Abstract
Vaccines containing mRNA with the capacity to self-amplify represent an alternative to the mRNA vaccines that came to prominence during the COVID-19 pandemic. To gain further insights on the safety profile of self-amplifying mRNA- (SAM-) vaccines, this preclinical toxicology study in rats evaluated the effect of (i) the type of delivery system (lipid nanoparticle [LNP] vs cationic nano-emulsion [CNE]); (ii) antigen-encoding sequence (rabies glycoprotein G vs SARS-CoV-2 Spike); and (iii) RNA amplification. Further analyses also evaluated gene expression in peripheral blood after vaccination, and the biodistribution of vaccine RNA. The SAM vaccines administered as two doses 2-weeks apart had acceptable safety profiles in rats, with respect to clinical signs, blood biochemistry, and macroscopic and microscopic pathology. A transient increase in ALT/AST ratio occurred only in female rats and in the absence of muscle and liver damage was dependent on RNA amplification and appeared related to the greater quantities of vaccine RNA in the muscle and livers of female rats vs male rats. The RNA and delivery-vehicle components, but not the nature of the antigen-coding sequence or the requirement for RNA amplification, affected aspects of the stimulation of innate-immune activity, which was consistent with the transient activation of type I and type II interferon signaling. The delivery vehicle, LNP, differed from CNE as vaccine RNA in CNE compositions appeared independently to stimulate innate-immune activity at 4 hours after vaccination. Our analysis supports further studies to assess whether these differences in innate-immune activity affect safety and efficacy of the SAM vaccine.
Introduction
RNA-based vaccines came into prominence during the COVID-19 pandemic because of two vaccines, one from Moderna and one from Pfizer/BioNTech, that have been, and continue to be, at the forefront of the global vaccination campaign. 1 Those vaccines contain mRNA coding for the SARS-CoV-2 Spike (S) antigen, and have demonstrated the feasibility, safety and effectiveness of delivering mRNA into the cells of the vaccine recipient such that it can be translated into antigen. Both vaccines are formulated with lipid nanoparticles (LNPs) to deliver the mRNA in the host cells.
Vaccines based on self-amplifying mRNA (SAM) technology represent an alternative mRNA vaccine platform, in that they consist of an engineered alphavirus genome encoding the genes of four non-structural proteins (nsP1-4) that form an RNA-dependent RNA polymerase and the gene for target antigen (Figure 1A).2,3 The vaccine is also formulated with a delivery system to support its stability and its uptake by host cells upon intramuscular injection.2,4-7 After uptake by host cells, the RNA genome is translated to produce the RNA-dependent RNA polymerase, which in turn produces (iterative negative- and positive-strand) copies of the RNA genome and RNA subgenome fragments coding only for the antigen. The theoretical advantage of the SAM vaccine is that the quantity of RNA in a given dose can be lower than the dose in a conventional mRNA vaccine, because of the amplification of the antigen encoding mRNA.2,8-13 Vaccine mechanism of self-amplification and antigen expression, and study details. (A) Schematic description of self-amplifying mRNA (SAM) vaccine cellular uptake, amplification and antigen expression in a host cell. In this scheme, SAM complexed with the delivery vehicle is endocytosed by the host cell and then released into the cytoplasm. The non-structural protein (nsP) coding regions, nsP1 to nsP4, of SAM are translated by the host cell to produce the RNA-dependent RNA polymerase (RDRP). The RDRP transcribes negative-strand copies of the SAM which then serve as templates for the same enzyme complex to produce multiple copies of antigen (Ag) coding mRNA. The antigen-coding mRNA is then translated by the host cell to produce the antigen. (B) Schematic description of the study based on the standard repeat-dose toxicology design with additional analyses of RNA in peripheral blood, and of the biodistribution of the CoV2 SAM (LNP) RNA. Male and female Sprague-Dawley rats were dosed on two occasions 14-days apart. Each dose was administered as two intramuscular injections, one for each hind limb, with the Day 15 injections more proximal on the hind limb than the Day 1 injections. Each group was divided into two subgroups (A and B) to allow for different sampling regimes and two different times for necropsy (3 days or 14 days after the second dose, respectively). CRP is C-reactive protein.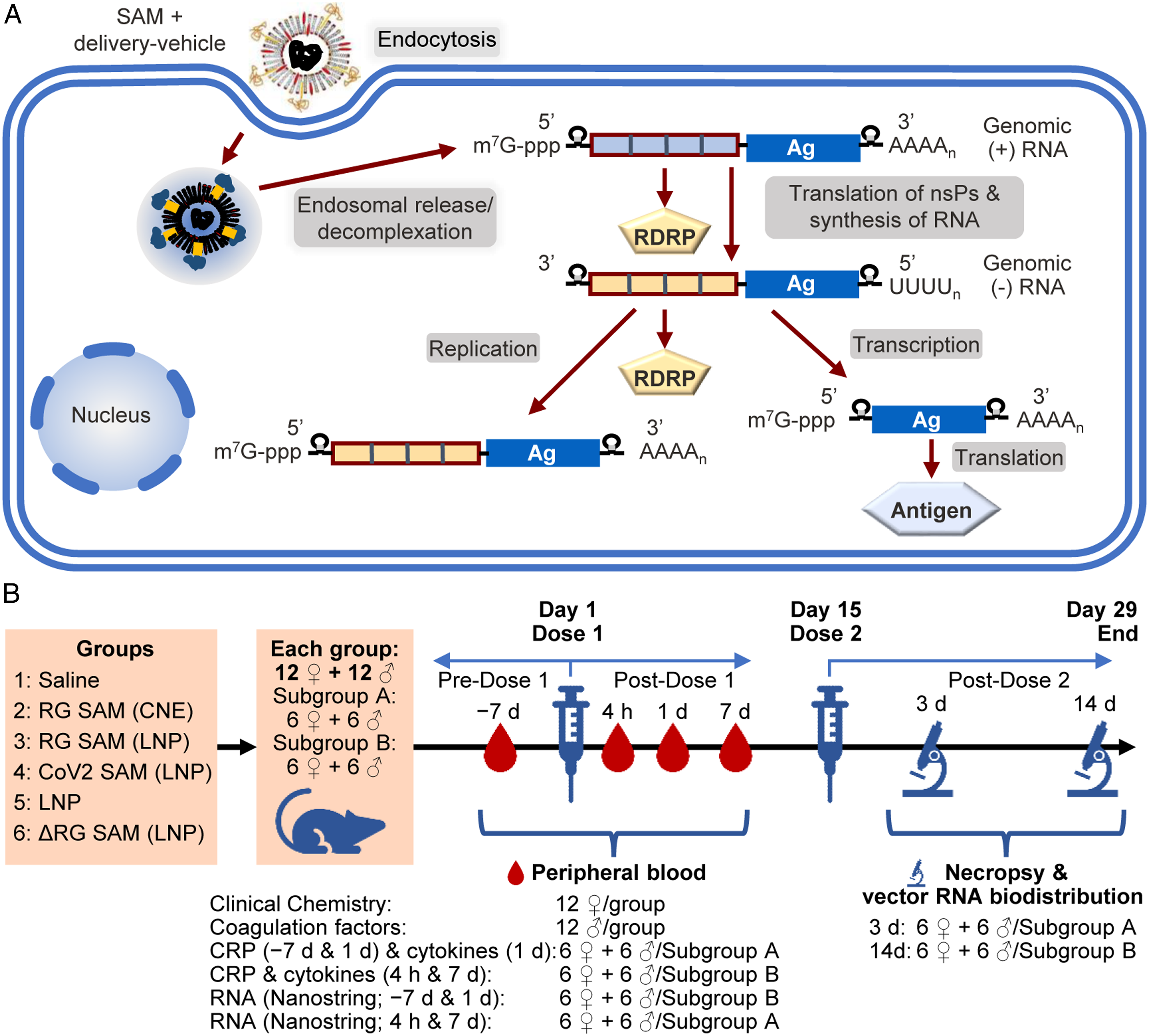
The results from previous independent preclinical toxicology studies in rats of the SAM vaccine coding for the rabies glycoprotein G (RG) antigen (RG SAM [CNE]) and of the self-amplifying mRNA vaccine coding for SARS-CoV-2 pre-fusion stabilized S glycoprotein (CoV2 SAM [LNP]) showed that these vaccines have acceptable safety profiles, with vaccine-related findings consistent with a transitory inflammatory response at the injection sites and lymph nodes draining the injection sites.14,15 The delivery systems used in those two studies were cationic nano-emulsion (CNE) and LNP, respectively, both of which potentially have immunostimulatory properties by themselves.2,4,6,14-18 Those studies were performed in the rat model, because in contrast to rabbits, rats (and humans) express TLR7, one of the pattern-recognition receptors that recognizes exogenous RNA and activates innate immunity.2,8-13,19,20 Antigen-coding RNA was detected by RT-PCR at the injection site and draining lymph nodes one day after injection, after which levels declined. Antigen-coding RNA was also transiently detected in blood, lungs, spleen and liver. Moreover, a vaccine using a similar self-amplifying mRNA platform, formulated with LNP and coding for pre-fusion stabilized SARS-CoV-2 S, raised no short-term safety concerns in a dose-escalation Phase 1 clinical trial of nearly 200 adult individuals. 21
To gain further insights on the safety profile of SAM-based vaccines, this preclinical toxicology study in rats was undertaken to evaluate the effect of (i) the type of delivery system (LNP vs CNE); (ii) antigen-encoding sequence (RG vs SARS-CoV-2 S); and (iii) the effect of RNA amplification (by comparing with a vector engineered to block self-amplification). Further analyses were also performed to evaluate changes in gene expression in peripheral blood after vaccination, and to evaluate the biodistribution of two vaccine-derived RNA species that were potentially subject to different rates of amplification and degradation.
Methodology
Study ethics and regulatory principles
The study was performed in accordance with the OECD Principles of Good Laboratory Practice and as accepted by Regulatory Authorities throughout the European Union, United States of America (FDA), Japan (MHLW), and other countries that are signatories to the OECD Mutual Acceptance of Data Agreement.
GSK is committed to the replacement, reduction and refinement of animal studies (3Rs). Non-animal models and alternative technologies are part of our strategy and employed where possible. When animals are required, application of robust study design principles and peer review minimizes animal use, reduces harm and improves benefit in studies.
Animal Husbandry
Sprague Dawley rats were obtained from Charles River Labs International Inc. (Raleigh, NC, USA), were 10 weeks old and weighed 306-483 g and 200-286 g for males and females, respectively, at the start of treatment. Twenty-four animals (12/sex) were randomly allocated to each of the six groups (Figure 1). Animals were group housed (up to 3 animals of the same sex and same treatment group) in polyethylene cages containing appropriate bedding, with the temperature set at 18°C to 24°C, humidity at 30% to 70%, and under a cycle of 12 hours light and 12 hours dark. Control group animals were housed on a separate rack from the animals of the test-item groups. Animals were fed with Envigo Teklad Certified Global Rodent Diet #2018C and had ad libitum access to municipal tap water, treated by reverse osmosis and ultraviolet irradiation. For psychological/environmental enrichment, animals were provided with items such as a hiding device, a chewing object, and occasional treats.
Study Design and Conduct
The study design followed a standard design for repeat dose toxicology and similar to that conducted previously on the RG SAM (CNE) vaccine. 15 Animals received doses on Days 1 and 15 (Figure 1B). Each dose was administered as two 100-μl intramuscular injections in the right and left thigh muscles of the hind limbs, respectively. The injection sites in a given limb were separated, the Dose 2 site was more proximal than the Dose 1 site. The injection site was shaved not more than one day before administration and disinfected with 70% ethanol just prior to injection. All test items and the saline control were obtained from GSK, Rockville, MD USA.
Group 1 was administered saline; Group 2 was administered the SAM vaccine coding for rabies strain Flury-LEP optimized for human expression (RG SAM) formulated with CNE (RG SAM (CNE); 15 15 μg RNA/dose); Group 3 was administered SAM-RG formulated with LNP (RG SAM (LNP); 12 μg RNA/dose); Group 4 was administered SAM coding for pre-fusion stabilized SARS-CoV-2 S (CoV2) formulated with LNP (CoV2 SAM [LNP]; 14 12 μg RNA/dose); Group 5 was administered LNP without RNA (at the same concentration and quantity contained in the LNP-formulated vaccines); and Group 6 was administered amplification-defective RG SAM (LNP), where the SAM backbone had been engineered to suppress self-amplification by mutating the nsP2 gene (RG ΔSAM [LNP]; 12 μg RNA/dose). Male and female animals (1:1) in each group were randomly assigned (1:1) to two subgroups (Subgroups A and B; Figure 1B). Animals in Subgroup A were necropsied 3 days after the second dose, whereas animals in Subgroup B necropsied 14 days after the second dose.
The SAM vaccine dose administered to the rats (i.e., 12 μg) represented approximately one third of the highest planned human dose (30 μg), equating to an estimated exposure margin of approximately 66 times the highest planned human dose, considering a 50 kg human body weight and a 300 g rat body weight.
Clinical examinations were performed as previously described, 15 except that rectal temperature was measured prior to and 6, 12, 24 and 48 hours after each injection. Peripheral-blood samples were collected 7 days prior to the first dose, and 4 hours, 1 day and 7 days after the first dose for analyses on clinical chemistry, coagulation factors, C-reactive protein (CRP) and cytokine concentrations (interleukin- [IL] 1β, IL-6, IL-10, tumor necrosis factor- [TNF] α, interferon- [IFN] γ, and IL-4), as previously described 15 and for gene expression (Nanostring Technologies, Inc. Seattle WA, USA). Due to blood volume limitations, the collection of blood samples for the different analyses was dependent on the subgroup allocation (Figure 1B).
Tissue processing, histopathological examination, and RT-qPCR for vaccine-derived-RNA biodistribution analysis (only CoV2 SAM (LNP) – Group 4), were performed as previously described, except for the RT-qPCR primers and probes sequences. 15 The RT-qPCR analysis was limited to the CoV2 Sam (LNP) vaccine to focus the assessment and not introduce other variables like different antigen constructs.
For the RT-qPCR, the primers and probes sequences were as follows:
The two primer sets were used to take into account potential mRNA degradation that is more likely to occur from the 3’ end of capped transcripts. 22 The SAM vector is longer than a conventional RNA vector because of the inclusion of the coding sequences for the nsPs. Hence the nsP1 primers target a section of the mRNA that is ∼10.7 kb upstream from the sequence targeted by the SARS-CoV-2 S primers.
Statistical analyses for the comparison of mean body weight, organ weights and relative organ weights, mean body temperature, food consumption, hematology, coagulation, or clinical chemistry mean values were performed as previously described. 15 Briefly, groups were compared using an overall one-way ANOVA F-test. A Levene test was performed on residuals to assess the homogeneity of group variances. When the Levene test was not significant (P > .05) and the overall F-test was significant (P ≤ .05), a t-test on least-square means was used to perform the pairwise comparisons of interest. When the Levene test was significant, the F-test was discarded and an overall Kruskal-Wallis test was performed. When the Kruskal-Wallis test was significant (P ≤ .05), a Wilcoxon rank-sum test was used to perform the pairwise comparisons of interest.
Peripheral-Blood RNA Analysis
RNA was purified from PAXgene stabilized peripheral-blood samples using MagMAX total RNA isolation kit (Life Technologies) with slight modifications to the protocol. Briefly, blood-sample tubes were thawed for at least 30 minutes at room temperature and centrifuged at 3000g for 10 minutes to collect pellet of the crude RNA. Pellets were then washed with 100μL of nuclease-free water (Ambion) and resuspended by vortexing. Tubes were then centrifuged at 3000g for 10 minutes and supernatant was discarded. The pellet was mixed in resuspension solution (resuspension buffer + proteinase) and resuspended by vortexing. Resuspended solution was transferred to 1.5 mL Lo-DNA-bind Eppendorf tubes and placed in a thermomixer for 10 minutes at 55°C and 1000 rpm. Samples were then centrifuged at 16000 g for 6 minutes and the supernatants were transferred to a 96-well processing plate on semi-automated KingFisher Flex 96 purification instrument. Briefly, the lysate was treated with TURBO DNase, using MagMAX magnetic bead-based methodology. The quality of the total RNA was assessed using NanoDrop8000 spectrophotometer (ThermoScientific) and Qubit high-sensitivity RNA kit (Invitrogen), and concentration was measured using Agilent TapeStation4200 Bioanalyzer (Agilent Technologies).
RNA samples were processed using the NanoString multiplex system with the mouse immunology gene panel (NanoString Technologies, Inc., #115000052) and associated methodology. Briefly, 100 ng input RNA was mixed with reporter-capture probe pairs and incubated at 65°C for 16 hours using Thermocycler (Life Technologies). Raw data were analyzed using nSolver 4.0 software (NanoString Technologies, Inc.) and normalization of the counts was done using a standard method an geNorm algorithm. 23 The nine highest ranked genes for invariable expression were selected for reference gene normalization. After normalization, log2 gene ratios were used in the analysis of differentially expressed genes and gene set (pathway) analysis. Multivariate differential expression analysis was performed using optimal methods for each gene. A negative binomial mixture model for low expression probes or a simplified negative binomial model for high expression probes was used by defining experimental variables as predictor. Resulting P-values were adjusted for multiple comparisons by Benjamini-Hochberg adjustment method. 24 All data (all groups and all timepoints) were normalized together, and differential expression was assessed by group using time-point as predictor and using the baseline timepoint as the contrast. For pathway analysis, a gene set enrichment module was used. In this analysis, global significance for a covariate was determined by measuring the cumulative evidence for the differential expression of genes in a pathway and calculated as the square root of the mean squared t-statistics of the genes, whereas the directed global significance took into account the sign (negative and positive values representing down- or up-regulated genes, respectively) of the t-statistics. 25
Results
Clinical Signs, Including Temperature
No SAM-based vaccine or the LNP control was associated with adverse clinical signs. No deaths occurred, and clinical signs such as skin or hair conditions, swelling of paws, red feces, and salivation were considered incidental because they occurred across groups or in control animals, were transient, and/or were common in laboratory-housed rats.
There were no vaccine-related clinical signs or effects on food consumption or body weights. Slightly reduced body-weight gains were observed after the first and second dose across all groups, and were considered procedural-related because the changes did not affect mean body weights in comparison with the saline control group.
Very slight erythema with undefined edges was observed locally at the 4 injection sites across all groups including the saline control group. After the first dose, erythema and edema resolved by 24 to 48 hours, and in general, resolved more rapidly than after the second dose. However, the incidence of erythema was generally higher after the first dose.
Peak Body Temperatures After Dose 1 and Dose 2.
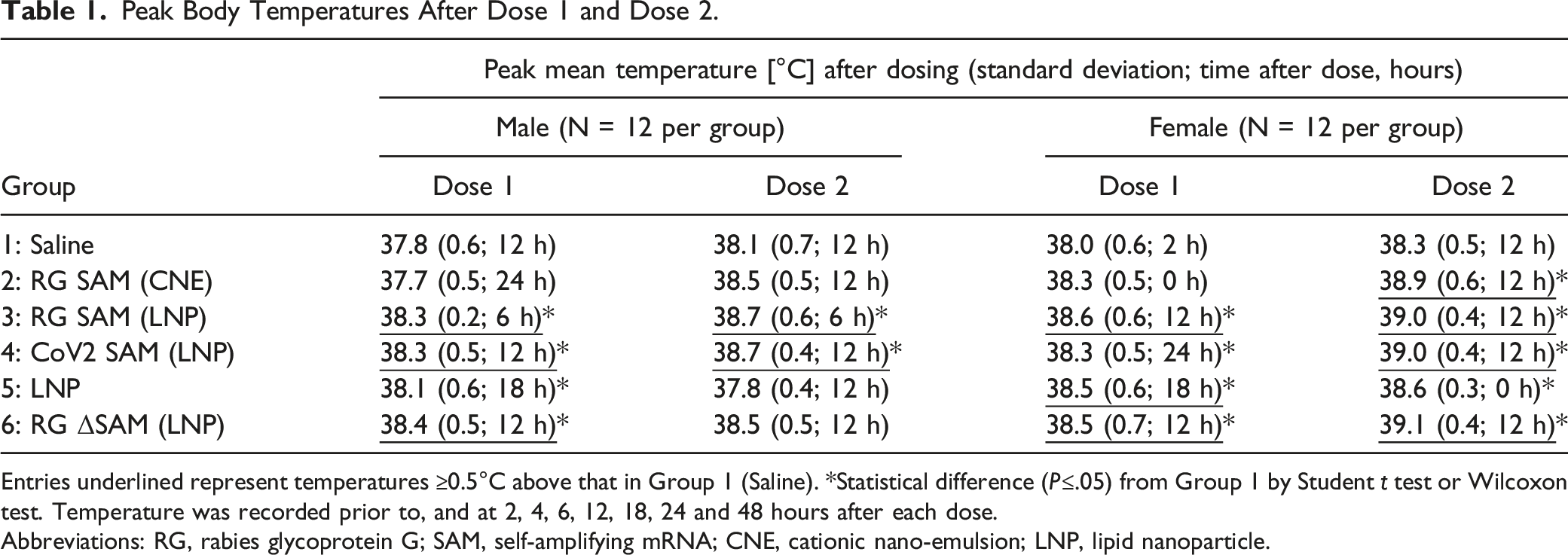
Entries underlined represent temperatures ≥0.5°C above that in Group 1 (Saline). *Statistical difference (P≤.05) from Group 1 by Student t test or Wilcoxon test. Temperature was recorded prior to, and at 2, 4, 6, 12, 18, 24 and 48 hours after each dose.
Abbreviations: RG, rabies glycoprotein G; SAM, self-amplifying mRNA; CNE, cationic nano-emulsion; LNP, lipid nanoparticle.
Peripheral-Blood Biochemistry and Cytokine Expression
Selected Clinical Chemical Parameters in Peripheral Blood Before and After Dose 1.
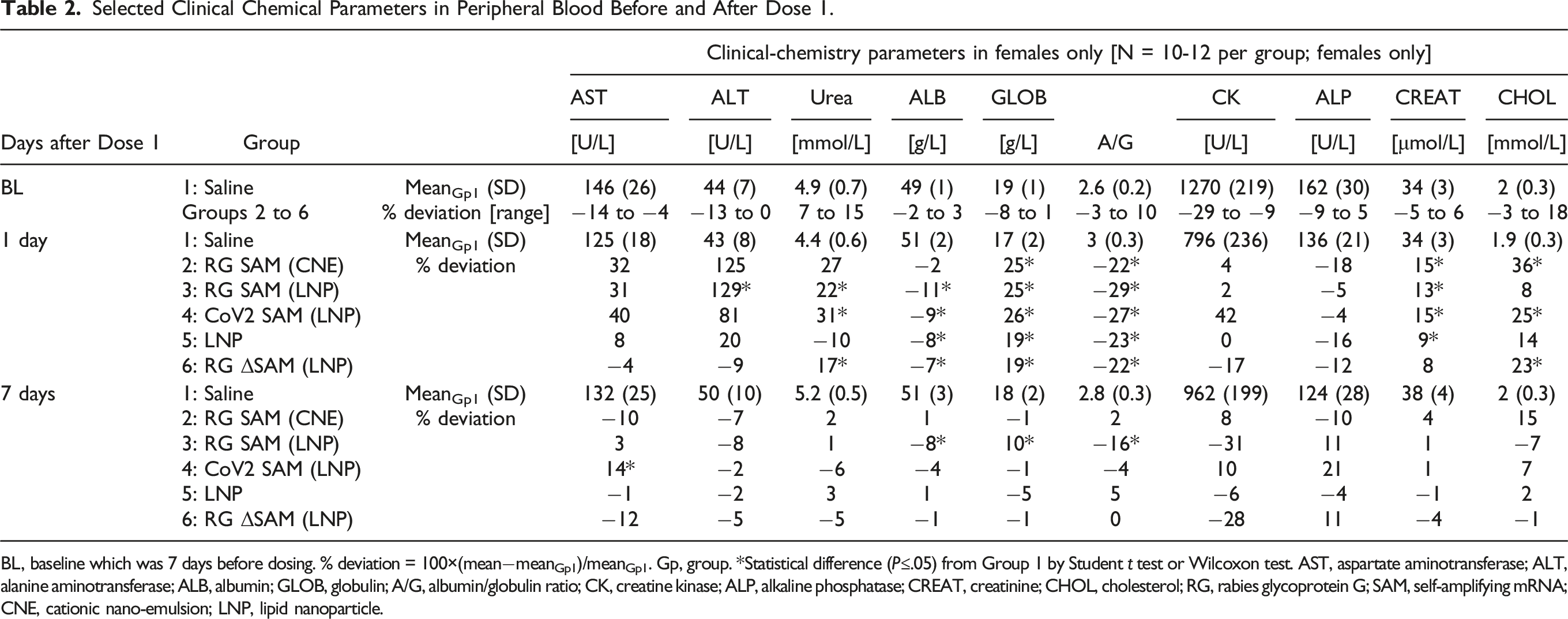
BL, baseline which was 7 days before dosing. % deviation = 100×(mean−meanGp1)/meanGp1. Gp, group. *Statistical difference (P≤.05) from Group 1 by Student t test or Wilcoxon test. AST, aspartate aminotransferase; ALT, alanine aminotransferase; ALB, albumin; GLOB, globulin; A/G, albumin/globulin ratio; CK, creatine kinase; ALP, alkaline phosphatase; CREAT, creatinine; CHOL, cholesterol; RG, rabies glycoprotein G; SAM, self-amplifying mRNA; CNE, cationic nano-emulsion; LNP, lipid nanoparticle.
C-Reactive Protein Concentration in Peripheral Blood Before and After Dose 1.
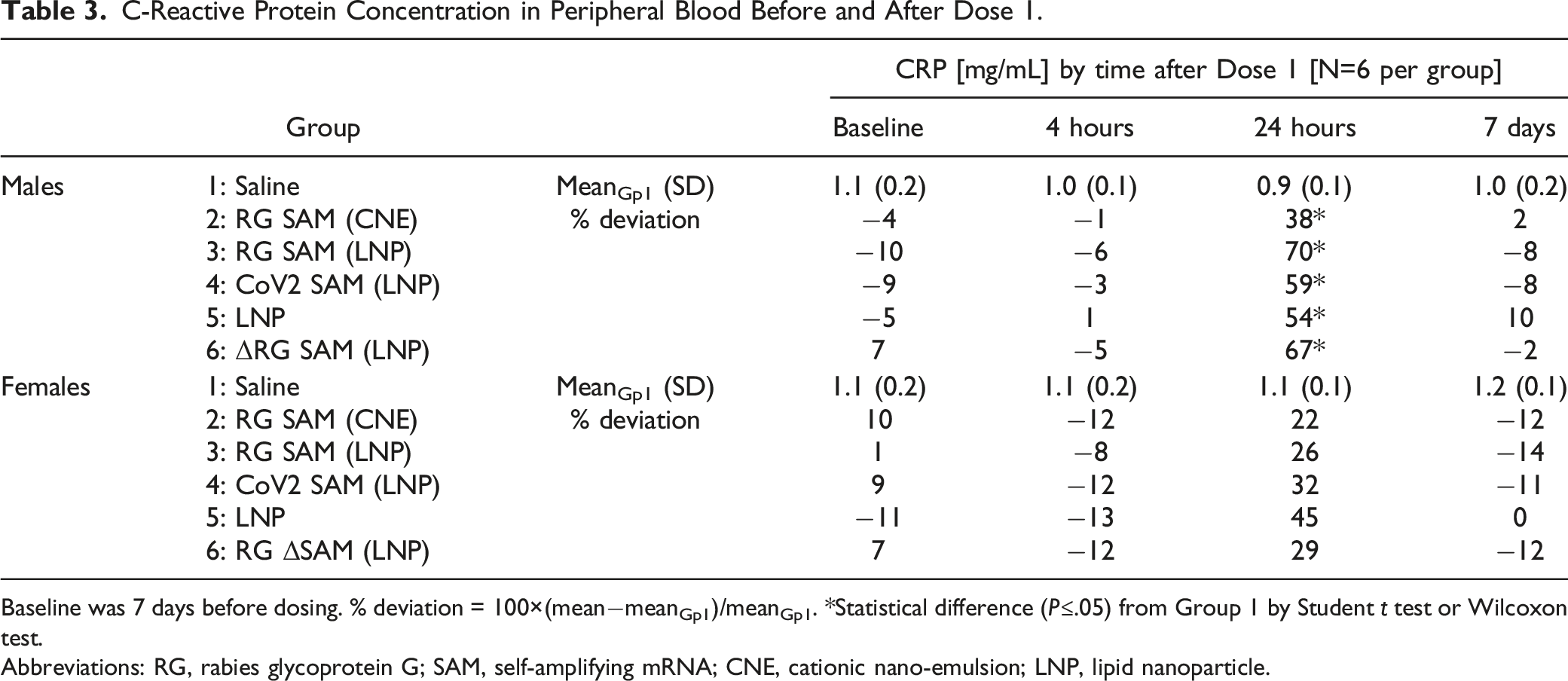
Baseline was 7 days before dosing. % deviation = 100×(mean−meanGp1)/meanGp1. *Statistical difference (P≤.05) from Group 1 by Student t test or Wilcoxon test.
Abbreviations: RG, rabies glycoprotein G; SAM, self-amplifying mRNA; CNE, cationic nano-emulsion; LNP, lipid nanoparticle.
Fibrinogen Concentration in Peripheral Blood Before and After Dose 1.
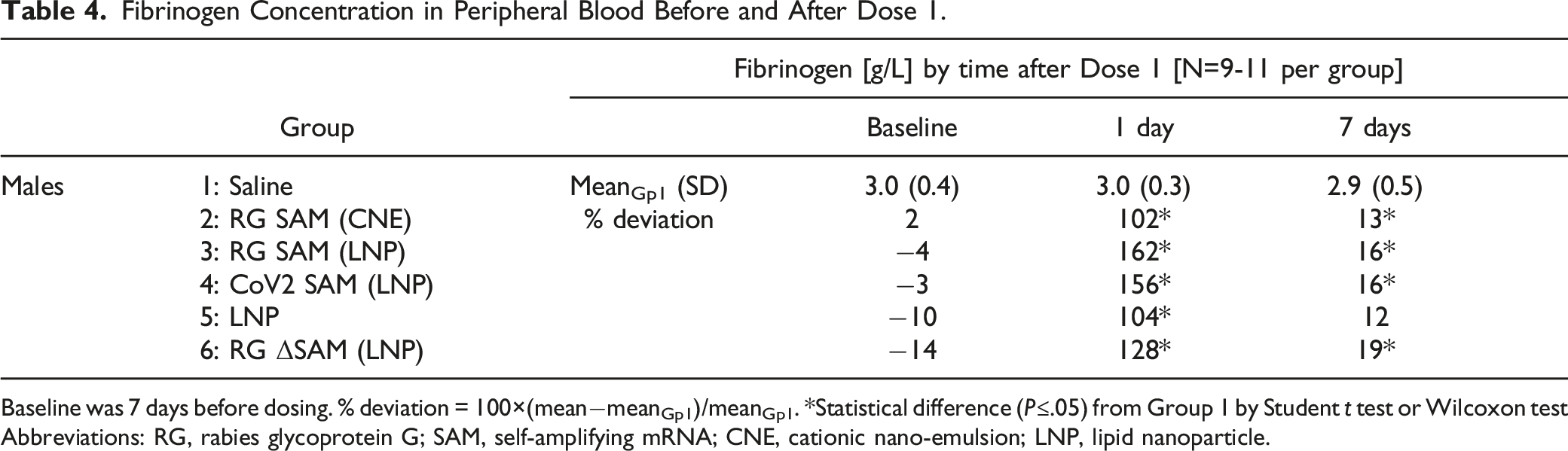

Baseline was 7 days before dosing. % deviation = 100×(mean−meanGp1)/meanGp1. *Statistical difference (P≤.05) from Group 1 by Student t test or Wilcoxon test
Abbreviations: RG, rabies glycoprotein G; SAM, self-amplifying mRNA; CNE, cationic nano-emulsion; LNP, lipid nanoparticle.
Cytokine Concentrations in Peripheral Blood After Dose 1.
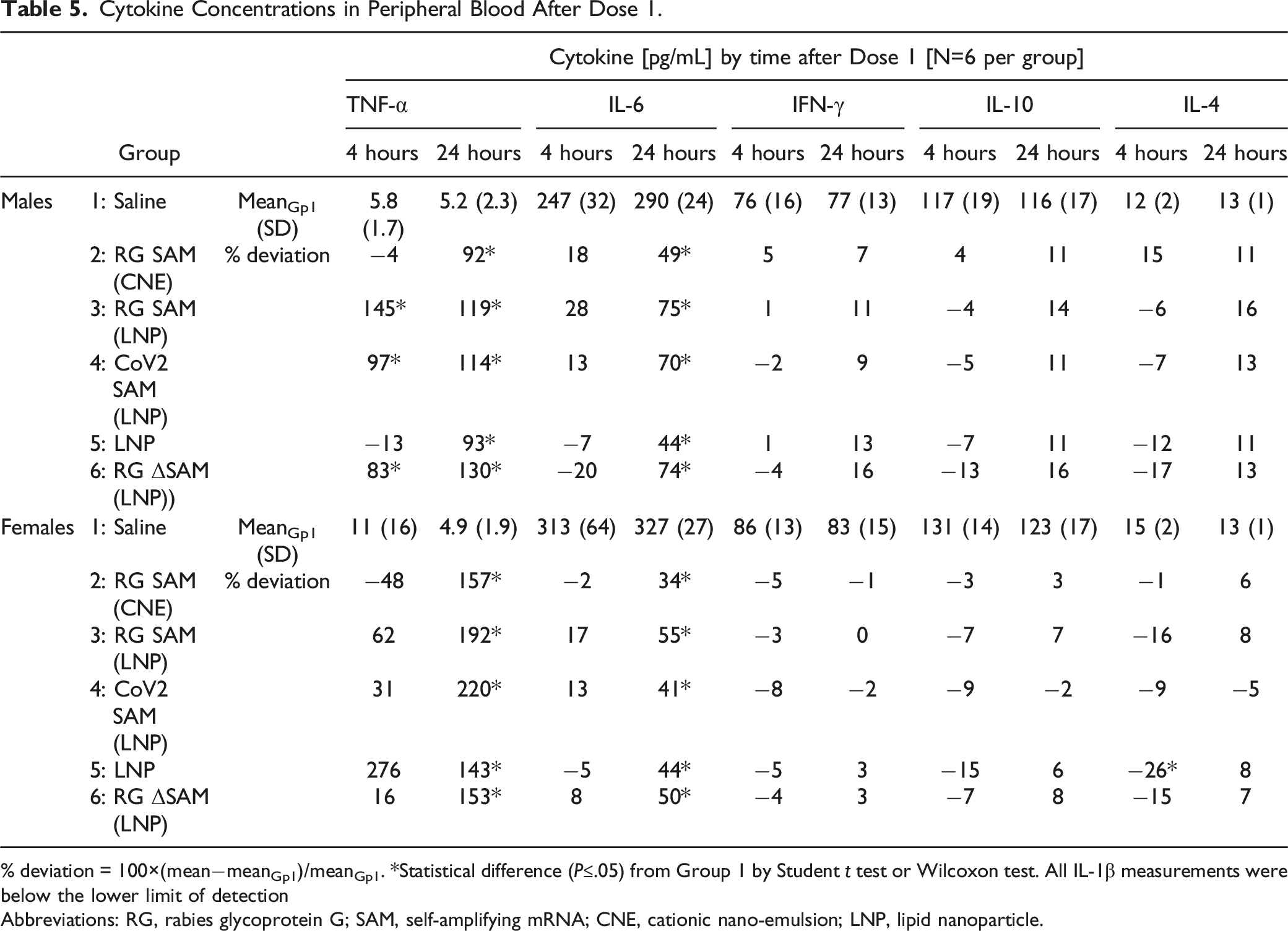
% deviation = 100×(mean−meanGp1)/meanGp1. *Statistical difference (P≤.05) from Group 1 by Student t test or Wilcoxon test. All IL-1β measurements were below the lower limit of detection
Abbreviations: RG, rabies glycoprotein G; SAM, self-amplifying mRNA; CNE, cationic nano-emulsion; LNP, lipid nanoparticle.
Greater concentrations of urea and creatinine compared with the saline control group were supportive of slight subclinical dehydration (Table 2).
Only amplification-competent SAM-based vaccines (i.e., Groups 2, 3, and 4) were associated with greater activities of ALT and AST, and only in female rats. No corresponding changes were observed in creatine kinase activity to suggest a muscular rather than a hepatic origin to the greater ALT/AST activities. However, as stated above, no signs of toxicity were detected microscopically in the liver.
Pathology
Based on the selected tissues analyzed microscopically, no SAM-based vaccine or the LNP control was associated with systemic toxicity (Dose 2 injection site, spleen, liver, the (iliac) lymph node draining the second dose injection site, and gross lesions). However, all SAM-based vaccines and the LNP control were associated with greater weights in the iliac lymph nodes draining the Dose 2 injection sites, 3 days after dosing, but not at 14 days after dosing, consistent with a transient inflammatory response to the vaccines and LNP alone, that was in proximity to the injection site. Other organ weight changes were not accompanied by corresponding microscopic findings and/or were considered variations in tissue trimming and therefore, were not considered treatment related.
The transient inflammatory response to SAM-based vaccines and the LNP control was supported by the histochemical analysis of injection-site and draining-lymph-node sections at 3 days and 14 days after the second dose (Figure 2). At the Dose 2 injection site on the right limb, minimal to moderate inflammation was generally observed in subcutaneous tissue and in the muscle. Minimal degeneration/necrosis of muscle myofibers was observed in 4/12 animals in the RG SAM (LNP) groups and moderate degeneration/necrosis was observed in RG SAM (CNE). In the draining (iliac) lymph node, minimal to mild cellularity was generally observed. Histochemical analysis of the right-side injection-site and draining-lymph-node (dLN) sections. (A) Representative injection-site sections from animals injected with saline (Group 1; upper images) or CoV2 SAM (LNP) (Group 4; lower images) 3 days after the second dose. Right-hand images represent high magnification images of the left-hand images demarcated by the rectangular outlines. Scale bars represent 1000 μm and 50 μm for the left and right images, respectively. Histograms describing the histology findings (B) 3 days and (C) 14 days after the second dose, by group and by sex. The severity of each finding was graded and depicted by differently shaded bars. At 14 days after the second dose, no muscle fiber necrosis or degeneration was detected. Although, immune-cell infiltrates were occasionally detected in the saline group (A; upper images), the frequency and extent were insufficient to be graded as minimal.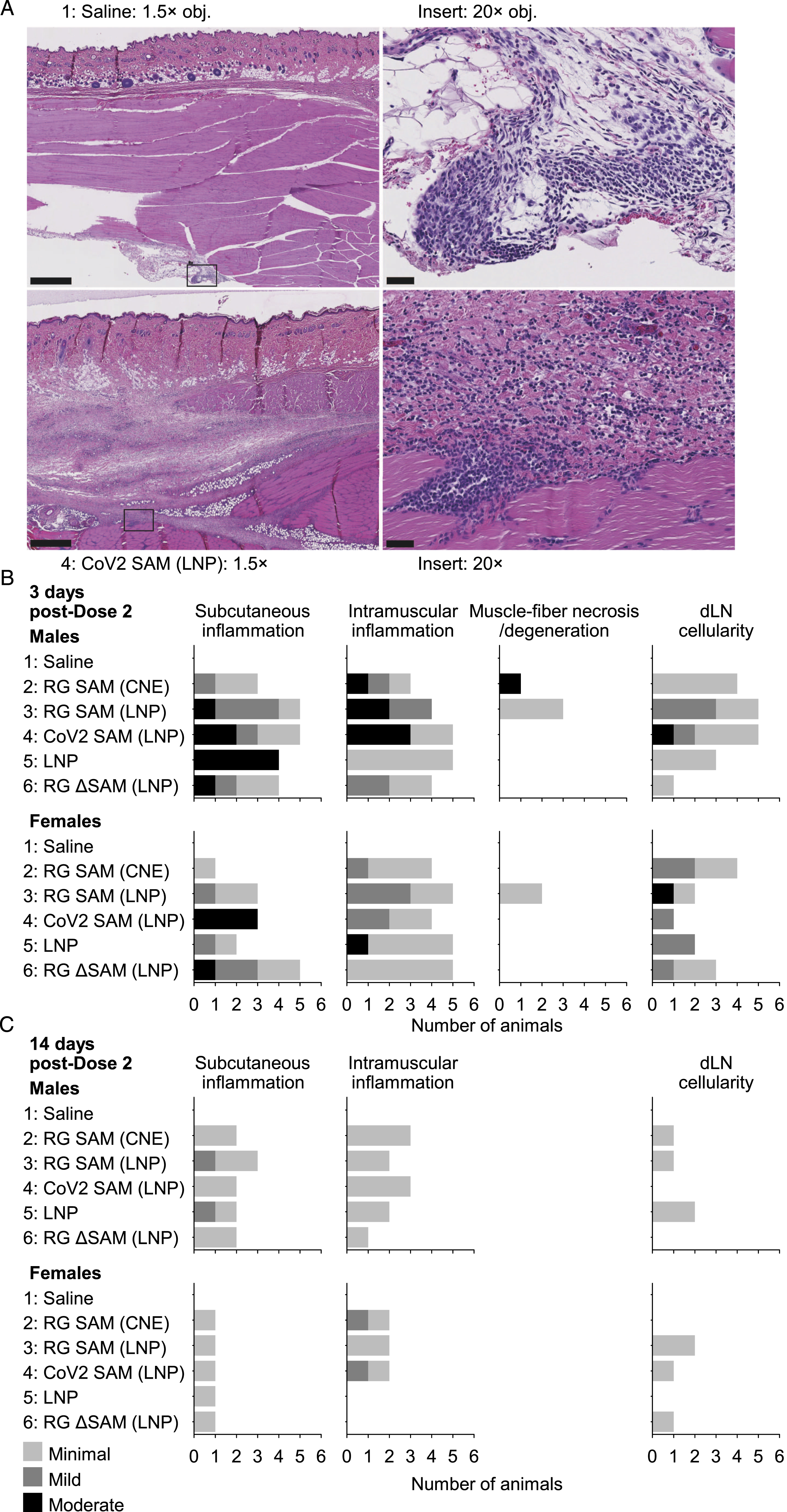
At 14 days after the second dose, signs of inflammation were less frequent than at 3 days after the second dose (Figure 2). No signs of degeneration or necrosis were detected in the muscle in any animal evaluated, and subcutaneous inflammation or intramuscular inflammation was detected in no more than half of the animals in any of the vaccine groups or in the LNP control group and was generally minimal. Only minimal signs of cellularity in the draining lymph nodes were detected in no more than 2/6 animals per group. Therefore, there was no difference in anatomic pathology between groups that were administered either the functional SAM vector or the amplification-defective SAM vector.
Peripheral-Blood Gene Expression (Nanostring Analysis)
The transient up regulation of innate-immunity genes was detected in peripheral blood in response to all SAM-based vaccines and the LNP control (Figure 3A-C). However, this transient up-regulation was observed earlier in the LNP-containing vaccines than in the CNE containing vaccine (Group 2). At 4 hours after the first dose, in the groups that were dosed with vaccines plus LNP (Groups 3, 4, and 6), 12 or 14 genes per group were upregulated from baseline (7 days before dosing), whereas no gene was differentially expressed in the saline control group and RG SAM (CNE) group (Group 2), and only 3 genes were upregulated in the LNP control group (Group 5). At 1 day after the first dose in the groups that were dosed with vaccines or LNP alone, 35 to 40 genes per group were upregulated and 6 to 18 genes per group were down regulated from baseline, whereas no gene was differentially expressed in the saline control group. Overall, 37 of those differentially expressed genes were common to all five groups. At 7 days after the first dose, no more than 6 genes per group were differentially expressed. At 4 hours and 1 day after the first dose, type I interferon signaling and type II interferon signaling were the two pathways most associated with the differentially expressed genes, for those groups with ≥12 up-regulated genes. Characterization of the expression of innate-immune RNA and the biodistribution of RNA after dosing. (A) Bar graph showing the up- and down-regulated genes in peripheral blood at 4 hours, 24 hours and 7 days after the first dose (with respect to baseline values, 7 days prior to the first dose), as measured by Nanostring gene expression. (B) Venn diagram distribution by group of the up- and down-regulated genes at 24 hours after the first dose. (C) Heatmap description of up- and down-regulated gene pathways (enrichment analysis) by group and with respect to significance (P-values) of the t statistics. The P-values were determined from within-group comparisons by timepoint with respect to baseline values (i.e. values from 7 days before the first dose); the heatmap presents reciprocal P-values on a log10 scale, where the positive (red) and negative (blue) signs of the derived values indicate overall up- and down-regulation of the pathways, respectively. Also, the pathways shown are only those for which the P-value ≤10−5 in at least one group. (D) RT-qPCR detection of vaccine-derived-RNA at the injection site (Inj. Site), at the draining lymph node (dLN), in peripheral blood, and in the liver 3 days (magenta symbols) and 14 days (cyan symbols) after the second dose in Group 4 (CoV2 SAM [LNP]). Two regions of the vaccine-derived RNA were targeted; (left graph) nsP1, representing genomic RNA, and (right graph) SARS-CoV-2 S, representing the antigen-coding RNA sequence. Each symbol represents a single animal (female; closed circle; and male open diamond), and the bar represents the geometric mean. The dotted lines represent the lower limit of quantification (LLOQ) and the limit of detection (LOD). Any undetected signal was assigned the value of 1. Note, one subgroup provided samples at 3 days after the second dose and a different subgroup provided samples 14 days after the second dose.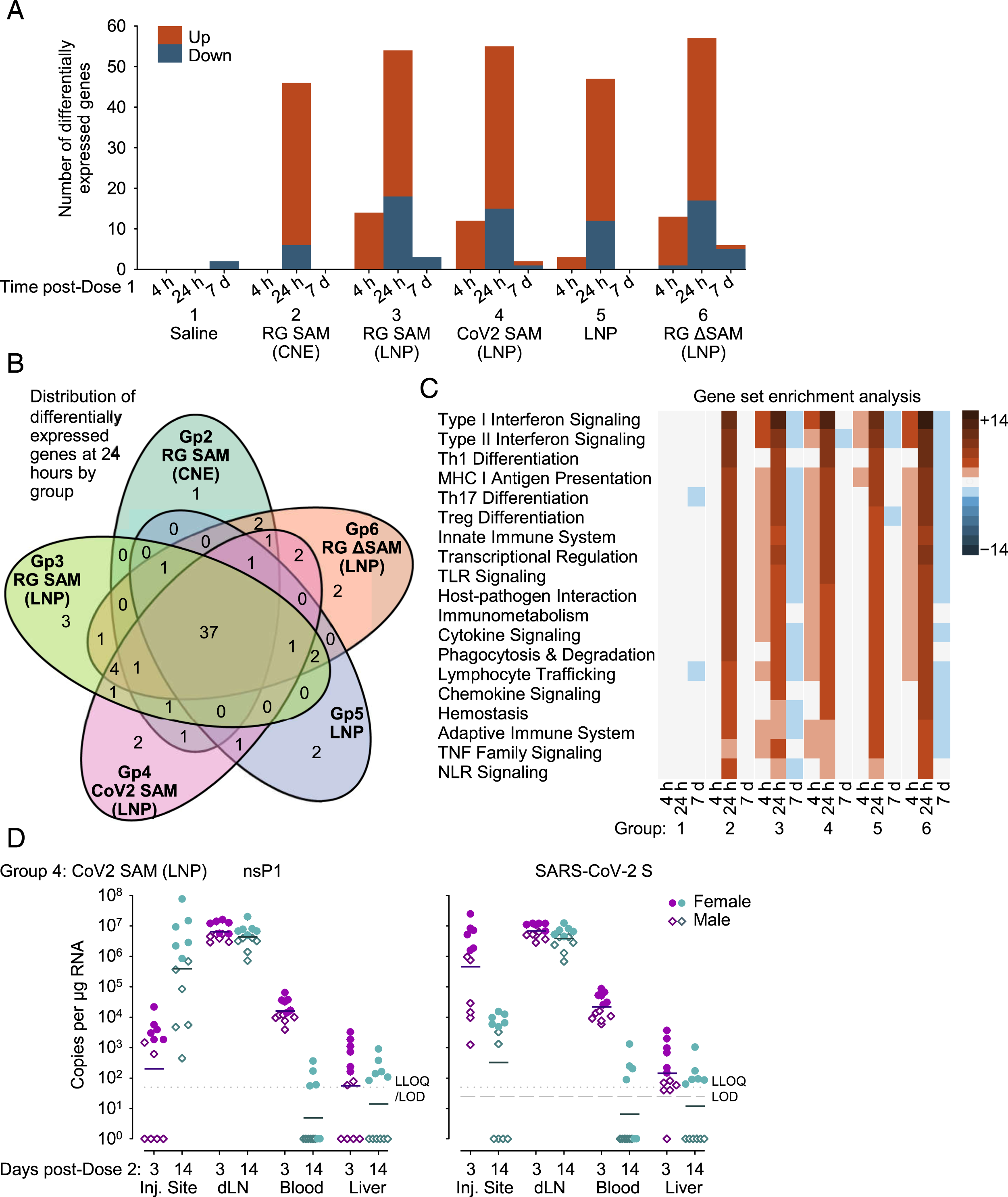
Limited Biodistribution of CoV2 SAM (LNP) RNA and Comparison of Two Distinct Primer Sets After the Second Dose
In a RT-qPCR assessment using distinct primers and samples from recipients of the CoV-2 SAM [LNP] vaccine, RNA sequences for nsP1 (the SAM construct backbone) and SARS-CoV-2 S (the antigen) were detected at the injection site, in the draining (iliac) lymph node, in the blood, and in the liver, 3 and 14 days after the second dose. Greater quantities of RNA were detected in females than in males. The greatest quantity of RNA was detected in the draining lymph nodes, followed by the injection site and blood. In the liver, nsP1 RNA and SARS-CoV-2 S RNA were not detected in all animals and the geometric mean quantities were near or below the lower limit of quantification (LLoQ).
Vaccine RNA appeared to persist in the draining lymph node up to 14 days after the second dose, but appeared to dissipate from the blood and liver by Day 14 after the second dose. At 14 days after the second dose in the draining lymph node, the geometric mean quantities of nsP1 RNA and SARS-CoV-2 S RNA were similar to those at 3 days after the second dose; whereas, in the blood and liver, the geometric mean quantities nsP1 RNA and SARS-CoV-2 S RNA were below the LLoQ. Moreover, at 14 after the second dose in the blood and liver, nsP1 RNA and SARS-CoV-2 S RNA was only detected in female animals (blood, 4/6; and liver 6/6), but not male animals (0/6).
The injection site was the only tissue that demonstrated a difference in the detection of nsP1 RNA and SARS-CoV-2 S RNA. At the injection site, the geometric mean quantity of nsP1 RNA was higher at 14 days than at 3 days after the second dose, whereas geometric mean quantity SARS-CoV-2 mRNA was lower at 14 days than at 3 days after the second dose; and the geometric mean quantity of nsP1 RNA at 14 days was similar to SARS-CoV-2 RNA at 3 days after the second dose. Overall, in the four tissues evaluated, there was no consistent evidence that the quantity of SARS-CoV-2 S RNA was generally lower than nsP1 RNA as a consequence of preferential 3’ to 5’ degradation of the SAM vector.
Discussion
This study compared the preclinical safety signals of different SAM-based vaccines in rats based on the standard repeat-dose toxicology design with additional analyses of cytokine levels and RNA expression in peripheral-blood, and a limited analysis of the biodistribution of SARS CoV-2 SAM (LNP) RNA. Five test items were used to assess (i) the nonclinical safety of two delivery vehicles (LNP vs CNE); (ii) the nature of the coded antigen (SARS-CoV-2 S vs RG); and (iii) the biodistribution of two regions of vaccine RNA that were potentially subject to different rates of amplification and degradation. Overall, no safety concerns were identified for any of the SAM-based vaccines, in agreement with previous studies.14,15 The changes that were observed were generally common to all test items and associated with a mild transient elevation in temperature within 1 day after dosing accompanied by a mild transient inflammation localized to the injection site and draining lymph node. Two weeks after the second dose, most signs of localized inflammation had resolved. However, vaccine RNA in the draining lymph node was more prominent than at the injection site and remained at a relatively high level two weeks after the second dose. In the blood, and liver, although vaccine-derived RNA was detected in some animals, the level was relatively low and at two weeks after the second dose was only detected in female animals.
Both the delivery vehicle and the vaccine-derived RNA were associated with transient inflammation, which was evaluated in detail after the first dose. LNP-containing vaccines appeared to activate innate-immune activity earlier than the CNE containing vaccine, manifested by more up-regulated innate-immune genes and generally higher concentrations of TNF-α at 4 hours. Also, LNP-containing vaccines appeared to be more associated with an elevation in body temperature than the CNE-containing vaccine. RNA in the LNP-containing vaccines appeared to contribute to this innate-immune activity because for LNP-containing vaccines, including the amplification-defective RNA vaccine vs LNP alone, more innate-immune genes were upregulated at 4 hours, and elevated temperatures were more frequent among the male-female subgroups. Hence, this early induction of innate-immune activity appeared primarily independent of RNA amplification and concurred with a previous study in mice which showed that amplification-defective self-amplifying mRNA induced an IFN-β reporter gene after intradermal electroporation to a similar degree to that induced by the amplification-competent vector. 26
By 24 hours after the first dose, the degree of activation and the nature of the transient inflammation was similar between all test-item groups including LNP alone. The nature of the inflammation coincided with similar serum concentrations of IL-6 and similar numbers of up-regulated genes primarily associated the type I and type II interferon signaling pathways. Hence the delivery vehicles (LNP and CNE) rather than vaccine RNA11,12 appeared to be the main factor behind the prominent and differential stimulation of interferon signaling pathways. Nevertheless, the activation was relatively mild, in that these pathways did not lead to the appearance of IFN-γ in the serum.
The capacity for RNA amplification in the vaccines appeared to be associated with a transient increase in ALT and AST and in the ALT/AST ratio in female rats only, and these findings were aligned with previous toxicology studies on RG SAM (CNE) and CoV2 SAM (LNP).14,15 As previously reported, the tissue source of the elevations in ALT and AST in female rats remains unclear, although vaccine RNA quantities in the muscle injection-site and liver were consistently higher in female than male animals. Muscle damage appeared not to be a factor because there was no elevation in CK or consistent histochemical signs of fiber degeneration or necrosis. Although there were no histochemical signs of liver toxicity, a hepatic origin of the ALT/AST signal in females could not be ruled out given that vaccine RNA was detected in the liver.
In the limited biodistribution evaluation after the second dose, there was no consistent evidence for a difference in RNA quantities from two genomic regions separated by ∼10.7 kb. It was reasoned that the RNA vector may be subjected to two countervailing processes which could affect the detection of these different regions; (i) a preferential 3’ to 5’ RNA degradation of the RNA vector (i.e. removing SARS-CoV-2 S before nsP1 sequences) and (ii) a preferential self-amplification, of subgenomic transcripts containing the SARS-CoV-2 S sequence. Although no remarkable differences were observed in the draining lymph node, blood and liver, inter animal variation (notably, between 3 days and 14 days after dosing) could not be ruled out as an explanation for the differences observed at the injection site.
In conclusion, the SAM vaccines administered as two doses 2-weeks apart had acceptable safety profiles in rats, in agreement with previous studies.14,15 The transient increase in ALT/AST ratio that occurred in female rats only in the absence of muscle and liver damage, has also been observed previously,14,15 was dependent on RNA amplification and appeared related to the greater quantities of vaccine RNA in the muscle and livers of female rats vs male rats. The RNA vector and delivery-vehicle components of the SAM vaccine, but not the nature of the antigen-coding sequence or the requirement for RNA amplification, affected aspects of the stimulation of innate-immune activity, which was consistent with the transient activation of type I and type II interferon signaling. The delivery vehicle, LNP, differed from CNE in that vaccine RNA in CNE compositions appeared to independently stimulate innate immune activity at 4 hours after vaccination. Therefore, further studies should assess whether the subtle differences in innate-immune activity related to the delivery vehicles have tangible effects on the safety and efficacy of the SAM vaccines in the clinic.
Footnotes
Author’s Note
All authors had full access to the data and gave final approval before it was submitted by the corresponding author.
Acknowledgments
The authors thank Jessica Firestone for providing the test materials (GSK, Rockville, USA), Shreshta Phogat for preparation of the RG ΔSAMRNA construct. Matthew Morgan (MG Science Communications, Belgium) provided scientific writing services and Ulrike Krause and Pascal Cadot (GSK, Belgium) provided editorial assistance and manuscript coordination.
Author Contributions
AHS , CB, GM, L-AR, MM were involved in the conception/design of the study. AHS, CB, DAD, and GM participated in the acquisition and/or analysis of the data. AHS and CB supervised the design and analysis of experiments. CG and MAR participated in review of the pathology data. NBA and MM conducted the biodistribution of nsP1 and Cov2 sequence, respectively. AA conducted the Nanostring analysis. All authors participated in the interpretation of data and were involved in drafting the manuscript or revising it critically for important intellectual content. DAD, GM, MAR, L-AR and AHS are, or were at the time of the study, employees of the GSK group of companies. GM and AHS reported ownership of GSK stocks. AA, CB, CG, MM, and NBA are, or were at the time of the study, employees of Charles River Laboratories, a Contract Research Organization contracted by GSK in the undertaking of the experiments contributing to this study. These have been stated in the manuscript.
Declaration of Conflicts Interest
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: DAD, GM, MAR, L-AR and AHS are, or were at the time of the study, employees of the GSK group of companies. GM and AHS reported ownership of GSK stocks. AA, CB, CG, MM, and NBA are, or were at the time of the study, employees of Charles River Laboratories, a Contract Research Organization contracted by GSK in the undertaking of the experiments contributing to this study.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was sponsored by GlaxoSmithKline Biologicals SA, which was involved in all stages of the study conduct and analysis. The costs associated with the development and publishing of the manuscript, including scientific writing assistance were provided by GlaxoSmithKline Biologicals SA.
