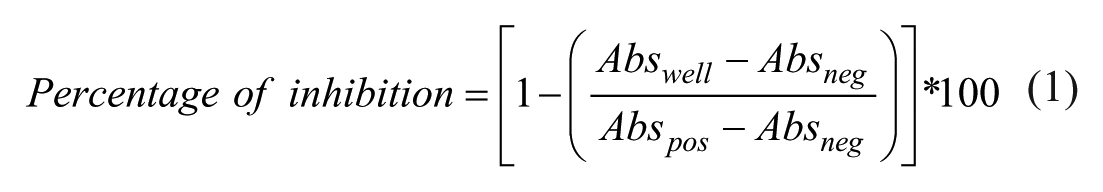Abstract
It is widely accepted that central nervous system inflammation and systemic inflammation play a significant role in the progression of chronic neurodegenerative diseases such as Alzheimer’s disease and Parkinson’s disease, neurotropic viral infections, stroke, paraneoplastic disorders, traumatic brain injury, and multiple sclerosis. Therefore, it seems reasonable to propose that the use of anti-inflammatory drugs might diminish the cumulative effects of inflammation. Indeed, some epidemiological studies suggest that sustained use of anti-inflammatory drugs may prevent or slow down the progression of neurodegenerative diseases. However, the anti-inflammatory drugs and biologics used clinically have the disadvantage of causing side effects and a high cost of treatment. Alternatively, natural products offer great potential for the identification and development of bioactive lead compounds into drugs for treating inflammatory diseases with an improved safety profile. In this work, we present a validated high-throughput screening approach in 96-well plate format for the discovery of new molecules with anti-inflammatory/immunomodulatory activity. The in vitro models are based on the quantitation of nitrite levels in RAW264.7 murine macrophages and interleukin-8 in Caco-2 cells. We have used this platform in a pilot project to screen a subset of 5976 noncytotoxic crude microbial extracts from the MEDINA microbial natural product collection. To our knowledge, this is the first report on an high-throughput screening of microbial natural product extracts for the discovery of immunomodulators.
Keywords
Introduction
Despite different triggering events, a common feature of neurodegenerative diseases is chronic immune activation, in particular of microglia, the resident macrophages of the central nervous system. 1 Therefore, several approaches have been made to explore ways to inhibit or modulate microglial activation through the major anti-inflammatory target proteins, such as inducible nitric oxide synthase (iNOS) and interleukin-8 (IL-8), among many others thought to contribute to neuronal damage. Nitric oxide (NO) regulates numerous physiological processes, including neurotransmission, smooth muscle contractility, platelet reactivity, and the cytotoxic activity of immune cells. Because of the ubiquitous nature of NO, inappropriate release of this mediator has been linked to the pathogenesis of a number of disease states. This provides the rationale for the design of therapies that modulate NO concentrations selectively. The inhibitors of iNOS are potentially beneficial agents in the treatment of conditions associated with an overproduction of NO, including septic shock, neurodegenerative disorders, and inflammation. In turn, the proinflammatory chemokine IL-8, produced by macrophages and other cell types, exposed to inflammatory stimuli, is also an attractive target for inflammation treatment. 2
Several classes of drugs, such as corticosteroids, nonsteroidal anti-inflammatory drugs, and biologics, are used to treat inflammatory disorders. However, all these drugs possess several adverse side effects, such as gastrointestinal ulceration, bleeding, and platelet dysfunction due to the inhibition of COX-1–derived prostanoids, or cardiovascular side effects due to inhibition of prostacyclin formation. Therefore, there is a continuous need to search for more effective and safer anti-inflammation agents, which is a key objective for the pharmaceutical industry. In this regard, natural products or natural product–derived compounds offer great structural diversity, which is not commonly seen in synthetic compounds. Several reviews have recently been published on natural anti-inflammatory compounds, underlining the potential that these products offer in identifying and developing novel bioactive compounds into drugs for the treatment of inflammatory diseases. Nevertheless, these reviews have been devoted exclusively to plants, and no mention is made of marine or microbial sources. 3 Interestingly, the 3 prominent categories of natural product compounds identified from these reviews (poliphenolics, terpenoids, and fatty acids) are commonly produced in microbial fermentations. 4 The bioactivities of these classes of compounds are exhibited through the down-regulation of different types of inflammatory mediators, highlighting the importance of microbial fermentations as potential biofactories for the production of natural anti-inflammatory metabolites of highly diversified chemical structures. The possibility of obtaining them by scaling up the fermentations of the producing microbial strain is an additional advantage that microbial natural products offer when compared with natural compounds from other sources. However, as with natural products, the search for active compounds from whole crude extracts is both time-consuming and capital intensive. For this reason, it is critical to develop quick, efficient, high-throughput primary screening assays integrated with chemistry platforms for early de-replication of known compounds to guarantee that only extracts with relevant pharmacological properties are selected for further study. 5 The purpose of this ongoing project is to identify, characterize, and develop potent natural product–based drug leads with anti-inflammatory/immunomodulatory activity. In this study, we report on a validated high-throughput screening (HTS) platform for the screening of microbial extracts to discover new molecules with anti-inflammatory/immunomodulatory activity ( Fig. 1 ). In vitro studies were performed to evaluate immunomodulatory properties of microbial extracts, both in bacterial lipopolysaccharide (LPS)–stimulated murine macrophage cells (RAW264.7) for NO production 6 and interleukin-1β (IL-1β)–stimulated intestinal epithelial cells (Caco-2) for IL-8 production. 7 NO production in LPS-activated macrophages is mainly due to the increased expression of iNOS, which is regulated by different cytokines via the transcription factor nuclear factor-kappaB (NF-κB)–dependent pathway. 8 The transcription factor NF-κB controls more than 500 different gene products, and it has emerged as a major mediator of inflammation. Thus, agents that can inhibit NF-κB have the potential to diminish the onset of chronic inflammation. Similarly, IL-8 expression and its production are increased in Caco-2 cells after stimulation with IL-1β, a process also mediated by activation of the transcription factor NF-κB. 9 Even though the involvement of NO in the production of IL-8 has been studied in different cell types and tissues, the mechanism of action has not been clarified yet. However, the results suggest that the production of IL-8 is regulated by similar transcription factors to those responsible for the increased expression of iNOS. 10 Therefore, the correlated measurements of NO and IL-8 productions provide not only information about the ability of the compounds to inhibit the activity of iNOS and thus the production of NO but also the mechanism of action by their potential to interfere with the expression of inducible inflammatory mediators. Consequently, by using this experimental design, we aim to discover compounds with a dual activity because they may indirectly interfere with common pathways involved in the inflammatory response. We have used this platform in a pilot project to screen a subset (5976) of noncytotoxic crude microbial extracts from the MEDINA microbial natural product collection library, one of the world’s most diverse and productive collections of filamentous fungi, actinomycetes, and unicellular bacteria. As a result, we found that 2 extracts contained potentially new compounds that are currently undergoing nuclear magnetic resonance (NMR) and high-resolution mass spectrometry (HRMS) de-replication for future isolation and characterization. In fact, based on the strong in vitro immunomodulatory activity displayed, we have isolated and identified an epimer mixture of a previously described molecule 11 with several reported biological activities in a human neutrophil-like cell line. 12 These results demonstrate that this natural product cell-based screening may provide a powerful means of identifying new potential drugs with anti-inflammatory/immunomodulatory activity.
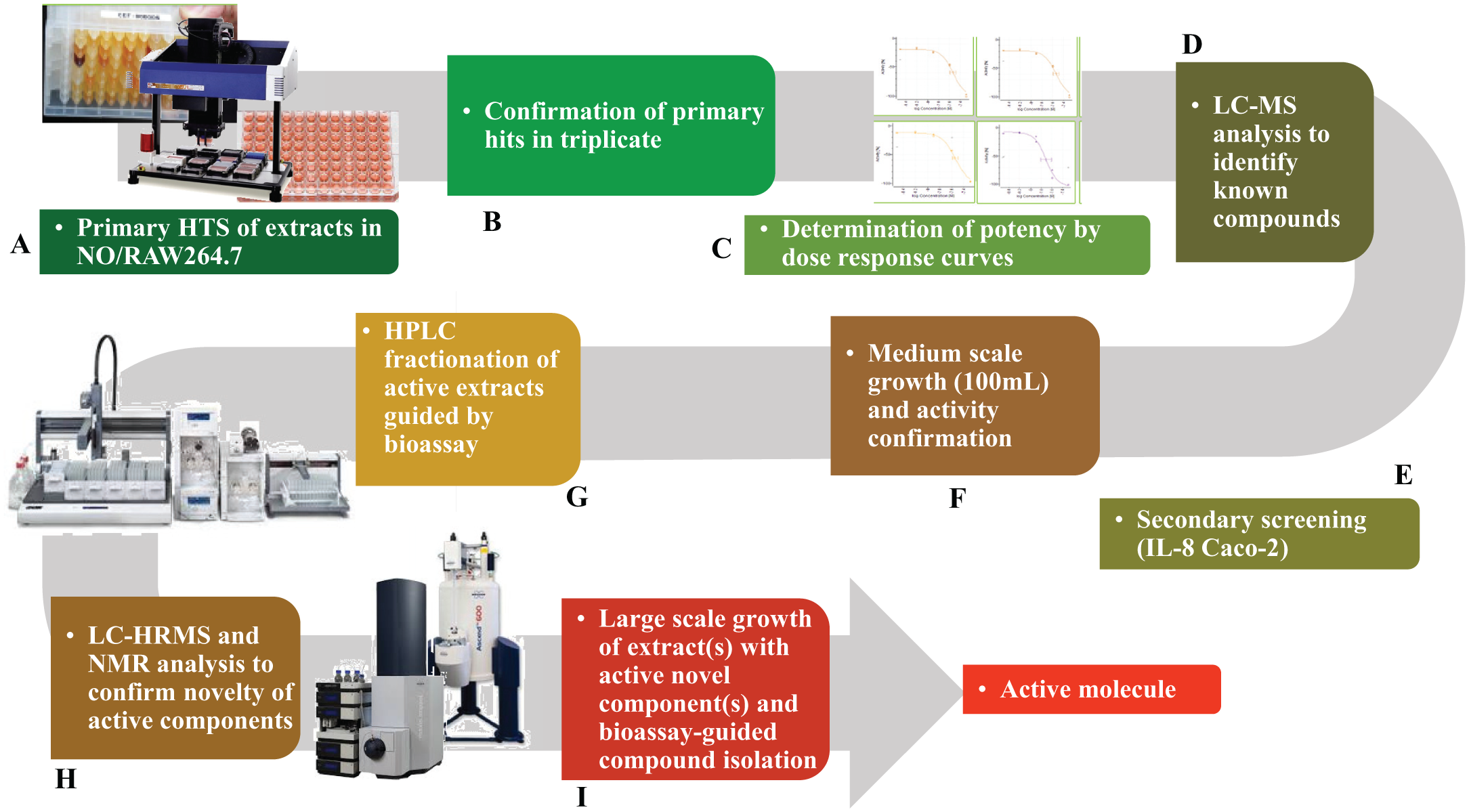
Schematic representation of the high-throughput screening and integrated de-replicating chemistry platform used in discovering immunomodulatory molecules from microbial extracts. (
Materials and Methods
Reagents
Fetal bovine serum (FBS), L-glutamine, sodium pyruvate, Modified Eagle Medium (MEM) nonessential amino acids, penicillin–streptomycin, and TrypLE Express were purchased from Invitrogen Gibco, Inc. (Life Technologies, Carlsbad, CA). NP40 was purchased from Thermo Scientific (Rockford, IL). Diphenyl tetrazolium bromide (MTT), LPS from Escherichia coli, and methyl methane sulfonate were obtained from Sigma Aldrich (St. Louis, MO). Human IL-1β was purchased from Miltenyi Biotec (Bergisch Gladbach, Germany).
Microbial Extracts Collection
For the primary screening campaign, a subset of the MEDINA microbial natural product collection consisting of 5976 noncytotoxic microbial extracts (35,683,568 from actinomycetes and 2408 from fungi) was used. This subset of extracts was previously found to inhibit by less than 20% the growth of HepG2 cells when tested at a final concentration of 1/40 × whole-broth equivalent (WBE) for 24 h. The microbial extracts were obtained from a selection of producing strains that were cultivated for 7 and 14 d at 28 °C in the case of actinomycetes and for 21 and 28 d at 22 °C for fungi. The secondary metabolites in the broths were extracted with acetone (1:1) by mixing 10 mL of culture broth with 10 mL of acetone and shaking in an orbital shaker (Adolf Kuhner AG, Birsfelden, Switzerland) for 1 h. The extracts were then centrifuged at 1500 × g for 15 min, and the supernatants were transferred to 16 mm glass tubes. DMSO was then added to each extract, and the organic solvent and part of the water were evaporated until half of the original fermentation volume under a warm nitrogen stream to attain a final crude concentration of 2 × WBE and 20% DMSO. Five copies of 100 µL (500 µL) of these crude extracts were stored at −20 °C in 96-well ABgene plates (AB-0756) until needed.
Cell Cultures
The mouse macrophage cells, RAW264.7 (ATCC TIB-71), and the human colon adenocarcinoma cells, Caco-2 (ATCC HTB-37), were obtained from ATCC and cultured in DMEM, supplemented with 10% FBS and 2 mM l-glutamine, in a humidified 5% CO2 atmosphere at 37 °C. All cell-handling steps were carried out using the SelecT automated cell culture system.
NO Assay
The following procedure is an adaptation to the 96-well format from a methodology previously described. 6 RAW264.7 cells were seeded into 96-well plates at a density of 20 × 103 cells per well and grown to approximately 50% of confluence (24 h). They were cultured for another 24 h with each of the extracts described above (5 µL of 2 × WBE at 20% DMSO were dispensed into 190 µL of fresh medium using Perkin Elmer Evolution P3 [Waltham, MA], obtaining a final extract concentration of 1/20 × WBE). Afterward, these cells were stimulated with LPS (5 µL of 6 µg/mL were dispensed using a Thermo Scientific Multidrop Combi dispenser [MTX Lab Systems, Vienna, VA]). Untreated (5 µL of 20% DMSO, microbial extract vehicle) and unstimulated (5 µL of purified water, LPS vehicle) cells were used as negative controls (n = 4), whereas untreated but stimulated cells were used as positive controls (n = 4) in each assay plate. After 24 h, supernatants were collected and centrifuged at 10,000 × g for 5 min, and nitrite levels were measured in 100 µL of supernatant according to the Griess 6 assay using a Thermo Scientific Multidrop Combi dispenser and PerkinElmer EnVision Multilabel Reader (Waltham, MA). After discarding the remaining supernatant, the cell viability was examined in the remaining cells through the MTT test described elsewhere 6 and also using a Thermo Scientific Multidrop Combi dispenser and PerkinElmer EnVision Multilabel Reader.
Assays for IL-8 Regulation Assessment in IL-1β–Stimulated Caco-2 Cells
The following procedure is an adaptation to the 96-well format from a methodology previously described. 6 Caco-2 cells were seeded into 96-well plates at a density of 20 × 103 cells per well and grown to approximately 50% confluence (24 h, 37 °C and 10% CO2). Then, they were preincubated for 24 h with each of the extracts described above (5 µL of 2 × WBE at 20% DMSO were dispensed into 190 µL of fresh medium using Perkin Elmer Evolution P3, obtaining a final extract concentration of 1/20 × WBE). Afterward, cells were stimulated with IL-1β (5 µL of 2 ng/mL were dispensed using a Thermo Scientific Multidrop Combi dispenser) for 24 h. Untreated (5 µL of 20% DMSO, microbial extract vehicle) and unstimulated (5 µL of purified water, IL-1β vehicle) cells were used as negative controls (n = 4), whereas untreated but stimulated cells were used as positive controls (n = 4) in each assay plate. The supernatants (100 µL) were then collected, centrifuged at 10,000 × g for 5 min, and stored at −80 °C until IL-8 determination by enzyme-linked immunosorbent assay (Biosource, Invitrogen) was performed. Cell viability was examined using the MTT test described elsewhere. 6
Data Analysis
Pure compounds and extract activities were calculated automatically using the Genedata Screener software (Genedata AG, Basel, Switzerland), and the percentage inhibition of each extract (or compound) was determined by eq 1, integrated in the Genedata Screener software: Percentage of inhibition for production of either NO or IL-8:
where Abswell is the absorbance measured per specific well and Abspos and Absneg are the average absorbance measured for the positive and negative controls, respectively.
Determination of IC50 Values for Pure Known Compounds
The IC50 values of commercially known compounds (dexamethasone, cortisol, and cyclosporine A) were obtained from dose-response curves in the NO assay described above. Compound concentrations were started at different levels to optimize IC50 calculation, and twofold serial dilutions were performed (DMSO concentration below 0.5%). The IC50 values were calculated as the compound concentration that inhibits the production of NO by 50% using the online Genedata Screener software.
Primary Screening and Dose-Response Experiments
All of the extracts were first screened against the macrophages model at a single concentration (1/20 × WBE) in duplicate ( Fig. 1A ), and those exhibiting inhibition of NO production ≥80% and cell viability ≥75% were selected as hits and confirmed in triplicate ( Fig. 1B ). The extracts selected from this stage were then tested in a five-point dose-response assay for potency determination ( Fig. 1C ); the first point was the initial dilution previously described for each assay (1/20 × WBE), and the four subsequent points were twofold serial dilutions from each point. Extracts that exhibited a minimum of 70% inhibition at initial dilution (1/20 × WBE) in dose-response experiments were selected for de-replication ( Fig. 1D ) by liquid chromatography low-resolution mass spectrometry (LC-MS).
LC-MS and Database Matching of Known Secondary Metabolites (De-Replication)
Two microliters of the selected extracts were analyzed by LC-MS. Analysis was performed on an Agilent (Santa Clara, CA) 1100 single quadrupole LC-MS, using a Zorbax SB-C8 column (2.1 × 30 mm), maintained at 40 °C with a flow rate of 300 µL/min. Solvent A consisted of 10% acetonitrile and 90% water with 1.3 mM trifluoroacetic acid and 1.3 mM ammonium formate, whereas solvent B was 90% acetonitrile and 10% water with 1.3 mM trifluoroacetic acid and 1.3 mM ammonium formate. The gradient was started at 10% B and went to 100% B in 6 min; it was kept at 100% B for 2 min and was returned to 10% B for 2 min to initialize the system. Full diode array ultraviolet (UV) scans from 100 to 900 nm were collected in 4 nm steps at 0.25 s/scan. The eluting solvent was ionized using the standard Agilent 1100 ESI source adjusted to a drying gas flow of 11 L/min at 325 °C and a nebulizer pressure of 40 psig. The capillary voltage was set to 3500 V. Mass spectra were collected as full scans from m/z 150 to 1500, with 1 scan every 0.77 s, in both positive and negative modes. Database matching was performed using an application developed in-house, in which the diode array detection, retention time, and positive and negative mass spectra of the active samples were compared with the UV-LC-MS data of known metabolites stored in a proprietary database and in which metabolite standard data were obtained using the exact same LC-MS conditions as the samples under analysis.
Secondary Screening Assay
Extracts with LC-MS profiles suggestive of not containing known compounds were tested against the Caco-2 model in a 5-point dose-response dilution pattern in duplicate for confirmation of activity ( Fig. 1E ). The first point was the initial dilution previously described for each assay (1/20 × WBE), and the four subsequent points were twofold serial dilutions from each point. Those exhibiting inhibition of IL-8 production ≥60% and cell viability ≥75% at initial dilution were selected for medium scale-up fermentation and bioactivity confirmation.
Medium Scale-Up Fermentation and Fractionation Guided by Bioassay
Extracts with confirmed activity in secondary screening were regrown in 100 mL volume ( Fig. 1F ). An equal volume of acetone was added to each 100 mL fermentation, and the mixture was homogenized and shaken in a Kühner shaker at 220 rpm for 3 h. From each extract, 13 mL were transferred into separate glass tubes and evaporated under a nitrogen stream until only 3.2 mL remained (extract concentration 2 × WBE). From each of these glass tubes, 5 × 520 µL extract volumes were transferred into 5 ABgene plates (AB-0756) using a Multiprobe II liquid handler (PerkinElmer). Aliquots from 1 of these 5 ABgene plates were bioassayed to confirm activity ( Fig. 1F ) before storing the plate at −20 °C as a reference plate. The remaining four ABgene plates of each extract identified as active were centrifuged, each of the extracts was pooled together in a glass vial and loaded onto a 10 g SP207ss charged column, and the flow-through was collected and stored at −20 °C. The column was then washed with an equal volume of Milli-Q water, and the aqueous flow-through was also collected and saved in a new glass vial at −20 °C. The column was finally eluted to dryness with 20 mL of acetone. The collected solutions were evaporated under a nitrogen gas to yield a dry crude extract. Each crude extract was then reconstituted in 400 µL DMSO, and half of this volume was injected and separated into 80 fractions ( Fig. 1G ) using a linear gradient of 5% to 100% acetonitrile–water with 0.1% trifluoroacetic acid in reverse-phase, semipreparative, high-performance liquid chromatography (HPLC) using Agilent column Zorbax RX-C8, 5 µm, 9.4 × 250 mm, with UV detection at 210 nm (Gilson, Middleton, WI). The 80 fractions collected were evaporated to dryness in an ABgene plate using a Genevac evaporator (Genevac, Ipswich, UK). These fractions were then reconstituted in 30 µL of DMSO and 150 µL of Milli-Q water to make a final solution at 20% DMSO concentration. The fractions were then tested for activity following the previously described screening paradigm. Active fractions from this stage were then analyzed by liquid chromatography high resolution mass spectrometry (LC-HRMS) and NMR ( Fig. 1H ). The extracts in which novel components are confirmed will be then regrown in 1 L volumes for the purification of the active compounds by bioassay-guided compound isolation ( Fig. 1I ).
Evaluation of Antimicrobial Activity
Evaluation of the antimicrobial activity of the fractions was performed using a broth microdilution technique against various clinically relevant strains (all from Fundación MEDINA’s collection). Antibacterial susceptibility was tested against Pseudomonas aeruginosa PAO1, Acinetobacter baumannii CL5973, Escherichia coli MB2884 (wild type), Escherichia coli MB5746 (envA/tolC), and methicillin-resistant Staphylococcus aureus MB5393, whereas antifungal susceptibility was tested against Candida albicans MY1055. Previously described methods were used to test for antimicrobial and antifungal activities. 13 Experiments were made in triplicate and on different days. The Genedata Screener software was used to process and analyze the data.
Growth Scale-Up Bioassay-Guided Purification
One of the hit extracts containing compounds of interest detected in the previous step was growth again at 1 L scale using the appropriate production conditions. The producing organism (CF-097474) was isolated from the surface-disinfected stems and leaves of Juniperus thurifera collected in Peralejos de las Truchas, Guadalajara, Spain, using a method for isolating fungal endophytes. Frozen stock cultures in 10% glycerol (–80 °C) are maintained in the collection of Fundación MEDINA. Total genomic DNA was extracted from mycelia grown on YM agar. The 28S rDNA fragments, containing D1-D2 and ITS regions, were PCR amplified and sequence aligned. 14 Sequence homology searches in public databases (www.fungalbarcoding.org; NCBI) indicated that the fungus had affinities to specie to Seimatosporium lichenicola CBS 506.71. Extensive taxonomical analysis placed this strain in the Seimatosporium genus (data not shown). Seimatosporium sp. (CF-097474) was fermented by inoculating 10 mycelial agar plugs into SMYA medium (Bactoneopeptone 10 g; maltose 40 g; yeast extract 10 g; agar 3 g; water 1 L) into an Erlenmeyer flask (50 mL medium in a 250 mL flask). The flask was incubated in a rotary shaker at 220 rpm at 22 °C with 70% relative humidity (RH). After growing the seed stage for 7 d, a 3 mL aliquot was used to inoculate each flask of the production medium CYS80 (sucrose 80 g, corn meal 50 g, yeast extract 1 g, water 1 L). Ten flasks (100 mL medium per 500 mL Erlenmeyer flask) were incubated at 22 °C with 70% RH in a rotary shaker at 220 rpm for 14 d. A 1 L fermentation was carried out for purifying the compounds. An equal volume of acetone was added to the growth broth, and the mixture was homogenized by shaking in a Kühner cabinet at 220 rpm for 3 h. The remaining solution was filtered and the acetone was evaporated under a nitrogen stream. This solution was loaded (with continuous 1:1 water dilution, discarding the flow-through) on a column packed with SP-207SS reversed-phase resin (brominated styrenic polymer, 65 g) previously equilibrated with water. The loaded column was further washed with water (1 L) and afterward eluted at 10 mL/min on an automatic flash-chromatography system (CombiFlash Rf, Teledyne Isco) using a 30 min step gradient from 10% to 100% acetone in water with a final 100% acetone step (for 12 min), collecting 19 fractions of 20 mL. Fractions were submitted to be tested in the primary screening, and fraction number 9 (gradient step at 40% acetone) showed the desired activity. Fraction number 9 was subjected to semipreparative HPLC (Agilent Zorbax RX-C8, 9.4 × 250 mm, 5 µm; 3.6 mL/min, UV detection at 210 nm, gradient water + 0.1% trifluoroacetic:acid:acetonitrile + 0.1% trifluoroacetic acid from 5% to 100% acetonitrile for 35 min), yielding a fraction eluting at 15.0 min that retained the target activity and that was further purified by semipreparative HPLC (Agilent Zorbax RX-C8, 9.4 × 250 mm, 5 µm: 3.6 mL/min, UV detection at 210 nm, gradient water + 0.1% trifluoroacetic acid:acetonitrile + 0.1% trifluoroacetic acid from 25% to 40% acetonitrile for 35 min) to yield 2.5 mg of an epimer mixture of TAN-2483A and TAN-2483B. 11
Results
Optimization of Assays in a 96-Well Format
The objective was to optimize the experimental model of NO production in mouse macrophages in response to LPS stimulation for HTS in a 96-well format. The influence of cell density, LPS concentration, LPS stimulation duration, and culture medium volume on NO production was investigated. As a result, the macrophage density and the LPS concentration were identified as the main factors affecting LPS-stimulated NO production in mouse macrophages. The density of macrophages produced the optimal effect on NO production at 1 × 105 cells/mL in a volume of 200 µL in each well (20 × 103 cells/well) of a 96-well plate (
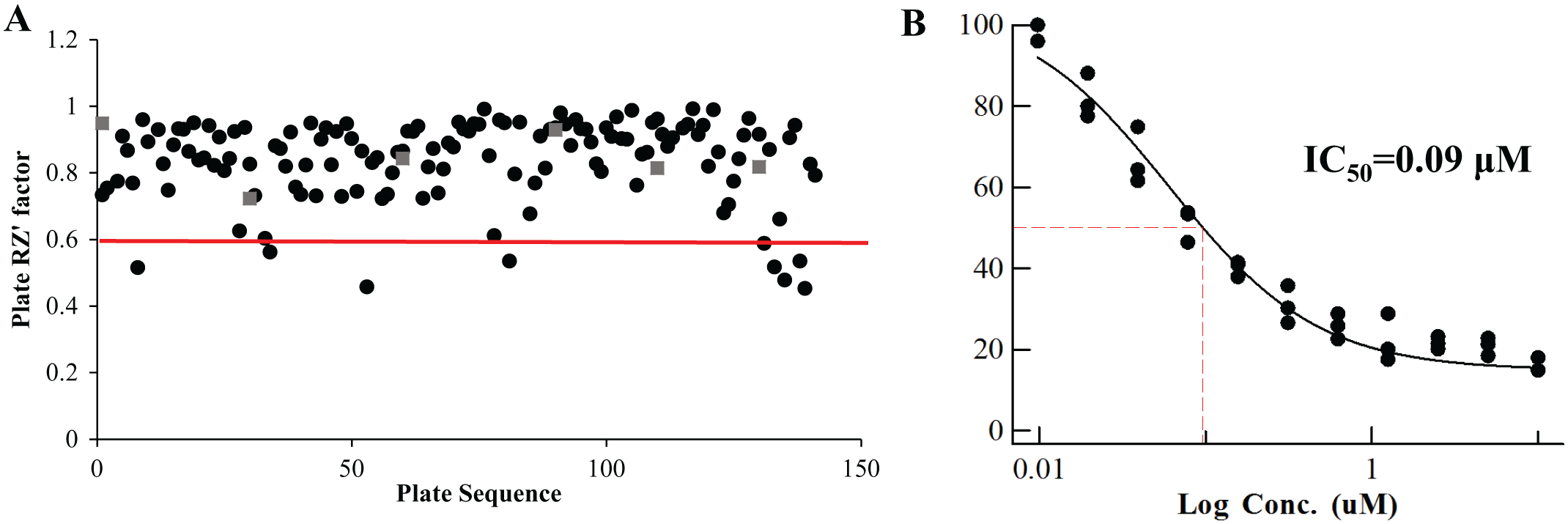
Quality control (QC) in high-throughput screening. To assess the quality of screening data in the screening campaign, each 96-well plate contained a number of control wells (four positive and four negative control wells), which were used for calculation of QC parameters (RZ′ factor and IC50 values of pharmacological standards). (
Screening Campaigns and Hit Identification
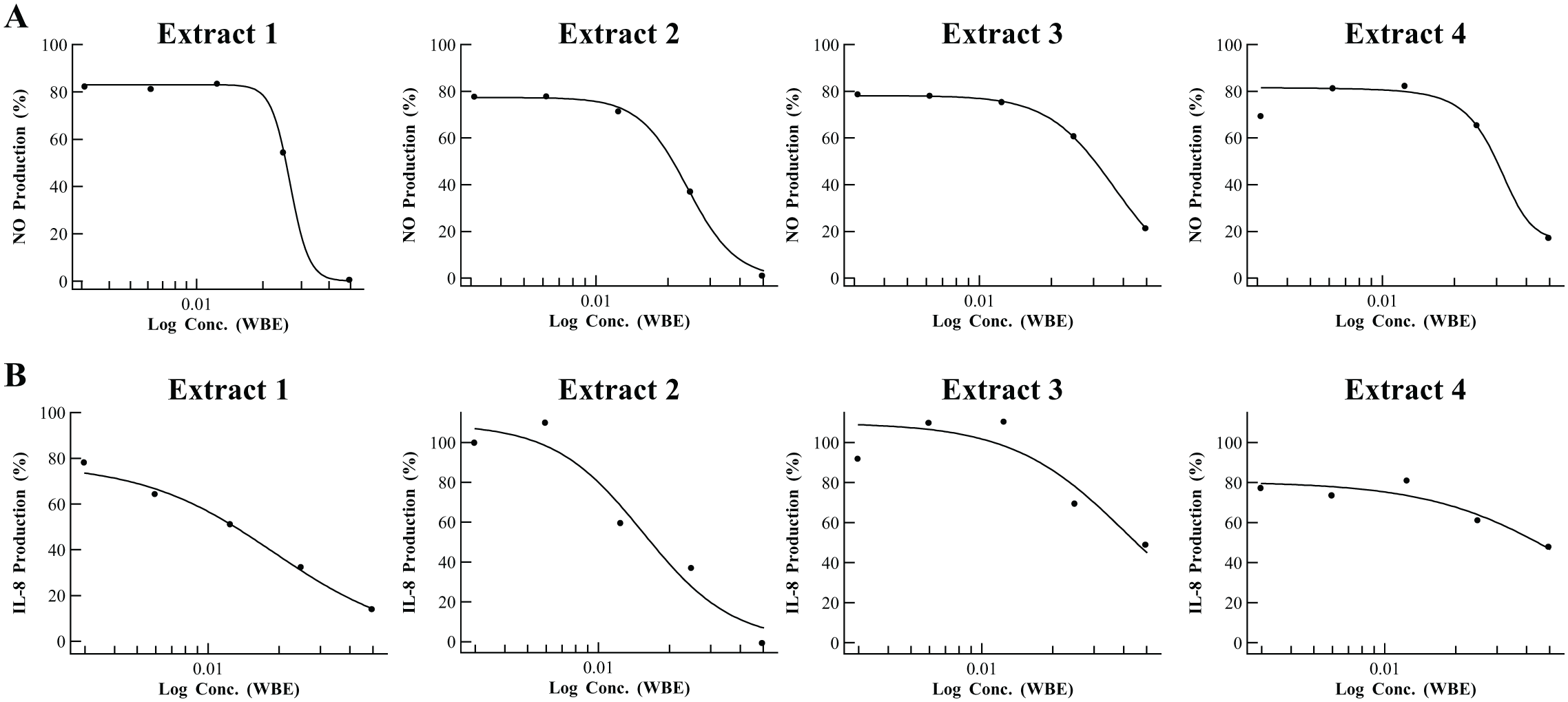
From the primary screening of 5976 noncytotoxic microbial extracts from the MEDINA microbial extract collection (general workflow and screening process shown in Fig. 1 and Table 1 , respectively), 79 extracts (1.32%) were initially identified as hits. We assessed the activities of the selected extracts again in triplicate by cherry-picking, and 34 extracts were confirmed as NO inhibition hits. Next, we used dose-response experiments to investigate the extract potency to select only those exhibiting at least 70% of NO inhibition at the initial extract dilution ( Fig. 3A ). Using this criterion, 14 inhibitor extracts were selected as hits for LC-MS de-replication. Results showed that none of them obtained a positive identification in the spectral metabolite database. Within these extracts, those showing steeper dose-response curve slopes (12 extracts) were selected for confirmation of immunomodulatory activity in the secondary screening assay. In these results, 4 extracts out of 12 showed inhibition of IL-8 production ≥60% at the initial extract dilution ( Fig. 3B ) and cell viability ≥75%, and they were selected for medium scale-up regrowths.
Number of Extracts Identified as Hits at Different Stages of the Screening Process. a
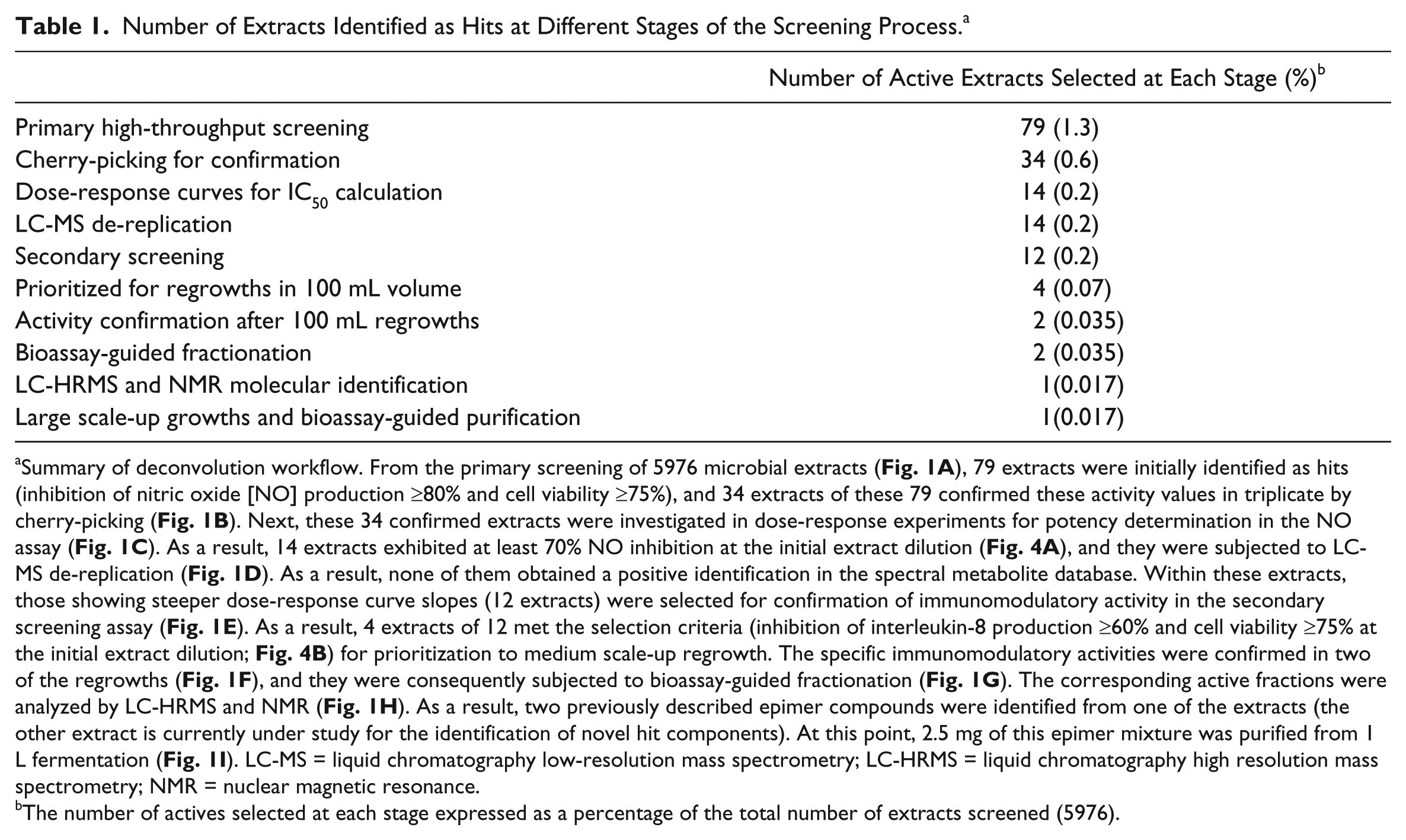
Summary of deconvolution workflow. From the primary screening of 5976 microbial extracts ( Fig. 1A ), 79 extracts were initially identified as hits (inhibition of nitric oxide [NO] production ≥80% and cell viability ≥75%), and 34 extracts of these 79 confirmed these activity values in triplicate by cherry-picking ( Fig. 1B ). Next, these 34 confirmed extracts were investigated in dose-response experiments for potency determination in the NO assay ( Fig. 1C ). As a result, 14 extracts exhibited at least 70% NO inhibition at the initial extract dilution ( Fig. 4A ), and they were subjected to LC-MS de-replication ( Fig. 1D ). As a result, none of them obtained a positive identification in the spectral metabolite database. Within these extracts, those showing steeper dose-response curve slopes (12 extracts) were selected for confirmation of immunomodulatory activity in the secondary screening assay ( Fig. 1E ). As a result, 4 extracts of 12 met the selection criteria (inhibition of interleukin-8 production ≥60% and cell viability ≥75% at the initial extract dilution; Fig. 4B ) for prioritization to medium scale-up regrowth. The specific immunomodulatory activities were confirmed in two of the regrowths ( Fig. 1F ), and they were consequently subjected to bioassay-guided fractionation ( Fig. 1G ). The corresponding active fractions were analyzed by LC-HRMS and NMR ( Fig. 1H ). As a result, two previously described epimer compounds were identified from one of the extracts (the other extract is currently under study for the identification of novel hit components). At this point, 2.5 mg of this epimer mixture was purified from 1 L fermentation ( Fig. 1I ). LC-MS = liquid chromatography low-resolution mass spectrometry; LC-HRMS = liquid chromatography high resolution mass spectrometry; NMR = nuclear magnetic resonance.
The number of actives selected at each stage expressed as a percentage of the total number of extracts screened (5976).
(
Early LC-MS De-Replication
A database search performed using our in-house application, which matches the UV-LC-MS data of the metabolites in the active extracts to the UV-LC-MS data of known metabolites stored in our proprietary database obtained using the exact same LC-MS conditions, did not identify any metabolites in the 14 previously mentioned active extracts, and therefore, these extracts were classified as potentially not containing known active components. As proof of concept to assess the ability of this HTS platform to effectively retrieve already known immunomodulatory compounds from microbial fermentations, we also subjected the remaining 20 active extracts from primary screening that did not meet the de-replication selection criteria described above (extracts that had lower than 70% inhibition at the initial extract dilution in the dose response experiment) to LC-MS de-replication. As a result, we identified profiles associated with known secondary metabolites, such as diketocoriolin B, a secondary metabolite of fungi (Coriolus consors) with immunostimulant activity described, or eremoxylarin A, a secondary metabolite of xylariaceous endophytic fungus strain YUA-026 with calcineurin inhibitor activity. 21
Medium and Large Scale-Up Growths for Bioassay-Guided Fractionation
At this stage, the four previously selected active extracts identified as potentially not containing known active components were prioritized, and the producing strains were regrown in 100 mL volumes. The specific immunomodulatory activities were confirmed in the regrowths for two of them ( Fig. 3 , extracts 1 and 2), and consequently, they were submitted to fractionation guided by bioassays. Simultaneously, we ran LC-HRMS and NMR analyses of the active semipreparative HPLC fractions of each of the two active extracts to assess the novelty of the active components. As a result, two previously described epimer compounds (1:3) were identified (TAN-2483A and TAN-2483B) 11 in a highly purified fraction (concentration 245 µg/mL) from extract 1 (CF-097474; Fig. 4 ). These compounds are members of the gamma-lactone family, and they were previously isolated by the Takeda Chemical Industries group22,23 from the filamentous fungus NF 2329,3, but no immunomodulatory activity has been reported for this compound so far. Therefore, it was tested for immunomodulatory activity. Determination of the IC50 value in NO production showed the compound to be active with an IC50 value of 2.26 µM ( Fig. 5A ). In addition, it also showed strong inhibition of IL-8 production (100%) at 29.1 µM. At this point, 2.5 mg of this epimer mixture was purified from 1L fermentation of Seimatosporium (CF-097474) and will be assessed for immunomodulatory activity in an animal inflammation model. Regarding extract 2, it is currently under study for the identification of novel hit components.
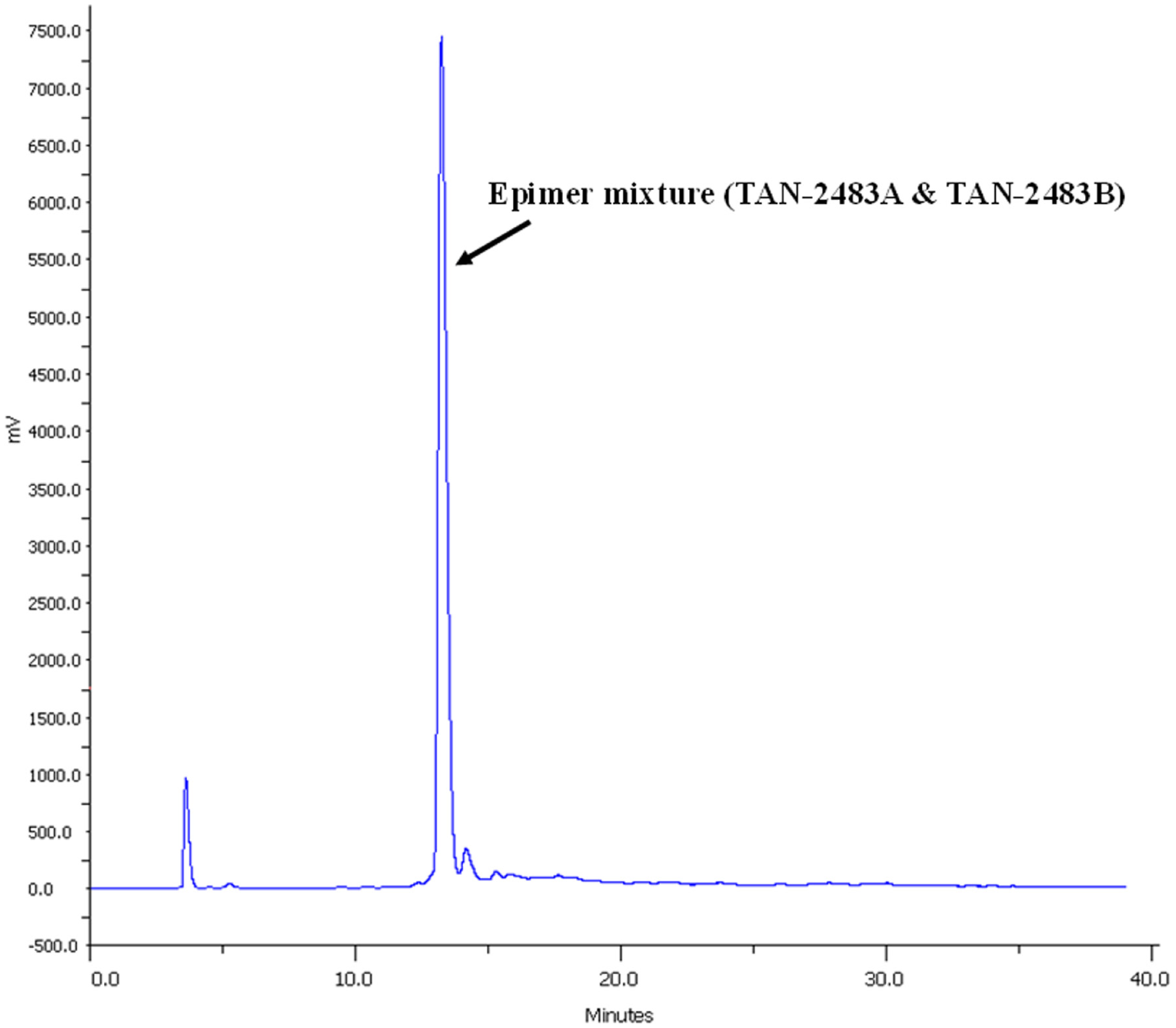
Ultraviolet (210 nm) chromatogram of high-performance liquid chromatography fractionation of active extract containing the epimer mixture (TAN-2483A and TAN2483B). A highly purified fraction of these compounds was collected at 13 min, corresponding to the most intense peak.
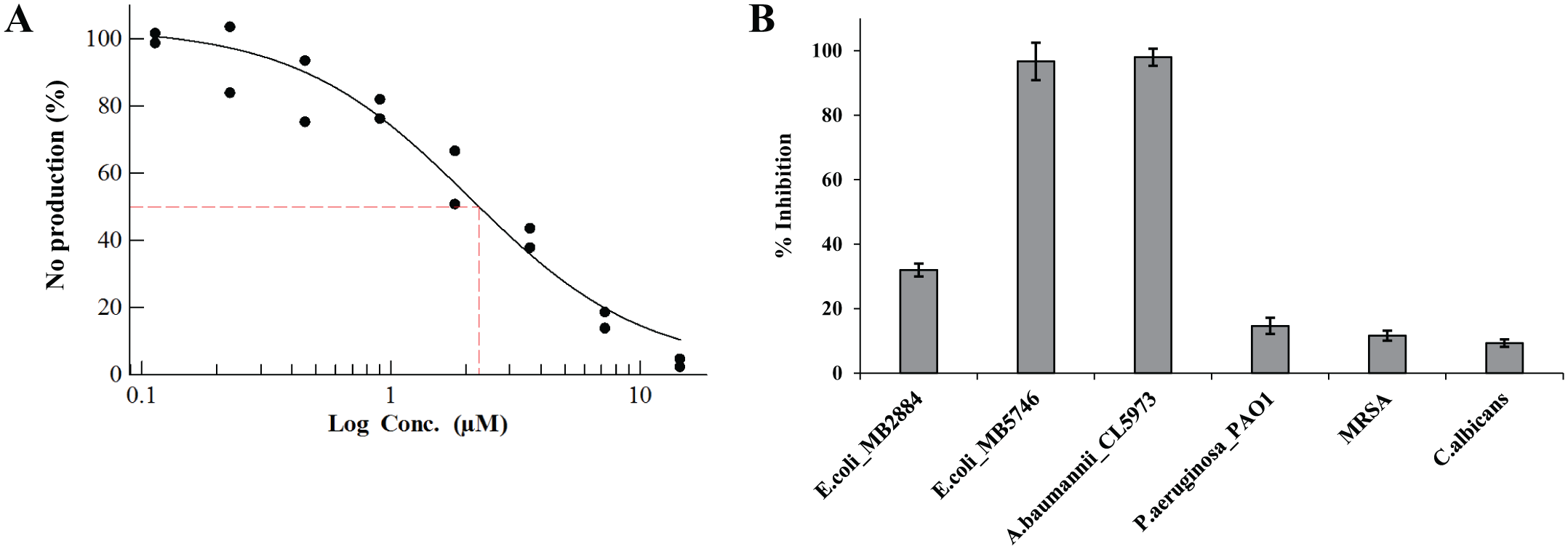
Bioactivity results of high-performance liquid chromatography fraction containing the epimer mixture (TAN-2483A and TAN2483B). (
Evaluation of Antimicrobial Activity
Considering the long and successful track record of microbial fermentations yielding new antibiotic compounds, the highly purified extract fraction containing TAN-2483A and TAN-2483B was evaluated for antimicrobial properties. As shown in Figure 5B , this fraction was able to inhibit strongly the growth of -gram negative A. baumannii CL5973 (100%) and E. coli MB5746 (100%), whereas moderate inhibition (37%) was observed for E. coli MB2884 and no effect was observed against methicillin-resistant S. aureus MB5393 or C. albicans MY1055 at this concentration (24 µg/mL).
Discussion
It has been long reported that neurodegenerative diseases (NDD), including Alzheimer’s disease, are characterized by redox imbalance, glial cell activation, neuroinflammation, and ultimately neuronal death. 24 In this context, NF-κB plays an important role in the pathogenesis of oxidative stress-associated neurodegenerative diseases, because this is one of the most important transcriptional factors of proinflammatory gene expression. It facilitates the synthesis of proinflammatory cytokines, such as tumor necrosis factor–α, IL-1β, IL-6, and IL-8, as well as of inducible enzymes, such as iNOS, which produces large amounts of NO, all of which can contribute to neurodegeneration. The suppression of the production of these proinflammatory mediators can therefore offer protection from neuronal damage in neurodegenerative conditions by interfering with the inflammatory cascade at different levels (mainly enzyme inhibition and down-regulation of gene transcription), as previously demonstrated in animal models. 25 For this reason, the measurements of NO or IL-8 in cultured cells are popular assays for identification of anti-inflammatory compounds of pharmacological interest. However, most applications previously described are devoted to a 24-well plate format or manual liquid dispensing, which, for whole cell assay–based HTS campaigns, is costly in terms of time and economic resources. In recent years, advances in spectrophotometry-based detection techniques and automation technologies have made feasible the adaptation of these assays to the 96-well microplate miniaturized format, in which assay volumes can be decreased and plate throughput increased several fold. This platform has provided a robust, reliable, and comprehensive methodology in the 96-well format for the discovery of natural product–based drugs with immunomodulatory properties meeting the demanding requirements of HTS drug discovery. This novel adaptation of NO and IL-8 assays to HTS format, using highly automated liquid handling and high-sensitivity signal detection systems, delivers a substantial reduction in the consumption of cells, sample volume, and other expensive reagents as well as a shorter turnaround time. This allows large screening campaigns in a reduced time frame, as typically required in natural product discovery. Moreover, this approach to microbial fermentations enables access to the unfathomable depths of chemical diversity in natural products, which consists of as many diverse chemotypes as there are species and strains in this realm. Useful information may be drawn from this experience. First, the general workflow used ( Fig. 1 ) ensures that only extracts that have been tested and proven, through rigorous methodology, to consistently contain active and potentially novel metabolites with immunomodulatory activity are selected for further development. Second, our efficiently integrated LC-MS de-replication system also enables us to pick up on active extracts containing already known compounds for which the immunomodulatory activity has not been previously reported. In this regard, the results confirming the immunomodulatory activity of the TAN-2483A and TAN-2483B epimer mixture (3:1) further validate the strategy used in the present screening approach. These known bioactive compounds have not been previously reported to exhibit immunomodulatory activity. However, TAN-2483A has displayed strong inhibitory action in Src kinase, a human protein kinase that plays a pivotal role in macrophage-mediated inflammatory responses. 26 In fact, a variety of inflammatory diseases are closely related to macrophage activation. The critical role of Src in macrophage activation has prompted researchers to consider that the inhibition of Src activity may be a useful therapeutic strategy for macrophage-mediated diseases; specifically, Src has been described as a suitable therapeutic target for the treatment of NDD involving the activation of pathological microglia. 27 Taking into consideration other works highlighting the anti-inflammatory potential of natural products inhibitors of Src activity, 28 we expect that novel and safe compounds exhibiting strong immunosuppressive and anti-inflammatory properties might be discovered and might contribute to the development of innovative therapies for the treatment of macrophage-mediated diseases. In addition, it has been demonstrated that certain antibiotics have potential as neurotherapeutics for treating neurological diseases such as amyotrophic lateral sclerosis, adult motor neuron disease, and ischemic injury. 29 In multiple sclerosis—the archetypal inflammatory response in the central nervous system—T cells and macrophages invade the brain and damage the myelin and neurons. These macrophages are primed by components of neuropathology but might be further activated by systemic infection, which in turn has pronounced effects on inflammation in the brain and perhaps on neurological function. There is emerging evidence to support the idea that non-specific systemic infection or inflammation in people with existing inflammation in the brain contributes to the disease progression rate through further activation of these already primed macrophages. 30 Therefore, the use of a compound with both immunomodulatory and antibacterial properties could be very useful in the treatment of inflammation in NDD. The results from antimicrobial in vitro activity of the purified epimer mixture (TAN-2483A and TAN-2483B) underline the ability and the specificity of these compounds against gram-negative bacteria. This is important when set against the difficulty in inhibiting these kinds of pathogens and suggests the potential of microbial extracts as a source of compounds with dual activity. Clearly, further investigations will be needed to determine the modes of action and establish the association of antibacterial and immunomodulatory properties. Furthermore, we can confidently infer from the LC-MS de-replication data that the screening platform can effectively determine known active immunomodulatory compounds (diketocoriolin B and eremoxylarin A) from crude extracts, thus proving that we have a working system that can be used to de-replicate extracts that contain compounds with immunomodulatory activities. Finally, the disadvantage of this de-replication step is that potentially active metabolites that may be present at undetectable levels in these crude extracts may be overshadowed by more abundant cytotoxic metabolites and consequently missed. It is our hope that, through this work, we will generate more interest in natural products as a source of novel chemotypes to meet global medical challenges.
Footnotes
Acknowledgements
We express our thanks to Alan Mathes (ITC/Grad. Dip. Phys.) for proofreading.
Authors’ Note
The results presented in this work will be compiled in the doctoral thesis (knowledge area code 32089 and subject code 320903) titled “Evaluación de la actividad de los metabolitos hepáticos derivados de compuestos inmunomodulares e inhibidores de GSK3,” carried out currently by José Pérez del Palacio at Fundación MEDINA in collaboration with the Department of Pharmacology at the University of Granada and whose thesis directors are Francisca Vicente (Fundación MEDINA) and Julio Gálvez (Universidad de Granada).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Spanish Ministry of Economy and Competitiveness (SAF2011-29648) and the Junta de Andalucía (AGR-6826 and CTS 164) with funds from the European Union. F. Algieri is a predoctoral fellow of the Junta de Andalucía; M. E. Rodríguez-Cabezas is a postdoctoral fellow of CIBER-EHD. The CIBER-EHD is funded by the Instituto de Salud Carlos III. The MEDINA authors disclosed the receipt of financial support from Fundación MEDINA, a public-private partnership of Merck Sharp & Dohme de España S.A./Universidad de Granada/Junta de Andalucía. Moreover, some of the equipment used in this work was supported by the Ministerio de Ciencia e Innovación and European Union (grant INP-2011-0016-PCT-010000-ACT6).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.