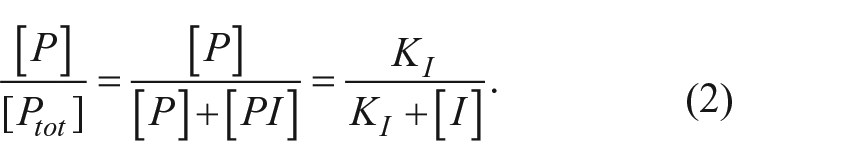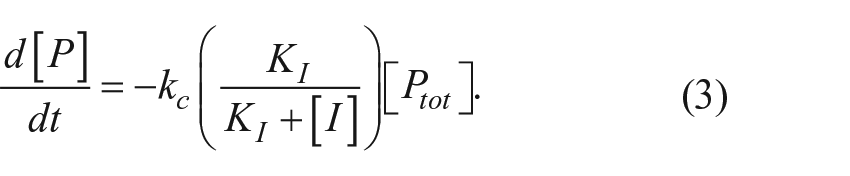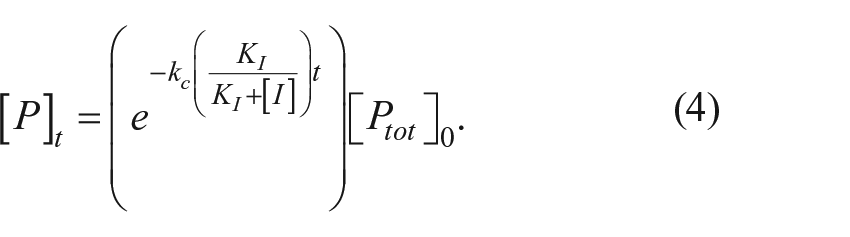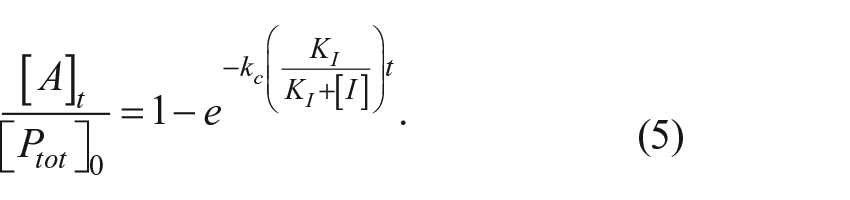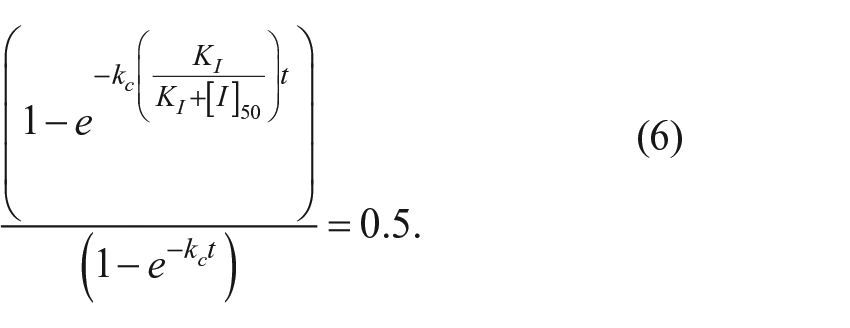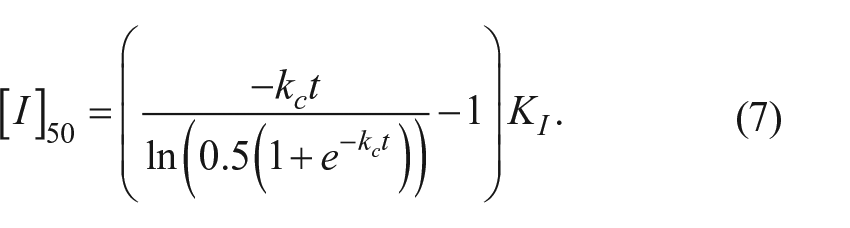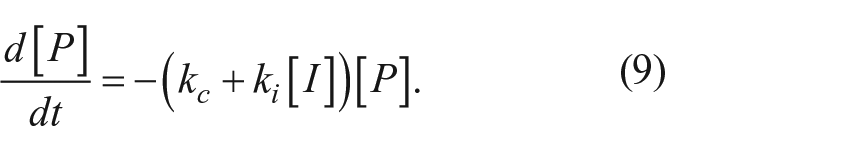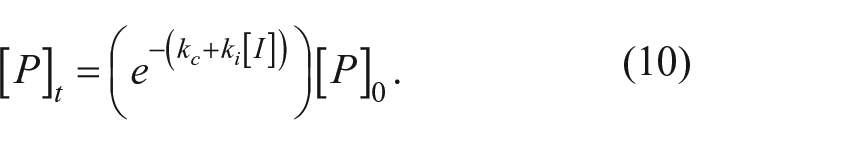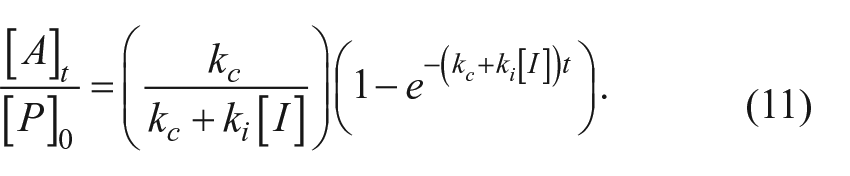Abstract
PCSK9 plays a significant role in regulating low-density lipoprotein (LDL) cholesterol levels and has become an important drug target for treating hypercholesterolemia. Although a member of the serine protease family, PCSK9 only catalyzes a single reaction, the autocleavage of its prodomain. The maturation of the proprotein is an essential prerequisite for the secretion of PCSK9 to the extracellular space where it binds the LDL receptor and targets it for degradation. We have found that a construct of proPCSK9 where the C-terminal domain has been truncated has sufficient stability to be expressed and purified from Escherichia coli for the in vitro study of autoprocessing. Using automated Western analysis, we demonstrate that autoprocessing exhibits the anticipated first-order kinetics. A high-throughput time-resolved fluorescence resonance energy transfer assay for autocleavage has been developed using a PCSK9 monoclonal antibody that is sensitive to the conformational changes that occur upon maturation of the proprotein. Kinetic theory has been developed that describes the behavior of both reversible and irreversible inhibitors of autocleavage. The analysis of an irreversible lactone inhibitor validates the expected relationship between potency and the reaction end point. An orthogonal liquid chromatography–mass spectrometry assay has also been implemented for the confirmation of hits from the antibody-based assays.
Introduction
Proprotein convertase subtilisin-like/kexin type 9 (PCSK9) is a major regulator of plasma low-density lipoprotein (LDL) cholesterol (LDL-C) and consequently an important therapeutic target for treating coronary heart disease. 1 PCSK9 binds to the LDL receptor (LDLR) on the cell surface of hepatocytes and directs the receptor to intracellular lysosomes for degradation. Abundant genetic data confirm that the level of PCSK9 function affects the level of LDL-C.2,3 Gain-of-function mutations in PCSK9, which increase its affinity for LDLR, result in decreased LDLR on the cell surface, increased serum LDL-C, and negative consequences for cardiovascular health. Conversely, loss-of-function mutations in PCSK9 lead to higher LDLR levels and lower LDL-C, which is associated with positive clinical outcomes. Therapeutic strategies that have demonstrated clinical efficacy in reducing LDL-C accomplish the loss of PCSK9 function by either blocking PCSK9’s LDLR binding activity with a monoclonal antibody or reducing the PCSK9 protein level with RNA interference (RNAi).4-6
PCSK9 belongs to the mammalian proprotein convertase family of serine proteases and contains an N-terminal signal sequence, prodomain, catalytic domain, and C-terminal domain. 7 While in the endoplasmic reticulum, PCSK9 performs its only catalytic activity, an autocleavage between residues Gln-152 and Ser-153.8,9 The prodomain remains tightly associated with the catalytic domain during subsequent trafficking through the trans-Golgi network. The maturation via autocleavage has been demonstrated to be critical for PCSK9 secretion and subsequent extracellular function 10 and may thus present an opportunity for a small-molecule therapeutic intervention with this target. Herein we describe assays we have developed for the discovery and characterization of PCSK9 autoprocessing inhibitors that may be investigated as novel leads for hypercholesterolemic therapy.
Materials and Methods
Materials
Most of the reagents and supplies for the automated Western Sally instrument were contained in the Rabbit (12–180 kDa) Size Separation Master Kit (CBS-01-01) purchased from ProteinSimple (San Jose, CA). The hexahistidine monoclonal primary antibody was obtained from Clontech (Mountain View, CA; 631212). D2-labeled anti-His (61HISDL), EuK-labeled anti–human IgG (61HFCKL), and homogeneous time-resolved fluorescence (HTRF) detection buffer (62SDBRDF) were purchased from Cisbio (Bedford, MA). Merck PCSK9 antibody AX-1 was custom labeled with ~6 molar equivalents of EuK by Cisbio. Sodium malonate (3.4 M, pH 7.0) was purchased from Hampton Research (Aliso Viejo, CA). 3,4-Dichloroisocoumarin (DCI) and autosampler wash solutions were purchased from Sigma-Aldrich (St. Louis, MO): water solution (633321; 80% water/0.1% formic acid/20% acetonitrile) and acetonitrile solution (685461; 50% acetonitrile/0.5% formic acid/40% 2-propanol/10% acetone). Liquid chromatography–mass spectrometry (LC-MS) grade solvents for the mobile phases were purchased from Fisher Scientific (Waltham, MA): water/0.1% formic acid (LS118-4) and acetonitrile/0.1% formic acid (LB9823-4).
Preparation of proPCSK9ΔCT
A synthetic gene encoding residues 31 to 454 of human PCSK9 was designed and synthesized by CODA Genomics (Laguna Hills, CA). Using traditional cloning techniques, the gene was inserted into pET17b (EMD Millipore, Darmstadt, Germany) between the NdeI and XhoI sites. The plasmid was transformed into BL21(DE3). Colonies were used to inoculate 2YT media and were grown at 30 °C. In mid-log phase, the incubator temperature was reset to 4 °C. When the culture media reached 12 °C, recombinant protein expression was induced with the addition of isopropyl β-D-1-thiogalactopyranoside (IPTG) to a final concentration of 0.5 mM. Cell growth continued overnight. In the morning, cells were harvested by centrifugation. All purification steps were performed as rapidly as possible at 4 °C. Cell pellets were resuspended in 50 mM sodium phosphate (pH 8.0)/0.3 M NaCl and lysed by two passes through a microfluidizer at 17,000 psi. The lysate was clarified by centrifugation and applied to Ni-NTA resin column (HisTrap FF). After washing with 10 column volumes each of 50 mM HEPES (pH 7.5)/10% glycerol/1 M NaCl/25 mM imidazole and 50 mM HEPES (pH 7.5)/10% glycerol/0.3 M NaCl/50 mM imidazole, the protein was step eluted with 50 mM HEPES (pH 8.0)/10% glycerol/0.3 M NaCl/200 mM imidazole. The eluate was loaded onto a Superdex 75 (GE Healthcare, Marlborough, MA) size exclusion column equilibrated in 25 mM HEPES (pH 7.5)/0.15 M NaCl. Pooled fractions were concentrated by ultrafiltration to 0.5 mg/mL.
Automated Western Assays
Time courses were initiated by 10-fold dilution of the cold 0.5-mg/mL stock solution of proPCSK9ΔCT into 25 mM HEPES (pH 7.5)/300 mM sodium malonate at room temperature. At time zero and various additional time points, a 48-µL aliquot was removed and mixed with 2 µL of 125 mM Tris(2-carboxyethy1)phosphine (TCEP) at room temperature. At the completion of the time course, the samples were diluted 10-fold with water, and then 11.25 µL of each diluted sample was combined with 3.75 µL of 4× MasterMix (ProteinSimple) prepared according to kit instructions. After heating to 95 °C for 5 min to denature the proteins, the samples were added to the assay plate along with primary antibody (diluted 1:50), secondary antibody, biotinylated ladder, stacking matrix, separation matrix, streptavidin–horseradish peroxidase (HRP), luminol-S, peroxide, and water following kit instructions. The Sally instrument running conditions adhered to vendor recommendations for the Simple Western Sally. All analyses were done with multiple image exposures to accommodate different signal strengths. Signals were converted to electropherograms by ProteinSimple’s Compass software. The catalytic domain and uncleaved proprotein were identified by peaks with molecular weights of 33 and 46 kDa, respectively. The percent conversion was determined as 100 × (peak area of catalytic domain)/(peak area of catalytic domain + peak area of proprotein).
Time-Resolved Fluorescence Resonance Energy Transfer Assays
The autoprocessing time course was initiated by 10-fold dilution of the cold 0.5-mg/mL stock solution of proPCSK9ΔCT into 25 mM HEPES (pH 7.5)/300 mM sodium malonate at room temperature and rapidly distributing 10 µL/well into a black 384-well plate (3676; Corning, Corning, NY). The reaction was stopped at various times by the addition of 5 µL/well of 80 µM DCI in HTRF detection buffer. At the end of the time course, 5 µL/well of detection reagents was added (40 nM AX-1, 120 nM D2-labeled anti-His, 8 nM EuK-labeled anti–human IgG) and incubated for 1 h at room temperature. The fluorescence resonance energy transfer (FRET) signal was determined by ratioing the emission signals at 665 and 620 nm (excitation 337 nm) from a BMG PheraStar (BMG Labtech, Ortenberg, Germany) microplate reader.
To screen for inhibitors of autoprocessing, 100 nL of test compounds in DMSO was distributed to plate wells using an acoustic dispense system (Labcyte Echo; Labcyte, Sunnyvale, CA) followed by 10 µL proPCSK9ΔCT at 50 µg/mL in 25 mM HEPES (pH 7.5)/300 mM sodium malonate. After a 4-h incubation at room temperature, the reaction was stopped by the addition of 10 µL quench/detection reagents (80 µM DCI, 60 nM D2-labeled anti-His, 2.4 nM EuK-labeled AX-1 in HTRF detection buffer). After an additional 1-h incubation, the FRET signal was measured as above.
For the experiment examining the impact of an inhibitor upon the autoprocessing time course, 100 nL of various concentrations of the test compound was dispensed to plate wells via acoustic dispense before initiating reactions with 10 µL of 50 µg/mL proPCSK9ΔCT. The reaction wells were stopped at various times by the addition of 5 µL of 160 µM DCI. After the final time point, all wells received 5 µL of detection reagents (120 nM D2-labeled anti-His, 4.8 nM EuK-labeled AX-1 in HTRF detection buffer).
LC-MS Assay
Time courses were initiated by 10-fold dilution of the cold 0.5-mg/mL stock solution of proPCSK9ΔCT into 25 mM HEPES (pH 7.5)/300 mM sodium malonate at room temperature. At time zero and various additional time points, a 78-µL aliquot was removed and mixed with 2 µL of 200 mM TCEP at room temperature. Samples were analyzed on a Transcend LX4-TSQ Vantage triple quadrupole LC-MS system (Thermo Scientific, Waltham, MA) using a Zorbax 3.5 micron 300SB-C8 2.1 × 50-mm column (Agilent Technologies, Santa Clara, CA). Samples were maintained in a thermostatted autosampler set to 4 °C, and the LC column was maintained at ambient temperature. A sample injection volume of 20 µL and a binary solvent system composed of water/0.1% formic acid (v/v, mobile phase A) and acetonitrile/0.1% formic acid (v/v, mobile phase B) was used for the separations. The column was initially conditioned with 2% mobile phase B for 20 s before linearly increasing mobile phase B to 60% over 5 min. The composition was then immediately stepped to 100% mobile phase B and held for an additional 80 s to wash the column before returning to the initial conditions of 2% mobile phase B. The flow rate was maintained at 0.75 mL/min throughout. The TSQ Vantage was operated in positive electrospray ionization mode, with a spray voltage of 3000 V, vaporizer temperature of 400 °C, sheath gas pressure of 45 arbitrary units, auxiliary gas pressure of 10 arbitrary units, capillary temperature of 280 °C, S-lens setting of 116 V, and collision energy of 10 eV. The abundance of the proPCSK9ΔCT prodomain (residues 31–152) was determined using selected (pseudo)reaction monitoring of a specific ion from the protein envelope (
Kinetics Theory
For the reversible inhibition mechanism depicted in Figure 1a , the binding of the inhibitor to the proprotein is assumed to be in rapid equilibrium relative to the autocleavage reaction with a dissociation constant defined by
Using that definition, it can be shown that the fraction of total proprotein that is free to autoprocess is
The rate of disappearance of the proprotein is dependent on the amount of free proprotein and the first-order rate constant kc:
The time course for proprotein disappearance is
The time course for appearance of the mature autoprocessed protein as a fraction of the total original proprotein is
The concentration of [I] that reduces the amount of mature protein by 50% relative to an uninhibited reaction at a given time point is
Solving for [I]50 gives
At one half-life (t = 0.693/kc), the [I]50 = 1.41 * KI, while at four half-lives (t = 2.772/kc), the [I]50 = 3.4 * KI.
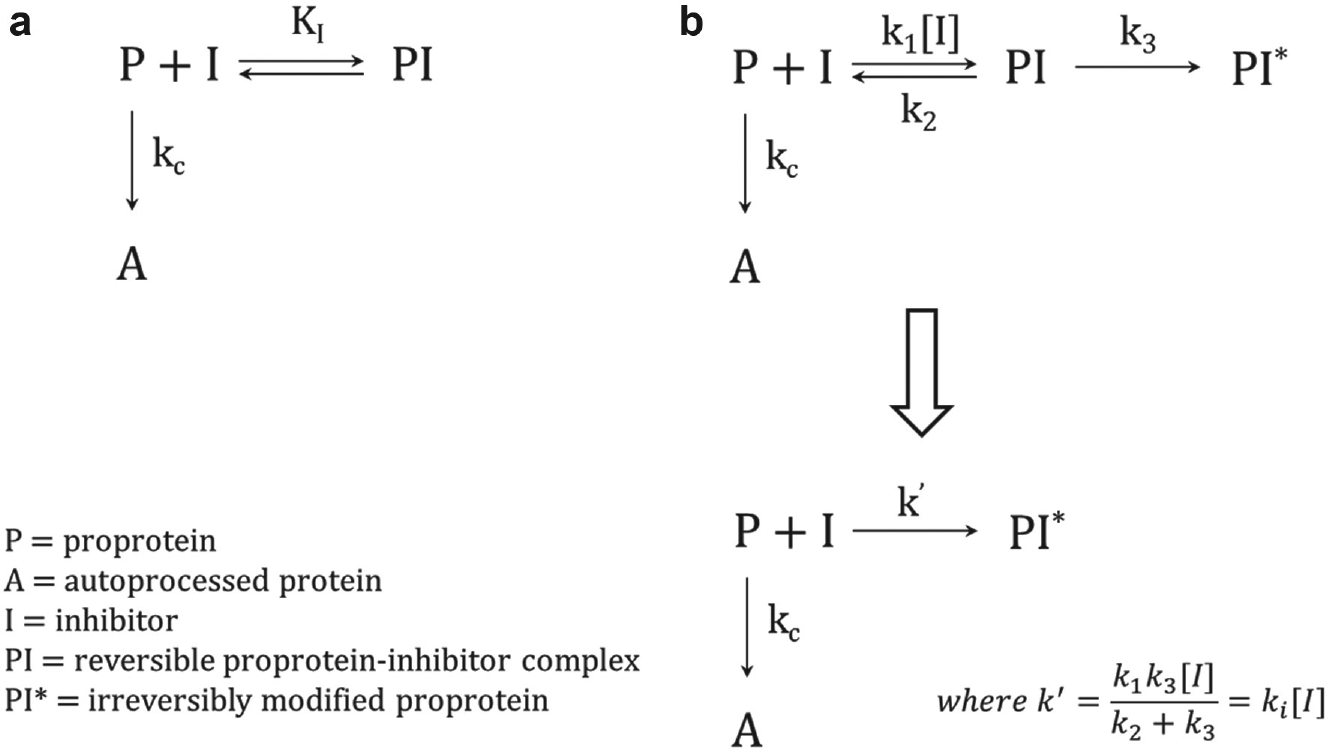
Outline of the (
For the irreversible inhibition mechanism depicted in Figure 1b , the fate of the proprotein is determined by the ratio of the competing processes of autocleavage (with rate constant kc) or inactivation (with rate constant k′). For a mechanism involving a reversible binding step for the inhibitor prior to inactivation, k′ is a net rate constant that varies with the concentration of the inhibitor:
The rate of disappearance of the proprotein is the sum of two first-order processes:
The time course for proprotein disappearance is
The time course for the appearance of the mature autoprocessed protein as a fraction of the total original proprotein is
The fractional reaction end point (when t = ∞) decreases hyperbolically with increasing inhibitor concentration:
The inhibitor IC50 after complete reaction is where
Results
The study of PCSK9 autoprocessing requires an isolable form of the PCSK9 proprotein. Recombinant PCSK9 secreted from mammalian cells undergoes complete processing, yielding the tightly associated pro- and catalytic domains. Efforts to prepare a soluble full-length version of the proprotein in Escherichia coli have been unsuccessful due to poor expression. A construct of PCSK9 lacking the C-terminal domain had been expressed and purified from E. coli previously, but it also was in a mature (i.e., fully cleaved) form. 11 We have found, however, that the intact C-terminal truncated proprotein can be isolated at >90% purity when expression and purification are carried out rapidly in the cold. The resulting proprotein, referred to here as proPCSK9ΔCT, is metastable toward autocleavage. A series of salts were empirically screened for their effect upon the maturation reaction, and a buffer containing 25 mM HEPES and 300 mM sodium malonate at pH 7.5 was found to be optimal for promoting efficient autoprocessing (~80% completion). The autoprocessing could be monitored by automated Western as described in the Materials and Methods. Using an antibody directed toward the His-tag at the C-terminus, the disappearance of the proprotein and appearance of the catalytic domain could be quantitatively monitored over time. TCEP was found to be an effective stop reagent for this experiment, presumably via reduction of the three disulfide bonds in the catalytic domain of proPCSK9ΔCT. A plot of the percent conversion as a function of time ( Fig. 2 ) was fit well by an exponential consistent with first-order kinetics for autocleavage with a rate constant of 0.014 min–1 (half-life 50 min).
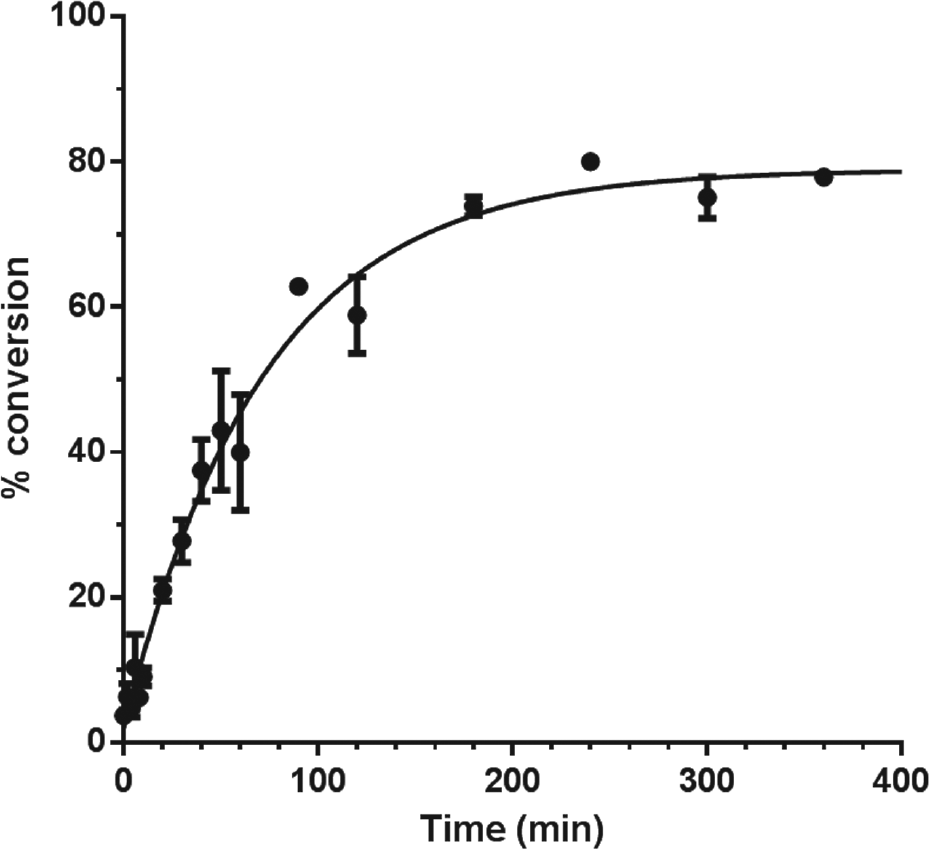
Time course for proPCSK9ΔCT autoprocessing as determined by automated Western analysis. The average of duplicate time points with range is shown.
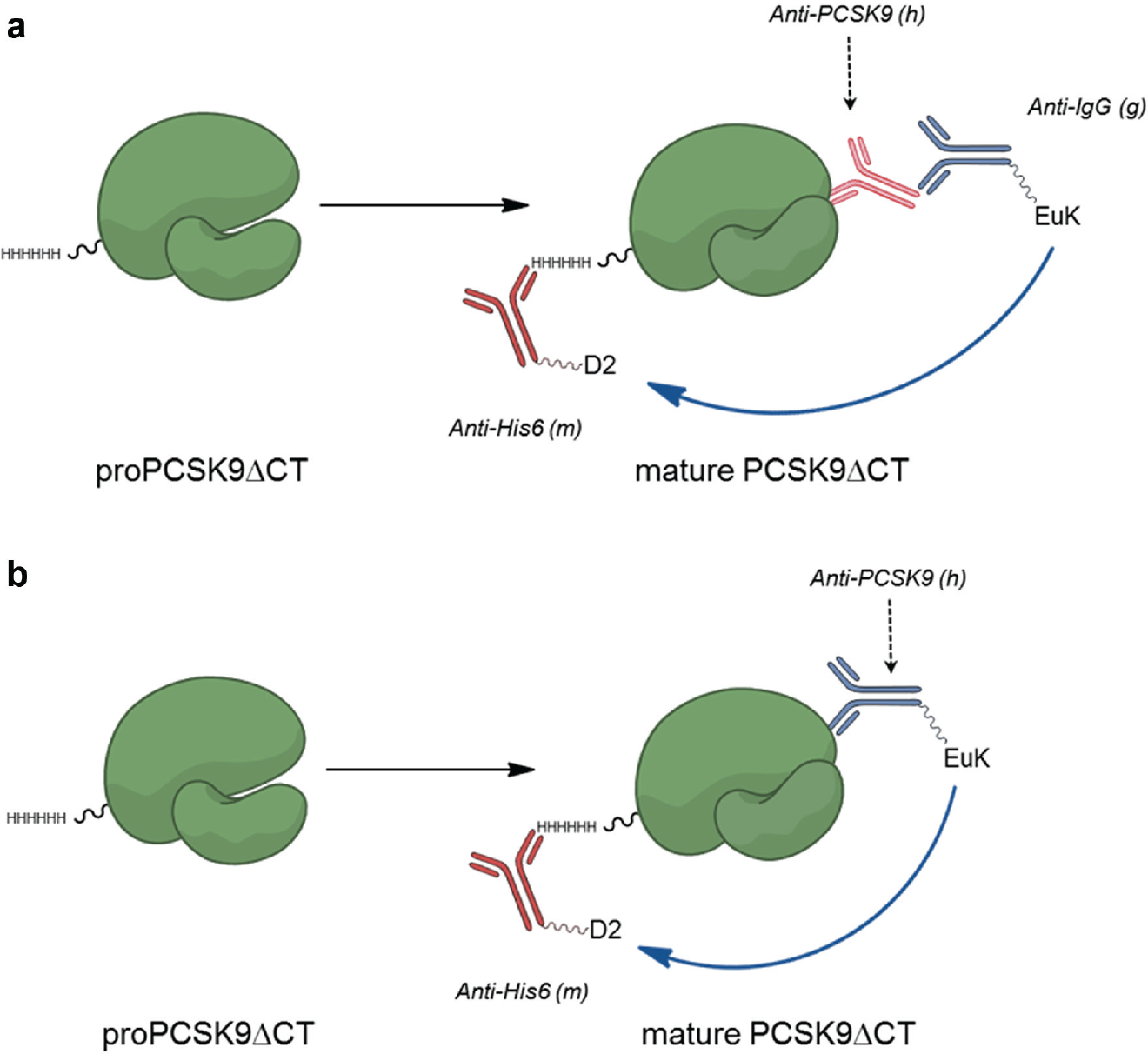
While use of the automated Western instrument provided significant improvements in quantitation and throughput over conventional Western blots, we sought a genuine high-throughput assay that would enable screening for inhibitors of autoprocessing. We thought that a conformation-sensitive antibody might provide us the opportunity to develop such an assay in a time-resolved FRET (TR-FRET) format. We had a number of human anti-PCSK9 monoclonal antibodies from our internal programs that we tested for suitability with the proximity assay diagrammed in Figure 3a . In this format, the poly-His epitope tag on PCSK9 was recognized by a mouse monoclonal antibody labeled with a fluorescence acceptor while a goat polyclonal anti–human IgG antibody labeled with a fluorescent donor was used to sandwich the anti-PCSK9 antibody being tested. Several antibodies exhibited the desired conformational selectivity, yielding a differential in the TR-FRET signal between the proprotein and mature protein. We focused on AX-1, which appeared to have the greatest utility for our assay development.
(
We wished to examine the time course of autocleavage with the new assay format, but the denaturation of the protein by TCEP was incompatible with the subsequent need for antibody recognition of the native state. Instead we turned to DCI, a reactive acylation agent used for serine protease inhibition. 12 As seen before with the automated Western assay, the autocleavage time course of proPCSK9ΔCT using the TR-FRET assay was first order with a rate constant of 0.012 min–1 (half-life 57 min) ( Fig. 4 ). To further simplify the TR-FRET format, we directly labeled the AX-1 antibody with the europium cryptate fluorescent donor reagent, eliminating the need to employ a sandwich antibody ( Fig. 3b ). This modification enabled a streamlined protocol with an improved signal-to-background (S/B) ratio that could be readily implemented in high-density plates for screening purposes. For example, using a 30-µL assay volume in 384-well plates with a 4-h reaction, we observed S/B >2.5 with Z′ > 0.6.
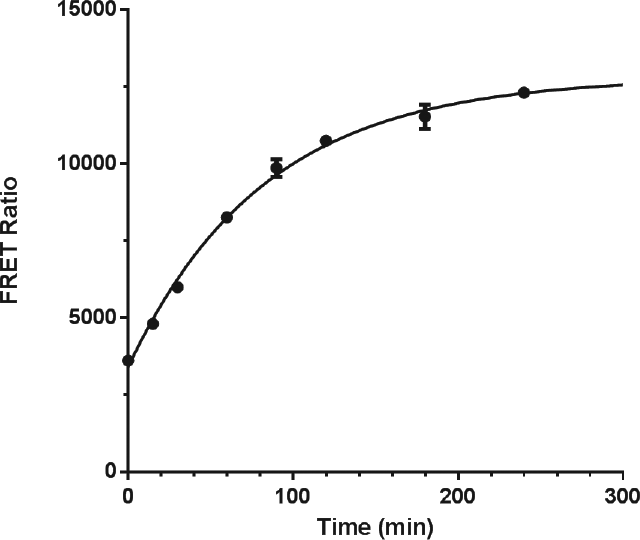
Time course for proPCSK9ΔCT autoprocessing as determined by the time-resolved fluorescence resonance energy transfer (TR-FRET) assay. The average of duplicate time points with range is shown.
The potential to test many thousands of compounds for inhibition of proPCSK9ΔCT autoprocessing prompted a theoretical examination of the kinetics for different inhibition mechanisms, as conventional enzyme kinetics do not apply to the single-turnover autocleavage where there is no external substrate. We broadly considered two possibilities, reversible and irreversible mechanisms, in Figure 1 . For the reversible mechanism, the inhibitor complexes with the proPCSK9ΔCT; the cis cleavage event cannot occur during the lifetime of this complex. Inhibition via this mechanism can slow down autoprocessing, but since the cis cleavage is irreversible, complete maturation will ultimately occur. The absence of a difference in the final end point for inhibited and uninhibited reactions has the practical consequence that a reversible inhibitor will appear most potent and is thus most easily detected at “early” time points (within a few half-lives). This means that a balance needs to be struck between maximizing the signal/background (which increases with time of reaction) and the sensitivity to inhibition (which decreases with time of reaction).
For the irreversible inhibition mechanism, an initial reversible complex is posited to form with subsequent trapping of the proprotein into a noncleavable form. In this scenario, two time-dependent processes are in competition with each other, and the partition between autocleavage and inactivation is determined by the relative magnitude of their rate constants. While the rate constant for autocleavage is fixed, the rate constant of inactivation, and thus the final partition ratio, is a function of the inhibitor concentration. The dependence of the compound IC50 as a function of time is a transcendental equation without a closed-form solution, but simulations indicate that the apparent IC50 of the inhibitor should decrease with the time of reaction. This results in the greatest sensitivity for detecting inhibition at “late” time points approaching the end point of the autocleavage reactions where signal/background for the uninhibited control is also maximal. Thus, a reaction time of several half-lives improves both assay performance and detection of irreversible inhibition.
To illustrate the behavior of an irreversible inhibitor, we provide characterization of a representative of a lactone series of autoprocessing inhibitors described elsewhere. 13 Figure 5a shows the time courses for proPCSK9ΔCT maturation at various concentrations of the compound. It is apparent that the progress curve for each compound concentration reaches a different end point plateau as expected for an irreversible inhibition mechanism. The entire data set was globally fit to a version of equation (11) to determine a value of 0.0059 min–1 for kc, the intrinsic autocleavage rate constant, and 0.0025 µM−1 min–1 for ki, the rate constant for inactivation. Figure 5b presents the data in the form of inhibition titration curves for each time point. Consistent with the kinetic theory presented, as the time of reaction increased, the apparent IC50 decreased to approach the predicted theoretical limit of 2.4 µM (= kc/ki).
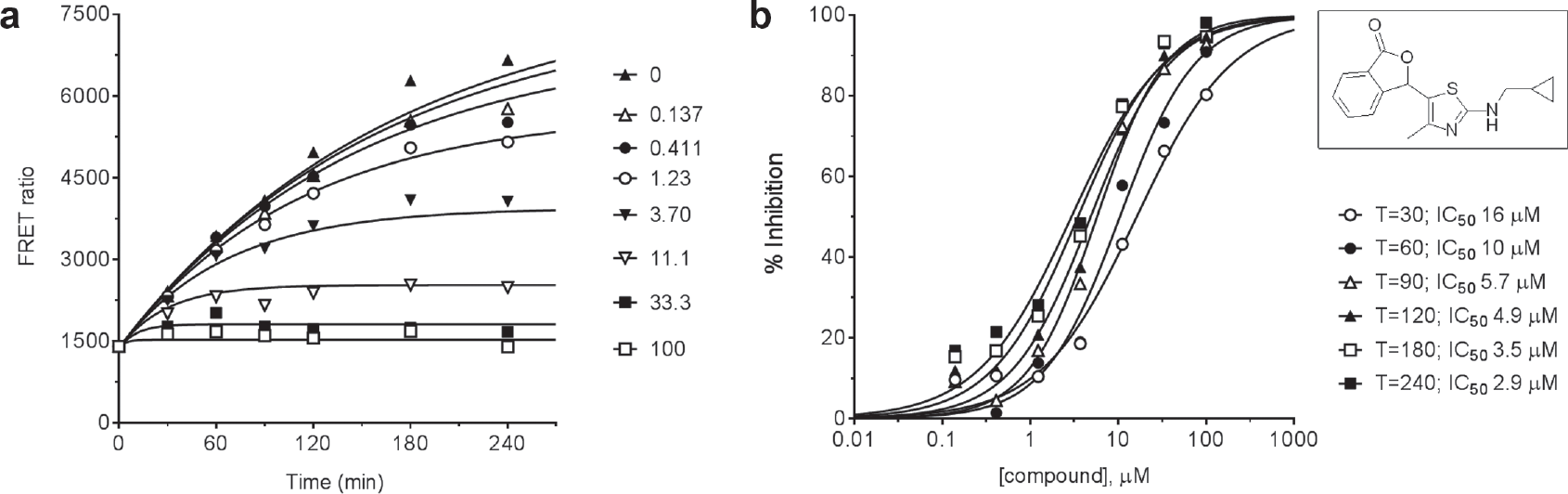
(
As part of our triage plan for screening for autoprocessing inhibitors, we also wanted an orthogonal assay that would directly observe a product of cis cleavage without reliance on antibodies. A fit-for-purpose LC-MS assay was developed where the appearance of the prodomain (residues 31–152) could be quantitated after separation from the catalytic domain (residues 153–454) and intact proprotein (
Discussion
The objective of our study was to delineate methods for the discovery and characterization of inhibitors of the critical autocleavage event that is required for secretion of PCSK9 to the extracellular space where it is active in removing LDLR from the cell surface. Essential to this effort was the successful isolation of the proenzyme form of PCSK9, which could be used in autoprocessing assays. While there have not been any reports of the preparation of a recombinant full-length PCSK9 proprotein, we found that truncating the C-terminal domain led to the successful bacterial expression of a soluble form of the proprotein. Following rapid isolation of proPCSK9ΔCT, we used three orthogonal assays (automated Western, TR-FRET, LC-MS) to show that the truncated proprotein does in fact undergo a first-order autoprocessing maturation. The in vitro half-life we measured at room temperature is similar to that recently observed at 37 °C in HepG2 cells or HEK293 cells stably overexpressing full-length PCSK9 13 but appears to be somewhat longer than an earlier report. 14 The evidence thus supports the use of proPCSK9ΔCT as a surrogate for studying autoprocessing of the full-length proprotein.
We considered both reversible and irreversible scenarios for the inhibition of autoprocessing. The maturation of proPCSK9 via autocleavage is a single-turnover irreversible event, and the enzyme kinetics literature does not provide an adequate theoretical framework for the analysis of its inhibition. Our derivations show that reversible inhibitors can slow the time course for maturation in proportion to their binding affinity but cannot alter the final end point of complete autocleavage. In contrast, irreversible inhibitors exhibit the greatest fractional reduction of mature protein at the reaction end point. For this type of compound, the efficacy of autoprocessing inhibition, as measured by IC50, is a function of both the binding affinity of the initial reversible complex and the rate constant for the chemical modification step itself. The analysis of the time course does not enable the determination of these individual parameters; additional methods are required to determine the separate contributions of inhibitor binding affinity and reactivity.
The characterization of the lactone tool compound validated the theoretical predictions for the behavior of an irreversible inhibitor, particularly the inhibitor concentration dependence of the reaction end point and the increase in potency with increasing extent of reaction. Since longer reaction times also maximize the signal/background for the assay, the detection of irreversible inhibitors has an advantage over reversible inhibitors whose inhibition diminishes with increasing extent of conversion. The properties of irreversible inhibitors might also bestow practical advantages over reversible inhibitors in vivo. In the case of the irreversible inhibitor, the reduction of secreted PCSK9 requires the presence of drug only long enough to divert a substantial fraction of the proprotein pool into a form that cannot mature. With a reversible inhibitor, an efficacious drug level must be continuously maintained to minimize the otherwise inevitable maturation of proprotein. These considerations could be a rationale for selecting chemical libraries to screen that are known to contain, or are even enriched in, reactive functionalities that would be avoided with other targets. While there are potential risks with such an approach, there has been an increasing acceptance of irreversible inhibition as a viable mechanism for drug development.15,16
Irreversible inhibition of serine proteases by compounds that react with active site nucleophiles has abundant precedent. 17 Although PCSK9 is a serine protease, access to its active site is restricted after autoprocessing by the continued presence of the prodomain. While S386 in the catalytic triad was initially considered a likely site of modification by the lactone tool compound, peptide mapping revealed the primary labeling site to be a structural element in the protein remote from the active site that unexpectedly affects autoprocessing of the proprotein. 13 There may thus be several routes to intercepting autocleavage via small-molecule inhibition. The assays described in this work should aid the screening process and subsequent characterization of new chemical matter targeting PCSK9.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.