Abstract
Cathine is the stable form of cathinone, the major active compound found in khat (
Introduction
Cathine (d-norpseudoephedrine (NPE)) is one of the major constituents found in the
Over the past decades, cathine has received little experimental attention. Cathine acts as a central stimulant (indirect sympathomimetic) and an inhibitor or monoamine oxidase. 7 As compared to cathinone, cathine has a slower onset of action and may not be as lipophilic to enter the central nervous system, which to some extent explains the slower metabolism of cathine. 10 Oral administration of cathine (6.5 mg/kg) may affect adrenocortical function and was found to produce rapid synchronized cell death in human leukaemia cell lines and peripheral blood leucocytes. 10 With its milder psychostimulatory effects, a higher dose of cathine is needed to exert its effects, and this causes severe adverse systemic effects and the inhibition of noradrenaline uptake.11,12 In recent years, khat use was reported in England, Wales, Rome, Amsterdam, Canada, Israel, Australia, New Zealand and the United States. 13 Khat is chewed by individuals idling on streets in Europe accompanied sometimes by alcoholic beverages and other drugs. 3
It was found that a majority of khat users used more than one other psychoactive substances in which cigarettes were mainly used to maximize the stimulation power of khat with a combination of alcohol to break the aftereffect. 14 Polysubstance users are also found to be using synthetic cathinone 15 and cathine concurrently with alcohol leading to haemorrhage and intoxication. 16 Moreover, khat use have been found to hinder antipsychotic 17 and tuberculosis 18 medication effects on patients who are khat users. With the widespread global use of khat together with the co-administration with clinically used drugs, the detrimental effects of khat-drug interactions are pushed towards the centre of the attention. According to the National Drug Intelligence Center, a component of the U.S. Department of Justice, the potency of harvested khat fades after 48hours as cathinone degrades into cathine (https://www.justice.gov/archive/ndic/pubs31/31482/index.htm). Thus, the effects of cathine on human drug metabolizing enzymes and the possible herb-drug interactions caused by cathine warrant further exploration.
Cytochrome P450 (CYP) are a crucial family of enzymes for producing cholesterol, steroids, prostacyclins and thromboxane A2 besides playing an essential role in detoxification of foreign substances and drugs.
19
CYPs are predominantly expressed in the liver, besides also occurring in the small intestines, lungs, placenta and kidneys. In mammals, CYP family 1, 2, 3 and 4 are involved in detoxification, steroid and eicosanoids metabolism.
20
In humans, CYP family 1, 2, 3 and 4 and to a lesser extent CYP family 5, 8, 19, 21 and 26 are involved in xenobiotic metabolism.
20
The CYP enzymes are predisposed to inhibition or induction by xenobiotics including herbal medications that comprise mixtures of phytochemicals.
21
Phytochemicals such as resveratrol and quercetin have been reported to significantly inhibit CYP3A4 which translates to reduced clearance and subsequent toxicity from other CYP3A4 substrates in humans.22,23 There are few well characterized examples including resveratrol and quercetin which were classified as clinically significant perpetrators as numerous herbal preparations interact with CYPs
Investigations into the inhibitory effects of cathine on CYPs are generally lacking in the literature as most studies about khat-drug interactions have focused on cathinone, which is deemed to be the major active compound in khat. Our earlier study found that khat ethanol extract (KEE) inhibited CYP2C9, CYP2D6 and CYP3A4 significantly, but cathinone showed negligible inhibition on these CYP isoforms
Materials and Methods
Chemicals and Reagents
Cathine or d-Cathine.HCl (d-Norpseudoephedrine.hydrochloride; (
Determination of CYPs Activity Using Vivid P450 Assay Kits and Time Curve
Fluorescence readings (RFU) produced by blue (3-cyano-7-hydroxycoumarin), green (fluorescein) and cyan (7-hydroxy-4-trifluoromethylcoumarin) standards against a range of concentrations of the respective standard was plotted to derive the standard curves. The standard curve equation was used in succeeding assays to enumerate the fluorescent metabolites produced. Time curves were plotted afterwards to define the incubation time for single CYP assays namely 60 minutes for CYP1A2, CYP2A6, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP2J2, CYP3A4 and 120 minutes for CYP3A5, respectively, as determined previously.25,26
The inhibitory effects of cathine on human CYP enzymes activities were determined using Vivid® CYP450 Screening Kits 27 for all CYP isoforms including Vivid® EOMCC CYP1A2 Blue, Vivid® CC CYP2A6 Blue, Vivid® BOMCC CYP2B6 Blue, Vivid® DBOMF CYP2C8 Green, Vivid® BOMCC CYP2C9 BLUE Vivid® EOMCC CYP2C19 Blue, Vivid® EOMCC CYP2D6 BLUE Vivid® EOMCC CYP2E1 Blue, Vivid® MOBFC CYP2J2 Cyan Vivid® BOMCC CYP3A4 BLUE and Vivid® BOMCC CYP3A5 Blue, according to manufacturer’s instructions. Using the 96-well black plates, in each well, 40 μL of respective reaction buffers (Buffer I for CYP1A2, CYP2B6, CYP2D6, CYP3A4, CYP3A5; Buffer II for CYP2A6, CYP2C8, CYP2C9, CYP2C19, CYP2J2; Buffer III for CYP2E1) was added with 50 μL of master premix (including CYP450 BACULOSOMES® Plus, human CYP reductase, potassium phosphate buffer, NADPH regeneration system containing glucose-6-phosphate buffer (333 mM) and 0.3 μ/mL glucose-6-phosphate dehydrogenase in 100 mM potassium phosphate at pH 8.0) for incubation with shaking at room temperature for 30 minutes. 10 μL per well of a mixture of respective substrates and NADP+ were added to kick start the reaction. Total volume per well was 100 μL in the 96-well plate.
The CYP enzyme specific substrates added were as follows: Vivid® BOMCC (7-benzyloxymethyloxy-3-cyanocoumarin) for CYP2B6, CYP2C9, CYP3A4 and CYP3A5; Vivid® CC (3-cyanocoumarin) for CYP2A6; Vivid® DBOMF (dibenzylmethylfluorescein) for CYP2C8; Vivid® EOMCC (ethoxymethyloxy-3-cyanocoumarin) for CYP1A2, CYP2C19, CYP2D6 and CYP2E1; Vivid® MOBFC (7-p-methoxy-benzyloxy-4-trifluorocoumarin) for CYP2J2; and 0.03 mM NADP+. The mixture was shaken at room temperature, 60 minutes for CYP1A2, CYP2A6, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP2J2, CYP3A4 and 120 minutes for CYP3A5. 50 μL of 0.5 M tris-base solution was added lastly to halt the reaction. The enzyme activities were measured by using the Varioskan® Fluorescence Spectrophotometer (Thermo Fisher Scientific®, Waltham, MA, USA) at excitation/emission wavelengths of 415/460 nm (blue – CYP1A2, CYP2A6, CYP2B6, CYP2C9, CYP2C19, CYP2E1, CYP2D6, CYP3A4 and CYP3A5), 415/520 nm (cyan – CYP2J2) and 490/520 nm (green – CYP2C8).
Reversible Inhibition
Cathine was dissolved in water to attain the 2.5 mM of cathine stock. The assay conditions were as described above. The assay conditions were as described above, with the exception that 40 μL of buffer was substituted using 40 μL of cathine with concentrations ranging from 0 to 1000 μM, which was obtained following two times serial dilution of the cathine stock. Cathine inhibited CYP2A6 and CYP3A4 according to their IC50 values which were less than 100 μM, and then their Ki values were further assessed. Various concentrations of cathine (0, 31.25, 62.5, 125, 250, 500, 1000 μM) were incubated with Vivid® Fluorogenic Probe Substrates CC (5, 10, 20, 40 μM) for CYP2A6 and BOMCC (5, 10, 20, 40 μM) for CYP3A4.
Determination of Time-Dependent Inhibition
In mechanism-based or time-dependent inhibition, the master premix was added with NADP+ to produce NADPH during the pre-incubation. The mixture (containing CYP450 BACULOSOMES® Plus, human CYP reductase, potassium phosphate buffer, NADPH regeneration system and NADP+) was incubated for 30 minutes with shaking under similar range of concentrations of cathine as mentioned above for IC50 determination. Substrates were subsequently added to initiate the reaction, and fluorescence readings were taken after 60 minutes for CYP1A2, CYP2A6, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1 and CYP2J2, CYP3A4 and 120 minutes for CYP3A5 incubation. The IC50 shift was derived by calculating ratios of IC50 obtained from pre-incubation with and without NADPH. IC50 shift ratio >2 implies irreversible time-dependent inhibition. 28
Data Analysis
To determine IC50 values, the remaining enzyme activity of each cathine concentration was divided by the solvent control well (without cathine but replaced by water) and multiplied the value with 100% to obtain the percent control activity (%) for each cathine concentration. The resulting percent control activity (%) were plotted against different concentrations of cathine to attain the half maximal inhibitory concentrations (IC50) curve. IC50 values of reversible and irreversible inhibition were determined by non-linear regression analysis using GraphPad Prism 9 for Windows (GraphPad Software, San Diego, CA). Data for inhibition constant (Ki) analysis and mode of inhibition were evaluated using Excel spreadsheet (Microsoft, USA) and Lineweaver-Burk plots. The secondary plots of cathine concentrations against slopes of Lineweaver-Burk plots were plotted to determine the Ki values. All assays were carried out in triplicate and stated as mean ± SD.
Molecular Docking
The crystal structures of human cytochrome P450 CYP2A6 (PDB code: 2FDV) and CYP3A4 (PDB code: 4D75) were retrieved from Protein Data Bank. 29 Cathine conformations were generated using Open Babel GUI v2.3.1. 30 The protein was prepared using AutoDock Tools 1.5.6 while AutoDock 4.2 software 31 was used to carry out molecular docking. AutoGrid was used to create the grid box around co-crystallized ligand covering the active site of CYPs, which included the haem group. The grid size for specifying the search space was set at 60 × 60 × 60 Å (for CYP3A4) and 70 × 70 × 70 Å (for CYP2A6) with a default grid point spacing of 0.375 Å with a total of 10 docking runs. Docking simulations were performed using the Lamarckian genetic algorithm with 25 × 105 energy evaluations and 27 000 iterations per run. The best configuration and possible ligand binding was then determined based on the binding score. After docking, analysis of the ligand interactions at the binding site was performed and visualized using the PyMOL Molecular Graphics system (Schrödinger, LLC, New York, NY, USA).
Results
Standard Curves and Time Curve
Standard curves were plotted using fluorescence readings (RFU) against standard concentrations, and the following standard equations and
Inhibition of Cathine on CYPs
Cathine significantly inhibited CYP2A6 and CYP3A4 with IC50 values of 80 and 90 μM as shown in non-linear graphs (Figure 1). Cathine showed no inhibition on CYP1A2, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP2J2 and CYP3A5 with IC50 values more than 100 μM. Inhibitory effects of cathine on (A) CYP1A2, (B) CYP2A6, (C) CYP2B6, (D) CYP2C8, (E) CYP2C9, (F) CYP2C19, (G) CYP2D6, (H) CYP2E1, (I) CYP2J2, (J) CYP3A4 and (K) CYP3A5. IC50 values were determined by non-linear regression analysis using GraphPad Prism version 9 for Windows (GraphPad Software, La Jolla California, USA). Each point represents mean ± SD (n = 3). CYP: Cytochrome P450, IC50: 50% inhibitory concentration.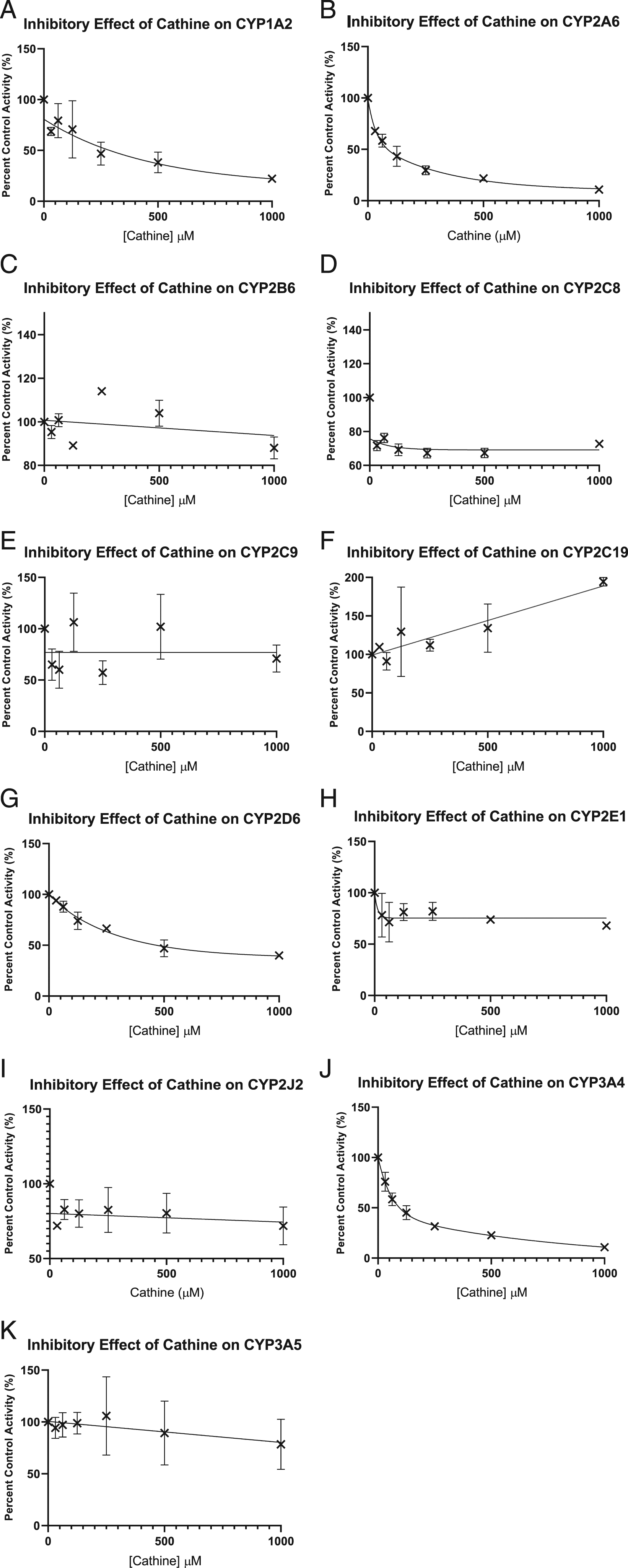
IC50 without/with NADPH in pre-incubation, and IC50 shift of CYP1A2, CYP2A6, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP2J2, CYP3A4 and CYP3A5 with cathine.
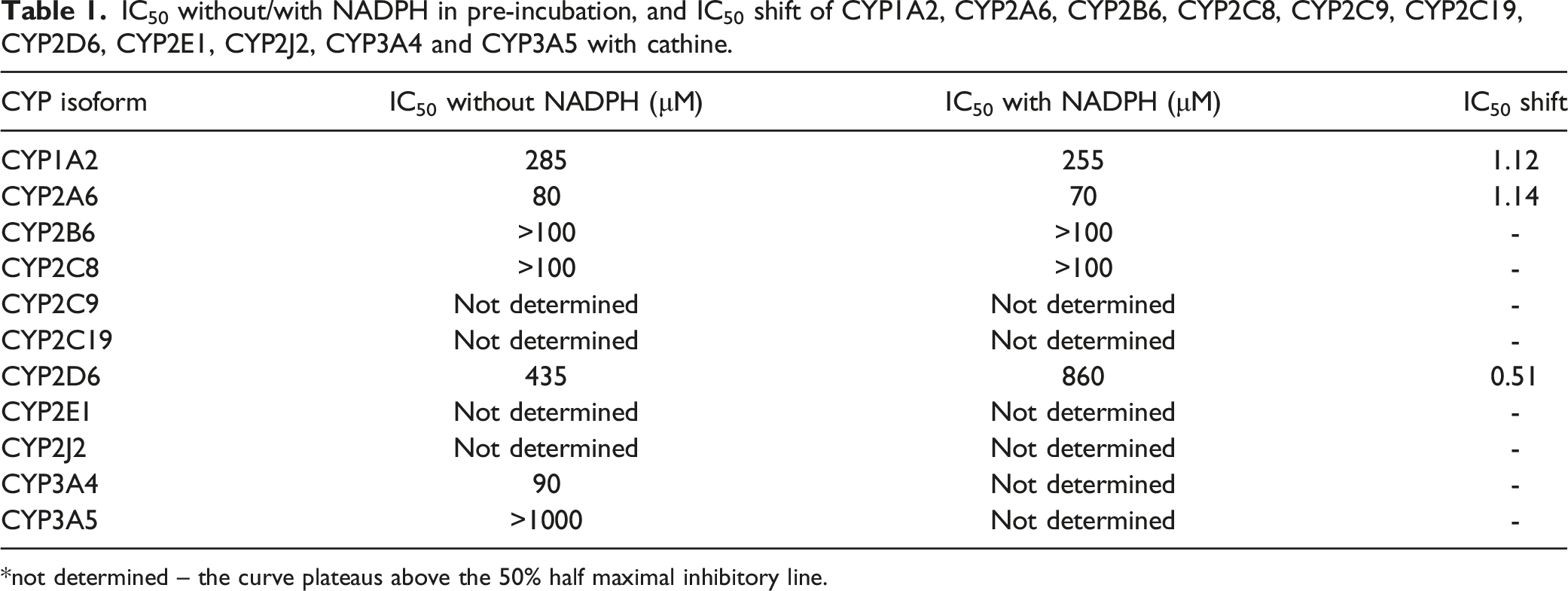
*not determined – the curve plateaus above the 50% half maximal inhibitory line.
Ki Analysis and Mode of Inhibition
The Ki values were obtained from secondary plots of each CYP isoforms. Cathine inhibited CYP2A6 and CYP3A4 via non-competitive or mixed mode with Ki of 63 μM, and non-competitive or mixed mode with Ki of 100 μM as shown in the Lineweaver-Burk plots (Figure 2). The Lineweaver-Burk plot or double reciprocal was plotted with inverse velocity (1/V) against the inverse of the substrate concentrations (1/[S]). Lineweaver-Burk plot of (A) inhibition of CYP2A6 by cathine and (B) inhibition of CYP3A4 by cathine at the indicated concentrations of cathine and substrate. The substrate concentration used were CC (5, 10, 20, 40 μM) for CYP2A6 and BOMCC (5, 10, 20, 40 μM) for CYP3A4. The secondary plots were plotted using slopes from Lineweaver-Burk plot against cathine concentrations which was used to derive the Ki, inhibition constant values. Each data point are triplicates that was represented by mean ± SD (n = 3).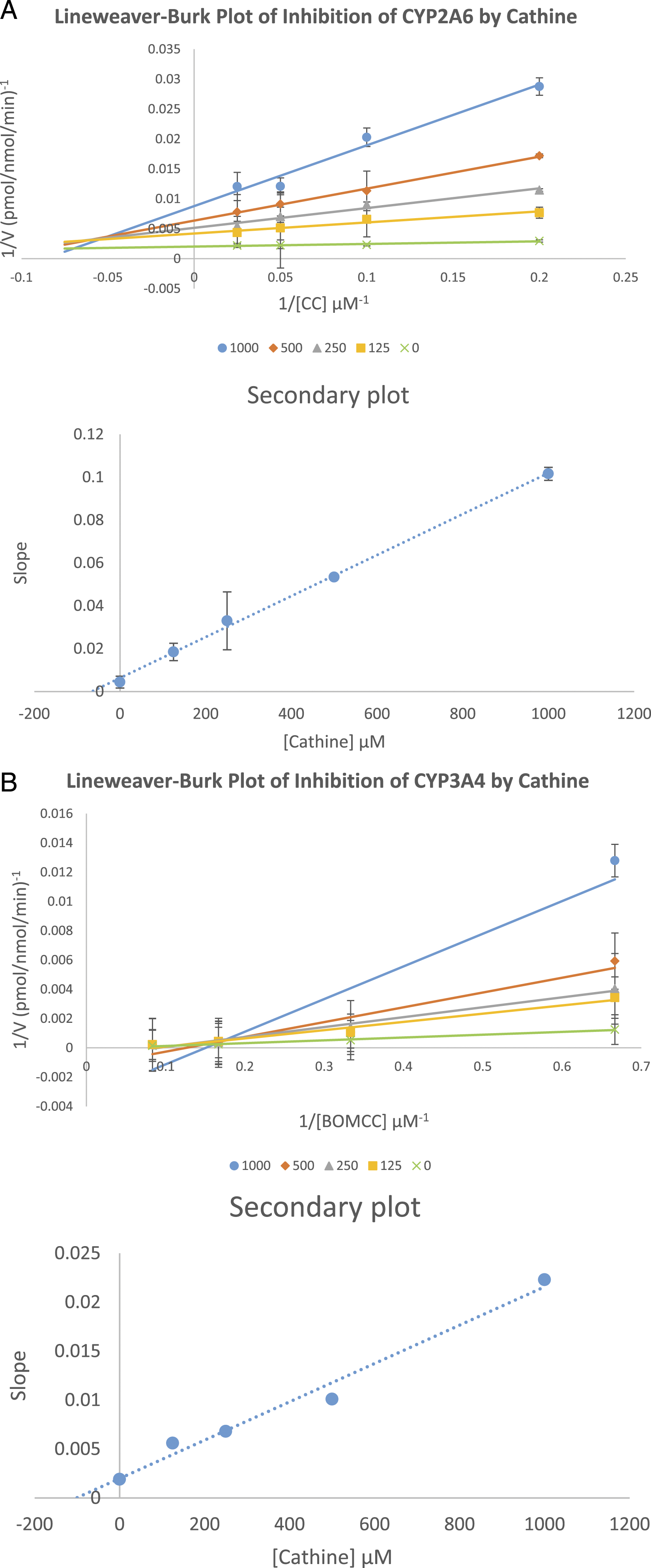
Docking of Cathine Into Human CYPs
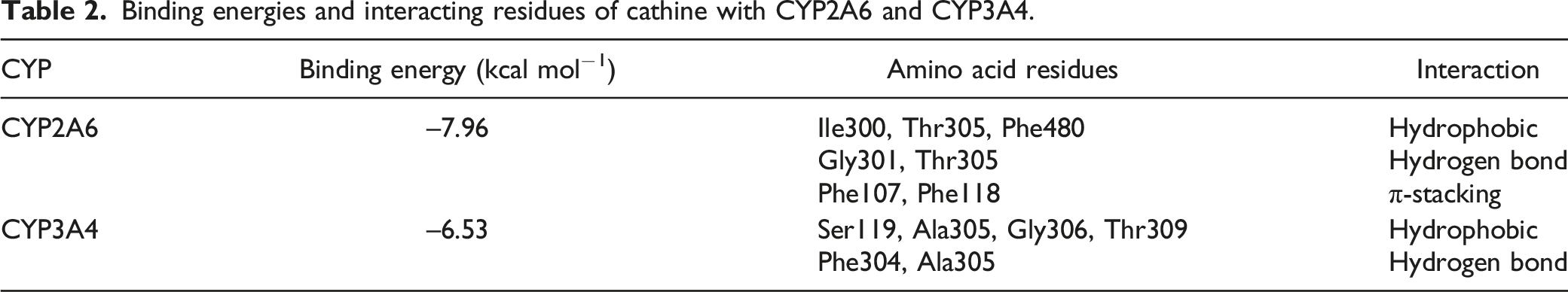
Cathine was docked into the active site of human CYP2A6 and CYP3A4. After inspection of the top-ranked poses in AutoDock, the potential binding sites were identified. The binding poses and key residues interacting with cathine are presented in Figure 3. The binding energies and the interacting residues for cathine with CYP2A6 and CYP3A4 are as shown in Table 2. Molecular docking demonstrating binding modes and key interactions of cathine (green) relative to the haem group (white) in the active sites of (A) CYP2A6 (PDB 2FDV) and (B) CYP3A4 (PDB 4D75). Hydrogen bonds are displayed as yellow dashed lines. Non-polar hydrogens have been removed for visual clarity. Binding energies and interacting residues of cathine with CYP2A6 and CYP3A4.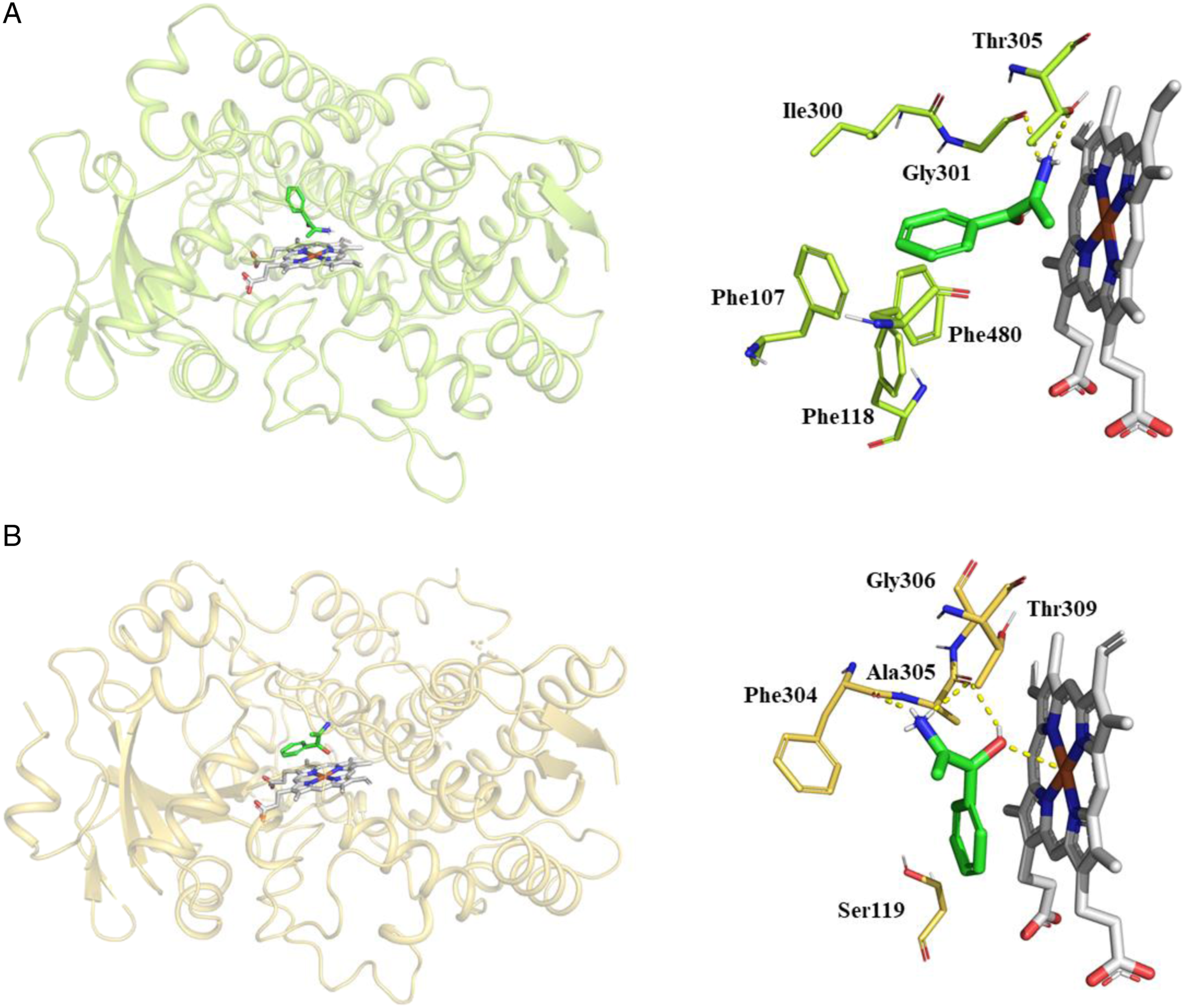
Discussion
Current knowledge on the mechanisms of action of khat and its active constituents with their short-term and long-term effects are not as extensive as other amphetamine-type stimulants.
32
Khat use remains a debatable topic whether it is an innocuous cultural tradition or drug of abuse.
32
Following the extensive use of khat with alcohol and clinically used drugs,
33
there is an urgent obligation to explore the possible inhibitory effects of the second major active compound in khat, cathine, on human drug metabolizing CYP enzymes.
With regard to herb-drug interaction via CYP inhibition by khat constituents, our previous study found that khat significantly inhibited CYP2D6, CYP2C9 and CYP3A4 while cathinone showed negligible inhibitory effects.
25
Besides, the subsequent studies in our laboratory found that khat ethanol extracts inhibited most of the major drug metabolizing CYPs including CYP2A6, CYP2B6, CYP2C8, CYP2C19, CYP2E1, CYP2J2 and CYP3A5 except CYP1A2.
26
On the other hand, our ongoing investigations demonstrated that the CYPs were inhibited by cathinone differently from that of khat extracts (unpublished data). A study by Bedada et al
35
using human volunteers on one week of daily khat use (400 g) showed khat significantly inhibited CYP2D6, marginally inhibited CYP3A4, CYP2C19 and CYP1A2.
35
An
Cathine showed no time-dependent (mechanism-based) inhibition on CYP1A2, CYP2A6, CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP2D6, CYP2E1, CYP2J2, CYP3A4 and CYP3A5 (Table 1). Cathine inhibited two CYPs out of the 11 CYP isoforms in this study (Table 2). Cathine inhibited CYP2A6 and CYP3A4 with IC50 values (Figure 2) of 80 and 90 μM, respectively. From the current
Based on our
To our best knowledge, pharmacokinetic studies on cathine are very limited. Khat chewing releases cathine and cathinone which was subsequently absorbed by the mucous membranes of the oral cavity and followed by the stomach lining. 44 The average content of cathine in 100 g of fresh khat leaves is 83-120 mg10. A 100-400 g of fresh khat leaves is consumed on a daily basis. 45 Hence, assuming that khat users who took 400 g of fresh khat leaves would have consumed about 332-480 mg of cathine, therefore, an average 70 kg adult with 5 L blood would have absorbed approximately 66.4 × 10−3 to 96 ×10−3 mg/ml of cathine or in other words, about 351.7-511.5 μM. Despite the low Ki values determined in our study that ranges from 63 to 100 μM, these values may be clinically relevant since khat users commonly take huge amount of khat on a daily basis. Toennes et al 45 reported that at a mean of 43.8 g of khat chewed, the mean ingested dose of cathine in milligrams was 32.4, while the absorbed proportion of cathine ranges from 16% to 84%. 45 The mucosa of the oral cavity is the first absorption segment, where the major fraction of the cathine (84 ± 6%) is absorbed. 45 Cathine could be detected in the urine up to 50-70 hours after ingestion which was about a day time more than cathinone at 22-26 hours. 46 In addition, a study found that cathine concentration in oral fluid was higher than that of cathinone because khat users did not chew fresh leaves and cathinone easily metabolized into cathine. 47 According to European Monitoring Centre for Drugs and Drug Addiction, hepatic first-pass metabolism of cathinone forms norephedrine and only 2% of cathinone is excreted in the urine (https://www.emcdda.europa.eu/publications/drug-profiles/khat). The elimination half-life of cathine (5.2 ± 3.4 hours) is longer than cathinone (1.5 ± 0.8 hours) (https://www.emcdda.europa.eu/publications/drug-profiles/khat). As compared to cathinone, cathine is less lipophilic and may also penetrate to the central nervous system. 48 Nevertheless, lipophilicity of cathine should also be considered during prescribing by healthcare professionals to khat users because lipophilicity impacts cellular uptake and ADMET (absorption, distribution, metabolism, excretion and toxicity). 49 High dosage accompanied by high lipophilicity compound is a detrimental combination. 50 Although less lipophilic than cathinone, the lipophilicity of cathine may still affect metabolic activity as lipophilic compounds showed greater affinity to metabolic enzymes. Khat compounds’ lipophilicity need to be known during drug administration as they may readily cross the blood brain barrier. 51
However, in the early 1945-1961, there were contradicting debates by researchers that cathine has low potency and the amount of cathine in khat is insufficient to exert symptoms shown after consumption versus cathine was the only khat alkaloid responsible to give pharmacological significance. 4 Dried khat leaves showed an increase in cathine amount from 0.172% to 0.192% (fresh khat) to 0.184-0.198%. 52 According to one case report, a male patient who suffered from haemorrhagic stroke with hypertension was found to have no chronic diseases history but instead daily khat chewing. 53 The patient’s urine samples showed positive for amphetamine-like substance while liquid chromatography-tandem mass spectrometry (LC-MS/MS) detected presence of cathine and cathinone. The serum concentration of cathine was 100-fold of cathinone, suggesting that the patient’s symptoms are most likely due to cathine. 53 The maximal plasma concentrations of cathine will be reached at 2.6 hours with a mean residence time (MRT) of 10.2 ± 2.6 hours, which was twice more than cathinone. 45 Cathine retain in the human body for a longer period of time as compared to cathinone and could be detected in the urine samples even after approximately ≥3 days after ingestion. 46 Since cathine is retained for a longer period of time within the human body, it may exert greater inhibitory effects on human CYPs and higher chances of causing khat-drug interactions as compared to cathinone. Nevertheless, the effects of cathine retaining within the body for a longer time have yet to be explored.
From the molecular docking analysis, the interactions between cathine and the CYPs consisted of hydrogen bonds, hydrophobic and to a lower extent, π-π stacking (Table 2). As our
We have performed docking using known inhibitors of CYP2A6 (PDB 2FDY) – aldrithiol and CYP3A4 (PDB 2V0M) – ketoconazole from Protein Data Bank to compare with cathine’s interactions. Cathine overlaps well with experimentally determined binding modes of aldrithiol and ketoconazole (at the same position above the haem group of CYP2A6 and CYP3A4 – as shown in Supplemental Figure 1). In CYP2A6, cathine overlaps very well with aldrithiol while ketoconazole is much bigger in size and has more interactions as compared to cathine in CYP3A4. In the active site of CYP2A6, aldrithiol formed hydrogen bonds with Asn297, π-stacking with Phe107, Phe111, and Phe118 and hydrophobic bonds with Phe209, Ile300, Gly301, Thr305 and Phe408. Ketoconazole formed π-stacking with Phe304, and strong ionic interaction with Fe2+ of the haem group of CYP3A4 and hydrophobic bonds with Phe57, Arg105, Ser119, Leu210, Phe241, Ile301, Ala305, Ala370, Arg372, Glu374, Gly481 and Leu482. Some similar interactions seen from our docking outcome such as (1) in CYP2A6, cathine and aldrithiol (4,4′-Dipyridyl disulfide) both have a series of π-stacking interactions with Phe107, Phe118 and Phe480 to stabilize them, hydrophobic interactions with Thr305 58 and (2) in CYP3A4, cathine and ketoconazole both have π-stacking with Phe304 formed. 59
The current study utilizes high-throughput fluorescence-based assays for detection of enzyme-drug interactions.26,60 In spite of that, a number of caveats exist in this
It is worth mentioning that
Conclusion
Cathine reversibly inhibited CYP2A6 and CYP3A4 activities. Docking results of CYP2A6 and CYP3A4 is aligned to the
Supplemental Material
Supplemental Material - Protein-Ligand Identification and In Vitro Inhibitory Effects of Cathine on 11 Major Human Drug Metabolizing Cytochrome P450s
Supplemental Material for Protein-Ligand Identification and
Footnotes
Author Contributions
Lim, Sharoen Yu Ming contributed to conception and design, contributed to acquisition, analysis, and interpretation, drafted manuscript, and critically revised manuscript; Loo, Jason Siau Ee contributed to conception and design, contributed to acquisition, analysis, and interpretation, and critically revised manuscript; Alshagga, Mustafa contributed to conception, contributed to interpretation, and critically revised manuscript; Alshawsh, Mohammed Abdullah contributed to conception, contributed to interpretation, and critically revised manuscript; Ong, Chin Eng contributed to conception and design and critically revised manuscript; Yan, Pan contributed to conception and design, contributed to acquisition, analysis, and interpretation, and critically revised manuscript. All authors gave final approval and agree to be accountable for all aspects of work ensuring integrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by University of Nottingham Pump-Priming, grant code: 54-ER-0-744019.
Supplemental Material
Supplemental Material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
