals
Imidacloprid (99.9%) and thiamethoxam (99.6%) were obtained from Sigma-Aldrich (St Louis, MO, USA). A stock solution (500 mM) for each compound was prepared by dissolving in dimethylsulfoxide (DMSO; Sigma-Aldrich) and kept at −20°C. Cell culture media and all supplements were purchased from Wisent Bioproducts (Saint-JeanBaptiste, QC, Canada). DNA and RNA isolation kits, cDNA synthesis kit, and SYBR-Green Master Mix were obtained from Roche Life Sciences (Penzberg, UpperBavaria, Germany). 5-Methylcytosine (5-mC) DNA ELISA kit was purchased from Epigentek (Farmingdale, NY). Primers for gene expression studies were obtained from Sentromer DNA Technologies (Istanbul, Turkey).
Cell Culture and Exposure
SH-SY5Y cell lines were obtained from the American Type Culture Collection (ATCC®). SHSY-5Y cells were maintained in Dulbecco’s Modified Eagle’s medium/Ham’s nutrient mixture F-12 (DMEM-F12) medium containing 10% fetal bovine serum and penicillin-streptomycin (100 U/mL) and incubated at 37°C in a humidified atmosphere with 5% carbon dioxide. Subculturing was performed when the cells reached 70– 80% confluence (3 days in general) using trypsinization. The cells were counted by the Lunacell counter (Annandale, VA, USA) and cell viability was determined by trypan blue dye exclusion. For the gene expression and DNA methylation analysis, cells (.5 × 106 in 2 mL) were seeded in a 6-well microplate, then exposed to the imidacloprid and thiamethoxam at concentrations of 100, 200, and 500 μM, and DMSO (1%) as a control, for 24 h. All experiments were repeated at least three independent times and each test was carried out in triplicate. The exposure concentrations of imidacloprid and thiamethoxam used in these experiments (100-500 μM) are similar to the concentrations used in the previous studies of toxicity mechanisms of imidacloprid and thiamethoxam. 29
Global DNA Methylation Analysis
For the global DNA methylation analysis, genomic DNA was isolated from SH-SY5Y cells using the High Pure PCR Template Preparation kit (Roche Applied Science, Mannheim, Germany) according to the instructions provided by the manufacturer. DNA concentration (optical density (OD): 260) and quality (OD: 260/280)) were evaluated by spectrophotometric measurement. Global DNA methylation analysis was performed by measuring the levels of 5-methylcytosine (5-mC) in total DNA using Elisa kit (Epigentek, Farmingdale, NY) according to the instructions provided by the manufacturer. Briefly, 100 ng of input genomic DNA samples and standard solutions, or the control solution were treated with binding solution, added to the kit plate, and incubated at 37°C. After 90 minutes the plate was washed with buffer. The plate-bonded DNA samples were incubated with an anti-5-mC monoclonal antibody and detection antibody. After the addition of enhancer and developer solution, the absorbance was read at 450 nm using a microplate spectrophotometer system (Biotek-Epoch, Winooski-Vermont, USA). Positive control (DNA containing 5% 5-mC) and negative control (100% unmethylated DNA) were used for preparation of the standard calibration curve. To calculate percentage of 5-mC, first we generated a standard calibration curve (in the range of .1-5% of 5-mC) and plotted the OD values of samples vs the standard calibration curve at each percentage point. The calculated 5-mC% is expressed as 5-mC/total DNA (A + G + C + T).
Gene Expression Analysis
The primer Sequences and the Annealing Temperatures of DNA Methyl Transferase Related Genes.
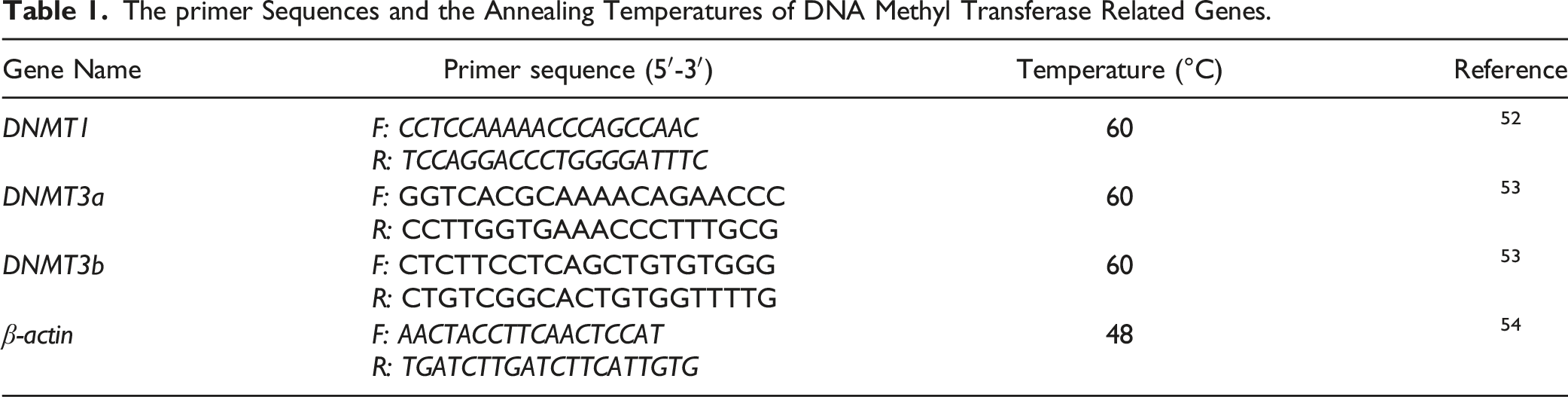
Statistical Analysis
5-mC% levels were expressed as the mean ± standard deviation (SD). The DNMT gene expression was represented as the relative expression compared to that for the β-actin gene at the same samples. The statistical analysis was performed by ANOVA followed by Dunnett’s multiple comparison test using SPSS version 21.0 for Windows (IBM Analytics, New York, USA). P values less than .05 and .001 were considered statistically significant.
Results
Cytotoxicity
Cytotoxicity studies of imidacloprid and thiamethoxam have been investigated in our previous study. 29 According to the methyl tetrazolium (MTT) test, the IC50 values was 3.69 mM and 1.33 mM for imidacloprid and thiamethoxam, respectively, after 24 h of exposure in SH-SY5Y cells.
Global DNA Methylation
Exposure to imidacloprid at 100, 200, and 500 µM for 24 h, significant increase in 5-mC% levels was observed only in the 100 µM group (1.5-fold, p < 0.05), compared to the control group, while there was no significant change at 200 and 500 µM groups (Figure 1). For the cells treated with 100, 200, and 500 μM of thiamethoxam, increases in 5-mC% levels was significant for 200 µM (1.62-fold, P < .05) and 500 μM (1.56-fold, P < .05) but not for 100 µM (1.4-fold, p > 0.05) (Figure 1). Effects of imidacloprid and thiamethoxam (100-500 μM) on levels of 5-mC% in SH-SY5Y cells after 24 h exposure. Data are presented as mean ± SD. (Statistical analysis was performed by ANOVA + Dunnett post hoc test. Statistically significant changes are indicated by *P < .05).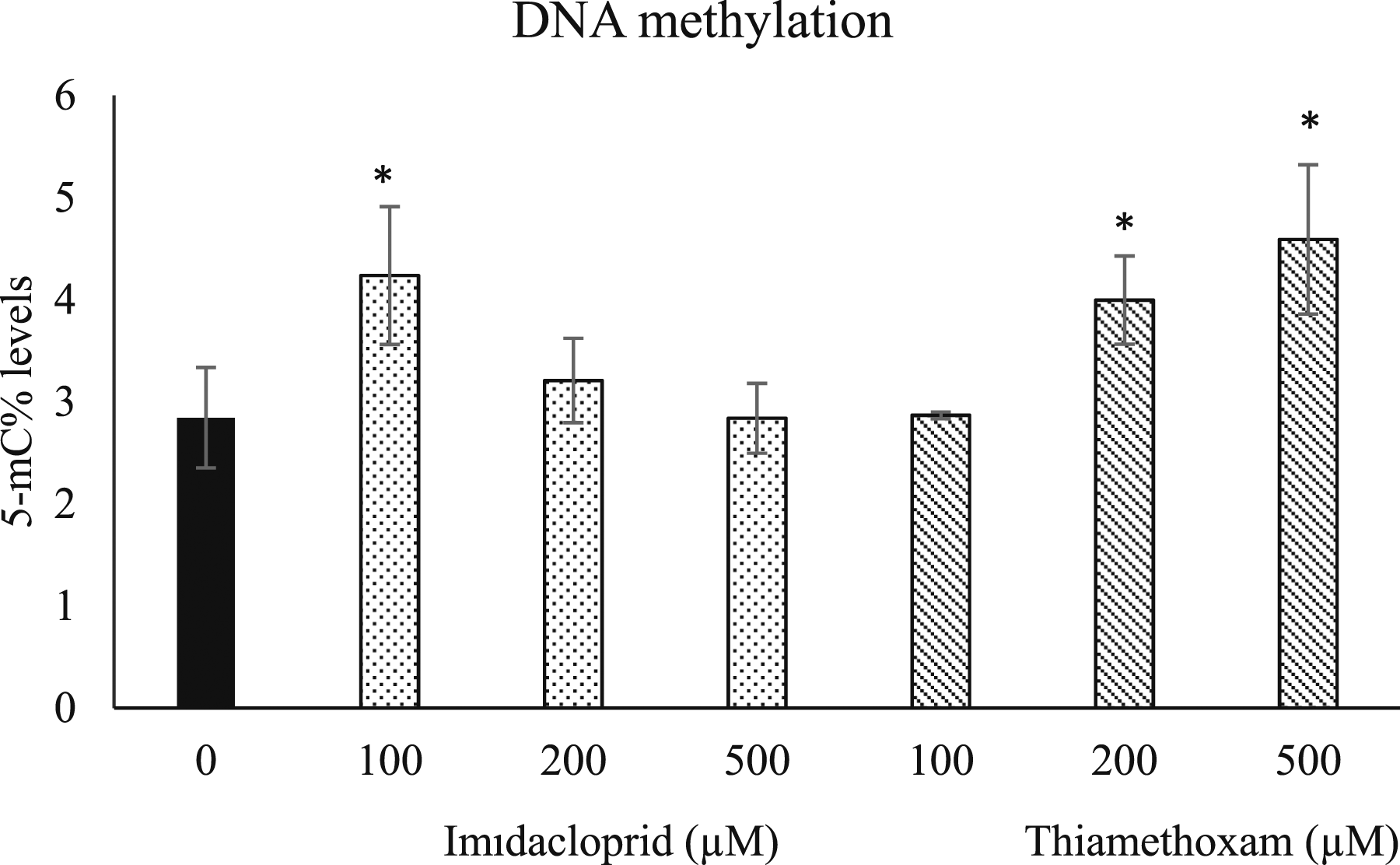
Gene Expression Levels
In SH-SY5Y cells exposed to imidacloprid at 100, 200, and 500 µM concentrations, a concentration-dependent decrease in gene expression of DNMT1 and DNMT3a was observed (Figure 2). For the DNMT1 gene expression the reduction was significant only at the highest exposure group (500 µM) of imidacloprid at 1.4-fold (P < .05) (Figure 2A). The reduction in DNMT3a gene expression was significant for all imidacloprid exposure groups and it was 1.5, 1.4 and 1.4-fold for 100, 200 and 500 µM, respectively (Figure 2B). Imidacloprid did not change the expression levels of DNMT3b compared to the control group (Figure 2C). Effects of imidacloprid and thiamethoxam (100-500 μM) on relative gene expression levels of DNMT1 (A), DNMT3a (B) and DNMT3b (C) in SH-SY5Y cells after 24 h exposure. Data are presented as mean ± SD. (Statistical analysis was performed by ANOVA + Dunnett post hoc test. Statistically significant changes are indicated by *P < .05).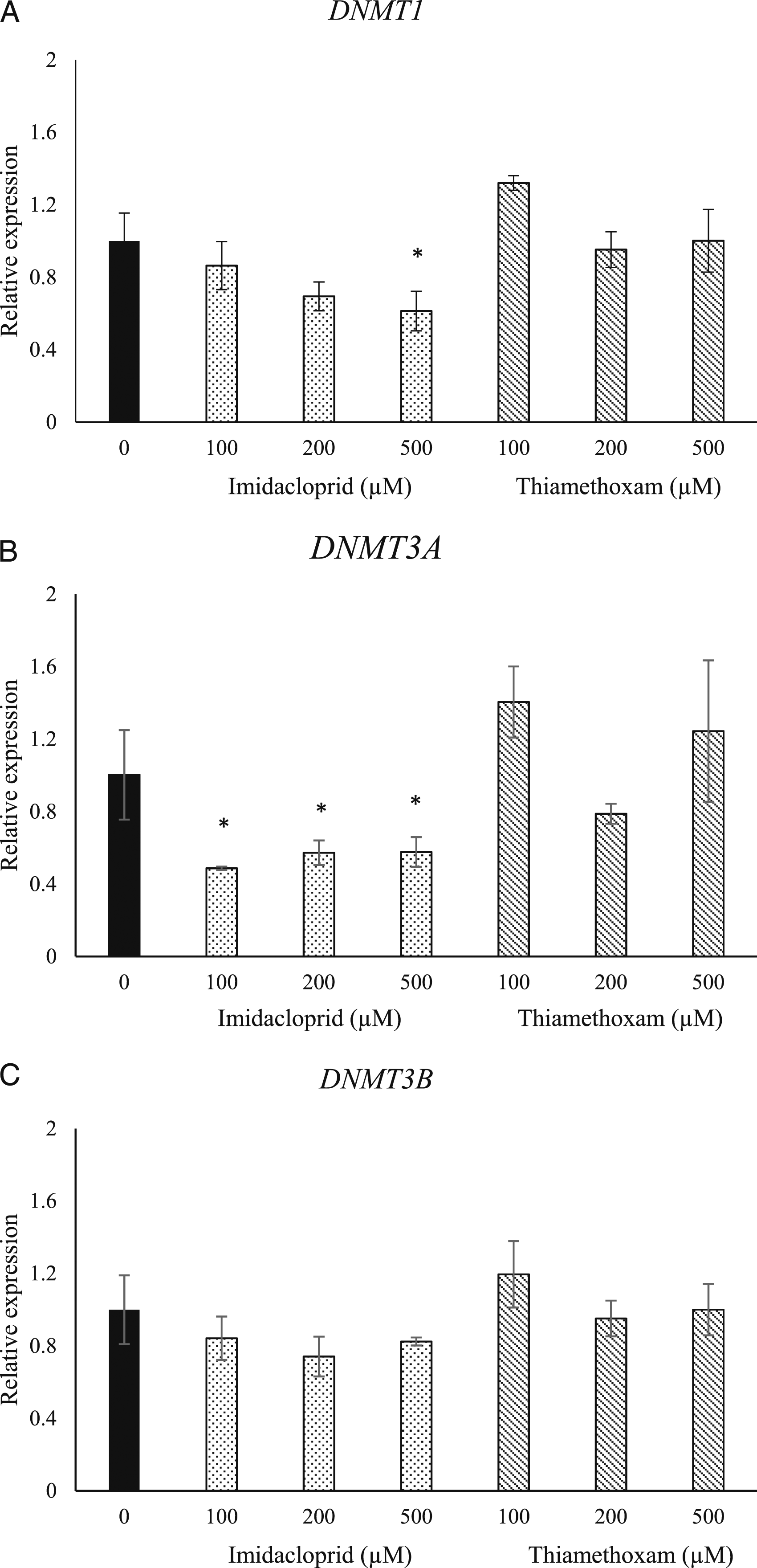
Exposure to thiamethoxam for 24 hours increased the expression of DNMT1 at 100 µM (1.3-fold) but decreased it at 200 µM (1.5-fold) and did not change the expression for the 500 µM group (Figure 2A). However, none of these changes were significant. Similarly, an increase in DNMT3a gene expression was noted when the cells were treated with 100 µM (1.4-fold), a decrease in the gene expression at 200 µM (1.21-fold) and an increase in 500 µM (1.24-fold) were observed; none of the changes in DNMT3a were significant (Figure 2B). An increase in DNMT3b gene expression level was observed in 100 µM (1.19-fold) but no changes were noticed in the 200 and 500 µM groups (Figure 2C). In conclusion, no significant changes in the expression of DNMT1, DNMT3a, and DNMT3b were observed after the exposure to thiamethoxam for 24 hours (Figure 2).
Discussion
Neonicotinoids are alternative insecticides to organophosphates with higher selectivity and efficacy toward insects.31,32 Although neonicotinoids are reported to have low mammalian toxicity, some studies have shown that neonicotinoids cause neurotoxicity, immunotoxicity, hepatotoxicity, nephrotoxicity, and reproductive system toxicity in humans, vertebrates, and invertebrate animals based on the mechanism of cytotoxicity and oxidative damage.33-37 A single intraperitoneal injection of imidacloprid (337 mg/kg, .75 × LD50, in corn oil) administered to pregnant Sprague-Dawley rats, resulted in significant neurobehavioral deficits and sensorimotor disturbances associated with increased AChE activity in the midbrain, cortex and brainstem, and plasma; histopathological change in the cerebellum of offspring exposed in utero were also observed. 16 In a 90 days oral toxicity study in rats, imidacloprid decreased body weight at 20 mg/kg/day and at necropsy the relative body weights of liver, kidney and adrenal was significantly increased, while spontaneous locomotor activity decreased and mild pathological changes in brain, liver and kidney tissues were observed at this dose level. 17 Thiamethoxam decreased the locomotor activity and acetylcholinesterase (AChE) activity in the striatum of rats in a dose-dependent manner. 27 Therefore, we used the neuroblastoma cell lines for the toxic effects of imidacloprid and thiamethoxam. Some studies stated that neonicotinoids might have genotoxic effects;8,38-47 however their effects on molecular and especially epigenetic mechanisms defining the regulation of such toxic effects remains to be elucidated. In our previous study we observed cell viability data that showed that 50% of inhibitory concentration values of imidacloprid and thiamethoxam were in 3.69 and 1.33 mM in SHSY-5Y cells for 24 h. 29 We also demonstrated imidacloprid and thiamethoxam caused DNA damage by comet assay only at 500 µM after 24 h and 48 h exposure in SH-SY5Y cells. Therefore, in present study we selected the exposure concentrations (in the range of 100-500 μM), which corresponded to approximately .02-.4 of LC50 in SH-SY5Y cells to represent a low exposure, to study how DNA methylation was affected in similar conditions as in our previous study. 29
There are a few studies on the effects of imidacloprid on DNA methylation changes especially in non-mammalian organisms. An elevation of 5-mC% levels status was reported in western honeybees in response to imidacloprid after exposure to 5 or 200 μg/kg concentration for 1-20 days 48 and high CpG methylation was observed especially in the axons after .01 μg/mL of imidacloprid in Bombus terrestris bee. 49 A decrease in the 5-mC% status and alterations in gene-specific DNA methylation were observed in Colorado potato beetle Leptinotarsa decemlineata after 10 replicate exposures to 1.0 and .01 mg/kg doses, which suggested an epigenetic mechanism in imidacloprid induced toxicity. 50 In the present study, imidacloprid significantly increased the 5-mC% status and decreased the expression of DNMT1 and DNMT3a genes that play a role in DNA methylation in human neuroblastoma SH-SY5Y cells. A previous study evaluated the effect of imidacloprid on the methylation status in the mouse embryonic stem cells in 1 × 10−8 to 1 × 10−6 M concentrations for 48 h, which showed an increase in the 5-mC% status, and decreased methylation of cell apoptosis, cell adhesion and tissue development genes such as Cdkn2, Apc, Cdh1, Timp3 and Rarb. 51 To the best of our knowledge, there are no studies evaluating the epigenetic effects of thiamethoxam in mammalian models or in the other ecosystem organisms. In the present study, exposure of SH-SY5Y cells to thiamethoxam for 24 hours elevated 5-mC% status; however, thiamethoxam induced non-significant changes in the expression of DNA methylation-related genes.
Conclusions
We observed that imidacloprid and thiamethoxam may cause alterations in 5-mC% levels and expression of DNMT genes, which suggests that epigenetic mechanisms may be involved in neonicotinoid induced toxicity.
Footnotes
Author Contributions
Guler, Zeynep Rana contributed to conception and design, contributed toacquisition, and drafted manuscript; Yilmaz, Umran contributed to conception,contributed to acquisition, and drafted manuscript; Uzunosmanoglu, Kubracontributed to design, contributed to acquisition, and drafted manuscript;Ozturk, Busra contributed to design, contributed to acquisition, and draftedmanuscript; Abudayyak, Mahmoud contributed to conception and design,contributed to analysis and interpretation, drafted manuscript, and criticallyrevised manuscript; Ozden, Sibel contributed to conception and design,contributed to analysis and interpretation, drafted manuscript, and criticallyrevised manuscript. All authors gave final approval and agree to be accountablefor all aspects of work ensuring integrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Scientific Research Projects Coordination Unit of Istanbul University (Project numbers: 35359).
